Abstract
Previous research indicated that hyphae of Aspergillus fumigatus (A. fumigatus) rather than conidia could successfully build a pulmonary aspergillosis model in immunocompetent mice. In this study, we compared the immune responses induced by hyphae and conidia to explore the possible mechanism of this striking phenomenon. Herein, a novel method was designed and adopted to quantify hyphal fragments. Murine macrophages RAW264.7 and human peripheral blood mononuclear cells were stimulated by A. fumigatus hyphae and conidia in vitro, respectively, and then immunological reactions were measured. Male C57BL/6 mice were challenged with conidia and hyphae through intratracheal inoculation. Dynamic conditions of mice were recorded, and RNA-seq measured corresponding immune responses. The results of the study confirmed that hyphae could induce more intensive inflammation than conidia in vitro and in vivo. However, macrophages revealed a higher production of ROS and M1 polarisation in response to conidia stimuli. Additionally, conidia could promote Th1 cell differentiation, while hyphae could increase the CD4/CD8 ratio. RNA-seq validated the fact that those multiple immunologically relevant pathways were more strongly activated by hyphae than conidia, which also promoted Th2 cell differentiation and suppressed Th1 signalling. Both hyphae and conidia could activate Th17 signalling. In general, conidia and hyphae induced distinctly different host immune responses, and the immune responses induced by conidia played a better protective effect. Therefore, the unique function of hyphae in the spread and infection of Aspergillus should be emphasised, and more research is required to clarify the underlying mechanisms for better understanding and management of aspergillosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pulmonary aspergillosis refers to a spectrum of clinical syndromes caused by airborne Aspergillus spp. These are a series of saprophytic moulds abundant in indoor and outdoor environments that mainly affect the respiratory tract of immunocompromised patients. Depending on the host characteristics, aspergillosis was categorised into invasive pulmonary aspergillosis (IPA), chronic pulmonary aspergillosis and allergic bronchopulmonary aspergillosis [1,2,3,4]. With the widespread use of broad-spectrum antibiotics and immunosuppressive agents, the high incidence of aspergillosis remains a public concern [5,6,7,8]. Moreover, the pandemic of the coronavirus disease 2019 (COVID-19) has shown a high frequency of co-infection with Aspergillus spp., leading to severe outcomes and heavy medical burdens [9,10,11,12,13,14].
Aspergillus fumigatus (A. fumigatus) is the most common etiologic species of aspergillosis. In its life cycle, numerous asexual conidia are produced and spread, which can be inhaled by humans and easily reach the alveoli owing to their tiny size [15]. For immunocompetent individuals, invading conidia will get cleared by airway immune systems, including epithelium, macrophages, neutrophils, and other immune cells. Once the host immune system gets compromised, the uneliminated conidia will swell and germinate to form invasive hyphae, resulting in different types of aspergilloses [15, 16]. The mechanism of host immunity against Aspergillus is stage-specific. During swelling and germination, the outermost hydrophobic rodlet layer of conidia gets lost, and the melanin layer is disorganised, thus resulting in the exposure of the inner layer β-1,3-glucan [17]. These molecular changes account for various pathogen-associated molecular patterns (PAMPs) in the surface of A. fumigatus conidia and hyphae, including chitin, β-glucan, galactomannan, and the like. As for the host immune system, various pathways are activated to defend against the pathogen via specific pattern recognition receptors (PRRs). These include Toll-like receptors (TLRs) and C-type lectin receptors (Clecs), which can recognise the PAMPs in the surface of conidia or hyphae and trigger a series of pro-inflammatory and anti-inflammatory reactions [16].
In the past decades, researchers focused much more on conidia as the inhalable causative agent of pulmonary aspergillosis. However, hyphal fragments can also be airborne and directly invade the respiratory tract [18, 19]. It is common to see nonsporulating moulds, especially A. fumigatus, be isolated and identified as the pathogenic microorganism in the respiratory tract samples [20]. We had previously obtained two clinical nonsporulating A. fumigatus strains from the pulmonary specimens of two immunocompetent patients. To prove their pathogenicity, Zhang Z et al. found an interesting phenomenon that hyphae of A. fumigatus rather than conidia could cause pulmonary aspergillosis in immunocompetent hosts [21]. Hyphal fragments have been reported that they could induce stronger inflammatory cytokines secretions than conidia, indicating that hyphae may have stronger immunogenicity [22]. However, the previous reports only exhibited the superficial secretion phenomenon in macrophages, which has not been generalised to the whole immune system and correspondingly short of conviction. Therefore, more research must be performed to determine hyphae’s unique effect on aspergillosis. This study's main objective was to compare host immune responses against airborne conidia and hyphae in vitro and in vivo, laying a foundation for future in-depth mechanistic research.
Materials and Methods
Mould Preparation
A. fumigatus wild-type strain Af293 (purchased from FGSC, the Fungal Genetics Stock Center, University of Missouri, USA) was cultured in rich media YAG containing 0.5% (w/v) yeast extract, 2% (w/v) glucose and 0.1% (v/v) trace elements at 37 °C. Conidia and hyphal fragments were cultured and harvested as described in a previous study [23].
Fungal Quantification
To better compare conidia and hyphal fragments, the ratio of surface area to volume was adopted as the quantitative parameter. At first, the cross-sectional areas of conidia and hyphae were measured, respectively, by Image J software after taking fluorescent pictures with Calcofluor white (Sigma, USA) staining. Further, a hemacytometer was used to keep an equal volume of each picture.
The cross-sectional areas of conidia and hyphal fragments measured by Image J software were, respectively, named \({\mathrm{S}}_{1}\) and \({\mathrm{S}}_{2}\). The total surface areas of conidia and hyphae were named \({\mathrm{S}}_{c}\) and \({\mathrm{S}}_{h}\), respectively. Then, the radius of the conidia was set to be \(\mathrm{r}\), while the radius and length of the hyphal fragments were set to be \(\mathrm{R}\) and \(\mathrm{L}\), respectively. The numbers of conidia and hyphal fragments were \(\mathrm{n}\) and \(\mathrm{N}\), respectively, while that of the original hyphae was \({\mathrm{N}}_{2}\) \({(\mathrm{N}}_{2}\ll \mathrm{N}).\) The process of whole conversion was as follows:
The final concentration of fungal suspension was calculated and standardised by the multiplicity of infection (MOI) ratios of conidia to cells. Conidia and hyphae were inactivated at 95 °C in the water bath for 30 min.
RAW264.7 Macrophages Culture and Stimulation
RAW264.7 macrophages were maintained in humidified air at 37 °C, 5% CO2 in DMEM supplemented with 10% fetal bovine serum (FBS; Gibco, USA). Macrophages were stimulated with gradient concentrations of conidia or hyphal fragments. They were then incubated with live conidia (LC) and live hyphal fragments (LH) for 6 h and with heat-inactivated conidia (HIC) and hyphal fragments (HIH) for 24 h (MOI ranging from 0.1 to 100). Supernatants were collected and stored at −80 °C until examination.
CCK-8 Assay
Cell Counting Kit-8 (CCK-8) assay was performed to measure cell viability. RAW264.7 macrophages were plated in 96-well flat-bottom culture plates at 5*103 cells/well density and incubated overnight. Thereafter, cells were stimulated with gradient live organisms for 6 h and inactivated ones for 24 h. After washing with PBS (Hyclone, USA) twice, it was added with fresh medium and 10 μL CCK-8 reagent in each well for 1-h incubation. Finally, the absorbance value was measured at the wavelength of 450 nm using an ELISA plate reader. The viability index of the cells was determined using the following equation:
control group: group without fungal stimuli, blank group: group without cells and fungal stimuli.
ROS Assay
RAW264.7 macrophages were washed and collected with PBS after fungal stimulation (MOI = 10), as mentioned above. Macrophages were then incubated with dihydroethidium (Beyotime, China) at 37 °C for 30 min. Next, fluorescence intensity was detected at 488 nm (excitation wavelength) and 535 nm (emission wavelength) through flow cytometry (Invitrogen, USA).
PBMC Isolation and Stimulation
With written informed consent upon approval of the ethics committee of Jinling Hospital, 20 mL EDTA blood samples were collected from healthy volunteers. Peripheral blood mononuclear cells (PBMCs) isolation was performed by Ficoll-Paque density-gradient centrifugation. After washing and centrifuging, PBMCs were resuspended in an RPMI 1640 (Gibco, USA) culture medium containing 10% FBS and then stimulated with live or heat-inactivated fungi for different durations. PMA (25 ng/mL) + ionomycin (1 μg/mL) stimulation for 6 h was used as a positive control. Supernatants were collected and stored at -80 °C until cytokine assays were performed. Cells from different donors were used for the biological replicates.
T Cell Subset Analysis
Live conidia and hyphal fragments stimulated PBMCs for 6 h and inactivated ones for 24 h. At the last 6 h of incubation, PMA (25 ng/mL), ionomycin (1 μg/mL) and brefeldin A (10 μg/mL) were added. Next, cells were collected and washed with PBS, followed by extracellular staining with anti-human CD3 PE-Cy7, CD4 APC-Cy7 and CD8 PE-Cy5.5 for 15 min. After that, PBMCs were centrifuged and resuspended in Fixation/permeabilisation reagents (Biogems, USA) for 30 min to perform the subsequent intracellular cytokine staining with anti-human IL-4 R-PE, IL-17a APC and IFN-γ Alexa Flour™ 700 (all antibodies: BioLegend, USA). Cell populations were analysed through flow cytometry. T lymphocyte was defined as CD3+ cells, and T cell subsets were defined as Th1: CD4+ IFN-γ+, Th2: CD4+ IL-4+ and Th17: CD4+ IL-17a +. CD4 (%) = CD3+CD4+/CD3+, CD8 (%) = CD3+CD8+/CD3+, Th1 (%) = CD3+CD4+IFN-γ+/CD3+CD4+, Th2 (%) = CD3+CD4+IL-4+/ CD3+CD4+, Th17 (%) = CD3+CD4+IL-17a+/ CD3+CD4+
Mice and Infection
Forty-eight male C57BL/6 mice (aged 6–8 weeks) were purchased from Qinglongshan animal centre (Jiangsu, China) and acclimatised. Mice were randomly divided into three groups, with 16 mice in each group. Ten mice from each group were used for survival observation, three for histopathology examination and three for RNA-seq. Chloral hydrate (350 mg/kg) was intraperitoneally injected into mice for anaesthesia. The three groups were challenged intratracheally with 50 μL conidia suspension (109/mL, conidia group), hyphal fragments suspension (hyphae group, equal concentration of conidia), or normal saline (control group), respectively. The mice were observed every 12 h for 10 days, and their lungs were immediately removed after anaesthesia or death. Lungs from three mice in each group were harvested, frozen and ground for RNA extraction 24 h after fungal inoculation, and the RNA products were stored for RNA-seq after quality testing. On the fourth day of the intervention, lungs from another three mice in each group were removed and fixed with 10% (v/v) formalin for periodic acid Schiff (PAS) staining. All animal experiments were carried out per the Institutional Animal Care and Use Committee of the Jinling Hospital.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
According to the instructions, total RNA was isolated from RAW264.7 cells or murine lung tissues using TRIzol reagent (Invitrogen, USA). OD260/280 ratio of RNA between 1.8–2.0 tested by spectrophotometer was qualified for the subsequent experiment. A total of 1 μg RNA was reverse-transcribed into a cDNA template, and cDNA products were diluted 1:5 with diethylpyrocarbonate-treated water without DNAse treatment. Quantitative real-time polymerase chain reaction (20 μL reaction volume) was performed in triplicate using forward and reverse primers (0.8 μL and 10 μmol/L, respectively), a 2 μL cDNA aliquot and 10 μL TB Green® Premix Ex TaqTM II (Takara, China). Housekeeping gene β-actin was used for the standardisation of mRNA, and relative expression analysis was performed using the 2−ΔΔCt method. All primers were validated by NCBI Primer-Blast (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to ensure the specificity and the details of the primers are shown in Table S1.
RNA-Seq
Total RNA was isolated from murine lung tissues harvested 24 h after fungal inoculation using the TRIzol reagent. mRNA was purified using Oligo(dT)-containing beads and fragmented into small pieces. Then, the products were reverse transcribed to first- and second-strand cDNA. Afterwards, A-Tailing Mix and RNA Index Adapters were added by incubating to end repair. cDNA fragments amplification was performed via PCR, and the products were purified by Ampure XP Beads. Subsequently, quality control was performed on the Agilent Technologies 2100 bioanalyser. The double-stranded PCR products were denatured and circularised, and the synthesised single-strand circle DNA (ssCir DNA) was formatted as the final library. The final library was amplified with phi29 to make a DNA nanoball (DNB) that had more than 300 copies of one molecule. DNBs were loaded into the patterned nanoarray, and 50 single-end base reads were generated on the BGIseq500 platform (BGI-Shenzhen, China). The sequencing data were filtered with SOAPnuke to obtain clean reads and stored in FASTQ format. The Dr. Tom Multi-omics Data mining system (https://biosys.bgi.com) was adopted for subsequent analysis and data mining. Then the obtained clean reads were aligned to the gene set built by BGI (Beijing Genomic Institute in ShenZhen) using software Bowtie2. The gene expression level was calculated by RSEM (v1.3.1), and the heatmap was drawn by pheatmap (v1.0.8). Essentially, DESeq2 (v1.4.5) was applied for differential expression analysis with Qvalue ≤ 0.05. Furthermore, KEGG (https://www.kegg.jp/) enrichment analysis of differentially expressed genes was performed using Phyper (https://en.wikipedia.org/wiki/Hypergeometric_distribution) based on the Hypergeometric test. Qvalue ≤ 0.05 was the significant threshold for the correction of terms and pathways.
ELISA
Supernatants of macrophages and PBMCs were collected after stimulation and centrifuged for enzyme-linked immunosorbent assay (ELISA). Commercially available ELISA kits for mouse TNF-α, IL-6 and human TNF-α, IL-6, IL-1β and IL-4 (Novusbio, USA) were used according to the manufacturers’ instructions.
Data Analysis
The analysis of flow cytometric samples was performed in FlowJo software (TreeStar, USA). Statistical analyses were performed in GraphPad Prism 7 (GraphPad Software Inc., USA). One- or two-way analysis of variance (ANOVA) was used to analyse data sets of CCK-8, qRT-PCR and ELISA, depending on the number of explanatory variables in the experimental design. Results were presented as mean ± standard deviation (SD) of triplicate experiments. Flow cytometry results about T cell populations were analysed by the Friedman test. P < 0.05 was considered statistically significant.
Results
Macrophages RAW264.7 Were Classically Activated by A. fumigatus Conidia and Produced More ROS than Hyphal Stimulation
Cells were stimulated with the A. fumigatus in different stages to elucidate the different responses of immune cells against conidia or hyphae. In vitro, immune cells were exposed to live fungi for 6 h to mimic the early dynamic stage of fungal invasion. Meanwhile, accumulated inflammatory responses were observed when exposed to heat-inactivated ones for 24 h.
Concerning macrophages, inducible nitric oxide synthase (iNOS) mRNA expression after the stimulation of Aspergillus was concentration and fungal state dependent. As shown in Figs. 1a and b, the iNOS mRNA expression is a marker of classical macrophage activation (M1 polarisation), which was increased in conidia-stimulated macrophages rather than hyphae-stimulated ones.
Immune responses of macrophages induced by A. fumigatus conidia and hyphae. a The iNOS mRNA expression of macrophages induced by A. fumigatus conidia and hyphae in different conditions (stimulation time and fungal activity). b The iNOS mRNA expression of macrophages induced by A. fumigatus conidia and hyphae at different concentrations. c Cell viability of macrophages after stimulation by A. fumigatus different at different concentrations. MOI ranging from 1:10 to 100:1. Data represent mean ± standard deviation (n = 3).*P < 0.05 vs. the controls. Live Conidia LC, Live Hyphal fragments LH, Heat-inactivated Conidia HIC, Heat-inactivated Hyphal fragments HIH
In host defence against fungi like alveolar macrophages, reactive oxygen species (ROS) play an essential role. In this study, intracellular superoxide (O2–) was detected by DHE staining. Macrophages stimulated by conidia revealed a higher ROS production than the control and hyphae group (Supporting Information Fig. S1 a & b), which was consistent with the cellular iNOS levels.
Fungal Stimulation Promotes the Proliferation of Macrophages RAW264.7
CCK-8 assay was used to measure the cell viability after fungal stimulation. Both conidia and hyphal fragments stimulation exhibited a concentration-dependent rise in the cell proliferation (P < 0.05), and the live fungus groups indicated a more obvious alteration than the inactivated ones (P < 0.05), though no significant difference was found between the two groups (P > 0.05) (Fig. 1c).
Cytokine Profile of A. fumigatus-Stimulated Macrophages RAW264.7 and PBMCs
The cytokine profile can intuitively reflect inflammatory reactions of macrophages after exposure to different fungal particles. As shown in Fig. S2, TNF-α and IL-6 indicated a concentration-dependent increase in secretion. Additionally, the releases of TNF-α and IL-6 were more remarkable after exposure to hyphal fragments for 6 h than exposure to conidia.
The cytokine profile of PBMCs induced by A. fumigatus conidia and hyphal fragments were detected to explore the host immune response. The human TNF-α, IL-6, IL-4 and IL-1β were directly proportional to stimuli concentration (Fig. 2), consistent with the cytokine profile of macrophages. In most cases, the human TNF-α, IL-6 and IL-4 concentrations of hyphae-stimulated PBMCs were higher than that of conidia-stimulated cells at the same concentration.
Cytokine profile of murine macrophages and PBMCs stimulated by A. fumigatus conidia and hyphae measured by ELISA. a human TNF-α, b human IL-6, c human IL-1β, and d human IL-4. Data represent mean ± standard deviation (n = 3). *P < 0.05 vs. the control group. Live Conidia LC, Live Hyphal fragments LH, Heat-inactivated Conidia HIC, Heat-inactivated Hyphal fragments HIH
Dynamic T Cell Subset Changes Induced by Conidia and Hyphae
Dynamic T cell subset changes were observed after exposure to conidia or hyphae (Fig. 3& Fig. S3). In general, hyphae-stimulated cells showed an obvious elevation of the CD4/CD8 ratio compared with conidia-stimulated ones (P < 0.05). The percentage of CD4+ cells was significantly increased (P < 0.05) and CD8+ slightly decreased compared to that of conidia-stimulated group (P > 0.05). The percentage of Th1 cells (Th1%) increased greatly in both LC and HIC groups (P < 0.05). In contrast, the hyphae group exhibited a higher proportion of Th2 cells than the corresponding conidia group without statistical significance (P > 0.05). Neither group showed any changes in Th17 cell differentiation (P > 0.05).
Survival of Immunocompetent C57BL/6 Mice Following Inhalation of A. fumigatus Conidia or Hyphal Fragments
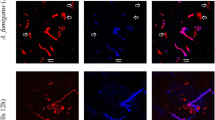
In the murine pulmonary aspergillosis model, the mortality of hyphae-stimulated mice reached 40% (P < 0.05). Meanwhile, the mice stimulated with conidia and normal saline all survived to the end point of the assay (Fig. 4a). Additionally, focal infection of Aspergillus was only observed on the pulmonary histopathology of hyphae-inoculated mice four days post-inoculation (Fig. 4b). The fungal culture of murine lungs further confirmed the aspergillosis.
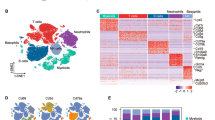
RNA-seq profile of the immunocompetent murine pulmonary aspergillosis model 24 h post-inoculation. a Unique gene expression. b Volcano map of DEGs. c KEGG pathway enrichment bubble chart. Note: The size of the bubble represents the number of genes annotated to the KEGG pathway. The colour represents the enriched significance. Rich Ratio = Term Candidate Gene Number/Term Gene Number. d KEGG pathway classification
The Immune Responses Against Hyphae and Conidia have Similarities and Differences in the Murine Pulmonary Aspergillosis Model
RNA-seq detected a total of 18,984 transcripts in three groups. Only 234 genes were significantly differentially regulated between the control and the conidia groups (|log2FC|≥ 2, Q-value ≤ 0.05). On the contrary, the number of differentially expressed genes (DEGs) between the hyphae and the control groups was up to 1,123, with 633 up-regulated and 490 down-regulated (Fig. 4a). However, 712 significant DEGs (Fig. 4a and b) between the conidia and the hyphae groups were analysed. KEGG pathway enrichment analysis showed a series of immunologically relevant pathways that were involved, including the IL-17 signalling pathway, TNF signalling pathway, Toll-like receptor signalling pathway, NOD-like receptor (NLR) signalling pathway, and others (Fig. 4c). Additionally, KEGG pathway classification revealed that signal transduction and immune system were common DEG-intensive categories (Fig. 4d).
Hyphae could induce more intensive inflammatory responses in murine lungs than conidia. The expression of genes associated with immunologically relevant pathways mentioned above was activated more strongly in the hyphae group than in the conidia group. We subsequently identified a series of differentially expressed cytokines as well as essential receptors, including TLRs, Clecs and NLRs, to gain insight into immune responses against Aspergillus (Fig. S5).
Thereafter, to better understand immune signalling patterns amongst the murine model, T cell differentiation pathway-associated genes were analysed by qRT-PCR. As shown in Fig. 5, both groups tended to activate the Th17 cell differentiation pathway after intratracheal mould injection. The expressions of IL-17a, IL-17f, and stat3 were up-regulated. Besides, expressions of IL-17 signalling-associated chemokines CXCL1, CXCL2, CXCL5, CXCL10 and CCL2 were likewise significantly increased. Regarding Th1 and Th2 cell differentiation, RNA-seq revealed that Th2 cell differentiation tended to get activated, and Th1 differentiation was relatively silenced in the hyphae group. The genes related to Th2 cell differentiation, including IL-13 and IL-4Ra, were significantly up-regulated, while the key genes in the Th1 cell differentiation pathway, such as stat1 and T-bet, were down-regulated. However, in the conidia group, the gene expressions of Th1 and Th2 cell differentiation pathways were not significantly altered except for the mild elevation of IL-4Ra.
T cell differentiation and inflammation in the murine pulmonary aspergillosis model. a The fold change of multiple immunological genes detected by RNA-seq and b The fold change of multiple immunological genes detected by qRT-PCR (b and c). Data represent mean ± standard deviation (n = 3)., *P < 0.05 vs. the control group
Discussion
A. fumigatus, the causative agent of a series of human infections, is a kind of ubiquitous fungus in the environment. IPA is the most dangerous and fatal type of aspergillosis. The current worldwide prevalence of COVID-19 leads to the elevated risk of IPA due to dysfunction of host immunity and disruption of normal lung structure. Therefore, more efforts should be made to clarify the interaction between A. fumigatus and host immunity.
This research is based on Zhang’s experimental results, which were published in 2020 [21]. As mentioned above, Zhang found A. fumigatus hyphae rather than conidia could successfully build a pulmonary aspergillosis model in immunocompetent mice. Previous studies generally chose swollen or germinated conidia to replace mature hyphae for experiments, which are convenient to quantify and easy to compare with the conidia group. However, it cannot well represent the natural state of mature hyphal infection. To simulate the inhalable hyphae infecting the respiratory tract naturally, we obtained properly sized hyphal fragments by grinding the mature hyphae rather than shortening the incubation time to get budding ones. This inevitably brought great difficulties in quantification. In the past reports, several methods for hyphal quantification showed advantages and disadvantages. Direct counting with a hemacytometer can be simple and fast to quantify newly germinated hyphae with nearly similar sizes, just like the common quantification of conidia [24]. However, this method cannot quantify mature hyphae or hyphal fragments because hyphal particles differ in size and growth activity, and thus, the systematic error of the process is difficult to measure. Colony-forming units have the same limitation as direct counting with a hemocytometer. Gravimetry has ever been adopted for fungal quantification, but the complicated operations of sample drying by freezing or heating, which often inevitably destroy the fungal activities limit its usage [25]. Meanwhile, the dried samples are easier to spread in the environment, leading to biological contamination for health risks. Additionally, the sample required for gravimetry is extremely large, causing great inconvenience for the prevalence of this method. Thus, this study made some improvements to Zhang’s protocol with a novel method for the quantification of mycelia. We first adopted the ratio of surface area to volume for mycelial and conidia quantification, which can be used to quantify other types of filamentous fungus. Admittedly, the fungal surface with different PAMPs is the first-line structure of Aspergillus to directly contact the host. Hence, the surface area is a good parameter to study the immune interaction between fungus and host.
Another critical light spot of this study is the successful establishment of in vitro and in vivo immunocompetent infection model of mature A. fumigatus hyphae. Innate and adaptive responses against Aspergillus conidia and hyphae were studied by co-incubating different immune cells and Aspergillus in vitro. Unlike most immunocompromised models, we constructed a murine immunocompetent aspergillosis model with hyphal fragments. Meanwhile, RNA-seq analysis was first performed to analyse the immune responses against respirable mature hyphal fragments in vivo.
In the study, we confirmed that hyphae of A. fumigatus could induce stronger host immune responses than conidia, as hyphae-stimulated cells secreted more inflammatory factors. The immunological differences of cytokine secretions and T cell differentiation in inactivated conidia and hyphae were larger than in active ones, probably attributed to extended incubation time. In the murine model, we again validated that immunologically relevant genes and pathways were activated more significantly in hyphae than conidia group. Consistently, faster germination of conidia has been reported to drive to greater lung damage and inflammation, which means stronger invasiveness and lethality of hyphae [26]. Meanwhile, germinating conidia could induce stronger TNF-α, IL-1, IL-1β, IL-6 and MIP-2 secretion by alveolar macrophages, which is correlated to the levels of surface-exposed beta-glucans via dendritic cell-associated C-type lectin-1 (dectin-1) receptor [27].
As for immune cells, macrophages are known to be a decisive part of innate immunity to resist the infection from A. fumigatus. Both iNOS and ROS levels are important markers of the functions of macrophages, which were significantly higher in conidia-stimulated cells than that in hyphae-stimulated cells. iNOS was mainly induced in M1 macrophages, and nitric oxide produced by iNOS would be scavenged to generate ROS, leading to extra ROS production [28]. Macrophage polarisation is an essential symbol to distinguish its functions. M1 macrophages are generally found to be polarised by lipopolysaccharide- or Th1-associated cytokines [29]. As feedback, M1 macrophages can direct T cells towards Th1 through IL-12 and antigen presentation. Subsequent flow cytometry analysis indicated that the conidia-stimulated PBMCs expressed higher levels of IFN-γ (Th1 signalling), which was consistent with the macrophage polarisation state.
T lymphocytes are an indispensable part of adaptive immunity. Regarding T cell subset change, the percentage of CD8+ T cells tended to get down after hyphal stimulation, which usually indicated a poor prognosis in IPA patients [30]. Therefore, we speculate that the decrease of CD8+ T cell proportion in severe IPA patients is likely to be owing to the overgrowth of invasive hyphae at the late stage of infection, which can hardly be eliminated by cytotoxic CD8+ T cells. Concerning CD4+ T cells, cell differentiation clearly reflected its functions. Th1 signalling drives to fight viruses, bacteria and other intracellular infections, such as A. fumigatus conidia while Th2 signalling drives to defend extracellular organisms. Th1/Th2 is in a dynamic balance, and over-activation of either pathway can down-regulate the other [31]. Besides, Th17 also plays an important role in host defence against micro-organisms, which is associated with neutrophil migration and increased inflammation [32]. However, there are contradictions and unities in the in vitro and in vivo experimental results. In the murine model, the expression of Th1 and Th2 cell differentiation-related genes were not significantly altered after inoculation of conidia, which was inconsistent with the result of an experiment in vitro. We speculate that the phenomenon might be ascribed to the effective anatomical elimination of conidia in an immunocompetent host. Only a small portion of escaped conidia can challenge the subsequent immune system [16, 33]. Besides, Th1 and Th17 cells showed antagonistic action against each other and activation of IL-17 signalling would suppress Th1 cell differentiation [34]. Overall, it concludes that conidia stimulation promotes Th1 cell differentiation, while hyphal stimulation leads to the opposite result and activates Th2 cell differentiation. Furthermore, both conidia and hyphae could activate the IL-17 signalling pathway, which may make a dual-directional regulation related to antifungal immune resistance [16].
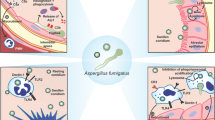
It is a sophisticated orchestration for innate and adaptive immunity to protect from fungal invasion. The host immune responses induced by A. fumigatus conidia and hyphae are distinctly different, mainly attributed to the different PAMPs on the surface of hyphae and conidia. Various PAMPs can be recognised by specific PRRs of immunocytes, activating downstream intracellular signal transduction pathways and different inflammatory responses [15]. For instance, hyphae and conidia of A. fumigatus can activate TLR2-related cytokine synthesis, but TLR4-related pathways can only be induced by conidia [35]. Additionally, both dectin-1 and dectin-2 receptors can recognise swollen conidia and hyphae to induce host immune responses, but neither can recognise resting conidia [36, 37]. Besides, hyphae can trigger the NLR pyrin domain-containing 3 inflammasome assembly, which cannot be effectively activated by resting conidia [38]. As for the explanation of the initial phenomenon in the study, it is a paradox that hyphal fragments successfully constructed the aspergillosis model rather than conidia despite that those hyphae induced more potent inflammation. In other words, the host immune responses induced by conidia played a better protective role. As reported, various effective ways were reported to eliminate conidia by host immunity, from anatomical barriers to the synergistic action of multiple immune cells. However, the hyphae are too large to get internalised and can hardly be cleared. Even neutrophil extracellular traps (NETs), an effective form of neutrophil-mediated antimicrobial defence, can mostly inhibit hyphal growth instead of killing them [16, 39]. Moreover, hyphal invasion tends to activate Th2 cell differentiation, which is known for its suppressive function in immunity and usually leads to unfavourable outcomes [40].
However, there are still some limitations of the study. Firstly, the live conidia would be swelling and germinating during the 6 h incubation with tremendous changes in PAMPs. However, if the incubation time was cut down, the immune responses against fungus would become indistinct or non-significant. For compensation, Aspergillus conidia and hyphae were inactivated to stop the growth, and the incubation time was extended to 24 h to explore the cumulative immune effects. However, the inactivated fungus could only keep partial immune characteristics from the live ones. The second limitation is the time point of lung tissue harvest for RNA-seq analysis in the murine model. In fact, 24 h post-inoculation was chosen based on Wang’s work that the early immune cells infiltration and differentiation in the bronchi and lung tissues can be clearly scanned 24 h after fungal inoculation intratracheally [41]. However, a single time point is insufficient to elucidate the progressive process of the early immune response against Aspergillus and setting more time points can make the study protocol more logical and convictive. Thirdly, the study did not include the comparison of neutrophils defending against Aspergillus conidia and hyphae. Germinating rather than resting conidia has been reported could recruit neutrophils to the airways [42]. Meanwhile, two distinct neutrophil-mediated killing mechanisms of A. fumigatus conidia and hyphae were characterised depending on CR3 binding or IgG binding, respectively [43]. However, the function and mechanism of NETs against Aspergillus need more study, and NETs-related Aspergillus capture experiments will be carried out in subsequent studies.
Conclusion
To summarise, the hyphae state is a neglected pathogenetic form of aspergillosis. We conducted a preliminary study about immune response against hyphae in vivo and in vitro and confirmed the different host defence against hyphae and conidia. However, the underlying mechanism of the immunological differences remains unclear and future research is needed.
Data Availability
The data of RNA-seq was submitted in GEO data sets (GSE183979). The data-sets used and analysed during the current study are available from the corresponding author upon reasonable request.
References
Gao Y, Soubani A (2020) Advances in the diagnosis and management of pulmonary aspergillosis. Adv Respir Med 87:231–243
Kosmidis C, Denning DW (2014) The clinical spectrum of pulmonary aspergillosis. Thorax 70:270–277
Kanj A, Abdallah N, Soubani AO (2018) The spectrum of pulmonary aspergillosis. Resp Med 141:121–131
Chabi ML, Goracci A, Roche N, Paugam A, Lupo A, Revel MP (2015) Pulmonary aspergillosis. Diagn Interv Imaging 96:435–442
Tarka P, Nitsch-Osuch A, Gorynski P, Tyszko P, Bogdan M, Kanecki K (2019) Epidemiology of Pulmonary Aspergillosis in Hospitalized Patients in Poland During 2009–2016. Advances in Pulmonary Medicine: Research and Innovations. Springer, Cham, pp 73–80
Tavakoli M, Charati JY, Hedayati MT, Moosazadeh M, Badiee P, Seyedmousavi S, Denning DW (2019) National trends in incidence, prevalence and disability-adjusted life years of invasive aspergillosis in Iran: a systematic review and meta-analysis. Expert Rev Resp Med 13:1121–1134
Bassetti M, Garnacho-Montero J, Calandra T, Kullberg B, Dimopoulos G, Azoulay E, Chakrabarti A, Kett D, Leon C, Ostrosky-Zeichner L et al (2017) Intensive care medicine research agenda on invasive fungal infection in critically ill patients. Intens Care Med 43:1225–1238
Shorr AF, Spalding JR, Harrington R, Nathanson BH, Zilberberg MD (2018) Epidemiology and outcomes of hospitalizations with invasive aspergillosis in the United States, 2009–2013. Clin Infect Dis 67:727–735
Wu Z, McGoogan JM (2020) Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese center for disease control and prevention. JAMA 323(13):1239–1242
Bartoletti M, Pascale R, Cricca M, Rinaldi M, Maccaro A, Bussini L, Fornaro G, Tonetti T, Pizzilli G, Francalanci E et al (2020) Epidemiology of invasive pulmonary aspergillosis among COVID-19 intubated patients: a prospective study. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa1065
Nasir N, Farooqi J, Mahmood SF, Jabeen K (2020) COVID-19-associated pulmonary aspergillosis (CAPA) in patients admitted with severe COVID-19 pneumonia: an observational study from Pakistan. Mycoses 63:766–770
Salehi M, Ahmadikia K, Badali H, Khodavaisy S (2020) Opportunistic fungal infections in the epidemic area of COVID-19: a clinical and diagnostic perspective from Iran. Mycopathologia 185:607–611
Lai CC, Yu WL (2020) COVID-19 associated with pulmonary aspergillosis: a literature review. J Microbiol Immunol Infect 54:46–53
Borman AM, Palmer MD, Fraser M, Patterson Z, Mann C, Oliver D, Linton CJ, Gough M, Brown P, Dzietczyk A et al (2020) COVID-19-associated invasive aspergillosis: data from the UK national mycology reference laboratory. J Clin Microbiol 59:1
van de Veerdonk FL, Gresnigt MS, Romani L, Netea MG, Latge JP (2017) Aspergillus fumigatus morphology and dynamic host interactions. Nat Rev Microbiol 15:661–674
Margalit A, Kavanagh K (2015) The innate immune response to Aspergillus fumigatus at the alveolar surface. FEMS Microbiol Rev 39:670–687
Latgé JP, Beauvais A, Chamilos G (2017) The cell wall of the human fungal pathogen Aspergillus fumigatus: biosynthesis, organization, immune response, and virulence. Annu Rev Microbiol 71:99–116
Afanou KA, Straumfors A, Skogstad A, Skaar I, Hjeljord L, Skare O, Green BJ, Tronsmo A, Eduard W (2015) Profile and morphology of fungal aerosols characterized by field emission scanning electron microscopy (FESEM). Aerosol Sci Technol 49:423–435
Afanou KA, Straumfors A, Skogstad A, Nilsen T, Synnes O, Skaar I, Hjeljord L, Tronsmo A, Green BJ, Eduard W (2014) Submicronic fungal bioaerosols: high-resolution microscopic characterization and quantification. Appl Environ Microbiol 80:7122–7130
Dufresne PJ, Moonjely SS, Ozaki K, Tremblay C, Laverdière M, Dufresne SF (2017) High frequency of pathogenic Aspergillusspecies among nonsporulating moulds from respiratory tract samples. Med Mycol 55:233–236
Zhang Z, Jiang Y, Chen J, Chen P, Kong Q, Lu L, Sang H (2020) In vitro and in vivo characterization of two nonsporulating Aspergillus fumigatus clinical isolates from immunocompetent patients. Med Mycol 58:543–551
Øya E, Solhaug A, Bølling AK, Øvstebø R, Steensen TB, Afanou AKJ, Holme JA (2019) Pro-inflammatory responses induced by A. fumigatus and A. versicolor in various human macrophage models. J Toxicol Environ Health, Part A 82:483–501
Zhang C, Kong Q, Cai Z, Liu F, Chen P, Song J, Lu L, Sang H (2015) The newly nonsporulated characterization of an Aspergillus fumigatus isolate from an immunocompetent patient and its clinic indication. Fungal Genet and Biol 81:250–260
Chryssanthou E, Loebig A, Sjolin J (2008) Post-antifungal effect of amphotericin B and voriconazole against germinated Aspergillus fumigatus conidia. J Antimicrob Chemother 61:1309–1311
Oya E, Afanou AKJ, Malla N, Uhlig S, Rolen E, Skaar I, Straumfors A, Winberg JO, Bang BE, Schwarze PE et al (2018) Characterization and pro-inflammatory responses of spore and hyphae samples from various mold species. Indoor Air 28:28–39
Caffrey-Carr AK, Kowalski CH, Beattie SR, Blaseg NA, Upshaw CR, Thammahong A, Lust HE, Tang YW, Hohl TM, Cramer RA et al (2017) Interleukin 1alpha is critical for resistance against highly virulent aspergillus fumigatus isolates. Infect Immun 85(12):e00661-e717
Steele C, Rapaka RR, Metz A, Pop SM, Williams DL, Gordon S, Kolls JK, Brown GD (2005) The beta-glucan receptor dectin-1 recognizes specific morphologies of Aspergillus fumigatus. PLoS Pathog 1:e42
Yan M, Hou M, Liu J, Zhang S, Liu B, Wu X, Liu G (2017) Regulation of iNOS-Derived ROS Generation by HSP90 and Cav-1 in porcine reproductive and respiratory syndrome virus-infected swine lung injury. Inflammation 40:1236–1244
Mills CD (2015) Anatomy of a discovery: m1 and m2 macrophages. Front Immunol 6:212
Cui N, Wang H, Long Y, Liu D (2013) CD8+ T-cell counts: an early predictor of risk and mortality in critically ill immunocompromised patients with invasive pulmonary aspergillosis. Crit Care 17(4):R157
Kidd P (2003) Th1/Th2 balance: the hypothesis, its limitations, and implications for health and disease. Altern Med Rev 8(3):223–246
Wilson NJ, Boniface K, Chan JR, McKenzie BS, Blumenschein WM, Mattson JD, Basham B, Smith K, Chen T, Morel F et al (2007) Development, cytokine profile and function of human interleukin 17-producing helper T cells. Nat Immunol 8:950–957
Latge JP, Chamilos G (2019) Aspergillus fumigatus and Aspergillosis in 2019. Clin Microbiol Rev 33(1):e00140-e218
Zelante T, De Luca A, Bonifazi P, Montagnoli C, Bozza S, Moretti S, Belladonna ML, Vacca C, Conte C, Mosci P et al (2007) IL-23 and the Th17 pathway promote inflammation and impair antifungal immune resistance. Eur J Immunol 37:2695–2706
Netea MG, Warris A, Van der Meer JWM, Fenton MJ, Verver-Janssen TJG, Jacobs LEH, Andresen T, Verweij PE, Kullberg BJ (2003) Aspergillus fumigatus evades immune recognition during germination through loss of toll-like receptor-4-mediated signal transduction. J Infect Dis 188(2):320–326
Gersuk GM, Underhill DM, Zhu L, Marr KA (2006) Dectin-1 and TLRs permit macrophages to distinguish between different Aspergillus fumigatus cellular states. J Immunol 176:3717–3724
Sun H, Xu XY, Tian XL, Shao HT, Wu XD, Wang Q, Su X, Shi Y (2014) Activation of NF-kappaB and respiratory burst following Aspergillus fumigatus stimulation of macrophages. Immunobiology 219:25–36
Said-Sadier N, Padilla E, Langsley G, Ojcius DM (2010) Aspergillus fumigatus stimulates the NLRP3 inflammasome through a pathway requiring ROS production and the Syk tyrosine kinase. PLoS ONE 5:e10008
McCormick A, Heesemann L, Wagener J, Marcos V, Hartl D, Loeffler J, Heesemann J, Ebel F (2010) NETs formed by human neutrophils inhibit growth of the pathogenic mold Aspergillus fumigatus. Microbes infect 12(12–13):928–936
Hebart H, Bollinger C, Fisch P, Sarfati J, Meisner C, Baur M, Loeffler J, Monod M, Latgé J, Einsele H (2002) Analysis of T-cell responses to Aspergillus fumigatus antigens in healthy individuals and patients with hematologic malignancies. Blood 100(13):4521–4528
Wang F, Zhang C, Jiang Y, Kou C, Kong Q, Long N, Lu L, Sang H (2017) Innate and adaptive immune response to chronic pulmonary infection of hyphae of Aspergillus fumigatus in a new murine model. J Med Microbiol 66:1400–1408
Hohl TM, Van Epps HL, Rivera A, Morgan LA, Chen PL, Feldmesser M, Pamer EG (2005) Aspergillus fumigatus triggers inflammatory responses by stage-specific beta-glucan display. PLoS Pathog 1:e30
Gazendam RP, van Hamme JL, Tool ATJ, Hoogenboezem M, van den Berg JM, Prins JM, Vitkov L, van de Veerdonk FL, van den Berg TK, Roos D et al (2016) Human neutrophils use different mechanisms to kill Aspergillus fumigatus conidia and hyphae: evidence from phagocyte defects. J Immunol 196(3):1272–1283
Acknowledgements
We thank Dr Weida Liu and his team in the Department of Medical Mycology, Institute of Dermatology, Chinese Academy of Medical Science and Peking Union Medical College, for their help with the experiment.
Funding
This work was financially supported by the National Natural Science Foundation of China (NSFC) [Grant Number 81871630 and 81330035].
Author information
Authors and Affiliations
Contributions
YL, HS and YS contributed to the study conception and design. YL, LD and JX performed the experiments; YL, FL and QK analysed and interpreted results; YL wrote the first draft of the manuscript; FL reviewed; and all authors contributed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no competing interest to declare.
Ethical approval
The study was approved by the ethics committee of Jinling Hospital (2015NJKY-035–02) and the Animal Care and Use Committee of Nanjing Normal University (20200703).
Informed Consent
All volunteers the blood samples obtained from knew their rights and gave informed consents.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
284_2022_3102_MOESM1_ESM.tif
Supplementary file1 The conidia of fumigatus induced more ROS production in macrophages than hyphae. RAW264.7 cells were stimulated by a living fungi and b heat-inactivated fungi (TIF 3007 KB)
284_2022_3102_MOESM2_ESM.tif
Supplementary file2 Cytokine profile of murine macrophages stimulated by A. fumigatus conidia and hyphae measured by ELISA. a murine TNF-α, b murine IL-6, Data represent mean ± standard deviation (n = 3). *P < 0.05 vs. the control group. Live Conidia LC, Live Hyphal fragments LH, Heat-inactivated Conidia HIC, Heat-inactivated Hyphal fragments HIH (TIF 1419 KB)
284_2022_3102_MOESM3_ESM.tif
Supplementary file3 T cell expression of human PBMCs after stimulation by A. fumigatus conidia and hyphae. a CD4, b CD8. Each geometric symbol represents a sponsor (n=4). Live Conidia LC, Live Hyphal fragments LH, Heat-inactivated Conidia HIC, Heat-inactivated Hyphal fragments HIH (TIF 792 KB)
284_2022_3102_MOESM4_ESM.tif
Supplementary file4 a The Kaplan–Meier survival curves of mice inoculated with normal saline, conidia, and hyphal fragments. b Histopathology of lung tissues 4 days post-inoculation with PAS staining. The difference of survival rate between hyphae group and conidia or control group reaches statistical significance (n = 10, P < 0.05). The histopathology results of lung tissue from the control and the conidia-treated groups were basically normal, and no evidence of fungal infection was found in the sections. In contrast, in the lung tissue from the hyphae-treated group, Aspergillus hyphae were positive for PAS staining. (TIF 31922 KB)
284_2022_3102_MOESM5_ESM.tif
Supplementary file5 Profile of TLR, Clec, and NLR gene expression. Colour indicates the level of log2(FPKM + 1). FPKM: Fragments per Kilobase of exon per Million of mapped reads (TIF 7079 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luo, Y., Liu, F., Deng, L. et al. Innate and Adaptive Immune Responses Induced by Aspergillus fumigatus Conidia and Hyphae. Curr Microbiol 80, 28 (2023). https://doi.org/10.1007/s00284-022-03102-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03102-1