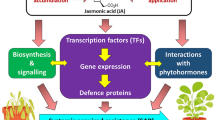
Abstract
Key message
Knocking out OsVQ1 in rice released OsMPK6 for activation and in turn promoted H 2 O 2 accumulation, which repressed the expression of flowering-promoting genes, thus delaying rice flowering but enhancing disease resistance.
Abstract
The valine–glutamine (VQ) protein family, which contains the conserved motif FxxxVQxLTG (“x” represents any amino acid), plays a crucial role in plant growth and immunity along with mitogen-activated protein kinase (MAPK) cascades. However, only a few rice VQ proteins have been functionally characterized, and the roles of the MAPK-VQ module in rice biological processes are not fully understood. Here, we investigated the role of OsVQ1 in rice disease resistance and the control of flowering time. The OsVQ1-knock out (KO) mutants exhibited increased resistance to Xanthomonas oryzae pathovars, accumulated high levels of hydrogen peroxide (H2O2), and showed a late flowering phenotype under natural long-day conditions, while the OsVQ1-overexpressing plants showed phenotypes similar to that of the wild type. Further studies revealed that OsVQ1 physically interacted with and inhibited OsMPK6 activity. In addition, OsVQ1 expression was downregulated by the pathogen-induced OsMPKK10.2-OsMPK6-OsWRKY45 cascade, suggesting a feedback loop between OsVQ1 and OsMPK6. Moreover, the OsVQ1-KO/osmpk6 double-mutant exhibited increased susceptibility to X. oryzae infection and showed an early flowering phenotype, which may partially be attributed to the reduced accumulation of H2O2 and the consequent up-expression of flowering-promoting genes. These results suggested that the OsVQ1-OsMPK6 module was involved in rice immunity and flowering.








Similar content being viewed by others
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Bi G, Zhou Z, Wang W et al (2018) Receptor-like cytoplasmic kinases directly link diverse pattern recognition receptors to the activation of mitogen-activated protein kinase cascades in Arabidopsis. Plant Cell 30:1543–1561. https://doi.org/10.1105/tpc.17.00981
Cao Y, Ding X, Cai M et al (2007) The expression pattern of a rice disease resistance gene Xa3/Xa26 is differentially regulated by the genetic backgrounds and developmental stages that influence its function. Genetics 177:52–533. https://doi.org/10.1534/genetics.107.075176
Chen H, Wang S, Zhang Q (2002) New gene for bacterial blight resistance in rice located on chromosome 12 identified from Minghui 63, an elite restorer line. Phytopathology 92:750–754. https://doi.org/10.1094/PHYTO.2002.92.7.750
Fan J, Bai P, Ning Y et al (2018) The monocot-specific receptor-like kinase SDS2 controls cell death and immunity in rice. Cell Host Microbe 23:498–510. https://doi.org/10.1016/j.chom.2018.03.003
Glander S, He F, Schmitz G et al (2018) Assortment of flowering time and immunity alleles in natural Arabidopsis thaliana populations suggests immunity and vegetative lifespan strategies coevolve. Genome Biol Evol 10:2278–2291. https://doi.org/10.1093/gbe/evy124
He Y, Zhang T, Yang N et al (2017) Self-cleaving ribozymes enable the production of guide RNAs from unlimited choices of promoters for CRISPR/Cas9 mediated genome editing. J Genet Genomics 44:469–472. https://doi.org/10.1016/j.jgg.2017.08.003
Hiei Y, Ohta S, Komari T et al (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282. https://doi.org/10.1046/j.1365-313x.1994.6020271.x
Huang X, Chen S, Li W et al (2021) ROS regulated reversible protein phase separation synchronizes plant flowering. Nat Chem Biol 17:549–557. https://doi.org/10.1038/s41589-021-00739-0
Ishimoto K, Sohonahra S, Kishi-Kaboshi M et al (2019) Specification of basal region identity after asymmetric zygotic division requires mitogen-activated protein kinase 6 in rice. Development 146:dev176305. https://doi.org/10.1242/dev.176305
Jones JD, Dangl JL (2006) The plant immune system. Nature 444:323–329. https://doi.org/10.1038/nature05286
Kazan K, Lyons R (2016) The link between flowering time and stress tolerance. J Exp Bot 67:47–60. https://doi.org/10.1093/jxb/erv441
Ke Y, Wu M, Zhang Q et al (2019) Hd3a and OsFD1 negatively regulate rice resistance to Xanthomonas oryzae pv. oryzae and Xanthomonas oryzae pv. oryzicola. Biochem Biophys Res Commun 513:775–780. https://doi.org/10.1016/j.bbrc.2019.03.169
Kishi-Kaboshi M, Okada K, Kurimoto L et al (2010) A rice fungal MAMP-responsive MAPK cascade regulates metabolic flow to antimicrobial metabolite synthesis. Plant J 63:599–612. https://doi.org/10.1111/j.1365-313X.2010.04264.x
Li Y, Dong O, Johnson K et al (2011) MOS1 epigenetically regulates the expression of plant Resistance gene SNC1. Plant Signal Behav 6:434–436. https://doi.org/10.4161/psb.6.3.14495
Li N, Li X, Xiao J et al (2014) Comprehensive analysis of VQ motif-containing gene expression in rice defense responses to three pathogens. Plant Cell Rep 33:1493–1505. https://doi.org/10.1007/s00299-014-1633-4
Li N, Yang Z, Li J et al (2021) Two VQ proteins are substrates of the OsMPKK6-OsMPK4 cascade in rice defense against bacterial blight. Rice 14:39. https://doi.org/10.1186/s12284-021-00483-y
Linden KJ, Callis J (2020) The ubiquitin system affects agronomic plant traits. J Biol Chem 295:13940–13955. https://doi.org/10.1074/jbc.REV120.011303
Liu J, Li W, Ning Y et al (2012) The U-Box E3 ligase SPL11/PUB13 is a convergence point of defense and flowering signaling in plants. Plant Physiol 160:28–37. https://doi.org/10.1104/pp.112.199430
Liu S, Hua L, Dong S et al (2015) OsMAPK6, a mitogen-activated protein kinase, influences rice grain size and biomass production. Plant J 84:672–681. https://doi.org/10.1111/tpj.13025
Liu H, Ding Y, Zhou Y et al (2017) CRISPR-P 2.0: An improved CRISPR-Cas9 tool for genome editing in plants. Mol Plant 10:530–532. https://doi.org/10.1016/j.molp.2017.01.003
Lyons R, Rusu A, Stiller J et al (2015) Investigating the association between flowering time and defense in the Arabidopsis thaliana-Fusarium oxysporum interaction. PLoS ONE 10:e0127699. https://doi.org/10.1371/journal.pone.0127699
Ma H, Chen J, Zhang Z et al (2017) MAPK kinase 10.2 promotes disease resistance and drought tolerance by activating different MAPKs in rice. Plant J 92:557–570. https://doi.org/10.1111/tpj.13674
Ma H, Li J, Ma L et al (2021) Pathogen-inducible OsMPKK10.2-OsMPK6 cascade phosphorylates the Raf-like kinase OsEDR1 and inhibits its scaffold function to promote rice disease resistance. Mol Plant 14:620–632. https://doi.org/10.1016/j.molp.2021.01.008
Meng X, Zhang S (2013) MAPK cascades in plant disease resistance signaling. Annu Rev Phytopathol 51:245–266. https://doi.org/10.1146/annurev-phyto-082712-102314
Nakagami H, Soukupova H, Schikora A et al (2006) A mitogen-activated protein kinase kinase kinase mediates reactive oxygen species homeostasis in Arabidopsis. J Biol Chem 281:38697–38704. https://doi.org/10.1074/jbc.M605293200
Pecher P, Eschen-Lippold L, Herklotz S et al (2014) The Arabidopsis thaliana mitogen-activated protein kinases MPK3 and MPK6 target a subclass of ‘VQ-motif’-containing proteins to regulate immune responses. New Phytol 203:592–606. https://doi.org/10.1111/nph.12817
Pitzschke A, Djamei A, Bitton F et al (2009) A major role of the MEKK1-MKK1/2-MPK4 pathway in ROS signalling. Mol Plant 2:120–137. https://doi.org/10.1093/mp/ssn079
Qiu D, Xiao J, Ding X et al (2007) OsWRKY13 mediates rice disease resistance by regulating defense-related genes in salicylate- and jasmonate-dependent signaling. Mol Plant Microbe Interact 20:492–499. https://doi.org/10.1094/MPMI-20-5-0492
Qiu JL, Fiil BK, Petersen K et al (2008) Arabidopsis MAP kinase 4 regulates gene expression through transcription factor release in the nucleus. EMBO J 27:2214–2221. https://doi.org/10.1038/emboj.2008.147
Shen X, Yuan B, Liu H et al (2010) Opposite functions of a rice mitogen-activated protein kinase during the process of resistance against Xanthomonas oryzae. Plant J 64:86–99. https://doi.org/10.1111/j.1365-313X.2010.04306.x
Sheng P, Wu F, Tan J et al (2016) A CONSTANS-like transcriptional activator, OsCOL13, functions as a negative regulator of flowering downstream of OsphyB and upstream of Ehd1 in rice. Plant Mol Biol 92:209–222. https://doi.org/10.1007/s11103-016-0506-3
Stanko V, Giuliani C, Retzer K et al (2014) Timing is everything: highly specific and transient expression of a MAP kinase determines auxin-induced leaf venation patterns in Arabidopsis. Mol Plant 7:1637–1652. https://doi.org/10.1093/mp/ssu080
Tang Q, Guittard-Crilat E, Maldiney R et al (2016) The mitogen-activated protein kinase phosphatase PHS1 regulates flowering in Arabidopsis thaliana. Planta 243:909–923. https://doi.org/10.1007/s00425-015-2447-5
Tao Z, Liu H, Qiu D et al (2009) A pair of allelic WRKY genes play opposite roles in rice-bacteria interactions. Plant Physiol 151:936–948. https://doi.org/10.1104/pp.109.145623
Ueno Y, Yoshida R, Kishi-Kaboshi M et al (2015) Abiotic stresses antagonize the rice defence pathway through the tyrosine-dephosphorylation of OsMPK6. PLoS Pathog 11:e1005231. https://doi.org/10.1371/journal.ppat.1005231
Uji Y, Kashihara K, Kiyama H et al (2019) Jasmonic acid-induced VQ-motif-containing protein OsVQ13 influences the OsWRKY45 signaling pathway and grain size by associating with OsMPK6 in rice. Int J Mol Sci 20:2917. https://doi.org/10.3390/ijms20122917
Vega-Sanchez ME, Zeng L, Chen S et al (2008) SPIN1, a K homology domain protein negatively regulated and ubiquitinated by the E3 ubiquitin ligase SPL11, is involved in flowering time control in rice. Plant Cell 20:1456–1469. https://doi.org/10.1105/tpc.108.058610
Xiong L, Yang Y (2003) Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid-inducible mitogen-activated protein kinase. Plant Cell 15:745–759. https://doi.org/10.1105/tpc.008714
Yamada K, Yamaguchi K, Shirakawa T et al (2016) The Arabidopsis CERK1-associated kinase PBL27 connects chitin perception to MAPK activation. EMBO J 35:2468–2483
Zeng LR, Qu S, Bordeos A et al (2004) Spotted leaf11, a negative regulator of plant cell death and defense, encodes a U-box/armadillo repeat protein endowed with E3 ubiquitin ligase activity. Plant Cell 16:2795–2808. https://doi.org/10.1105/tpc.104.025171
Zhang H, Tao Z, Hong H et al (2016) Transposon-derived small RNA is responsible for modified function of WRKY45 locus. Nat Plants 2:16016. https://doi.org/10.1038/nplants.2016.16
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31772145 to S.W. and 31901865 to H.M.).
Author information
Authors and Affiliations
Contributions
WP, MH, and WS designed the study; WP, LJ, ZZ, and MH performed the experiments and analyzed the data; ZQ, LX, and XJ provided experiments support; MH wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Consent to participate
Not applicable.
Consent for publication
The authors confirm that the work described has not been published before; that it is not under consideration for publication anywhere else; that its publication has been approved by all co-authors.
Additional information
Communicated by Ying-Tang Lu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
299_2021_2766_MOESM1_ESM.pptx
Supplementary file1 Supplementary Fig. 1 The sequencing results of off-target sites of target site 1 (a) and 2 (b) in OsVQ1-KO plants. The putative off-target sites are indicated with rectangles. Supplementary Fig. 2 OsVQ1 expression in OsVQ1 overexpressing (oe) transgenic plants. Bars represent mean (3 replicates) ± SE. (PPTX 155 KB)
299_2021_2766_MOESM2_ESM.doc
Supplementary file2 Supplementary Table 1 Primers used for vector construction and gene expression analysis. Supplementary Table 2 The putative off-target sites of CRISPR/Cas9 system in OsVQ1-KO plants. (DOC 61 KB)
Rights and permissions
About this article
Cite this article
Wang, P., Li, J., Zhang, Z. et al. OsVQ1 links rice immunity and flowering via interaction with a mitogen-activated protein kinase OsMPK6. Plant Cell Rep 40, 1989–1999 (2021). https://doi.org/10.1007/s00299-021-02766-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02766-6