Abstract
Key message
We have constructed and annotated a web-based database of over 280 Arabidopsis genes that have characterized mutants associated with Arabidopsis acyl lipid metabolism.
Abstract
Mutants have played a fundamental role in gene discovery and in understanding the function of genes involved in plant acyl lipid metabolism. The first mutant in Arabidopsis lipid metabolism (fad4) was described in 1985. Since that time, characterization of mutants in more than 280 genes associated with acyl lipid metabolism has been reported. This review provides a brief background and history on identification of mutants in acyl lipid metabolism, an analysis of the distribution of mutants in different areas of acyl lipid metabolism and presents an annotated database (ARALIPmutantDB) of these mutants. The database provides information on the phenotypes of mutants, pathways and enzymes/proteins associated with the mutants, and allows rapid access via hyperlinks to summaries of information about each mutant and to literature that provides information on the lipid composition of the mutants. In addition, the database of mutants is integrated within the ARALIP plant acyl lipid metabolism website (http://aralip.plantbiology.msu.edu) so that information on mutants is displayed on and can be accessed from metabolic pathway maps. Mutants for at least 30 % of the genes in the database have multiple names, which have been compiled here to reduce ambiguities in searches for information. The database should also provide a tool for exploring the relationships between mutants in acyl lipid-related genes and their lipid phenotypes and point to opportunities for further research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
New tools in science such as genome sequencing, RNASeq, microarrays, and metabolomics have provided enormous new data resources. Researchers are increasingly challenged to keep up with the avalanche of data and often feel overwhelmed. Fortunately, there are powerful search tools available to ‘mine’ many datasets, and there are excellent efforts to organize these data and to provide websites where specific data can be queried with sophisticated searches (Geneinvestigator, to name just one). However, the accumulation of new data almost always out-paces the ability to curate it, to integrate it with other databases, and to provide manual annotations by experts in the field.
This review describes the results of an effort to organize, annotate, update, and curate key information about characterized mutants that impact Arabidopsis acyl lipid metabolism. The database/catalog (ARALIPmutantDB) has evolved out of the ARALIP website (http://aralip.plantbiology.msu.edu) and its underlying data and annotations. In addition, information from the excellent database published by Lloyd and Meinke (2012) has been incorporated. Approximately 20–30 studies on mutants of Arabidopsis acyl lipid metabolism have been published every year in recent years (Fig. 1). We have updated ARALIPmutantDB to include literature up to the summer of 2014. The database is available in Supplement Table 1, and updated versions can also be downloaded at http://aralip.plantbiology.msu.edu/downloads.
Arabidopsis lipid mutants reported in publications by year. Publications in which acyl lipid-related mutants were characterized in terms of function and phenotype were sorted by year. An increase in the number of new publications characterizing novel mutants may have peaked in 2012. The survey for 2014 included publications up to summer of that year
The goals of this project have been to produce a catalog of mutants that is annotated by experts, that is linked to metabolic pathways, and that provides a resource to aid researchers in the field of plant lipid metabolism. This review provides a brief background on the identification of mutants in acyl lipid metabolism, and presents an overview of the database of these genes, and highlights some aspects of the distribution of mutants in different pathways and protein classes. The database provides information on the phenotypes of mutants, pathways, and enzymes/proteins associated with the mutants and allows rapid access via hyperlinks to summaries of information about each mutant and to references that provide information on the phenotypes and, in most cases, the lipid composition of the mutants. In addition, the database of mutants is integrated within the ARALIP plant acyl lipid metabolism website so that information about mutants is displayed on and can be accessed from over 15 metabolic pathway maps. At least 30 % of the genes in the database are associated with multiple mutants or mutants that have multiple names (Fig. 2). These have been compiled in the database to reduce confusion in searches for information. Overall, this database can provide a tool for exploring the relationships between mutants in genes and their lipid phenotypes. It is also hoped that knowledge of the lack of mutants for reactions, proteins, or pathways will help uncover opportunities for new insights.
Prevalence of alternate names associated with Arabidopsis mutants of acyl lipid metabolism. 30 % of all mutants associated with a single gene in this survey have more than one name in the literature, with the highest number of names being seven. An alternative name was defined as a different name for mutants which have the same gene locus identification. Alternate names can arise for several reasons: from different alleles or because of changes in names (e.g., act1 = ats1) or because mutants in the same gene were given different names when discovered/characterized in different laboratories. Therefore, the same mutant could be described with several alternate names in different publications
Forward and reverse genetic approaches for identification of mutants
The identification of mutants in Arabidopsis acyl lipid metabolism began with forward genetic screens pioneered by Chris Somerville. In 1985, Browse and Somerville reported in Science on the identification of a mutant (fad4, At4g27030) lacking trans-3 hexadecenoic acid (Browse et al. 1985). The mutant was identified by generating a population of EMS mutants and screening the fatty acid composition of leaves of approximately two thousand of these mutants by gas chromatography. It was initially very surprising to lipid biochemists that the fad4 mutant had no visible phenotypes even though it lacked a fatty acid that had been conserved throughout more than 200 million years of plant evolution. After this landmark publication, using the same and similar approaches, the Somerville lab identified mutants that represented all major desaturases responsible for chloroplast and cytoplasmic glycerolipid unsaturation as well as acyl transferases and other enzymes (for review, see Wallis and Browse 2002).
Some mutants (e.g. fad4, fad5) impacted only the ‘prokaryotic’ lipids in the plastid (e.g., MGDG), whereas others changed the composition of extraplastidial lipids together with those lipids that were imported into the plastid from the ER (e.g., fad2, fad3). Other key mutants were also identified in these early studies. The act1/ats1 mutant in the plastid acyl-ACP glycerol-3-phosphate acyltransferase was particularly illuminating because it resulted in a shift of flux from the prokaryotic to the eukaryotic pathway (Kunst et al. 1988). Thus, a major contribution of these mutants was that they provided crucial support and confirmation of the ‘two-pathway hypothesis’ of plant glycerolipid metabolism.
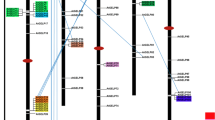
Progress as of 1991 with the forward genetics approach to Arabidopsis lipid metabolism was summarized in perhaps the most influential figure published in the field of plant acyl lipid metabolism (Fig. 3) (Browse and Somerville 1991). None of the genes that were responsible for the early mutant lipid phenotypes represented in Fig. 3 were known at the time of their publication. This changed with the historic first success in plant map-based cloning by Arondel et al. (1992), which resulted in the identification of the FAD3 gene. Map-based cloning continues to be a major method for the identification of genes responsible for glycerolipid phenotypes. For example, it is interesting to note that the gene encoding FAD4 was not identified until more than 20 years after the mutant was first identified, and this success was achieved by map-based cloning in the lab of former Somerville student, Christoph Benning (Gao et al. 2009). In addition to the analysis of leaves, forward genetic screening was applied to Arabidopsis seeds, leading to major breakthroughs in the identification of genes crucial to the biosynthesis of seed oil. The wrinkled1 mutant, with an 80 % reduction in seed oil, was identified by forward genetics and described by Focks and Benning (1998). The identification by map-based cloning of the WRI1 gene as an AP2-type transcription factor was a landmark in understanding the control of seed oil biosynthesis (Cernac and Benning 2004). Rod1 (reduced oleate desaturation1) was identified in a GC screen of seed fatty acids of EMS mutants (Lemieux et al. 1990). In 2009, map-based cloning of rod1 led to the discovery of a new enzyme reaction, phosphatidylcholine: diacylglycerolcholine transferase (PDCT) (Lu et al. 2009).
Mutants in Arabidopsis acyl lipid metabolism identified by forward genetics as of 1991. This figure is perhaps the most influential figure published in the field of plant acyl lipid metabolism. In addition to illustrating the mutants associated with acyl lipid metabolism, the width of arrows indicates flux through the prokaryotic and eukaryotic pathways of Arabidopsis leaves. Figure reproduced from Wallis and Browse (2002), and is based on an earlier version from Browse and Somerville (1991)
Reverse genetics (identification of genes based on knowledge of DNA sequences) became possible at a large scale with the advent of high-throughput DNA sequencing and now is the most frequent method for establishing gene-mutant-phenotype relationships (Fig. 4). For Arabidopsis, a key advance was the sequencing of several thousand randomly selected cDNA clones [‘expressed sequence tags’ (ESTs)] reported in 1993 and 1994 (Höfte et al. 1993; Newman et al. 1994). This advance opened up the ability to ‘knock-down’ the expression of genes using RNA interference techniques and, equally important, to more readily identify candidate genes corresponding to mapped mutations. The number of ESTs expanded rapidly to over 100,000 by 2000, and these ESTs were estimated to represent approximately two-third of expressed genes of Arabidopsis. The publication of the Arabidopsis genome in 2000, in addition to greatly facilitating identification of genes by map-based cloning, encouraged the development of new reverse genetic tools, including the hugely valuable collection of T-DNA insertion mutants (Alonso et al. 2003), which later included sequence information at the site of insertion (O’Malley et al. 2007). Large-scale forward genetic screens of the T-DNA population, including many projects referred to as functional genomics, led to the discovery of plant acyl lipid metabolism genes (e.g., Ajjawi et al. 2010, 2011). Over the past 15 years, reverse genetics has become increasingly facile and has accounted for over 80 % of gene-mutant characterizations since 2005 (Fig. 4).
Changes in methods leading to identification for Arabidopsis mutants. Key publications which characterize acyl lipid-related mutants were sorted based on whether or not they were identified using forward or reverse genetic techniques. Designations of methods from publications before 2012 are based on Lloyd and Meinke (2012). More recent data is based on analysis of the literature from 2012 until July 2014. Note that many mutants were identified by a combination of forward and reverse genetics, which is not indicated here
The identification of the gene encoding diacylglycerol acyltransferase (DGAT1) was a major contribution to the understanding of seed oil metabolism, and illustrates an example of forward genetics being aided by DNA sequence information from other organisms and from cDNA clones. The tag1mutant (as11) was obtained by EMS mutagenesis and GC screening of seed fatty acid composition. Further characterization of as11 indicated a reduction in diacylglycerol acyltransferase activity (Katavic et al. 1995). In 1999, two labs reported the sequence of the gene responsible for loss of DGAT activity in tag1-1 (as11) (Zou et al. 1999) and in tag1-2 (abx45), an independently identified low oil mutant from a T-DNA insertion population (Routaboul et al. 1999).
Although gene-mutant connections are now most frequently made via reverse genetics (Fig. 4), forward genetic approaches have continued to provide crucial new breakthroughs, particularly in the discovery of genes where candidate sequences could not be predicted based on the previous work. Thus, forward genetics has led to many of the most novel discoveries. Examples include discovery of four genes required for transport of lipids from the ER to the plastid (Benning 2009), and the discovery that cer7 is a subunit of a RNA processing and degrading exosome (Hooker et al. 2007).
This survey undoubtedly has missed many mutants in acyl lipid metabolism that have been studied. In some cases, authors of ARALIP or of this review simply were not aware of the published work. In many more cases, mutants have displayed little or no phenotype and this information is either not published, or information on the mutants is presented in ways that it is not easily found. For example, Zhang et al. (2009a) describe a number of acyltransferase mutants that have no seed oil phenotype, but these negative results only appear in one sentence. The database is therefore biased (as is the literature) toward mutants with clear biochemical phenotypes. There are also datasets from ‘functional genomic’ screens in which large populations of mutants have been analyzed for phenotypes, including lipids. These include, for example, a survey of leaf fatty acid composition of more than 5000 T-DNA insertion mutants (Ajjawi et al. 2010). Because most results from such surveys need to be further validated, we have not included these results unless they are described in full publications. To improve the database, we request our colleagues in the plant lipid community to submit corrections and additions to the authors or via the comment form at: http://aralip.plantbiology.msu.edu/forms.
Results and discussion
Development of the database
To create a central database that includes almost all acyl lipid-associated mutants and related information, we consolidated several sources. We started with data from the Arabidopsis Acyl-Lipid Metabolism website (ARALIP: http://aralip.plantbiology.msu.edu), an annotated database of lipid-related genes known or suspected to participate in Arabidopsis acyl lipid metabolism. Specific annotations in this database were provided by 27 authors of the Acyl Lipid Metabolism chapter of The Arabidopsis Book (Li-Beisson et al. 2013). 152 gene loci that were associated with a characterized mutant were identified from the ARALIP database.
We further identified mutants by comparing the ARALIP database to a dataset of Arabidopsis genes with a loss-of-function mutant phenotype that is provided in a key publication by Lloyd and Meinke (2012). By comparing the two lists, we identified 70 additional mutants that were curated by Lloyd and Meinke but that were not included in ARALIP. The Lloyd and Meinke list included details on the phenotypic classifications and descriptions of mutants that were not already in ARALIP, and some of this information has been incorporated into ARALIPmutantDB. Comparing these sets also indicated that the original ARALIP database includes approximately 60 mutants not in the Lloyd and Meinke set.
The ARALIP website and the Lloyd and Meinke database were based on the data up to 2012 and 2011, respectively, with publication dates of 2013 and 2012. To provide more recent information, we searched the literature to identify additional mutants described between January 2012 and summer of 2014. The major search tool was the ISI Web of Science, using the search words “Arabidopsis,” “lipid,” and “mutant.” Full text searches with Google Scholar retrieved too many unrelated references and Textpresso had not been updated to include references from 2014 at the time of the search; these were therefore less effective than ISI. We also consulted colleagues in the field for relevant publications from 2012 onward. In this way, we added an additional 68 mutants identified from publication in 2012–2014. Over 280 genes with characterized mutants are now included in ARALIPmutantDB compared to the original list of 152 in ARALIP. Thus, mutants have been characterized for approximately 30 % of the more than 900 genes in the 2013 version of the ARALIP database. This contrasts with the fact that, as of November 2014, approximately 90 % of all Arabidopsis loci are associated with T-DNA insertions (The Arabidopsis Information Resource). We searched a TAIR listing of loci without T-DNA insertions and found that only 20, or approximately 2 % of ARALIP genes, do not have an associated T-DNA insert (Table S2). Thus, there is a ‘reservoir’ of T-DNA insertion mutants that may provide information on the functions of many more ARALIP genes. Of the 20 ARALIP genes without known T-DNA insertions, 8 have associated mutants that were identified without the benefit of T-DNA insertions. Intriguingly, 6 of the remaining 12 loci are annotated as lipid transfer protein (LTP) and 4 as oleosins. These might represent attractive targets for further investigation.
We identified key references for each mutant in this database by using both ARALIP and Lloyd and Meinke’s (2012) list of references as a starting point. We then searched the literature for references that characterized the mutant phenotype. The earliest reference listed for each mutant is useful because it often provides the best information on lipid composition; information that is often difficult to find with available search tools, particularly when much of the data are presented in figures or tables in older publications. The earlier references also provide the best tool for tracking new publications based on citations. Other related references are also linked to the gene locus in the ARALIP databases.
It is very common for mutants or different mutant alleles of the same gene to have multiple names, and this occurs for 30 % of genes in ARALIPmutantDB, with the highest number of names being seven (Fig. 2). An alternative name was defined as a different name for mutants which have the same gene locus identification. Alternate names can arise for several reasons: from different alleles or because of changes in names (e.g., act1 was updated to ats1) or because mutants in the same gene were given different names when discovered/characterized in different laboratories. The use of different names can cause confusion in discussions, in the literature, and can lead to ambiguities in searches for information on the mutants which results in researchers missing key information. This is particularly true for new researchers entering the field. Thus, by compiling the names we hope to improve understanding of the large amount of literature on plant acyl lipid mutants.
The ARALIP website and database are organized with genes that are classified as belonging to different metabolic pathways. Some genes fit into more than one category. Figure 5 displays the proportion of genes in a few selected pathways that have characterized mutants. Eukaryotic phospholipid and TAG synthesis have the highest number of mutants at 50, which account for 45 % of the total records of genes assigned to these pathways. Other categories are striking in the low proportion of mutants. For example, there are 74 genes annotated as lipid transfer proteins (LTP) but mutants have been characterized for only 10 of these. In addition, over 300 genes in Arabidopsis include the term “lipase” in their annotation. However, we could find literature describing mutants for only 50 of these genes (discussed below).In the following sections we present a brief analysis of two areas of plant lipid research where the proportion of mutants identified compared to the number of genes is particularly low, and thus may offer opportunities for new discoveries. Our hope is that by highlighting these areas, other workers in the field will be attracted to the opportunities they present for new discoveries. In these sections, the genes which are associated with mutants are indicated by italicized locus IDs.
Mutants characterized by pathway. 293 lipid-related mutants in ARALIPmutantDB were sorted based on their involvement in different ARALIP pathways or protein type. The LTP family is not included in wax biosynthesis but instead is shown separately. Eukaryotic phospholipid and TAG synthesis has the highest number of mutants at 54, which account for 47 % of the total records of genes in that pathway. In contrast, a very small proportion of genes annotated as LTP or lipase have characterized mutants. Note that some genes are associated with multiple pathways
Arabidopsis putative lipases: why are there so many?
A large number of Arabidopsis genes that are annotated with the term “lipase” were originally not included in the ARALIP database because their function and catalyzed reaction is usually poorly characterized, if at all. However, to contribute to the general need for greater understanding of these genes, we expanded ARALIPmutantDB to include genes that include “lipase” in TAIR annotations [selected based on the lists presented in Troncoso-Ponce et al. (2013)]. Supplement Table 3 provides a summary of the distribution of characterized mutants within the different classes of lipases.
A major enigma in the study of plant lipid metabolism is why there may be so many genes involved in lipid breakdown or turnover. It is striking that there are more putative lipase genes than genes for the central anabolic pathways for fatty acid synthesis and glycerolipid assembly. However, experimental evidence confirming the in planta roles of these genes as lipases is available for only approximately 30–40 genes (Li-Beisson et al. 2013). The further exploration of these putative lipases may be a path toward new insights. Indeed major recent discoveries have included the role of a GDSL lipase (Yeats et al. 2012, 2014) and of an α/β hydrolase (Jakobson et al. 2014) in cutin synthesis. It is important to note that the proposed role of these genes is to catalyze acyltransferase reactions, rather than lipase reactions.
Lipases which have been annotated to date are grouped here into phospholipases (PL), phosphatidic acid phosphatase (PAP), triacylglycerol lipases (TAGL), monoacylglycerol lipases (MAGL), GDSL lipases, and proteins harboring an α/β hydrolase motif. A large number of these annotations are based only on sequence homology to enzymer from other organizms and have not been confirmed to be correct for the Arabidopsis genes.
Phospholipids are not only major structural components of membranes, but also a reservoir for lipid-signaling molecules. Phospholipases (PLs) are enzymes that catalyze the hydrolysis of phospholipids either at the acyl ester bond, to release fatty acids, or at the polar head group. There are four major classes, termed A, B, C and D, which are distinguished by the type of reaction which they catalyze. PLA is divided into PLA1 and PLA2 that cleave the sn-1 and sn-2 acyl chains, respectively. PLB hydrolyzes both sn-1 and sn-2 acyl chains and is also known as a lysophospholipase. PLC hydrolyzes the O–P bond adjacent to the glycerol, releasing diacylglycerol and a phosphate-containing head group. PLD hydrolyzes the O–P bond adjacent to the head group, releasing PA and a head group.
PLA is divided into two groups, which are secreted and patatin-related PLAs. The first putative PLA1 targeted into the plastid was identified from characterization of a defective in anther dehiscence 1(dad1, At2g44810) mutant, which is defective in the biosynthesis of jasmonic acid (JA) in flower buds (Hyun et al. 2008; Ishiguro et al. 2001). A homolog of DAD1, DONGLE (DGL, At1g05800), which is expressed in vegetative tissues was identified to be involved in the early phase of wound-inducible JA biosynthesis, whereas DAD1 functions in the late phase of wound-inducible JA production. DAD1 has a preferential substrate specificity for DGDG, suggesting that the major source of JA is the acyl chains derived from DGDG (Hyun et al. 2008). Shoot gravitropism 2 (SGR2, At1g31480) encodes a PLA1 involved in shoot gravitropism, and it has been suggested that the SGR2 might be important in the formation of membrane structure (Kato et al. 2002). The PLA2 subfamily is known to have four genes, PLA2α, β, γ, and δ. PLA2β (At1g53940) as like SGR2 is also involved in cell elongation and shoot gravitropism (Lee et al. 2003). The role of PLA2β, γ (At4g29460), and δ in the formation of membrane structure was discovered during pollen development and germination by the suppression of PLA2 (Kim et al. 2011). The suppressor of AvrBsT-elicited resistance1 (SOBER1, At4g22300) regulates PA levels generated in Arabidopsis in response to biotic stress (Kirik and Mudgett 2009). In addition, the patatin-related PLAs (pPLAs) are grouped into three subfamilies; pPLAI, pPLAII (α, β, γ, δ, and ɛ), and pPLAIII (α, β, γ, and δ). pPLAs preferentially hydrolyze membrane glycerolipids such as monogalactosyl monoacylglyceride (MGDG) and phosphatidyl-glycerol (PG). pPLAs and the released fatty acids have been implicated in plant growth and stress responses. pPLAI was implicated in maintaining the levels of basal JA that plays a positive role in defense responses (Yang et al. 2007). pPLAIIα-deficient plants were observed to have higher levels of JA, methyl-JA, and the oxylipin-biosynthetic intermediates than in wild type, indicating that pPLAIIα (At2g26560) might function in the removal of oxidatively modified fatty acids in membranes for membrane repair or remodeling (Yang et al. 2012). pPLAIIβ (AtPLAIVB/PLP5, At4g37060) is involved in root elongation under phosphate deficiency, and pPLAIIγ and δ were implicated in auxin responses (Rietz et al. 2010). Among four pplaIIIα (At5g43590), β (At2g39220), γ (At3g54950), and δ (At3g63200) knock-out mutants, only pplaIIIδ-KO seeds exhibited significant increase in seed oil contents with 20- and 22-carbon fatty acids, suggesting that pplaIIIδ, which is able to hydrolyze PC, may play a role in fatty acyl flux from plastid to the ER and/or PC fatty acyl remodeling for TAG synthesis (Li et al. 2013).
Possible functions of PLC have been identified in response to abiotic stresses. Non-specific phospholipase C5 (NPC5, At3g03540) is reported to be involved in galactolipid accumulation by providing DAG intermediates in leaves under phosphate limitation condition (Gaude et al. 2008). The npc4 mutant (At3g03530) shows decreased DAG levels, decreased ABA sensitivity in seed germination, root elongation and stomatal movement, and decreased resistance to drought and salt stresses, suggesting that NPC4 and DAGs promote stomatal opening under well-watered conditions and PA promotes plant growth under water-deficit conditions (Peters et al. 2010). Phosphoinositide (PI)-specific PLC9 (At2g40116) is involved in thermotolerance of Arabidopsis, suggesting that the released IP3 may regulate intracellular Ca2+ concentration, which could ultimately induce genes encoding heat shock proteins (Zheng et al. 2012).
Among 12 PLDs including PLDα (3), β (2), γ (3), δ, ɛ, and ξ (2), in planta roles of 8 PLDs have been characterized. PLD and its product PA are implicated in many cellular processes including plant growth and development, plant responses to abiotic and biotic stresses, hormone responses, and vesicle trafficking. PA produced by PLDα1 (At3g15730) binds to NADPH oxidase, which stimulates ROS production in guard cells, showing that PA is a signaling molecule for ABA-mediated stomatal closure (Zhang et al. 2009b). PLDα1 (At3g15730) and PLDδ (At4g35790) cooperatively function in ABA-induced stomatal closure and seed germination (Uraji et al. 2012; Distefano et al. 2012). PLDα3 (At5g25370) is involved in plant growth and development, particularly flowering, under water-deficit conditions (Hong et al. 2008). The pldβ 1 mutant (At2g42010) showed an increased resistance to the bacterial infection (Pseudomonas syringae pv. DC3000), but was more susceptible to fungal infection (Botrytis cinerea) compared with wild type, suggesting that PLDβ1 and associated lipid changes are involved in the SA-dependent and JA/ethylene-dependent plant defense (Zhao et al. 2013). PA produced by PLDδ (At4g35792) is involved in cell wall-based defense signaling in non-host resistance against powdery mildew fungi via hydrogen peroxide production (Pinosa et al. 2013). PLDɛ (At1g55180) was found to enhance plant growth under high salinity and water deficiency, suggesting that PLDɛ and PA may play a positive role in nutrients such as nitrogen signaling (Hong et al. 2009). PLDξ (At3g05630) and PA enhance vesicle trafficking and positively regulate auxin responses by regulating the activities of RCN1 (a protein phosphatase 2A regulatory subunit) and PID1 (a Ser/Thr protein kinase), which are a potential target protein of PA and a regulator of polar auxin transporter, respectively (Li and Xue 2007).
PA phosphatase (PAP) catalyzes the dephosphorylation of PA to diacylglycerol (DAG). Arabidopsis contains four genes encoding membrane-bound PAP (also called lipid phosphate phosphatase, LPP), two genes encoding soluble and cytosolic PAP (also called PA phosphohydrolase1 and 2), and three genes encoding plastidic PAP. Among 4 LPPs, disruption of LPP2 (At1g15080) caused hypersensitivity to ABA and significant PA accumulation during seed germination, suggesting that PA is involved in ABA signaling (Katagiri et al. 2005). The pah1pha2-1 mutant exhibited a significant increase in phospholipid content, up-regulation of genes encoding enzymes involved in phospholipid synthesis, and alteration in ER morphology, suggesting that PAH/2 may play a role in repressing phospholipid synthesis at the ER or may modulate phospholipid synthesis through changes in the level of PA or DAG (Eastmond et al. 2010). Based on sequence homology searches of LPP genes from Cyanobacterium synechosystis sp. PCC6803, 3 putative plastidic Arabidopsis LPP genes {LPPγ (At5g03080), LPPɛ1 (At3g50920), and LPPɛ2 (At5g66450)} were identified. The lppɛ1 lppɛ2 mutant showed no significant changes in lipid composition, whereas loss of LPPγ may cause a lethal effect on plant viability (Nakamura et al. 2007).
TAG hydrolysis is required for the mobilization of storage reserves during seed and pollen germination, and may be involved in membrane lipid remodeling during leaf senescence, and in plant defense against bacterial pathogen or insect attack. Arabidopsis PAD4 (At3g52430) and MLP1 (At5g14180), which display sequence similarity to TAG lipase, have been identified to be involved in plant defense (Jirage et al. 1999; Louis et al. 2010). Overexpression of SAG101 (At5g14930), a leaf senescence-associated gene causes premature leaf senescence, whereas its antisense lines showed delayed leaf senescence. The SAG101 fusion proteins showed an acyl hydrolase activity for triolein (He and Gan 2002). AtLip1 (At2g15230) with the best homology to human gastric lipase and SDP1 (Sugar-dependent 1, At5g04040) encoding a patatin-domain TAG lipase were reported to be involved in storage oil breakdown during seed germination (Karim et al. 2005; Eastmond 2006). In addition, monoacylglycerol lipase (MAGL) catalyzes the hydrolysis of MAG to fatty acid and glycerol, the last step of TAG breakdown. Among 16 Arabidopsis genes annotated as MAGL, only 1 gene, lysoPL2/AtMGAT (Arabidopsis monoacylglycerol acyltransferase, At1g52760) has been characterized by mutant analysis. lysoPL2 promotes the degradation of lysoPC in response to cadmium-induced oxidative stress (Gao et al. 2010) and AtMGAT was reported to have both monoacylglycerol acyltransferase and acyl hydrolase activities (Vijayaraj et al. 2012). Expression patterns, subcellular localizations, and enzyme activities of 16 Arabidopsis MAGLs have been recently characterized (Kim et al. 2012).
GDSL lipase containing a GDSL motif is a >100 member gene superfamily of possible lipolytic enzymes that collectively appear to display very broad substrate specificity. GDSL lipase 1 (GLIP1), GLIP2, Tomato Cutin Deficient1 (CD1), and Arabidopsis cutin synthase1 (AtCUS1) genes belong to this very large family. In the analysis of Arabidopsis secretome in response to salicylic acid, GLIP1 (At5g40990) was reported to be involved in plant immunity against fungal and necrotropic pathogens, suggesting that GLIP1 may play a role in ethylene-associated systemic immunity (Oh et al. 2005; Kim et al. 2014). GLIP2 (At1g53940) was also involved in resistance to necrotropic bacteria, Erwinia carotovora via negative regulation of auxin signaling (Lee et al. 2009). After a tomato GDSL lipase, Cutin Deficient 1 (CD1) was identified to be an acyltransferase in cutin synthesis (Yeats et al. 2012), putative Arabidopsis homologues (also called cutin synthase, CUS) of CD1/SlCUS1 (Solanum lycopersicum cutin synthase 1) were searched and AtCUS1/LTL1 demonstrated cutin synthase activity in vitro (Yeats et al. 2014).
Other lipases that contain a α/β-hydrolase motif include EDS1, BDG, CGI-58, and PES1 and 2. Enhanced disease susceptibility1 (EDS1, At3g48090) is involved in a disease-resistance process conditioned by TIR-NB-LRR type R gene that encodes a leucine-rich repeat protein (Falk et al. 1999). Bodyguard (BDG, At1g64670) is associated with cuticle development and morphogenesis (Jakobson et al. 2014; Kurdyukov et al. 2006). A mutant defective in the Arabidopsis CGI-58 homologue (also called ABHD5, or α/β-hydrolase-5, At4g24160) caused accumulation of lipid droplet in leaves, suggesting that CGI-58 may play a role in neutral lipid homeostasis in plants (James et al. 2010). Phytyl Ester Synthase1 and 2 (PES 1 and 2, At1g54570 and At3g26840) containing both α/β-hydrolase and acyltransferase motifs function in the formation of phytyl esters in chloroplasts (Lippold et al. 2012).
Several of the examples presented above make it clear that the original annotations of genes as “lipase”, PLA, etc. need to be revised/updated as new information on their actual in vivo enzymatic activity becomes available.
Mutants of transcription factors and other regulators of acyl lipid metabolism
Over the past several decades, considerable information has become available on regulation of lipid metabolism by transcription factors (TFs) and other regulators, and is well summarized in recent reviews (Lee and Suh 2013; Li-Beisson et al. 2013; Marchive et al. 2014; Borisjuk et al. 2014). However, considering there are over 1800 total TFs in Arabidopsis, it is perhaps surprising that such a small number have been shown to regulate acyl lipid metabolism. Some regulators have been identified in other plants but not in Arabidopsis, such as SHOOTMERISTEMLESS (STM), TT16, bZIP123 (Deng et al. 2012; Song et al. 2013). Possible reasons for the low number are that the functions of regulators may be redundant, such that the phenotypes in single mutants are masked by complementation by other genes, such as two negative regulators of oil, the High-level expression of Sugar-Inducible gene 2 (HSI2, At2g30470) and HSI2-Like1 (HSL1, At4g32010) (Tsukagoshi et al. 2007), or that mutations in regulation of critical primary metabolic pathways are lethal.
With more than 35 % oil in its seeds, Arabidopsis provides one of the best model plants for seed oil biosynthesis. Several transcription regulators involved in TAG accumulation have been identified. Forward genetic methods were used in early studies to find mutants with altered oil contents. Focks and Benning (1998) reported the isolation of a mutant named wrinkled1 (At3g54320), which has 80 % reduction in oil content and in flux of carbon through glycolysis and fatty acid synthesis. The identification of the WRI1 gene represents a landmark advance in studies of plant lipid metabolism (Cernac and Benning 2004) and its role is now by far the best understood of any TF in acyl lipid metabolism (Marchive et al. 2014). WRI1 binds to an upstream AW-box region of genes involved in fatty acid synthesis, such as pyruvate kinase (Pl-PKb1) and acetyl-CoA carboxylase (BCCP2) (Maeo et al. 2009). It is now known that WRI1 directly binds to a large number of target genes in the glycolytic and fatty acid biosynthesis pathways (Marchive et al. 2014). WRI3 (At1g16060) and WRI4 (At1g79700), two other WRI1-like proteins can complement the wri1 mutant, are expressed in flower and stem, and may regulate cutin synthesis indirectly through control of fatty acid synthesis (To et al. 2012).
LEC1 (At1g21970), LEC2 (At1g28300), FUSCA3 (FUS3,At3g26790) and ABI3 (At3g24650) are master regulators in embryo development and deficiencies of oil were found in their mutants (Finkelstein and Somerville 1990; Keith et al. 1994; Lotan et al. 1998; Stone et al. 2001). PKL (At2g25170), a chromatin-remodeling factor, represses LEC1 expression in vegetative tissues and oil accumulates in the primary roots of pkl mutant (Ogas et al. 1997, 1999). ABI4 (At2g40220), another member of APETALA2 TFs, plays a role in the degradation of storage oil and the expression of DGAT1 is down-regulated in its mutant (Penfield et al. 2006; Yang et al. 2011). GLABRA2 (GL2, At1g79840), a homeodomain (HD) TF, acts as a negative regulator of seed oil (Shen et al. 2006). Several other negative regulators of TAG accumulation have been recognized by reverse genetic strategies in recent studies. TT2 and TT8, regulators of proanthocyanidin and flavonoid synthesis in testa, also affect oil contents and fatty acid compositions in seeds. FUS3, CAC2 (At5g35360), KASII (At1g74960), FAD2 (At3g12120) and FAE1 (At4g34520) are up-regulated in tt2 (At5g35550) and tt8 (At4g09820) mutants (Chen et al. 2012, 2014). ASIL (At1g54060) belongs to the trihelix family of DNA-binding TFs and depresses the expressions of genes related to embryo-specific lipids as well as embryo development (Gao et al. 2011). Additionally, several TFs have been identified as regulators of fatty acid compositions of seed oil. The bZIP67 (At3g44460) targets to FAD3 (At2g29980) and 18:3 levels are reduced in seed oil of bzip67 mutant (Mendes et al. 2013), while total fatty acid compositions are changed differentially in the seeds of crc (At1g69180) and ap1 (At1g69120) mutants (Han et al. 2012). PII (GLB1, At4g01900), directly controlled by WRI1, finely adjusts fatty acid composition in seeds (Baud et al. 2010).
The plant cuticle is the protective layer coating the aerial surface of higher plants, which mainly consists of cutin and cuticular wax. Several transcription factors have been confirmed to control the synthesis of cuticular lipids. SHINE1/WAX INDUCER1 (SHN1/WIN1, At1g15360), an AP2-domain protein, was first identified to regulate cutin synthesis by coordinate induction or by direct interaction with genes known to be involved in cutin deposition, such as long chain acyl-CoA synthetase2 (LACS2, At1g49430) (Aharoni et al. 2004; Broun et al. 2004; Kannangara et al. 2007). SHN2 (At5g11190) and SHN3 (At5g25390), two other members of the SHINE clade of AP2 TFs, also regulate cutin- and suberin-associated genes, such as CYP86A cytochrome P450 s and fatty acyl-CoA reductases (Shi et al. 2011). Additional AP2 TFs affecting cuticle deposition include GLABRA1 and GLABRA3 (Xia et al. 2010). TTG1, a WD40 repeat protein also has been shown to affect cuticle deposition (Xia et al. 2010). The Curly Flag Leaf1 (CFL1, At2g33510), a WW domain TF, negatively regulates cuticle development by modulating the function of HDG1 (At3g61150) (Wu et al. 2011). As noted above, WRI3 and WRI4, two other WRI1-like proteins expressed in non-seed tissues, are likely indirect regulators of cutin synthesis through fatty acid synthesis (To et al. 2012). DEWAX (At5g61590), an AP2/ERF-type TF, negatively regulates the expressions of genes related to surface lipid synthesis. More wax accumulates on the surface of stems and leaves of dewax mutant (Go et al. 2014). After MYB30 (At3g28910) and MYB41 (At4g28110) were identified as regulators of surface lipids (Cominelli et al. 2008; Raffaele et al. 2008; Kosma et al. 2014), more members of MYB family have been confirmed in recent studies. MYB106 (At3g01140) and MYB16 (At5g15310), cooperate with SHN1 to regulate epicuticular wax crystals and cutin nanoridges (Oshima et al. 2013). MYB96 (At5g62470) activates a group of wax biosynthesis genes, such as very long chain fatty acid-condensing enzymes (Seo et al. 2011). MYB41 positively regulates the steps necessary for aliphatic suberin synthesis and deposition of cell wall-associated suberin-like lamellae (Kosma et al. 2014). Cer7 (At3g60500) is a core subunit of the RNA processing/degrading exosome and influences wax biosynthesis by controlling the expression of CER3/WAX2/YRE (Hooker et al. 2007). A screen for suppressors of the cer7 mutant revealed regulatory roles for RNA-DEPENDENT RNA POLYMERASE1 (RDR1) and SUPPRESSOR OF GENE SILENCING3 (SGS3) in CER7-mediatied RNA silencing of CER3 (Lam et al. 2012). Similarly, SERRATE (At2g27100) encodes a protein of RNA–processing multi-protein complexes thought to mediate “RNA signaling in the cuticle integrity pathway” (Voisin et al. 2009). HUB1 (At2g44950) and HUB2 (At1g55250) encode two orthologous RING E3 ligases and wax and cutin compositions are altered in hub mutants (Menard et al. 2014). Similarly, CER9 (At4g34100) “encodes an E3 ubiquitin ligase homologous to yeast Doa10 (previously shown to target endoplasmic reticulum proteins for proteasomal degradation)” thought to serve as a negative regulator of cuticle synthesis and stress responses (Lu et al. 2012). More general regulators of embryo epidermal differentiation have been implicated in regulating cuticle development including ZHOUPI (At1g49770), ALE1 (At1g62340), GASSHO1 (At4g20140), GASSHO2 (At5g44700) and ACR4 (At3g59420). (Tsuwamoto et al. 2008; Tanaka et al. 2002; Watanabe et al. 2004; Cao et al. 2005; Xing et al. 2013).
Although a few regulators have been described to impact membrane lipids, many of these may do so indirectly. Bax inhibitor-1 (At5g47120), an endoplasmic reticulum protein, regulates sphingolipid synthesis by interacting with four related enzymes (Nagano et al. 2014). GOLDEN2-LIKE1 (GLK1, At2g20570) and GLK2 (At5g44190), regulators of chlorophyll synthesis, also regulate the expressions of galactolipid-synthesis genes, particularly DGD1 under continuous light (Kobayashi et al. 2014). ARF7 (At5g20730) and ARF19 (At1g19220) are auxin-response TFs and the accumulation of DGDG and SQDG is suppressed in arf7arf19 mutant during phosphate starvation (Narise et al. 2010).
In addition to WRI1 and bZIP67 mentioned above, a small number of studies have identified direct targets of TFs by DNA binding assays. LEC2 directly binds with RY motifs in the 5′ flanking regions of some LEC2 induced genes (Braybrook et al. 2006). ChIP-tiling array results demonstrate that FUS3 directly targets to many embryogenesis related TFs and microRNAs (Wang and Perry 2013). ChIP assay suggests that TT8 binds to the promoter regions of LEC1, LEC2, FUS3 and Cytidinediphophate diacylglycerol synthase2 (CDS2) (Chen et al. 2014). MYB96 (At5g62470), confirmed by both in vivo and in vitro assays, binds to the promoters of 3-ketoacyl-CoA synthase 1 (KCS1, At1g01120), KCS2 (At1g04220), KCS6 (At1g68530), KCR1 (At1g67730), and lipid transfer protein3 (LTP3, At5g59320) and positively adjusts their expression (Seo et al. 2011; Guo et al. 2013). ABI4 binds the coupling element1 (CE1) element [CACC (G)] in the promoter of its target genes in maize and Arabidopsis (Niu et al. 2002; Yang et al. 2011). LACS2 was reported as a direct target of WIN1 by immunoprecipitation assay (Kannangara et al. 2007). Clearly, much more is needed to clarify the mechanism of the other regulators mentioned above.
Summary/conclusions
A few comments and observations derived from ARALIPmutantDB.
-
1.
There are some large gene families where a surprisingly small number of mutants have been identified. The lipid transfer protein (LTP) gene family is one intriguing example. There are 73 genes in this family, but fewer than 10 % of these genes have characterized mutants. The GDSL lipase family, with over 100 members has only 2 genes in ARALIPmutantDB. Of course, it is important to remember that many genes annotated as lipases may not in fact act on lipids or catalyze hydrolase reactions.
-
2.
Desaturases were some of the first enzymes where mutants were characterized and now all major desaturase genes in central pathways of plant glycerolipid desaturation likely have been identified and characterized with mutants. However, several genes in the ‘ADS’ or FAD5-like class have uncertain functions and may be attractive candidates for further focus.
-
3.
Although WRI1, WRI3 and WRI4 are clearly the master regulators of fatty acid synthesis in seed and non-seed tissues, there are a number of remaining questions. Triple wri1 wri3 wri4 mutants still accumulate oil at 20 % of WT levels, raising the question of whether there are other transcription factors in play for oil synthesis of developing seeds. These three transcription factors are expressed at low levels in tissues such as leaves; although young, rapidly expanding leaves synthesize fatty acids at rates approaching those of seeds. Thus, a related question is: what controls production of fatty acids/lipids in leaves, where WRI is not highly expressed?
-
4.
Although WRI1 clearly regulates the expression of a number of genes in glycolysis and fatty acid synthesis and this determines oil content of seeds, very little is known about control of expression of genes later in the TAG assembly pathway. DGAT1, PDAT1 and other acyltransferases have patterns of expression during seed development that are strikingly different than those of the fatty acid biosynthesis pathway (e.g., Troncoso-Ponce et al. 2011). What TF or other factors are responsible for controlling their expression?
-
5.
How close are we to having identified most genes that have a lipid phenotype in single mutants? The number of publications on new mutants in Arabidopsis acyl lipid metabolism may have peaked (Fig. 1). This might reflect the fact that most genes with readily detectable mutant phenotypes have been characterized or that gene redundancies ‘conceal’ the gene function in mutants.
Author contribution statement
JO and MCS conceived and designed the project. KM and VS collected and analyzed information and created the figures and database. WY collected information, added annotations, and edited the manuscript. BS designed and created the web version of the database. KM, MZ, RJK, MCS, and JO wrote the manuscript. All authors read and approved the manuscript.
References
Aharoni A, Dixit S, Jetter R, Thoenes E, van Arkel G, Pereira A (2004) The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties, and confers drought tolerance when overexpressed in Arabidopsis. Plant Cell 16:2463–2480
Ajjawi I, Lu Y, Savage LJ, Bell SM, Last RL (2010) Large-scale reverse genetics in Arabidopsis: case studies from the chloroplast 2010 project. Plant Physiol 152:529–540
Ajjawi I, Coku A, Froehlich JE, Yang Y, Osteryoung KW, Benning C, Last RL (2011) A J-Like protein influences fatty acid composition of chloroplast lipids in Arabidopsis. PLoS One 6:e25368
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen HM, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Arondel V, Lemieux B, Hwang I, Gibson S, Goodman HM, Somerville CR (1992) Map-based cloning of a gene controlling Omega-3-fatty-acid desaturation in Arabidopsis. Science 258:1353–1355
Baud S, Bourrellier ABF, Azzopardi M, Berger A, Dechorgnat J, Daniel-Vedele F, Lepiniec L, Miquel M, Rochat C, Hodges M, Ferrario-Mery S (2010) PII is induced by WRINKLED1 and fine-tunes fatty acid composition in seeds of Arabidopsis thaliana. Plant Journal 64:291–303
Benning C (2009) Mechanisms of lipid transport involved in organelle biogenesis in plant cells. Annual review of cell and developmental biology. Annual Reviews, Palo Alto, pp 71–91
Borisjuk N, Hrmova M, Lopato S (2014) Transcriptional regulation of cuticle biosynthesis. Biotechnol Adv 32:526–540
Braybrook SA, Stone SL, Park S, Bui AQ, Le BH, Fischer RL, Goldberg RB, Harada JJ (2006) Genes directly regulated by LEAFY COTYLEDON2 provide insight into the control of embryo maturation and somatic embryogenesis. Proc Natl Acad Sci USA 103:3468–3473
Broun P, Poindexter P, Osborne E, Jiang CZ, Riechmann JL (2004) WIN1, a transcriptional activator of epidermal wax accumulation in Arabidopsis. Proc Natl Acad Sci USA 101:4706–4711
Browse J, Somerville C (1991) Glycerolipid synthesis-biochemistry and regulation. Annu Rev Plant Physiol Plant Molec Biol 42:467–506
Browse J, McCourt P, Somerville CR (1985) A mutant of Arabidopsis lacking a chloroplast-specific lipid. Science 227:763–765
Cao XY, Li KJ, Suh SG, Guo T, Becraft PW (2005) Molecular analysis of the CRINKLY4 gene family in Arabidopsis thaliana. Planta 220:645–657
Cernac A, Benning C (2004) WRINKLED1 encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J 40:575–585
Chen MX, Wang Z, Zhu YN, Li ZL, Hussain N, Xuan LJ, Guo WL, Zhang GP, Jiang LX (2012) The effect of TRANSPARENT TESTA2 on seed fatty acid biosynthesis and tolerance to environmental stresses during young seedling establishment in Arabidopsis. Plant Physiol 160:1023–1036
Chen MX, Xuan LJ, Wang Z, Zhou LH, Li ZL, Du X, Ali E, Zhang GP, Jiang LX (2014) TRANSPARENT TESTA8 inhibits seed fatty acid accumulation by targeting several seed development regulators in Arabidopsis. Plant Physiol 165:905–916
Cominelli E, Sala T, Calvi D, Gusmaroli G, Tonelli C (2008) Over-expression of the Arabidopsis AtMYB41 gene alters cell expansion and leaf surface permeability. Plant J 53:53–64
Deng W, Chen GQ, Peng F, Truksa M, Snyder CL, Weselake RJ (2012) Transparent Testa16 plays multiple roles in plant development and is involved in lipid synthesis and embryo development in Canola. Plant Physiol 160:978–989
Distefano AM, Scuffi D, Garcia-Mata C, Lamattina L, Laxalt AM (2012) Phospholipase D delta is involved in nitric oxide-induced stomatal closure. Planta 236:1899–1907
Eastmond PJ (2006) SUGAR-DEPENDENT1 encodes a patatin domain triacylglycerol lipase that initiates storage oil breakdown in germinating Arabidopsis seeds. Plant Cell 18:665–675
Eastmond PJ, Quettier AL, Kroon JTM, Craddock C, Adams N, Slabas AR (2010) PHOSPHATIDIC ACID PHOSPHOHYDROLASE1 and 2 regulate phospholipid synthesis at the endoplasmic reticulum in Arabidopsis. Plant Cell 22:2796–2811
Falk A, Feys BJ, Frost LN, Jones JDG, Daniels MJ, Parker JE (1999) EDS1, an essential component of r gene-mediated disease resistance in arabidopsis has homology to eukaryotic lipases. Proc Natl Acad Sci USA 96:3292–3297
Finkelstein RR, Somerville CR (1990) 3 Classes of Abscisic-Acid (ABA)-insensitive mutations of arabidopsis define genes that control overlapping subsets of ABA responses. Plant Physiol 94:1172–1179
Focks N, Benning C (1998) wrinkled1: a novel, low-seed-oil mutant of Arabidopsis with a deficiency in the seed-specific regulation of carbohydrate metabolism. Plant Physiol 118:91–101
Gao JP, Ajjawi I, Manoli A, Sawin A, Xu CC, Froehlich JE, Last RL, Benning C (2009) FATTY ACID DESATURASE4 of Arabidopsis encodes a protein distinct from characterized fatty acid desaturases. Plant J 60:832–839
Gao W, Li HY, Xiao S, Chye ML (2010) Acyl-CoA-binding protein 2 binds lysophospholipase 2 and lysopc to promote tolerance to cadmium-induced oxidative stress in transgenic Arabidopsis. Plant J 62:989–1003
Gao M-J, Li X, Lui H, Gropp GM, Lydiate DD, Wei S, Hegedus DD (2011) ASIL1 is required for proper timing of seed filling in Arabidopsis. Plant Signal Behav 6:1886–1888
Gaude N, Nakamura Y, Scheible WR, Ohta H, Dormann P (2008) Phospholipase C5 (NPC5) is involved in galactolipid accumulation during phosphate limitation in leaves of Arabidopsis. Plant J 56:28–39
Go YS, Kim H, Kim HJ, Suh MC (2014) Arabidopsis cuticular wax biosynthesis is negatively regulated by the DEWAX gene encoding an AP2/ERF-Type transcription factor. Plant Cell 26:1666–1680
Guo L, Yang HB, Zhang XY, Yang SH (2013) Lipid transfer protein 3 as a target of MYB96 mediates freezing and drought stress in Arabidopsis. J Exp Bot 64:1755–1767
Han XX, Yin LL, Xue HW (2012) Co-expression analysis identifies CRC and AP1 the regulator of Arabidopsis fatty acid biosynthesis. J Integr Plant Biol 54:486–499
He YH, Gan SS (2002) A gene encoding an acyl hydrolase is involved in leaf senescence in Arabidopsis. Plant Cell 14:805–815
Höfte H, Desprez T, Amselem J, Chiapello H, Rouze P, Caboche M, Moisan A, Jourjon MF, Charpenteau JL, Berthomieu P, Guerrier D, Giraudat J, Quigley F, Thomas F, Yu DY, Mache R, Raynal M, Cooke R, Grellet F, Delseny M, Parmentier Y, Demarcillac G, Gigot C, Fleck J, Philipps G, Axelos M, Bardet C, Tremousaygue D, Lescure B (1993) An inventory of 1152 expressed sequence tags Obtained by partial sequencing of CDNAs from Arabidopsis thaliana. Plant J 4:1051–1061
Hong YY, Pan XQ, Welti R, Wang XM (2008) Phospholipase D alpha 3 is involved in the hyperosmotic response in Arabidopsis. Plant Cell 20:803–816
Hong YY, Devaiah SP, Bahn SC, Thamasandra BN, Li MY, Welti R, Wang XM (2009) Phospholipase D epsilon and phosphatidic acid enhance Arabidopsis nitrogen signaling and growth. Plant J 58:376–387
Hooker TS, Lam P, Zheng HQ, Kunst L (2007) A core subunit of the RNA-processing/degrading exosome specifically influences cuticular wax biosynthesis in Arabidopsis. Plant Cell 19:904–913
Hyun Y, Choi S, Hwang HJ, Yu J, Nam SJ, Ko J, Park JY, Seo YS, Kim EY, Ryu SB, Kim WT, Lee YH, Kang H, Lee I (2008) Cooperation and functional diversification of two closely related galactolipase genes for jasmonate biosynthesis. Dev Cell (2):183–192
Ishiguro S, Kawai-Oda A, Ueda J, Nishida I, Okada K (2001) The DEFECTIVE IN ANTHER DEHISCENCE1 Gene Encodes A novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 13:2191–2209
Jakobson L, Lindgren LO, Verdier G, Laanemets K, Moldau H, Brosché M, Beisson F, Kollist H (2014) The α/β-hydrolase fold protein BODYGUARD is involved in the biosynthesis pathway of the cutin polymer in Arabidopsis. 21th International Symposium on Plant Lipids ISPL, Ontario, Canada 27
James CN, Horn PJ, Case CR, Gidda SK, Zhang DY, Mullen RT, Dyer JM, Anderson RGW, Chapman KD (2010) Disruption of the Arabidopsis CGI-58 homologue produces Chanarin-Dorfman-like lipid droplet accumulation in plants. Proc Natl Acad Sci USA 107:17833–17838
Jirage D, Tootle TL, Reuber TL, Frost LN, Feys BJ, Parker JE, Ausubel FM, Glazebrook J (1999) Arabidopsis thaliana PAD4 encodes a lipase-like gene that is important for salicylic acid signaling. Proc Natl Acad Sci USA 96:13583–13588
Kannangara R, Branigan C, Liu Y, Penfield T, Rao V, Mouille G, Hofte H, Pauly M, Riechmann JL, Broun P (2007) The transcription factor WIN1/SHN1 regulates cutin biosynthesis in Arabidopsis thaliana. Plant Cell 19:1278–1294
Karim EK, Stephanie B, Emilia O, Anne-Marie G, Natalie F, Vincent A (2005) Identification and characterization of a triacylglycerol lipase in Arabidopsis homologous to mammalian acid lipases. FEBS Lett 579:6067–6073
Katagiri T, Ishiyama K, Kato T, Tabata S, Kobayashi M, Shinozaki K (2005) An important role of phosphatidic acid in ABA signaling during germination in Arabidopsis thaliana. Plant J 43:107–117
Katavic V, Reed DW, Taylor DC, Giblin EM, Barton DL, Zou JT, Mackenzie SL, Covello PS, Kunst L (1995) Alteration of seed fatty-acid composition by an ethyl methanesulfonate-induced mutation in Arabidopsis thaliana affecting diacylglycerol acyl transferase activity. Plant Physiol 108:399–409
Kato T, Morita MT, Fukaki H, Yamauchi Y, Uehara M, Niihama M, Tasaka M (2002) SGR2, a phospholipase-like protein, and ZIG/SGR4, a SNARE, are involved in the shoot gravitropism of Arabidopsis. Plant Cell 14:33–46
Keith K, Kraml M, Dengler NG, McCourt P (1994) fusca3—a heterochronic mutation affecting late embryo development in Arabidopsis. Plant Cell 6:589–600
Kim HJ, Ok SH, Bahn SC, Jang J, Oh SA, Park SK, Twell D, Ryu SB, Shin JS (2011) Endoplasmic reticulum- and golgi-localized phospholipase a (2) plays critical roles in Arabidopsis pollen development and germination. Plant Cell 23:94–110
Kim RJ, Kim HJ, Kim HJ, Shim D, Carlson JE, Suh MC (2012) Protein structure modeling, expression patterns, subcellular localization, and enzymatic analysis of monoacylglycerol lipase (MAGL) gene family in Arabidopsis thaliana. 20th International Symposium on Plant Lipids ISPL, Seville, Spain 142
Kim HG, Kwon SJ, Jang YJ, Chung JH, Nam MH, Park OK (2014) GDSL lipase 1 regulates ethylene signaling and ethylene-associated systemic immunity in Arabidopsis. FEBS Lett 588:1652–1658
Kirik A, Mudgett MB (2009) SOBER1 phospholipase activity suppresses phosphatidic acid accumulation and plant immunity in response to bacterial effector AvrBsT. Proc Natl Acad Sci USA 106:20532–20537
Kobayashi K, Fujii S, Sasaki D, Baba S, Ohta H, Masuda T, Wada H (2014) Transcriptional regulation of thylakoid galactolipid biosynthesis coordinated with chlorophyll biosynthesis during the development of chloroplasts in Arabidopsis. Front Plant Sci 5: Article 272
Kosma DK, Murmu J, Razeq FM, Santos P, Bourgault R, Molina I, Rowland O (2014) AtMYB41 activates ectopic suberin synthesis and assembly in multiple plant species and cell types. Plant J 80(2):216–229
Kunst L, Browse J, Somerville C (1988) Altered regulation of lipid biosynthesis in a mutant of Arabidopsis deficient in chloroplast glycerol-3-phosphate acyltransferase activity. Proc Natl Acad Sci USA 85:4143–4147
Kurdyukov S, Faust A, Nawrath C, Bar S, Voisin D, Efremova N, Franke R, Schreiber L, Saedler H, Metraux JP, Yephremov A (2006) The epidermis-specific extracellular BODYGUARD controls cuticle development and morphogenesis in Arabidopsis. Plant Cell 18:321–339
Lam P, Zhao LF, McFarlane HE, Aiga M, Lam V, Hooker TS, Kunst L (2012) RDR1 and SGS3, components of RNA-mediated gene silencing, are required for the regulation of cuticular wax biosynthesis in developing inflorescence stems of Arabidopsis. Plant Physiol 159:1385–1395
Lee SB, Suh MC (2013) Recent advances in cuticular wax biosynthesis and its regulation in Arabidopsis. Mol Plant 6:246–249
Lee HY, Bahn SC, Kang YM, Lee KH, Kim HJ, Noh EK, Palta JP, Shin JS, Ryu SB (2003) Secretory low molecular weight phospholipase A (2) plays important roles in cell elongation and shoot gravitropism in Arabidopsis. Plant Cell 15:1990–2002
Lee DS, Kim BK, Kwon SJ, Jin HC, Park OK (2009) Arabidopsis GDSL lipase 2 Plays a role in pathogen defense via negative regulation of auxin signaling. Biochem Biophys Res Commun 379:1038–1042
Lemieux B, Miquel M, Somerville C, Browse J (1990) Mutants of Arabidopsis with alterations in seed lipid fatty-acid composition. Theor Appl Genet 80:234–240
Li G, Xue HW (2007) Arabidopsis PLD zeta 2 regulates vesicle trafficking and is required for auxin response. Plant Cell 19:281–295
Li MY, Bahn SC, Fan CC, Li J, Phan T, Ortiz M, Roth MR, Welti R, Jaworski J, Wang XM (2013) Patatin-related phospholipase pPLAIII delta increases seed oil content with long-chain fatty acids in Arabidopsis. Plant Physiol 162:39–51
Li-Beisson Y, Shorrosh B, Beisson F, Andersson MX, Arondel V, Bates PD, Baud S, Bird D, Debono A, Durrett TP, Franke RB, Graham IA, Katayama K, Kelly AA, Larson T, Markham JE, Miquel M, Molina I, Nishida I, Rowland O, Samuels L, Schmid KM, Wada H, Welti R, Xu C, Zallot R, Ohlrogge J (2013) Acyl-Lipid Metabolism. Arabidopsis Book/Am Soc Plant Biol 11:e0161
Lippold F, vom Dorp K, Abraham M, Holzl G, Wewer V, Yilmaz JL, Lager I, Montandon C, Besagni C, Kessler F, Stymne S, Dormann P (2012) Fatty acid phytyl ester synthesis in chloroplasts of Arabidopsis. Plant Cell 24:2001–2014
Lloyd J, Meinke D (2012) A comprehensive dataset of genes with a loss-of-function mutant phenotype in Arabidopsis. Plant Physiol 158:1115–1129
Lotan T, Ohto M, Yee KM, West MAL, Lo R, Kwong RW, Yamagishi K, Fischer RL, Goldberg RB, Harada JJ (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93:1195–1205
Louis J, Lorenc-Kukula K, Singh V, Reese J, Jander G, Shah J (2010) Antibiosis against the green peach aphid requires the Arabidopsis thaliana MYZUS PERSICAE-INDUCED LIPASE1 gene. Plant J 64:800–811
Lu CF, Xin ZG, Ren ZH, Miquel M, Browse J (2009) An enzyme regulating triacylglycerol composition is encoded by the ROD1 gene of Arabidopsis. Proc Natl Acad Sci USA 106:18837–18842
Lu SY, Zhao HY, Des Marais DL, Parsons EP, Wen XX, Xu XJ, Bangarusamy DK, Wang GC, Rowland O, Juenger T, Bressan RA, Jenks MA (2012) Arabidopsis ECERIFERUM9 involvement in cuticle formation and maintenance of plant water status. Plant Physiol 159:930–944
Maeo K, Tokuda T, Ayame A, Mitsui N, Kawai T, Tsukagoshi H, Ishiguro S, Nakamura K (2009) An AP2-Type transcription factor, WRINKLED1, of Arabidopsis thaliana binds to the aw-box sequence conserved among proximal upstream regions of genes involved in fatty acid synthesis. Plant J 60:476–487
Marchive C, Nikovics K, To A, Lepiniec L, Baud S (2014) Transcriptional regulation of fatty acid production in higher plants: molecular bases and biotechnological outcomes. Eur J Lipid Sci Technol 116:1332–1343
Menard R, Verdier G, Ors M, Erhardt M, Beisson F, Shen WH (2014) Histone H2B monoubiquitination is involved in the regulation of cutin and wax composition in Arabidopsis thaliana. Plant Cell Physiol 55:455–466
Mendes A, Kelly AA, van Erp H, Shaw E, Powers SJ, Kurup S, Eastmond PJ (2013) bZIP67 regulates the omega-3 fatty acid content of Arabidopsis seed oil by activating FATTY ACID DESATURASE3. Plant Cell 25:3104–3116
Nagano M, Ishikawa T, Ogawa Y, Iwabuchi M, Nakasone A, Shimamoto K, Uchimiya H, Kawai-Yamada M (2014) Arabidopsis bax inhibitor-1 promotes sphingolipid synthesis during cold stress by interacting with ceramide-modifying enzymes. Planta 240:77–89
Nakamura Y, Tsuchiya M, Ohta H (2007) Plastidic phosphatidic acid phosphatases identified in a distinct subfamily of lipid phosphate phosphatases with prokaryotic origin. J Biol Chem 282:29013–29021
Narise T, Kobayashi K, Baba S, Shimojima M, Masuda S, Fukaki H, Ohta H (2010) Involvement of auxin signaling mediated by IAA14 and ARF7/19 in membrane lipid remodeling during phosphate starvation. Plant Mol Biol 72:533–544
Newman T, Debruijn FJ, Green P, Keegstra K, Kende H, McIntosh L, Ohlrogge J, Raikhel N, Somerville S, Thomashow M, Retzel E, Somerville C (1994) Genes galore-a summary of methods for accessing results from large-scale partial sequencing of anonymous Arabidopsis CDNA clones. Plant Physiol 106:1241–1255
Niu XP, Helentjaris T, Bate NJ (2002) Maize AB14 binds coupling element1 in abscisic acid and sugar response genes. Plant Cell 14:2565–2575
Ogas J, Cheng JC, Sung ZR, Somerville C (1997) Cellular Differentiation Regulated by Gibberellin in the Arabidopsis thaliana Pickle Mutant. Science 277:91–94
Ogas J, Kaufmann S, Henderson J, Somerville C (1999) PICKLE is a CHD3 chromatin-remodeling factor that regulates the transition from embryonic to vegetative development in Arabidopsis. Proc Natl Acad Sci USA 96:13839–13844
Oh IS, Park AR, Bae MS, Kwon SJ, Kim YS, Lee JE, Kang NY, Lee SM, Cheong H, Park OK (2005) Secretome analysis reveals an Arabidopsis lipase involved in defense against Alternaria brassicicola. Plant Cell 17:2832–2847
O’Malley RC, Alonso JM, Kim CJ, Leisse TJ, Ecker JR (2007) An adapter ligation-mediated PCR method for high-throughput mapping of t-dna inserts in the Arabidopsis genome. Nat Protoc 2:2910–2917
Oshima Y, Shikata M, Koyama T, Ohtsubo N, Mitsuda N, Ohme-Takagi M (2013) MIXTA-Like transcription factors and WAX INDUCER1/SHINE1 coordinately regulate cuticle development in Arabidopsis and Torenia fournieri. Plant Cell 25:1609–1624
Penfield S, Li Y, Gilday AD, Graham S, Graham IA (2006) Arabidopsis ABA INSENSITIVE4 regulates lipid mobilization in the embryo and reveals repression of seed germination by the endosperm. Plant Cell 18:1887–1899
Peters C, Li MY, Narasimhan R, Roth M, Welti R, Wang XM (2010) Nonspecific phospholipase C NPC4 promotes responses to abscisic acid and tolerance to hyperosmotic stress in Arabidopsis. Plant Cell 22:2642–2659
Pinosa F, Buhot N, Kwaaitaal M, Fahlberg P, Thordal-Christensen H, Ellerstrom M, Andersson MX (2013) Arabidopsis phospholipase D delta is involved in basal defense and nonhost resistance to powdery mildew fungi. Plant Physiol 163:896–906
Raffaele S, Vailleau F, Léger A, Joubès J, Miersch O, Huard C, Blée E, Mongrand S, Domergue F, Roby D (2008) A MYB transcription factor regulates very-long-chain fatty acid biosynthesis for activation of the hypersensitive cell death response in Arabidopsis. Plant Cell 20:252–267
Rietz S, Dermendjiev G, Oppermann E, Tafesse FG, Effendi Y, Holk A, Parker JE, Teige M, Scherer GFE (2010) Roles of Arabidopsis patatin-related phospholipases a in root development are related to auxin responses and phosphate deficiency. Mol Plant 3:524–538
Routaboul JM, Benning C, Bechtold N, Caboche M, Lepiniec L (1999) The TAG1 locus of Arabidopsis Encodes for a diacylglycerol acyltransferase. Plant Physiol Biochem 37:831–840
Seo PJ, Lee SB, Suh MC, Park MJ, Go YS, Park CM (2011) The MYB96 transcription factor regulates cuticular wax biosynthesis under drought conditions in Arabidopsis. Plant Cell 23:1138–1152
Shen B, Sinkevicius KW, Selinger DA, Tarczynski MC (2006) The homeobox gene GLABRA2 affects seed oil content in Arabidopsis. Plant MolBiol 60:377–387
Shi JX, Malitsky S, De Oliveira S, Branigan C, Franke RB, Schreiber L, Aharoni A (2011) SHINE transcription factors act redundantly to pattern the archetypal surface of Arabidopsis flower organs. PLoS Genet 7:e1001388
Song QX, Li QT, Liu YF, Zhang FX, Ma B, Zhang WK, Man WQ, Du WG, Wang GD, Chen SY, Zhang JS (2013) Soybean GmbZIP123 gene enhances lipid content in the seeds of transgenic Arabidopsis plants. J Exp Bot 64:4329–4341
Stone SL, Kwong LW, Yee KM, Pelletier J, Lepiniec L, Fischer RL, Goldberg RB, Harada JJ (2001) LEAFY COTYLEDON2 encodes a B3 domain transcription factor that induces embryo development. Proc Natl Acad Sci USA 98:11806–11811
Tanaka H, Watanabe M, Watanabe D, Tanaka T, Machida C, Machida Y (2002) ACR4, a putative receptor kinase gene of Arabidopsis thaliana, that is expressed in the outer cell layers of embryos and plants, is involved in proper embryogenesis. Plant Cell Physiol 43:419–428
To A, Joubes J, Barthole G, Lecureuil A, Scagnelli A, Jasinski S, Lepiniec L, Baud S (2012) WRINKLED transcription factors orchestrate tissue-specific regulation of fatty acid biosynthesis in Arabidopsis. Plant Cell 24:5007–5023
Troncoso-Ponce MA, Kilaru A, Cao X, Durrett TP, Fan JL, Jensen JK, Thrower NA, Pauly M, Wilkerson C, Ohlrogge JB (2011) Comparative deep transcriptional profiling of four developing oilseeds. Plant J 68:1014–1027
Troncoso-Ponce MA, Cao X, Yang ZL, Ohlrogge JB (2013) Lipid turnover during senescence. Plant Sci 205:13–19
Tsukagoshi H, Morikami A, Nakamura K (2007) Two B3 domain transcriptional repressors prevent sugar-inducible expression of seed maturation genes in Arabidopsis seedlings. Proc Natl Acad Sci USA 104:2543–2547
Tsuwamoto R, Fukuoka H, Takahata Y (2008) GASSHO1 and GASSHO2 encoding a putative leucine-rich repeat transmembrane-type receptor kinase are essential for the normal development of the epidermal surface in Arabidopsis embryos. Plant J 54:30–42
Uraji M, Katagiri T, Okuma E, Ye W, Hossain MA, Masuda C, Miura A, Nakamura Y, Mori IC, Shinozaki K, Murata Y (2012) Cooperative function of PLDδ and PLDα1 in abscisic acid-induced stomatal closure in Arabidopsis. Plant Physiol 159:450–460
Vijayaraj P, Jashal CB, Vijayakumar A, Rani SH, Rao DKV, Rajasekharan R (2012) A bifunctional enzyme that has both monoacylglycerol acyltransferase and acyl hydrolase activities. Plant Physiol 160:667–683
Voisin D, Nawrath C, Kurdyukov S, Franke RB, Reina-Pinto JJ, Efremova N, Will I, Schreiber L, Yephremov A (2009) Dissection of the complex phenotype in cuticular mutants of Arabidopsis Reveals a role of SERRATE as a mediator. PLoS Genet 5:e1000703
Wallis JG, Browse J (2002) Mutants of Arabidopsis reveal many roles for membrane lipids. Prog Lipid Res 41:254–278
Wang FF, Perry SE (2013) Identification of direct targets of FUSCA3, a key regulator of arabidopsis seed development. Plant Physiol 161:1251–1264
Watanabe M, Tanaka H, Watanabe D, Machida C, Machida Y (2004) The ACR4 receptor-like kinase is required for surface formation of epidermis-related tissues in Arabidopsis thaliana. Plant J 39:298–308
Wu RH, Li SB, He S, Wassmann F, Yu CH, Qin GJ, Schreiber L, Qu LJ, Gu HY (2011) CFL1, a WW domain protein, regulates cuticle development by modulating the function of HDG1, a class IV homeodomain transcription factor, in rice and Arabidopsis. Plant Cell 23:3392–3411
Xia Y, Yu KS, Navarre D, Seebold K, Kachroo A, Kachroo P (2010) The glabra1 mutation affects cuticle formation and plant responses to microbes. Plant Physiol 154:833–846
Xing Q, Creff A, Waters A, Tanaka H, Goodrich J, Ingram GC (2013) ZHOUPI controls embryonic cuticle formation via a signalling pathway involving the subtilisin protease ABNORMAL LEAF-SHAPE1 and the receptor kinases GASSHO1 and GASSHO2. Development 140:770–779
Yang WY, Devaiah SP, Pan XQ, Isaac G, Welti R, Wang XM (2007) AtPLAI is an acyl hydrolase involved in basal jasmonic acid production and arabidopsis resistance to Botrytis cinerea. J Biol Chem 282:18116–18128
Yang Y, Yu XC, Song LF, An CC (2011) ABI4 activates DGAT1 expression in Arabidopsis seedlings during nitrogen deficiency. Plant Physiol 156:873–883
Yang WY, Zheng Y, Bahn SC, Pan XQ, Li MY, Vu HS, Roth MR, Scheu B, Welti R, Hong YY, Wang XM (2012) The patatin-containing phospholipase A pPLAII alpha modulates oxylipin formation and water loss in Arabidopsis thaliana. Mol Plant 5:452–460
Yeats TH, Martin LBB, Viart HMF, Isaacson T, He YH, Zhao LX, Matas AJ, Buda GJ, Domozych DS, Clausen MH, Rose JKC (2012) The identification of cutin synthase: formation of the plant polyester cutin. Nat Chem Biol 8:609–611
Yeats TH, Huang WL, Chatterjee S, Viart HMF, Clausen MH, Stark RE, Rose JKC (2014) Tomato cutin deficient 1 (CD1) and putative orthologs comprise an ancient family of cutin synthase-like (CUS) proteins that are conserved among land plants. Plant Journal 77:667–675
Zhang M, Fan JL, Taylor DC, Ohlrogge JB (2009a) DGAT1 and PDAT1 acyltransferases have overlapping functions in Arabidopsis triacylglycerol biosynthesis and are essential for normal pollen and seed development. Plant Cell 21:3885–3901
Zhang YY, Zhu HY, Zhang Q, Li MY, Yan M, Wang R, Wang LL, Welti R, Zhang WH, Wang XM (2009b) Phospholipase D alpha 1 and phosphatidic acid regulate NADPH oxidase activity and production of reactive oxygen species in ABA-mediated stomatal closure in Arabidopsis. Plant Cell 21:2357–2377
Zhao J, Devaiah SP, Wang CX, Li MY, Welti R, Wang XM (2013) Arabidopsis phospholipase D1 modulates defense responses to bacterial and fungal pathogens. New Phytol 199:228–240
Zheng SZ, Liu YL, Li B, Shang ZL, Zhou RG, Sun DY (2012) Phosphoinositide-specific phospholipase C9 is involved in the thermotolerance of Arabidopsis. Plant J 69:689–700
Zou JT, Wei YD, Jako C, Kumar A, Selvaraj G, Taylor DC (1999) The Arabidopsis thaliana TAG1 mutant has a mutation in a diacylglycerol acyltransferase gene. Plant J 19:645–653
Acknowledgments
We thank Johnny Lloyd (Michigan State University) and David Meinke (Oklahoma State University) for helpful discussions and Dylan Kosma for suggestions on the manuscript. We are grateful for the helpful advice of Tanya Beradini and Eva Huala of The Arabidopsis Information Resource on accessing information on mutants, and to Tanya for providing an updated list of Arabidopsis loci without T-DNA insertions. We thank Sebastian Baud, Martine Miquel, Phil Bates, Vincent Arondel, ChangCheng Xu and Fred Beisson for reviewing sections of the data. This work was supported in part by Department of Energy–Great Lakes Bioenergy Research Center Cooperative Agreement DE-FC02-07ER64494 to JBO.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Neal Stewart.
V. Shaw, M. Zhang, R. J. Kim, and W. Yang contributed equally to this publication.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
McGlew, K., Shaw, V., Zhang, M. et al. An annotated database of Arabidopsis mutants of acyl lipid metabolism. Plant Cell Rep 34, 519–532 (2015). https://doi.org/10.1007/s00299-014-1710-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1710-8