Abstract
Positioning the nucleus to a specific cellular location is a prerequisite for high-fidelity transmission of the genetic material to daughter cells. The cellular location of the nucleus just before its division is variable in budding yeast species which rely on a variety of mechanisms for nuclear division. Dynamic growth and shrinkage kinetics of microtubules (MTs) and forces exerted by the MT plus- and minus-end-directed motor proteins empower nuclear movement. Even though the overall process of nuclear migration is largely conserved across budding yeasts, in-depth molecular analyses of newly emerging model budding yeasts began to reveal striking differences from the paradigms that have been established based on the studies performed in the well-characterized budding yeast Saccharomyces cerevisiae. Here, we highlight the molecular players involved in differential nuclear migration in diverse budding yeasts.

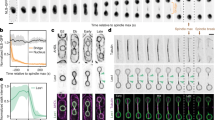
The U. maydis cartoons are adapted from Straube et al. 2005
Similar content being viewed by others
References
Adames NR, Cooper JA (2000) Microtubule interactions with the cell cortex causing nuclear movements in Saccharomyces cerevisiae. J Cell Biol 149:863–874
Alberti-Segui C, Dietrich F, Altmann-Johl R, Hoepfner D, Philippsen P (2001) Cytoplasmic dynein is required to oppose the force that moves nuclei towards the hyphal tip in the filamentous ascomycete Ashbya gossypii. J Cell Sci 114:975–986
Altamirano S, Fang D, Simmons C, Sridhar S, Wu P, Sanyal K, Kozubowski L (2017) Fluconazole-induced ploidy change in cryptococcus neoformans results from the uncoupling of cell growth and nuclear division. mSphere. https://doi.org/10.1128/mSphere.00205-17
Banuett F, Herskowitz I (2002) Bud morphogenesis and the actin and microtubule cytoskeletons during budding in the corn smut fungus, Ustilago maydis. Fungal Genet Biol 37:149–170
Byers B, Goetsch L (1975) Behavior of spindles and spindle plaques in the cell cycle and conjugation of Saccharomyces cerevisiae. J Bacteriol 124:511–523
Fink G, Steinberg G (2006) Dynein-dependent motility of microtubules and nucleation sites supports polarization of the tubulin array in the fungus Ustilago maydis. Mol Biol Cell 17:3242–3253. https://doi.org/10.1091/mbc.e05-12-1118
Finley KR, Berman J (2005) Microtubules in Candida albicans hyphae drive nuclear dynamics and connect cell cycle progression to morphogenesis. Eukaryot Cell 4:1697–1711. https://doi.org/10.1128/EC.4.10.1697-1711.2005
Finley KR, Bouchonville KJ, Quick A, Berman J (2008) Dynein-dependent nuclear dynamics affect morphogenesis in Candida albicans by means of the Bub2p spindle checkpoint. J Cell Sci 121:466–476. https://doi.org/10.1242/jcs.015172
Fischer R, Zekert N, Takeshita N (2008) Polarized growth in fungi–interplay between the cytoskeleton, positional markers and membrane domains. Mol Microbiol 68:813–826. https://doi.org/10.1111/j.1365-2958.2008.06193.x
Gladfelter A, Berman J (2009) Dancing genomes: fungal nuclear positioning. Nat Rev Microbiol 7:875–886. https://doi.org/10.1038/nrmicro2249
Harari Y, Ram Y, Kupiec M (2018) Frequent ploidy changes in growing yeast cultures. Curr Genet 64:1001–1004. https://doi.org/10.1007/s00294-018-0823-y
Hu G, Liu I, Sham A, Stajich JE, Dietrich FS, Kronstad JW (2008) Comparative hybridization reveals extensive genome variation in the AIDS-associated pathogen Cryptococcus neoformans. Genome Biol 9:R41. https://doi.org/10.1186/gb-2008-9-2-r41
Hwang E, Kusch J, Barral Y, Huffaker TC (2003) Spindle orientation in Saccharomyces cerevisiae depends on the transport of microtubule ends along polarized actin cables. J Cell Biol 161:483–488. https://doi.org/10.1083/jcb.200302030
Inoue S, Turgeon BG, Yoder OC, Aist JR (1998) Role of fungal dynein in hyphal growth, microtubule organization, spindle pole body motility and nuclear migration. J Cell Sci 111(Pt 11):1555–1566
Kopecka M, Gabriel M, Takeo K, Yamaguchi M, Svoboda A, Ohkusu M, Hata K, Yoshida S (2001) Microtubules and actin cytoskeleton in Cryptococcus neoformans compared with ascomycetous budding and fission yeasts. Eur J Cell Biol 80:303–311. https://doi.org/10.1078/0171-9335-00157
Kozubowski L, Yadav V, Chatterjee G, Sridhar S, Yamaguchi M, Kawamoto S, Bose I, Heitman J, Sanyal K (2013) Ordered kinetochore assembly in the human-pathogenic basidiomycetous yeast Cryptococcus neoformans. MBio 4:e00613–e00614. https://doi.org/10.1128/mBio.00614-13
Kubo K, Okada H, Shimamoto T, Kimori Y, Mizunuma M, Bi E, Ohnuki S, Ohya Y (2019) Implications of maintenance of mother-bud neck size in diverse vital processes of Saccharomyces cerevisiae. Curr Genet 65:253–267. https://doi.org/10.1007/s00294-018-0872-2
Lee WL, Oberle JR, Cooper JA (2003) The role of the lissencephaly protein Pac1 during nuclear migration in budding yeast. J Cell Biol 160:355–364. https://doi.org/10.1083/jcb.200209022
Legrand M, Jaitly P, Feri A, d’Enfert C, Sanyal K (2019) Candida albicans: an emerging yeast model to study eukaryotic genome plasticity. Trends Genet 35:292–307. https://doi.org/10.1016/j.tig.2019.01.005
Martin R, Walther A, Wendland J (2004) Deletion of the dynein heavy-chain gene DYN1 leads to aberrant nuclear positioning and defective hyphal development in Candida albicans. Eukaryot Cell 3:1574–1588. https://doi.org/10.1128/EC.3.6.1574-1588.2004
Mochizuki T, Tanaka S, Watanabe S (1987) Ultrastructure of the mitotic apparatus in Cryptococcus neoformans. J Med Vet Mycol 25:223–233
Morris NR (2000) Nuclear migration. From fungi to the mammalian brain. J Cell Biol 148:1097–1101
Raspelli E, Fraschini R (2019) Spindle pole power in health and disease. Curr Genet. https://doi.org/10.1007/s00294-019-00941-7
Reinsch S, Gonczy P (1998) Mechanisms of nuclear positioning. J Cell Sci 111(Pt 16):2283–2295
Roy B, Burrack LS, Lone MA, Berman J, Sanyal K (2011) CaMtw1, a member of the evolutionarily conserved Mis12 kinetochore protein family, is required for efficient inner kinetochore assembly in the pathogenic yeast Candida albicans. Mol Microbiol 80:14–32. https://doi.org/10.1111/j.1365-2958.2011.07558.x
Rusan NM, Fagerstrom CJ, Yvon AM, Wadsworth P (2001) Cell cycle-dependent changes in microtubule dynamics in living cells expressing green fluorescent protein-alpha tubulin. Mol Biol Cell 12:971–980. https://doi.org/10.1091/mbc.12.4.971
Sanyal K (2012) How do microbial pathogens make CENs? PLoS Pathog 8:e1002463. https://doi.org/10.1371/journal.ppat.1002463
Schwartz K, Richards K, Botstein D (1997) BIM1 encodes a microtubule-binding protein in yeast. Mol Biol Cell 8:2677–2691
Selmecki A, Bergmann S, Berman J (2005) Comparative genome hybridization reveals widespread aneuploidy in Candida albicans laboratory strains. Mol Microbiol 55:1553–1565. https://doi.org/10.1111/j.1365-2958.2005.04492.x
Shaw SL, Yeh E, Maddox P, Salmon ED, Bloom K (1997) Astral microtubule dynamics in yeast: a microtubule-based searching mechanism for spindle orientation and nuclear migration into the bud. J Cell Biol 139:985–994
Sheeman B, Carvalho P, Sagot I, Geiser J, Kho D, Hoyt MA, Pellman D (2003) Determinants of S. cerevisiae dynein localization and activation: implications for the mechanism of spindle positioning. Curr Biol 13:364–372
Steinberg G, Wedlich-Soldner R, Brill M, Schulz I (2001) Microtubules in the fungal pathogen Ustilago maydis are highly dynamic and determine cell polarity. J Cell Sci 114:609–622
Straube A, Enard W, Berner A, Wedlich-Soldner R, Kahmann R, Steinberg G (2001) A split motor domain in a cytoplasmic dynein. EMBO J 20:5091–5100. https://doi.org/10.1093/emboj/20.18.5091
Straube A, Brill M, Oakley BR, Horio T, Steinberg G (2003) Microtubule organization requires cell cycle-dependent nucleation at dispersed cytoplasmic sites: polar and perinuclear microtubule organizing centers in the plant pathogen Ustilago maydis. Mol Biol Cell 14:642–657. https://doi.org/10.1091/mbc.e02-08-0513
Straube A, Weber I, Steinberg G (2005) A novel mechanism of nuclear envelope break-down in a fungus: nuclear migration strips off the envelope. EMBO J 24:1674–1685. https://doi.org/10.1038/sj.emboj.7600644
Sudbery P, Gow N, Berman J (2004) The distinct morphogenic states of Candida albicans. Trends Microbiol 12:317–324. https://doi.org/10.1016/j.tim.2004.05.008
Suelmann R, Fischer R (2000) Nuclear migration in fungi–different motors at work. Res Microbiol 151:247–254
Sutradhar S, Yadav V, Sridhar S, Sreekumar L, Bhattacharyya D, Ghosh SK, Paul R, Sanyal K (2015) A comprehensive model to predict mitotic division in budding yeasts. Mol Biol Cell 26:3954–3965. https://doi.org/10.1091/mbc.E15-04-0236
Thakur J, Sanyal K (2011) The essentiality of the fungus-specific Dam1 complex is correlated with a one-kinetochore-one-microtubule interaction present throughout the cell cycle, independent of the nature of a centromere. Eukaryot Cell 10:1295–1305. https://doi.org/10.1128/EC.05093-11
Varshney N, Sanyal K (2019) Aurora kinase Ipl1 facilitates bilobed distribution of clustered kinetochores to ensure error-free chromosome segregation in Candida albicans. Mol Microbiol. https://doi.org/10.1111/mmi.14275
Varshney N, Som S, Chatterjee S, Sridhar S, Bhattacharyya D, Paul R, Sanyal K (2019) Spatio-temporal regulation of nuclear division by Aurora B kinase Ipl1 in Cryptococcus neoformans. PLoS Genet 15:e1007959. https://doi.org/10.1371/journal.pgen.1007959
Xiang X, Fischer R (2004) Nuclear migration and positioning in filamentous fungi. Fungal Genet Biol 41:411–419. https://doi.org/10.1016/j.fgb.2003.11.010
Yadav V, Sanyal K (2018) Sad1 Spatiotemporally Regulates Kinetochore Clustering To Ensure High-Fidelity Chromosome Segregation in the Human Fungal Pathogen Cryptococcus neoformans. mSphere. https://doi.org/10.1128/mSphere.00190-18
Yamaguchi M, Biswas SK, Ohkusu M, Takeo K (2009) Dynamics of the spindle pole body of the pathogenic yeast Cryptococcus neoformans examined by freeze-substitution electron microscopy. FEMS Microbiol Lett 296:257–265. https://doi.org/10.1111/j.1574-6968.2009.01643.x
Yeh E, Skibbens RV, Cheng JW, Salmon ED, Bloom K (1995) Spindle dynamics and cell cycle regulation of dynein in the budding yeast, Saccharomyces cerevisiae. J Cell Biol 130:687–700
Yeh E, Yang C, Chin E, Maddox P, Salmon ED, Lew DJ, Bloom K (2000) Dynamic positioning of mitotic spindles in yeast: role of microtubule motors and cortical determinants. Mol Biol Cell 11:3949–3961. https://doi.org/10.1091/mbc.11.11.3949
Yokoyama K, Kaji H, Nishimura K, Miyaji M (1990) The role of microfilaments and microtubules in apical growth and dimorphism of Candida albicans. J Gen Microbiol 136:1067–1075. https://doi.org/10.1099/00221287-136-6-1067
Acknowledgements
We thank members of the Molecular Mycology Laboratory, Jawaharlal Nehru Centre for Advanced Scientific Research (JNCASR), Bangalore, India for useful discussion and suggestions. We apologize to those whose work could not be mentioned because of space restrictions. This work was supported by Tata Innovation Fellowship ((BT/HRD/35/01/03/2017) and intramural support from JNCASR to KS and a research associate fellowship from the Council of Scientific and Industrial Research (CSIR) (09/733 (0253)/219-EMR-I) and (9/733 (0161)/2011-EMR-I) to NV.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Varshney, N., Sanyal, K. Nuclear migration in budding yeasts: position before division. Curr Genet 65, 1341–1346 (2019). https://doi.org/10.1007/s00294-019-01000-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-01000-x