Abstract
Key message
PeSNAC1 , a stress-related NAC1 from Phyllostachys edulis , was characterized. Ectopic expression in Arabidopsis indicated that PeSNAC1 together with ped -miR164b participated in the regulation of organ boundaries and stress tolerance.
Abstract
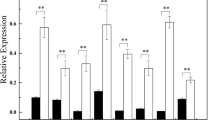
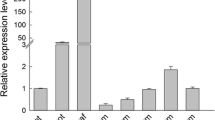
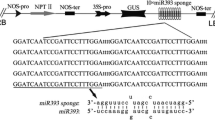
NAC (NAM, ATAF1/2 and CUC2) participates in many different processes regulating plant growth, development, and stress response. A total of 125 NAC genes have been predicted in moso bamboo (Phyllostachys edulis), but their roles are poorly understood. PeSNAC1 targeted by ped-miR164b was focused for further study. The cleavage of PeSNAC1 mRNA guided by ped-miR164b was validated using RLM-5′ RACE. Tissue-specific expression analysis demonstrated that ped-miR164b had a declining trend from root, sheath, leaf, to that of stem, which was opposite to that of PeSNAC1. Transgenic Arabidopsis plants overexpressing either PeSNAC1 (OX-PeSNAC1) or, ped-miR164b (OX-ped-miR164b) driven by the CaMV35S promoter were generated. OX-ped-miR164b plants showed similar phenotype of cuc2 mutants whose growth was seriously suppressed. Compared with Col-0, sense OX-PeSNAC1 plants grew rapidly and flowered earlier, whereas antisense plants grew slowly and exhibited delayed flowering. Sense OX-PeSNAC1 plants had the greatest number of lateral roots, while antisense OX-PeSNAC1 and OX-ped-miR164b plants had fewer lateral roots than Col-0. Under NaCl and PEG6000 stresses, survival rates were higher and F v/F m values declined more slowly in sense OX-PeSNAC1 plants than in Col-0, with lower survival rates and a more rapid decrease in F v/F m values conversely observed in antisense OX-PeSNAC1 and OX-ped-miR164b plants. These findings indicated that ped-miR164b-targeted PeSNAC1 may play key roles in plant development and tolerance to salinity and drought stresses.








Similar content being viewed by others
Abbreviations
- ATAF:
-
Arabidopsis transcription activation factor
- BLAST:
-
Basic local alignment search tool
- CaMV35S :
-
Cauliflower mosaic virus 35S
- cDNA:
-
Complementary DNA
- F v/F m :
-
Maximal quantum yield of PSII
- ORF:
-
Open reading frame
- CUC:
-
Cup-shaped cotyledon (CUC), a NAC protein
- miRNA:
-
MicroRNA, small regulatory RNAs
- NAC:
-
Abbreviation of NAM, ATAF1/2 and CUC2, a plant-specific TF family
- NAM:
-
No apical meristem (NAM), a NAC protein
- PCR:
-
Polymerase chain reaction
- qPCR:
-
Real time quantitative PCR
- RACE:
-
Rapid-amplification of cDNA ends
- RLM-5′ RACE:
-
RNA ligase-mediated 5′ RACE
- TF:
-
Transcription factor
References
Ahuja I, de Vos RC, Bones AM, Hall RD (2010) Plant molecular stress responses face climate change. Trends Plant Sci 15(12):664–674
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler K (2005) Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 33(20):e179
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743
Cui K, He CY, Zhang JG, Duan AG, Zeng YF (2012) Temporal and spatial profiling of internode elongation-associated protein expression in rapidly growing culms of bamboo. J Proteome Res 11(4):2492–2507
Dai X, Zhao PX (2011) psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res 39(Web Server issue):W155–W159
Demmig-Adams B, Adams WWI (1996) Chlorophyll and carotenoid composition in leaves of Euonymus kiautschovicus acclimated to different degrees of light stress in the field. Aust J Plant Physiol 23(5):649–659
Ding Y, Chen Z, Zhu C (2011) Microarray-based analysis of cadmium-responsive microRNAs in rice (Oryza sativa). J Exp Bot 62(10):3563–3573
Fan C, Ma J, Guo Q, Li X, Wang H, Lu M (2013) Selection of reference genes for quantitative real-time PCR in bamboo (Phyllostachys edulis). PLoS One 8(2):e56573
Fang Y, Xie K, Xiong L (2014) Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J Exp Bot 65(8):2119–2135
Gao ZM, Fan SH, Gao J, Li XP, Cai CJ, Peng ZH (2006) Extract genomic DNA from Phyllostachys edulis by CTAB-based method. For Res 19(6):725–728 (in Chinese)
Gruber AR, Lorenz R, Bernhart SH, Neuböck R, Hofacker IL (2008) The Vienna RNA websuite. Nucleic Acids Res 36(Web Server issue):W70–W74
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to down regulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
He CY, Cui K, Zhang JG, Duan AG, Zeng YF (2013) Next-generation sequencing-based mRNA and microRNA expression profiling analysis revealed pathways involved in the rapid growth of developing culms in Moso bamboo. BMC Plant Biol 13:119
Hibara K, Karim MR, Takada S, Taoka K, Furutani M, Aida M, Tasaka M (2006) Arabidopsis CUP-SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18(11):2946–2957
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297
Huang Q, Wang Y, Li B, Chang J, Chen M, Li K, Yang G, He G (2015) TaNAC29, a NAC transcription factor from wheat, enhances salt and drought tolerance in transgenic Arabidopsis. BMC Plant Biol 15:268
Jefferson RA (1987) Assaying for chimeric genes in plants: the GUS fusion system. Plant Mol Biol Rep 5(4):387–405
Jeong DH, Park S, Zhai J, Gurazada SG, De Paoli E, Meyers BC, Green PJ (2011) Massive analysis of rice small RNAs: mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 23(12):4185–4207
Laufs P, Peaucelle A, Morin H, Traas J (2004) MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 131(17):4311–4322
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18(4):263–276
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Li J, Guo G, Guo W, Guo G, Tong D, Ni Z, Sun Q, Yao Y (2012) MiRNA164-directed cleavage of ZmNAC1 confers lateral root development in maize (Zea mays L.). BMC Plant Biol 12:220
Li B, Hu S, Cao Y, Xu G (2015) Bioinformatics analysis of NAC gene family in moso bamboo. Genomics Appl Biol 34(8):1769–1777 (in Chinese)
Lin R, Zhao W, Meng X, Wang M, Peng Y (2007) Rice gene OsNAC19 encodes anovel NAC-domain transcription factor and responds to infection by Magnaporthe grisea. Plant Sci 172(1):120–130
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25(4):402–408
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14(12):1035–1046
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence–a practical guide. J Exp Bot 51(345):659–668
Meshi T, Iwabuchi M (1995) Plant transcription factors. Plant Cell Physiol 36(8):1405–1420
Moldovan D, Spriggs A, Yang J, Pogson BJ, Dennis ES, Wilson IW (2010) Hypoxia-responsive microRNAs and trans-acting small interfering RNAs in Arabidopsis. J Exp Bot 61:165–177
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465(1–2):30–44
Pastori GM, Foyer CH (2002) Common components, networks, and pathways of cross-tolerance to stress. The central role of “Redox” and abscisic acid-mediated controls. Plant Physiol 129(2):460–468
Peng Z, Lu Y, Li L, Zhao Q, Feng Q, Gao Z, Lu H, Hu T, Yao N, Liu K, Li Y, Fan D, Guo Y, Li W, Lu Y, Weng Q, Zhou C, Zhang L, Huang T, Zhao Y, Zhu C, Liu X, Yang X, Wang T, Miao K, Zhuang C, Cao X, Tang W, Liu G, Liu Y, Chen J, Liu Z, Yuan L, Liu Z, Huang X, Lu T, Fei B, Ning Z, Han B, Jiang Z (2013) The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nat Genet 45(4):456–461
Raman S, Greb T, Peaucelle A, Blein T, Laufs P, Theres K (2008) Interplay of miR164, CUP-SHAPED COTYLEDON genes and LATERAL SUPPRESSOR controls axillary meristem formation in Arabidopsis thaliana. Plant J 55(1):65–76
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110(4):513–520
Shen H, Yin YB, Chen F, Xu Y, Dixon RA (2009) A bioinformatic analysis of NAC genes for plant cell wall development in relation to lignocellulosic bioenergy production. Bioenergy Res 2:217–232
Song JB, Gao S, Sun D, Li H, Shu XX, Yang ZM (2013) miR394 and LCR are involved in Arabidopsis salt and drought stress responses in an abscisic acid-dependent manner. BMC Plant Biol 13:210
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The No Apical Meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85(2):159–170
Sunkar R, Zhou X, Zheng Y, Zhang W, Zhu JK (2008) Identification of novel and candidate miRNAs in rice by high throughput sequencing. BMC Plant Biol 8:25
Koyama T, Mistuda N, Seki M, Shinozaki K, Ohme-Takagi M (2010) TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis. Plant Cell 22(11):3574–3588
Unver T, Budak H (2009) Conserved microRNAs and their targets in model grass species Brachypodium distachyon. Planta 230(4):659–669
Wang N, Zheng Y, Xin H, Fang L, Li S (2013) Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep 32(1):61–75
Xie Q, Frugis G, Colgan D, Chua NH (2000) Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev 14(23):024–3036
Xu P, Mohorianu I, Yang L, Zhao H, Gao Z, Dalmay T (2014) Small RNA profile in moso bamboo root and leaf obtained by high definition adapters. PLoS ONE 9(7):e103590
Yang L, Lou Y, Peng Z, Zhao H, Sun H, Gao Z (2015) Molecular characterization and primary functional analysis of PeMPEC, a magnesium-protoporphyrin IX monomethyl ester cyclase gene of bamboo (Phyllostachys edulis). Plant Cell Rep 34(11):2001–2011
Yao Y, Guo G, Ni Z, Sunkar R, Du J, Zhu JK, Sun Q (2007) Cloning and characterization of microRNAs from wheat (Triticum aestivum L.). Genome Biol 8(6):R96
Zhao H, Chen D, Peng Z, Wang L, Gao Z (2013) Identification and characterization of microRNAs in the leaf of ma bamboo (Dendrocalamus latiflorus) by deep sequencing. PLoS One 8(10):e78755
Zhao H, Peng Z, Fei B, Li L, Hu T, Gao Z, Jiang Z (2014a) BambooGDB: a bamboo genome database with functional annotation and an analysis platform. Database (Oxford) 2014:bau006
Zhao H, Wang L, Dong L, Sun H, Gao Z (2014b) Discovery and comparative profiling of microRNAs in representative monopodial bamboo (Phyllostachys edulis) and sympodial bamboo (Dendrocalamus latiflorus). PLoS One 9(7):e102375
Zhu G, Chen G, Zhu J, Zhu Y, Lu X, Li X, Hu Y, Yan Y (2015) Molecular characterization and expression profiling of NAC transcription factors in Brachypodium distachyon L. PLoS One 10(10):e0139794
Acknowledgments
This work received financial support from the Special Fund for Forest Scientific Research in the Public Welfare from State Forestry Administration of China (No. 201504106) and the Sub-Project of National Science and Technology Support Plan of the Twelfth Five-Year in China (No. 2015BAD04B01 and No. 2015BAD04B03).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by B. Li.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, L., Zhao, H., Chen, D. et al. Characterization and primary functional analysis of a bamboo NAC gene targeted by miR164b. Plant Cell Rep 35, 1371–1383 (2016). https://doi.org/10.1007/s00299-016-1970-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-016-1970-6