Abstract
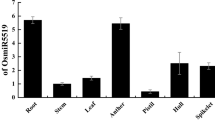
miR396 is a conservative microRNA family, which has been shown to be important for plant growth and development. To investigate the functions of miR396 in rice (Oryza sativa L.), we carried out a study on OsmiR396a. We found that OsmiR396a was expressed in various tissues with highest transcriptional level in leaves. Bioinformatic prediction suggested that OsmiR396a targets nine Growth Regulating Factor(GRF) genes, of which four (OsGRF1/2/6/8) were significantly down regulated by overexpression of OsmiR396a (aOE). aOE severely suppressed cell proliferation, leading to dwarf plants with smaller leaves. aOE also resulted in abnormal panicles and spikelets, especially a large proportion of rare conjoined-twin florets. Consistent with these defects, some related genes, including H4-2 and OsRAN2 (involved in cell division), FZP and LAX1 (controlling inflorescence architecture) and various floral organ identity genes (involved in floral development), exhibited significantly altered expressional levels in the aOE plants. Our findings further demonstrated the functional conservation of miR396 across dicot and monocot plants and suggested that OsmiR396a functions mainly by targeting OsGRF genes and indirectly regulating many downstream genes in rice.







Similar content being viewed by others
References
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Bao M, Bian H, Zha Y, Li F, Sun Y, Bai B, Chen Z, Wang J, Zhu M, Han N (2014) miR396a-Mediated basic helix-loop-helix transcription factor bHLH74 repression acts as a regulator for root growth in Arabidopsis seedlings. Plant Cell Physiol 55:1343–1353
Bartel B, Bartel DP (2003) MicroRNAs: at the root of plant development? Plant Physiol 132:709–717
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301:336–338
Carthew RW, Sontheimer EJ (2009) Origins and Mechanisms of miRNAs and siRNAs. Cell 136:642–655
Casadevall R, Rodriguez RE, Debernardi JM, Palatnik JF, Casati P (2013) Repression of growth regulating factors by the microRNA396 inhibits cell proliferation by UV-B radiation in Arabidopsis leaves. Plant Cell 25:3570–3583
Chen XM (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Chen X (2005) MicroRNA biogenesis and function in plants. FEBS Lett 579:5923–5931
Chen N, Xu Y, Wang X, Du C, Du J, Yuan M, Xu Z, Chong K (2010) OsRAN2, essential for mitosis, enhances cold tolerance in rice by promoting export of intranuclear tubulin and maintaining cell division under cold stress. Plant Cell Environ 34:52–64
Debernardi JM, Rodriguez RE, Mecchia MA, Palatnik JF (2012) Functional specialization of the plant miR396 regulatory network through distinct microRNA-target interactions. PLoS Genet 8:e1002419
Dreni L, Jacchia S, Fornara F, Fornari M, Ouwerkerk PBF, An G, Colombo L, Kater MM (2007) The D-lineage MADS-box gene OsMADS13 controls ovule identity in rice. Plant J 52:690–699
Duan CG, Wang CH, Guo HS (2006) Regulation of microRNA on plant development and viral infection. Chin Sci Bull 51:269–278
Duan Y, Xing Z, Diao Z, Xu W, Li S, Du X, Wu G, Wang C, Lan T, Meng Z, Liu H, Wang F, Wu W, Xue Y (2012) Characterization of Osmads6-5, a null allele, reveals that OsMADS6 is a critical regulator for early flower development in rice (Oryza sativa L.). Plant Mol Biol 80(4–5):429–442
Fang Y, Xie K, Xiong L (2014) Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J Exp Bot 65:2119–2135
Gao P, Bai X, Yang L, Lv D, Li Y, Cai H, Ji W, Guo D, Zhu Y (2010) Over-expression of osa-MIR396c decreases salt and alkali stress tolerance. Planta 231:991–1001
Gao F, Wang K, Liu Y, Chen Y, Chen P, Shi Z, Luo J, Jiang D, Fan F, Zhu Y, Li S (2015) Blocking miR396 increases rice yield by shaping inflorescence architecture. Nature Plants 2:15196
Hewezi T, Baum TJ (2012) Complex feedback regulations govern the expression of miRNA396 and its GRF target genes. Plant Signal Behav 7:749–751
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Horiguchi G, Kim GT, Tsukaya H (2005) The transcriptionfactor AtGRF5 and the transcription coactivator AN3cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J 43:68–78
Ikeda K, Sunohara H, Nagato Y (2004) Developmental course of inflorescence and spikelet in rice. Breed Sci 54:147–156
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53
Jung JH, Lee S, Yun J, Lee M, Park CM (2014) The miR172 target TOE3 represses AGAMOUS expression during Arabidopsis floral patterning. Plant Sci 215–216:29–38
Kamiuchi Y, Yamamoto K, Furutani M, Tasaka M, Aida M (2014) The CUC1 and CUC2 genes promote carpel margin meristem formation during Arabidopsis gynoeciumdevelopment. Front Plant Sci 5:165
Kater MM, Dreni L, Colombo L (2006) Functional conservation of MADS-box factorscontrolling floral organ identity in rice and Arabidopsis. J Exp Bot 57(13):3433–3444
Kim JH, Kende H (2004) A transcriptional coactivator, AtGIF1, is involved in regulating leaf growth and morphology in Arabidopsis. Proc Natl AcadSci USA 101:13374–13379
Kim JH, Choi DS, Kende H (2003) The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis. Plant J 36:94–104
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim VN, Chua NH, Park CM (2005) microRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42:84–94
Kim BH, Kwon Y, Lee BH, Nam KH (2014) Overexpression of miR172 suppresses the brassinosteroid signaling defects of bak1 in Arabidopsis. Biochem Biophys Res Commun 447:479–484
Komatsu M, Maekawa M, Shimamoto K, Kyozuka J (2001) The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev Biol 231:364–373
Komatsu M, Chujo A, Nagato Y, Shimamoto K, Kyozuka J (2003) FRIZZY PANICLE is required to prevent the formation of axillary meristems and to establish floral meristem identity in rice spikelets. Development 130:3841–3850
Liang G, He H, Li Y, Wang F, Yu D (2014) Molecular mechanism of microRNA396 mediating pistil development in Arabidopsis. Plant Physiol 164:249–258
Liu D, Song Y, Chen Z, Yu D (2009) Ectopic expression of miR396 suppresses GRF target gene expression and alters leaf growth in Arabidopsis. Physiol Plant 136:223–236
Liu X, Huang J, Wang Y, Khanna K, Xie Z, Owen HA, Zhao D (2010) The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in MicroRNA160a, in organogenesis and the mechanism regulating its expression. Plant J 62:416–428
Liu H, Guo S, Xu Y, Li C, Zhang Z, Zhang D, Xu S, Zhang C, Chong K (2014) Osa-miR396d-regulated OsGRFs function in floral organogenesis in rice through binding to their targets OsJMJ706 and OsCR4. Plant Physiol 165:160–174
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14:1035–1046
Mecchia MA, Debernardi JM, Rodriguez RE, Schommer C, Palatnik JF (2013) MicroRNA miR396 and RDR6 synergistically regulate leaf development. Mech Dev 130:2–13
Palatnik JF, Allen E, Wu XL, Schommer C, Schwab R, Carrington JC, Detlef W (2003) Control of leaf morphogenesis by MicroRNAs. Nature 425:257–263
Pelucchi N, Fornara F, Favalli C, Msiero S, Lago C, Pe MF, Colombo L, Kater MM (2002) Comparative analysis of rice MADS-box genes expressed during flower development. Sex Plant Reprod 15:113–122
Qin Y, Duan Z, Xia X, Yin W (2011) Expression profiles of precursor and mature microRNAs under dehydration and high salinity shock in Populus euphratica. Plant Cell Rep 30:1893–1907
Ramanjulu S, Guru J (2008) In silico identification of conserved microRNAs in large number of diverse plant species. BMC Plant Biol 8:37
Ramanjulu S, Thomas G, Pradeep KJ, Jian-Kang Z (2005) Cloning and characterization of microRNAs from rice. Plant Cell 17:1397–1411
Schommer C, Debernardi JM, Bresso EG, Rodriguez RE, Palatnik JF (2014) Repression of cell proliferation by miR319-regulated TCP4. Mol Plant 7:1533–1544
Tobina H, Uozu S, Matsuoka M, Kitano H, Hattori K (2003) Gene expression pattern of cell division/elongation factors in rice dwarf mutants. In: Khush GS, Brar DS, Hardy B (eds) In advances in rice genetics. International Rice Research Institute, Manila, pp 470–472
Tsuzuki M, Takeda A, Watanabe Y (2014) Recovery of dicer-like 1-late flowering phenotype by miR172 expressed by the noncanonical DCL4-dependent biogenesis pathway. RNA 20:1320–1327
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC (2005) Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132:3657–3668
Yamaguchi T, Nagasawa N, Kawasaki S, Matsuoka M, Nagato Y, Hirano HY (2004) The YABBY gene DROOPING LEAF regulates carpel specification and midrib development in Oryza sativa. Plant Cell 16:500–509
Yang C, Li D, Mao D, Liu X, Ji C, Li X, Zhao X, Cheng Z, Chen C, Zhu L (2013) Overexpression of microRNA319 impacts leaf morphogenesis and leads to enhanced cold tolerance in rice (Oryza sativa L.). Plant Cell Environ 36:2207–2218
Zhang B, Wang Q, Pan X (2007) MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol 210:279–289
Zhou S, Wang Y, Li W, Zhao Z, Ren Y, Wang Y, Gu S, Lin Q, Wang D, Jiang L, Su N, Zhang X, Liu L, Cheng Z, Lei C, Wang J, Guo X, Wu F, Ikehashi H, Wang H, Wan J (2011) Pollen semi-sterility1 encodes a kinesin-1-like protein important for male meiosis, anther dehiscence, and fertility in rice. Plant Cell 23(1):111–129
Acknowledgements
We thank Dr. Tao Lan, Dr. Mo Wang, Dr. Xiaofang Xie, Junjie Lin, Chunfen Lai, Binqing Lin, Qin Lin, Huan Chen and Owie Swithin Omosuwa for their help in the experiments. This work was supported by National Natural Science Foundation of China (Project No.: 31301003).
Author information
Authors and Affiliations
Contributions
ZD: Conceived and designed the experiments, performed the experiments, analyzed the data, contributed regents/materials/analysis tools and wrote the paper. WW: Conceived and designed the experiments, analyzed the data, contributed regents/materials/analysis tools and wrote the paper. MY: Performed the experiments, analyzed the data and wrote the paper. SB: Analyzed the data. YD: Contributed regents/materials/analysis tools. LZ: Performed the experiments.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Diao, Z., Yu, M., Bu, S. et al. Functional characterization of OsmiR396a in rice (Oryza sativa L.). Plant Growth Regul 85, 351–361 (2018). https://doi.org/10.1007/s10725-018-0406-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-018-0406-4