Abstract
Using an alternative bio-product is one of the most promising ways to control bovine mastitis and avoid new intra-mammary infections. The aims of this study were to ascertain the prevalence of biofilm-forming bacteria responsible for causing clinical mastitis in dairy herds and to assess the effectiveness of bacteriocins, produced by Bacillus subtilis, in controlling the growth of these bacteria in the milk of animals. A total of 150 milk samples were collected from cows and buffalos suffering from mastitis and the etiological agents were isolated and identified by the VITEK-2-COMPACT-SYSTEM®. Additionally, the capability of the bacterial isolates to produce biofilms was determined. RT-PCR was used to detect enterotoxin-producing genes (sed and seb), resistance genes (mecA and blaZ), and biofilm-associated genes (icaA and fnbA) in the isolated bacteria. The susceptibility patterns of the bacterial isolates to bacteriocins were assessed using an agar well-diffusion assay. S. aureus was significantly more capable of producing biofilms than coagulase-negative Staphylococcus isolates. S. ubris was the strongest biofilm producer among the Streptococcus species. The sensitivity profiles of the Staphylococcus spp. (S. aureus and coagulase-negative Staphylococcus) and their biofilm producers to bacteriocins were significantly higher (100% and 90%, respectively) at the same concentration. Bacteriocins had a lethal effect on Staphylococci, Streptococci, and biofilm development at a dose of 250 µg/mL. In dairy farms, bacteriocins are a viable alternative treatment for the prevention and control of bovine clinical mastitis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bovine mastitis is the most prevalent disease affecting dairy cattle, causing financial losses and detrimentally affecting animal welfare, production, food safety, and the quality of milk [1, 2]. It can be caused by various Gram-positive and Gram-negative bacteria, such as Staphylococcus aureus, Streptococcus agalactiae, and Mycoplasma species. They can be infectious or environmental, for example, Enterococcus spp., Escherichia coli, coagulase-negative Staphylococcus (CNS), and Streptococcus uberis. To manage new cases of mastitis, general sanitation practices, such as disinfecting the teats post-milking, improving milking hygiene, and maintaining milking equipment, must be employed. Thus, natural remedies, especially alternative medicines, are particularly important for the prevention and treatment of bovine mastitis [3].
S. aureus is the most common Gram-positive bacteria associated with different types of clinical and subclinical mastitis [4]. It is primarily found in persistently infected mammary glands; therefore, keeping the udder clean during milking can help shield healthy cows from diseased cows, lowering the infection rate [5]. S. agalactiae, found in the environment of dairy cows and the gastrointestinal tract of cattle, causes infectious mastitis [6]. It can be transferred via milking machines and through the fecal–oral route, notably through contaminated drinking water. In fact, the mammary gland may become infected by germs found in the cow’s bedding area and on the milking apparatus [7].
Biofilms are organized bacterial populations attached to biotic or abiotic surfaces that constitute a self-produced matrix, which includes exopolysaccharides, proteins, teichoic acids, enzymes, and extracellular DNA [8]. Before adhesion is facilitated by cell wall-associated structures (flagella, fimbriae, and pili), biofilm formation begins with bacterial attachment to an abiotic surface via hydrophobic or electrostatic interactions. In conjunction with this adherence, polymer bridges between bacteria and the surface are frequently formed [9, 10]. Teat dips used nowadays in commercial pre- and post-milking processes include chlorine, hydrogen peroxide, and iodine. Despite being efficient, these substances could seriously irritate the skin [11]. Therefore, it is necessary to seek about more natural substitutes that can be utilized in combination or in addition to the current chemical materials.
Gram-positive rod- or cocci-shaped facultative anaerobes called lactic acid bacteria are being increasingly investigated for their ability to create inhibitory compounds resembling bacteriocins, small antimicrobial peptides that are active against various bacteria [12, 13]. Several bacteriocins have been characterized in terms of their structure, mode of action, and range of inhibitory activity [14]. Bacteriocins of lactic acid bacteria are classified as extracellularly produced primary or modified products of bacterial ribosomal synthesis, which have a bactericidal activity [15]. The action of bacteriocins based on disrupting membranes of bacteria and it has a net positive charge that, despite their diversity as peptides, allows them to fold into an amphiphilic shape when they come into contact with bacterial membranes [16]. The production of bacteriocins is normally performed in complex growth media: De Man, Rogosa, and Sharpe (MRS) broth [17]. This study was aimed at controlling the bacterial populations that cause clinical mastitis in dairy animals. We determined the prevalence of biofilm-producing mastitis-causing bacteria in dairy herds, identified biofilm-associated genes, and assessed the efficacy of bacteriocins produced by Bacillus subtilis, as natural alternatives to antimicrobial agents, against all isolated bacterial strains.
Materials and Methods
Study Site and Period
The study was conducted on both lactating cows and buffalos at private dairy farms in El-Faiyum Governorate between May 2019 and March 2021. Lactating animals were housed in an earthen-floored cow house system. Following the recommendations of the “National Mastitis Council,” the udder of each lactating animal was examined for the presence of clinical signs of mastitis, such as asymmetry, hotness, swelling, or any physical changes prior to sampling. This was followed by palpation to look for injury, atrophy, fibrosis, or inflammatory swelling. The dairy farms under investigation had minimal to moderate hygiene measures.
Study Design
The study’s protocol was designed to estimate the prevalence of clinical mastitis in various lactating animals. In addition to examining their capacity to create biofilms, the most prevalent Gram-positive cocci that caused clinical mastitis in dairy farms were isolated and identified. Next, we assessed the effectiveness of bacteriocins against mastitis-causing, biofilm-forming bacteria to examine if they could be used as natural therapeutics to treat bovine clinical mastitis.
Collection of Milk Samples
Milk samples were obtained aseptically as described earlier [18]. The udder and tips of the teat orifice were thoroughly cleaned with water and soap and dried with a sterilized cloth. The teats were cleaned with 70% alcohol. The first few streams of milk were excluded and milk samples (n = 150) were collected in sterile, screw-capped McCartney bottles, labeled, serialized, and transported immediately to the lab on ice for microbial analysis.
Isolation and Identification of Clinical Mastitis-Causing Pathogenic Bacteria
After being incubated at 37 ℃ for 18–24 h, milk samples were centrifuged at 3000 rpm for 15 minutes. The fluid supernatant and the cream layer were discarded. A small amount of the sediment was extracted and cultured in tryptone soya broth for 18–24 h at 37 ℃. Loopfuls of broth were cultured on mannitol salt agar, Baird–Parker agar (to examine Staphylococcus spp.) and modified Edwards medium (to examine Streptococcus spp.) for 24–48 h at 37 ℃. For identification, bacteriological films were prepared, stained by Gram’s stain, and studied under a microscope. While suspected Streptococcus isolates were identified as Gram-positive cocci that were arranged either singly or in chains, suspected Staphylococcus isolates were identified as Gram-positive cocci occurring as singles, pairs, or mostly as irregular clusters (like bunches of grapes). Pure colonies that had been confirmed were transferred to tryptone soya agar and cultured for 24–48 h at 37 °C. Before biochemical identification of the isolates, their colony morphology and purity were assessed a second time according to Quinn et al. [18]. Additionally, VITEK-2-COMPACT-SYSTEM® (BioMérieux) was used to confirm the identity of the bacterial isolates. RT-PCR was also performed on different Staphylococcus and Streptococcus isolates to determine four genes: two enterotoxin-producing genes (sed and seb) and two resistance genes (mecA and blaZ). The primer sequences and sizes of PCR amplicons are illustrated in (S1) [19,20,21].
Detection of Biofilm-Forming Bacteria on Yeast Extract–Casamino Acid Agar Supplemented with Congo Red (YESCA CR)
Pure colonies of the bacterial isolates were streaked on Luria–Bertani agar and incubated for 48 h at 37 ℃. A single colony was selected using a sterilized bacteriological loop, streaked onto YESCA CR agar, and cultured for 48–72 h at 25 ℃. The development of biofilms was investigated according to Zhou et al [22]. Pink or white color of the bacterial colonies indicated a failure to uptake the stain (negative for biofilm formation), while a red color indicated successful uptake of the dye (positive for biofilm formation).
Detection of Biofilm-Associated Genes
RT-PCR was conducted on many distinct Streptococcus and Staphylococcus isolates to identify two biofilm-related genes: icaA, which encodes an N-acetylglucosaminyltransferase, and fnbA, which encodes fibronectin-binding protein A. The primer sequences used and amplicon sizes are summarized in S1 [23, 24].
The published sequence of the icaA and fnbA locus in GenBank was used to designate the primers for the icaA and fnbA genes. For icaA amplification, AF (5′-CCT AAC TAA CGA AAG GTA G-3') and AR (5′-AAG ATA TAG CGA TAA GTG C-3′) and for fnbA amplification, DF (5′-CAT AAA TTG GGA GCA TCA -3′) and DR (5′-ATC AGC AGC TGA ATT CCC ATT -3′) primers were used. The icaA and fnbA genes were used in the PCR to produce some products of 1315 bp and 127 bp, respectively. Ten μl of the rapidly extracted DNA were used as a template in a 50-μl PCR mixture containing 1X PCR buffer (50-mm KCl, 20-mM Tris–HCl), 5 μl of 25-mM MgCl2, 5 μl of 10-mM deoxynucleoside triphosphate (dNTP) mix, 1 μl of 20-μM each primers, and 1U of Taq DNA polymerase. The buffers and enzymes used in the assay were obtained from Fermentas Inc. The amplification of DNA was performed as follows: 92 ℃ for 5 min of initial denaturation; 30 cycles of 92 ℃ for 1 min, 49 ℃ for 1 min. and 72 ℃ for 1 min; and a final extension at 72 ℃ for 7 min. Amplicons were loaded onto 1.5% Agarose Gel containing 1-μg/ml ethidium bromide. The presence and molecular weight of the amplified DNA fragments were confirmed by agarose gel electrophoresis and visualized under UV light.
Synthesis and Purification of Bacteriocins Produced by Lactic Acid-Fermenting Bacteria
B. subtilis stock was prepared by inoculating 10 mL of De Man, Rogosa, and Sharpe (MRS) broth with 0.1 mL of fresh lactic acid bacteria cultures and incubating for 12 h at 37 ℃. Next, 1 mL of this pre-culture was inoculated into 100 mL of MRS broth and incubated for 24 h at 37 ℃ [25]. The isolates grown on MRS broth for 48 h at 37 ℃ were centrifuged at 8000 rpm for 30 min at 4 ℃ to extract bacteriocins. The crude extract, labeled as the cell-free supernatant, was purified by filter sterilization through a 0.22-µm filter (Merck Millipore Ltd., Cork, Ireland) [26, 27]. The critical dilution method in 10-mM phosphate-buffered saline at pH 6.5 was used to recover the bacteriocins. The cell-free supernatant was adjusted to pH six with 1-M NaOH and heated at 80 ℃ for l0 min to deactivate extracellular proteases and hydrogen peroxide.
Determining the Antibacterial Efficacy of Bacteriocins Using Agar Well-Diffusion Assay
The sensitivity profiles of biofilm-producing strains (n = 43) and pathogenic bacterial isolates (n = 49), obtained from animals suffering from mastitis, were examined in the presence of bacteriocins using an agar well-diffusion assay. All bacterial strains were freshly isolated and inoculated into brain heart infusion medium comprising 1.5% agar (w/v) at 1 × 105 CFU/mL using a pour-plate method. Bacteriocin extract (25 µL), prepared as previously described, was poured into wells in the agar that had been perforated to a diameter of 5 mm. Each well carried a different concentration of bacteriocins: 50, 100, 150, or 250 g/mL. The plates were incubated for 24 h at 37 ℃ after the bacteriocins had been allowed to diffuse overnight at 4 ℃. According to Godoy-Santos et al. [28], all plates were inspected for the presence of zones of clearing, identified from the greatest dilution displaying an inhibition zone with a diameter ≥ 9 mm. The procedure was carried out in triplicates.
Microscopic Analysis of Biofilm and Anti-Biofilm Activity of the Pathogenic Strains Using FESEM
On a sterilized acrylic strip, biofilm-producing bacteria were cultivated overnight. Thereafter, the samples were rinsed in 0.1-M buffer sodium cacodylate (Merck KGaA, Darmstadt, Germany) and then dehydrated through serial transfers in ethyl alcohol solutions of various concentrations for 30 min each. The specimens were mounted on metal stubs, after being left at room temperature for 24 h and a sputter coating equipment covered them with a layer of gold, while they were under vacuum (JEOL, JPC 1600, JEOL companies, Japan). At the National Research Center (Cairo, Egypt), the specimens were seen using field emission scanning electron microscope (FESEM; JEOL, JEOL Ltd., Japan) after being coated with gold.
Data Analysis
All data were assembled for statistical analyses using the Statistical Package for the Social Sciences software. A non-parametric test (chi-squared test) was used to evaluate the antibacterial effectiveness of B. subtilis bacteriocins against all bacterial isolates. Meanwhile, one-way ANOVA test was used to determine the diameter of inhibition zone (mm) of testing bacteriocins against gram-positive cocci isolates. P ≤ 0.05 was considered statistically significant.
Results
Prevalence and Distribution Rate of Clinical Mastitis-Causing Gram-Positive Cocci Among the Dairy Farms
The prevalence of clinical mastitis-causing gram-positive cocci on dairy farms is shown in Table 1. Staphylococcus spp. were significantly more prevalent in the milk of buffalo with mastitis (29/36, 80.55%) than in that of cows with mastitis (52/78, 66.67%) at P ≤ 0.05; the opposite trend was observed for Streptococcus spp. (7/36; 19.44% vs26/78, 33.33%, respectively). Furthermore, enterotoxin- and resistance-related genes were detected by RT-PCR (Fig. 1). The sed, seb, mecA, and blaZ genes were amplified to give 278-bp-, 164-bp-, 310-bp-, and 173-bp-long amplicons, respectively (Fig. 1a–d).
The sed gene (a) was amplified by PCR to obtain a 278-bp amplicon. Lanes (2, 3) show a positive result, LD stands for molecular size ladder. For the Seb gene (b), a 164-bp amplicon was expected. Lanes (1–3, 5, 8–10) show a positive result. For the mecA gene (c), a 310-bp amplicon was expected. Lanes (1, 3–10) show a positive result. For the blaZ gene (d), a 173-bp amplicon was expected. Lanes (2, 3, 5–9) show a positive result
The distribution of Staphylococcus and Streptococcus spp. isolates among the animals investigated is displayed in Table 2. The most common Staphylococcus spp. isolate found in cow and buffalo milk was CNS (63.46% and 62.07%, respectively) followed by S. aureus (36.54% and 37.93%, respectively). For Streptococcus spp., S. agalactiae was most prevalent in cow milk (34.61%) followed by S. dysgalactiae and E. faecalis (19.23% each), while S. dysgalactiae and E. faecalis were most abundant in buffalo milk (28.57% each) followed by S. agalactiae, S. ubris, and S. lactarius.
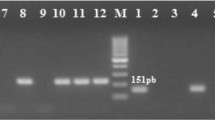
Detection of Biofilm-Related Genes of Gram-Positive Cocci Using RT-PCR
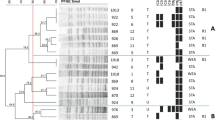
The ability of S. aureus to form biofilms was significantly higher than that of CNS isolates (86.67% and 74.51%, respectively) (Table 3). S. ubris was the most potent Streptococcus species producing 100% of the biofilms, followed by E. faecalis, S. agalactiae, and S. dysgalactiae (71.43%, 70.0%, and 57.14%, respectively). Additionally, biofilm-related genes were found by RT-PCR (Fig. 2a–b): the icaA and fnbA genes were amplified to give 131-bp- and 127-bp-long amplicons. Moreover, the red coloration of the bacterial colonies on YESCA CR agar confirmed the presence of Staphylococcus and Streptococcus spp. (Fig. 3a–b) that form biofilms on the agar. The antimicrobial activity of bacteriocins against Staphylococcus spp. is shown in Table 4. The sensitivity of S. aureus to bacteriocins was significantly high (100%) at a concentration of 250 µg/mL compared with the other tested concentrations at P ≤ 0.01, while that of CNS was 90% at the same concentration. At 150 µg/mL, their sensitivity to bacteriocins did not exceed 80%. In the case of Streptococcus spp., bacteriocins had a 100% lethal effect against S. ubris at 250 µg/mL, followed by S. agalactiae, S. dysgalactiae, and E. faecalis (90%, 80%, and 80%, respectively). Comparatively, the sensitivity of S. lactarius to bacteriocins did not exceed 75% at the same concentration
Biofilm-producing Staphylococcus spp. (a) and Streptococcus spp. (b) on YESCA CR agar. The red-colored colonies tested positive for biofilm formation. The procedure was carried out in triplicates. In addition, the sensitivity profile of the biofilm-producing strains of Staphylococcus and Streptococcus spp. to bacteriocins at different concentrations (1, 2, 3, and 4; 50, 100, 150, and 250 µg/mL, respectively) (Fig. 3c–d). The diameter of the inhibition zone was determined using one-way ANOVA test. The inhibition zone of tested bacteriocins at 250 µg/mL against Staphylococcus spp. was significant 27.6 ± 0.21 mm (c) and 30.4 ± 0.15 mm for Streptococcus spp. (d)
Evaluating the Efficacy of Bacteriocins Against Biofilm-Producing Bacteria Using Agar Well-Diffusion Assay
The sensitivity pattern of biofilm-producing Staphylococcus spp. to different doses of bacteriocins is shown in Table 5. Biofilm-producing S. aureus and CNS were highly sensitive (90%) to bacteriocins at 250 µg/mL compared with the other tested concentrations while their sensitivity did not exceed 80% toward 150-µg/mL bacteriocins. On the other hand, 250 µg/mL bacteriocins produced a 100% lethal effect against biofilm-producing S. ubris followed by S. agalactiae, E. faecalis, and S. dysgalactiae (85.71%, 80%, and 75%, respectively). The sensitivity of biofilm-producing S. lactarius did not exceed 75% at the same bacteriocin concentration. The sensitivity profile of the biofilm-producing strains of Staphylococcus and Streptococcus spp. to different concentrations of bacteriocins is displayed in Fig. 3c–d. The diameter of the inhibition zone at 250 µg/mL of bacteriocins was 27.6 ± 0.21 mm for Staphylococcus spp. (Fig. 3c) and 30.4 ± 0.15 mm for Streptococcus spp. (Fig. 3d).
Characterization of Biofilm-Forming Bacteria and Anti-Biofilm Activity of Bacteriocins Using FESEM
FESEM was used to characterize the biofilm-producing pathogenic strains and clarify the anti-biofilm activity of bacteriocins on tested biofilm-forming bacteria (Fig. 4). The FESEM image of biofilm-producing Staphylococcus spp. showed the spherical (cocci) form in grape-like clusters (Fig. 4a), while Streptococcus spp. appeared in pairs or chains (Fig. 4b). The bacteriocins exhibited their action on Staphylococcus spp. (Fig. 4c), leading to rupture and damage of the bacterial cell membrane. Oppositely, the Streptococcus spp. bulged, and the content of bacteria was destroyed (Fig. 4d).
Field emission scanning electron microscopy of biofilm-producing bacteria. The FESEM image of biofilm-producing Staphylococcus spp. displayed the normal morphological shape (spherical) as grape-like clusters (a). Streptococcus spp. appeared as several chains (b). The efficacy of bacteriocins against Staphylococcus spp. exhibited its action on the bacterial cell (c), leading to rupture and damage of the bacterial cell wall. As well, the Streptococcus spp. appeared bulged, and the content of bacteria was destroyed (d)
Discussion
Effective programs to manage mastitis are focus more on prevention than on therapy. Antibiotic therapy is still an established part of mastitis prevention regimens today. Even though antibiotics are frequently used in conjunction with other treatments, their effectiveness is still unsatisfactory. Hence, finding novel therapeutics is necessary. Numerous natural remedies derived from plants, animals, and microorganisms have been found capable of controlling bovine mastitis [3].
The prevalence rate of clinical mastitis-causing gram-positive cocci on dairy farms clarified that Staphylococcus spp. were the most prevalent isolates in milk of buffalo with mastitis than in milk of cow with mastitis. Contrarily, isolates of Streptococcus spp. were more common in the milk of mastitic cows compared to that of mastitic buffalo, as revealed in Table 1. These findings are consistent with Teklemariam et al. [29], who discovered that this variation may be related to the variations in herd management techniques. Some procedures, such as pre- and post-milking teat dipping and pre- and post-milking hand cleaning, have been shown to reduce the incidence of intra-mammary infections. Furthermore, Staphylococci and streptococci isolates were also examined for enterotoxin-related genes using RT-PCR. The sed and seb genes were amplified at 278 bp and 194 bp, respectively, while the resistance genes (mecA and blaZ genes) were amplified at 310 bp and 173 bp, respectively. Previously, El-nomrousey [30] discovered that sed and seb genes were the most common in all S. aureus isolates, while the mecA gene was found in 28.5% of them. Awad et al. [31] found the blaZ and mecA genes in 95.7% and 50% of the S. aureus isolates, respectively. Raheel et al. [32] used reverse transcription-PCR to demonstrate that the sed and seb genes were found in 20% and 80% of Streptococcus isolates, respectively, while mecA and blaZ were found in 90% and 70%, respectively.
The distribution of Staphylococcus spp. and Streptococcus spp. isolated among the mastitis animals is showed in the current text. The most prevalent Staphylococcus spp. isolates discovered in the tested mastitis animals were CNS and S. aureus, respectively. On the other hand, S. agalactiae, S. dysgalactiae, and E. faecalis were distributed differentially in mastitic cow milk. S. dysgalactiae and E. faecalis, followed by S. agalactiae, S. ubris, and S. lactarius, were recorded at the highest concentration in mastitic buffalo milk as displayed in Table 2. El-jakee et al. [33] reported CNS to be the most common pathogens isolated from both clinical and subclinical bovine mastitis in many different countries. According to Wente and Krömker [34], S. dysgalactiae is an intermediate pathogen since it may endure both inside as well as outside the host. Zhang et al. [35] isolated S. dysgalactiae from 7.5% of their total tested milk samples. Cheng and Han [3] pointed out that contagious bacteria like S. aureus can spread quickly and broadly. Environmental infections, on the other hand, can persist without the host and are a natural component of the area around the cow’s microbiota.
The capability of S. aureus to produce biofilm had a much higher capacity than that of CNS isolates among all biofilm-producing Staphylococcus spp. and Streptococcus spp. isolates. As well, S. ubris, followed by E. faecalis and S. agalactiae, was the most potent Streptococcus species that produced 100% of the biofilms as shown in Table 3. Additionally, isolated bacteria were shown to contain the genes (ica A and fnb A) associated with biofilms. Contrarily, Streptococcus species that produce biofilms are said to have significant virulence factors. Interestingly, biofilm-producing Streptococcus species are said to harbor significant virulence factors [30, 36, 37] in addition to the biofilm-associated genes (icaA and fnbA), reported in 90% and 70% of the isolates, respectively.
Both Gram-positive and Gram-negative bacteria can release bacteriocins. Those released by lactic acid bacteria are of special relevance to Gram-positive bacteria [38]. A diverse range of facultative anaerobes, acid-tolerant, and fermentative organisms, including lactic acid bacteria, have a ‘qualified presumption of safety’ status, which means the U.S. Food and Drug Administration considers these bacteriocins to be safe [39, 40]. All bacteriocins are amphiphilic and extremely hydrophobic, and they can be purified using a variety of techniques. The standard process includes high-performance liquid chromatography (HPLC-MS), ion exchange chromatography, hydrophobic chromatography on octyl sepharose, and precipitation of the bacteriocins from the culture phase using ammonium sulfate. For instance, low-molecular-weight proteins cannot be precipitated using the widely used ammonium sulfate technique. Even at 75% to 80% saturation, these proteins do not precipitate well, and as they pass through the dialysis sacs, they are completely or partially removed. Because of this, this approach cannot be used to purify low-molecular-weight bacteriocins [41, 42].
Bacteriocins’ antibacterial efficacy against various Staphylococcus species was clarified. The sensitivity profile of S. aureus to bacteriocins was notably high (100%) and CNS (90%) at a concentration of 250 g/mL. Meanwhile, at a concentration of 150 g/mL, its sensitivity to bacteriocins did not exceed 80%. Oppositely, the sensitivity pattern of Streptococcus spp. to bacteriocins showed that bacteriocins had a 100% fatal effect on S. ubris at 250 g/mL, followed by S. agalactiae, S. dysgalactiae, and E. faecalis, in that order. S. lactarius’ sensitivity to bacteriocins was also lower at the same concentration as clarified in Table 4. According to Perez et al. [43], bacteriocin targets may have a broad spectrum of activity just like antibiotics, blocking multiple biologically significant cell stages. The formation of pores in the cytoplasmic membrane, associated with the mechanism of bacteriocins, is likely caused by dissipation of the proton-motive force, changes in the membrane potential, and changes in the gradient of the proton-positive ion (H+). Bacteriocins can also be used in vitro to kill or suppress harmful, multidrug-resistant germs [44, 45]. Although bacteriocins have gained popularity as antibacterial peptides against food-borne pathogens [46], they may work better and more broadly when combined with conventional antibiotics [47].
The sensitivity profile of biofilm-producing Staphylococcus spp. to bacteriocins at various tested concentrations is showed that biofilm-producing S. aureus and CNS were highly sensitive to bacteriocins at 250-µg/mL concentration compared to other tested concentrations. Oppositely, the sensitivity profile of Streptococcus spp. that produce biofilms revealed that bacteriocins had the most lethal effect against biofilm-producing S. ubris, followed by S. agalactiae, E. faecalis, and S. dysgalactiae at 250-µg/mL concentration. In addition, the sensitivity of S. lactarius, which produces biofilm, to bacteriocins was not exceeded by 75% at the same concentration. Moreover, the diameter of the inhibitory zone at the greatest dilution (250 µg/mL) was 27.6 ± 0.21 mm for Staphylococcus spp. and 30.4 ± 0.15 mm for Streptococcus spp. as shown in Table 5. The skin of the teats acts as a habitat for a rich and diverse microbial community [48]. Bédard et al. [49] and Bennett et al. [50] have showed that bactofencin A has in vitro antibacterial activity against Listeria monocytogenes and S. aureus. Therefore, bactofencin A would be expected to decrease the Staphylococcus count more than the Streptococcus count and the total viable count. However, in an in vivo study, Bennett et al. [7] found that it did not significantly affect any of these counts in comparison to saline. This could be explained by the fact that bacteriocins are susceptible to degradation by proteolytic enzymes [51]. Consequently, the peptide may be vulnerable to the proteases found in teat skin, leading to its degradation.
Conclusion
This is the first in vitro study examining the efficacy of bacteriocins produced by lactic acid bacteria (B. subtilis) against biofilm-producing Staphylococcus and Streptococcus species. Higher bacteriocin concentrations (250 µg/mL) are required to inhibit bacterial growth and control the spread of clinical mastitis-causing bacteria in dairy farms. Bacteriocins are promising candidates for preventing new intra-mammary infections. Further in vivo studies are required to more holistically determine the efficacy of bacteriocins in suppressing mammary infections
Data Availability
All data are included in the main manuscript and are freely accessible.
Code Availability
Not applicable.
Abbreviations
- CNS :
-
Coagulase-negative Staphylococcus
- E. faecalis :
-
Enterococcus faecalis
- fnbA :
-
Fibronectin-binding protein A gene
- FESEM:
-
Field emission scanning electron microscopy
- icaA :
-
Intercellular adhesion A gene
- S. aureus :
-
Staphylococcus aureus
- S. agalactiae :
-
Streptococcus agalactiae
- S. dysgalactiae :
-
Streptococcus dysgalactiae
- S. lactarius :
-
Streptococcus lactarius
- S. ubris :
-
Streptococcus ubris
References
Ruegg PL (2017) A 100-year review: mastitis detection, management, and prevention. J Dairy Sci 100:10381–10397. https://doi.org/10.3168/jds.2017-13023
Algharib SA, Dawood A, Xie S (2020) Nanoparticles for treatment of bovine Staphylococcusaureus mastitis. Drug Delivery 27(1):292–308. https://doi.org/10.1080/10717544.2020.1724209
Cheng WN, Han SG (2020) Bovine mastitis: risk factors, therapeutic strategies, and alternative treatments—a review. Asian-Australas J Anim Sci 33(11):1699–1713. https://doi.org/10.5713/ajas.20.0156
Vasudevan P, Nair MKM, Annamalai T et al (2003) Phenotypic and genotypic characterization of bovine mastitis isolates of Staphylococcus aureus for biofilm formation. Vet Microbiol 92:179–185. https://doi.org/10.1016/S0378-1135(02)00360-7
Rainard P, Foucras G, Fitzgerald JR et al (2018) Knowledge gaps and research priorities in Staphylococcus aureus mastitis control. Transbound Emerg Dis 65(Suppl 1):149–165. https://doi.org/10.1111/tbed.12698
Jørgensen H, Nordstoga A, Sviland S et al (2016) Streptococcus agalactiae in the environment of bovine dairy herds–rewriting the textbooks? Vet Microbiol 184:64–72. https://doi.org/10.1016/j.vetmic.2015.12.014
Bennett S, Fliss I, Said LB et al (2022) Efficacy of bacteriocin-based formula for reducing staphylococci, streptococci, and total bacterial counts on teat skin of dairy cows. J Dairy Sci 105:4498–4507. https://doi.org/10.3168/jds.2021-21381
Gomes F, Saavedra MJ, Henriques M (2016) Bovine mastitis disease/pathogenicity: evidence of the potential role of microbial biofilms. Pathog Dis 74:ftw006. https://doi.org/10.1093/femspd/ftw006
Melchior M, Vaarkamp H, Fink-Gremmels J (2006) Biofilms: a role in recurrent mastitis infections? Vet J 171:398–407. https://doi.org/10.1016/j.tvjl.2005.01.006
Garrett TR, Bhakoo M, Zhang Z (2008) Bacterial adhesion and biofilms on surfaces. Prog Nat Sci 18:1049–1056. https://doi.org/10.1016/j.pnsc.2008.04.001
Sadakane K, Ichinose T (2015) Effect of the hand antiseptic agents benzalkonium chloride, povidone-iodine, ethanol, and chlorhexidine gluconate on atopic dermatitis in NC/Nga mice. Int J Med Sci 12:116–125. https://doi.org/10.7150/ijms.10322
Hammami R, Fernandez B, Lacroix C et al (2013) Anti-infective properties of bacteriocins: an update. Cell Mol Life Sci 70:2947–2967. https://doi.org/10.1007/s00018-012-1202-3
Ng ZJ, Zarin MA, Lee CK et al (2020) Application of bacteriocins in food preservation and infectious disease treatment for humans and livestock: a review. RSC Adv 10(64):38937–38964. https://doi.org/10.1039/D0RA06161A
Soltani S, Hammami R, Cotter PD et al (2021) Bacteriocins as a new generation of antimicrobials: Toxicity aspects and regulations. FEMS Microbiol Rev 45:fuaa039
Caplice E, Fitzgerald GF (1999) Food fermentations: role of microorganisms in food production and preservation. Int J Food Microbiol 50(1–2):131–149
Torres MD, Sothiselvam S, Lu TK, de la Fuente-Nunez C (2019) Peptide design principles for antimicrobial applications. J Mol Biol 431(18):3547–3567. https://doi.org/10.1016/j.jmb.2018.12.015
De Man JC, Rogosa M, Sharpe ME (1960) A medium for the cultivation of lactobacilli. J Appl Bacteriol 23:130–135
Quinn PJ, Makey BK, Carter ME et al (2002) Veterinary microbiology and microbial diseases. Blackwell Scince Ltd, USA
Mehrotra MG, Wang G, Johnson WM (2000) Multiplex PCR for detection of genes for Staphylococcus aureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1 and methicillin resistance. J Clinic Microbiol. https://doi.org/10.1128/JCM.38.3.1032-1035.2000
McClure JA, Conly JM, Lau V, Elsayed S, Louie T, Hutchins W, Zhang K (2006) Novel multiplex PCR assay for detection of the staphylococcal virulence marker Panton-Valentine leukocidin genes and simultaneous discrimination of methicillin-susceptible from - resistant staphylococci. J Clin Microbiol 44(3):1141–1144. https://doi.org/10.1128/JCM.44.3.1141-1144,2006
Duran N, Ozer B, Duran GG, Onlen Y, Demir C (2012) Antibiotic resistance genes and susceptibility patterns in staphylococci. Indian J Med Res 135:389–396
Zhou Y, Smith DR, Hufnagel DA et al (2013) Experimental manipulation of the microbial functional amyloid called curli. Methods Mol Biol 966:53–75. https://doi.org/10.1007/978-1-62703-245-2_4
Ciftci A, Findik A, Onuk A et al (2009) Detection of methicillin resistance and slime factor production of Staphylococcus aureus in bovine mastitis. Braz J Microbiol 40:254–261. https://doi.org/10.1590/S1517-83822009000200009
Vancraeynest D, Hermans K, Haesebrouck F (2004) Genotypic and phenotypic screening of high and low virulence Staphylococcus aureus isolates from rabbits for biofilm formation and MSCRAMMs. Vet Microbiol 103:241–247. https://doi.org/10.1016/j.vetmic.2004.09.002
Al-Zahrani SH, Al-Zahrani FS (2006) Production of bacteriocin(s) by four lactic acid bacteria isolated from raw milk on organic waste. World Appl Sci J 1(2):135–143
Dhewa T (2012) Screening, production purification and potential use of Bacteriocins from lactic acid bacteria of meat and dairy food origin. IPCBEE 39:35–41
Bromberg R, Moreno ZCL et al (2004) Isolation of bacteriocin-producing lactic acid bacteria from meat and meat products and its spectrum of inhibitory activity. Braz J Microbiol 35:137–144. https://doi.org/10.1590/S1517-83822004000100023
Godoy-Santos F, Pinto MS, Barbosa AAT et al (2019) Efficacy of a ruminal bacteriocin against pure and mixed cultures of bovine mastitis pathogens. Indian J Microbiol 59(3):304–312. https://doi.org/10.1007/s12088-019-00799-w
Teklemariam AD, Nigussie H, Tassew A et al (2015) Isolation and phenotypic characterization of Streptococcus uberis from mastitic cows in and around Batu town, Ethiopia. J Anim Plant Sci 26(3): 4124–4137. http://www.m.elewa.org/JAPS; ISSN 2071–7024 4124.
El-Nomrousy SMR (2014) Sequence analysis of enterotoxigenic Staphylococcus aureus isolated from milk and some milk products. MVSc thesis Faculty of Veterinary Medicine Alexandria University, Egypt
Awad A, Ramadan H, Nasr S et al (2017) Genetic characterization, antimicrobial resistance patterns and virulence determinants of Staphylococcus aureus isolated form bovine mastitis. Pak J Biol Sci 20:298–305. https://doi.org/10.3923/pjbs.2017.298.305
Raheel I, Mohammed AN, Mohamed AA (2022) Phenotypic and genotypic characterization of Streptococci associated with clinical bovine mastitis. JVMR 29(1):13–20. https://doi.org/10.21608/jvmr.2022.121460.1050
El-jakee JK, Aref NE, Gomaa A et al (2013) Emerging of coagulase negative staphylococci as a cause of mastitis in dairy animals: an environmental hazard I. Int J Vet Sci Med 1:74–78. https://doi.org/10.1016/j.ijvsm.2013.05.006
Wente N, Krömker V (2020) Streptococcus dysgalactiae—contagious or environmental? Animals 10:2185. https://doi.org/10.3390/ani10112185
Zhang SH, Piepers S, Shan R et al (2018) Phenotypic and genotypic characterization of antimicrobial resistance profiles in Streptococcus dysgalactiae isolated from bovine clinical mastitis in 5 provinces of China. J Dairy Sci 101(4):3344–3355. https://doi.org/10.3168/jds.2017-14031
Conley J, Olson ME, Cook LS et al (2003) Biofilm formation by group A streptococci: is there a relationship with treatment failure? J Clin Microbiol 41(9):4043–4048
Shome BR, Mitra SD, Bhuvana M et al (2011) Multiplex PCR assay for species identification of bovine mastitis pathogens. J Appl Microbiol 1(11):1349–1356
Lopetuso LR, Giorgio ME, Saviano A et al (2019) Bacteriocins and bacteriophages: therapeutic weapons for gastrointestinal diseases? Int J Mol Sci 20(1):183. https://doi.org/10.3390/ijms20010183
Mokoena MP (2017) Lactic acid bacteria and their bacteriocins: classification, biosynthesis and applications against uropathogens: a mini-review. Molecules 22(8):1255. https://doi.org/10.3390/molecules22081255
Dicks LM, Dreyer L, Smith C et al (2018) A review: the fate of bacteriocins in the human gastro-intestinal tract: do they cross the gut–blood barrier? Front Microbiol 9:2297
Borzenkov V, Surovtsev V, Dyatlov I (2014) Obtaining bacteriocins by chromatographic methods. Adv Biosci Biotechnol 5:446–451. https://doi.org/10.4236/abb.2014.55054
Carolissen-Mackay V, Arendse G, Hastings J (1997) Purification of bacteriocins of lactic acid bacteria: problems and pointers. Int J Food Microbiol 34:1–16
Perez RH, Zendo T, Sonomoto K (2018) Circular and leaderless bacteriocins: biosynthesis, mode of action, applications, and prospects. Front Microbiol 9:2085. https://doi.org/10.3389/fmicb.2018.02085
Perez RH, Zendo T, Sonomoto K (2014) Novel bacteriocins from lactic acid bacteria (LAB): various structures and applications. Microb Cell Fact 13:S3. https://doi.org/10.1186/1475-2859-13-S1-S3
Newstead LL, Varjonen K, Nuttall T et al (2020) Staphylococcal-produced bacteriocins and antimicrobial peptides: their potential as alternative treatments for Staphylococcus aureus infections. Antibiotics 9:40. https://doi.org/10.3390/antibiotics9020040
Ahmad V, Khan MS, Jamal QMS et al (2017) Antimicrobial potential of bacteriocins: in therapy, agriculture and food preservation. Int J Antimicrob Agents 49(1):1–11
Darbandi A, Asadi A, Ari MM et al (2022) Bacteriocins: properties and potential use as antimicrobials. Clin Lab Anal 36:e24093. https://doi.org/10.1002/jcla.24093
Verdier-Metz I, Gagne G, Bornes S et al (2012) Cow teat skin, a potential source of diverse microbial populations for cheese production. Appl Environ Microbiol 78:326–333. https://doi.org/10.1128/AEM.06229-11
Bédard F, Fliss I, Biron E (2019) Structure-activity relationships of the bacteriocin bactofencin A and its interaction with the bacterial membrane. ACS Infect Dis 5:199–207. https://doi.org/10.1021/acsinfecdis.8b00204
Bennett S, Ben Said L, Lacasse P et al (2021) Susceptibility to nisin, bactofencin, pediocin and reuterin of multidrug resistant Staphylococcus aureus, Streptococcus dysgalactiae and Streptococcus uberis causing bovine mastitis. Antibiotics 10(11):1418. https://doi.org/10.3390/antibiotics10111418
Durack E, Mallen S, O’Connor PM et al (2019) Protecting bactofencin A to enable its antimicrobial activity using mesoporous matrices. Int J Pharm 558:9–17. https://doi.org/10.1016/j.ijpharm.2018.12.035
Acknowledgements
The authors are grateful to all the workers at dairy farms that helped us to collect samples and complete the work.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). The authors did not receive a grant from any funding agency.
Author information
Authors and Affiliations
Contributions
IR and ANM performed the study design and planning for this work. AAM contributed to the samples’ collection and preparation. IR and AAM contributed to microbial isolation and identification. ANM performed statistical analysis of data and wrote the manuscript text. All authors reviewed the manuscript and approved the work’s publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interest.
Ethical Approval
The Institutional Animal Care and Use Committee (IACUC) of Beni-Suef University accepted the protocol with approval number (021–187) after confirming that the welfare of the animals was maintained throughout this investigation.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
284_2023_3324_MOESM1_ESM.docx
Supplementary file1 (DOCX 15 KB) Supplementary Table 1 Oligonucleotide primer sequences of enterotoxins, resistance, and biofilm genes of isolated bacteria
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raheel, I., Mohammed, A.N. & Mohamed, A.A. The Efficacy of Bacteriocins Against Biofilm-Producing Bacteria Causing Bovine Clinical Mastitis in Dairy Farms: A New Strategy. Curr Microbiol 80, 229 (2023). https://doi.org/10.1007/s00284-023-03324-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-023-03324-x