Abstract
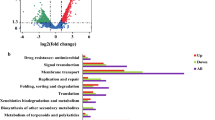
The application of sulfate-reducing bacteria (SRB) shows great potential in the anaerobic biological treatment of acid mine wastewater; therefore, it has attracted much attention. The low pH in acidic wastewater affects the growth and reducing power of SRB. To uncover the mechanism underlying the reduction efficiency of SRB under acidic conditions, in this study, transcriptomic analysis was performed with Desulfovibrio vulgaris ATCC 7757 under three different pH conditions (pH 4.0, 5.5 and 7.0) and in the initial inoculation, logarithmic growth and plateau phases. Our results showed that ATCC 7757 still had biological activity at pH 4.0 and exhibited gene expression patterns at pH 4.0 that were different from those at pH 5.5 and pH 7. Importantly, the gene expression pattern was similar between pH 5.5 and pH 7. Transcriptomic analysis identified differentially expressed genes that affected the growth of ATCC 7757 under pH 7.0 at 22 h compared to 15 h; 196 of these genes were upregulated and 575 were downregulated. These differentially expressed genes were mainly enriched in genetic information processing and metabolism. Additionally, we identified 57 candidate genes associated with low-pH tolerance. Adaptation to low pH was reflected by an increase in the expression of genes involved in cell membrane structure and proton transport. The expression of genes involved in the reduction process decreased, including the genes DVU0499 and sat, which encode proteins that affect the sulfate reduction process. Both gene activities were validated by qPCR. Our results will contribute to further promoting the reducing power of SRB in acid mine wastewater and the development of successful bioremediation strategies.




Similar content being viewed by others
References
Klein R, Tischler JS, Muhling M, Schlomann M (2014) Bioremediation of mine water. Adv Biochem Eng Biotechnol 141:109–172. https://doi.org/10.1007/10_2013_265
Tabak HH, Scharp R, Burckle J, Kawahara FK, Govind R (2003) Advances in biotreatment of acid mine drainage and biorecovery of metals: 1. Metal precipitation for recovery and recycle. Biodegradation 14(6):423–436. https://doi.org/10.1023/a:1027332902740
Karlsson K, Viklander M, Scholes L, Revitt M (2010) Heavy metal concentrations and toxicity in water and sediment from stormwater ponds and sedimentation tanks. J Hazard Mater 178(1–3):612–618. https://doi.org/10.1016/j.jhazmat.2010.01.129
Macías F, Caraballo MA, Rötting TS, Pérez-López R, Nieto JM, Ayora C (2012) From highly polluted Zn-rich acid mine drainage to non-metallic waters: implementation of a multi-step alkaline passive treatment system to remediate metal pollution. Sci Total Environ 433:323–330. https://doi.org/10.1016/j.scitotenv.2012.06.084
Chen A, Lin C, Lu W, Wu Y, Ma Y, Li J, Zhu L (2007) Well water contaminated by acidic mine water from the Dabaoshan Mine South China: chemistry and toxicity. Chemosphere 70(2):248–255. https://doi.org/10.1016/j.chemosphere.2007.06.041
Cozzolino D, Chandra S, Roberts J, Power A, Rajapaksha P, Ball N, Gordon R, Chapman J (2018) There is gold in them hills: predicting potential acid mine drainage events through the use of chemometrics. Sci Total Environ 619–620:1464–1472. https://doi.org/10.1016/j.scitotenv.2017.11.063
Neamtiu IA, Al-Abed SR, McKernan JL, Baciu CL, Gurzau ES, Pogacean AO, Bessler SM (2017) Metal contamination in environmental media in residential areas around Romanian mining sites. Rev Environ Health 32(1–2):215–220. https://doi.org/10.1515/reveh-2016-0033
Kefeni KK, Msagati TAM, Mamba BB (2017) Acid mine drainage: prevention, treatment options, and resource recovery: a review. J Clean Prod 151:475–493. https://doi.org/10.1016/j.jclepro.2017.03.082
Hallberg KB, Johnson DB (2005) Microbiology of a wetland ecosystem constructed to remediate mine drainage from a heavy metal mine. Sci Total Environ 338(1–2):53–66. https://doi.org/10.1016/j.scitotenv.2004.09.005
Diao Z, Shi T, Wang S, Huang X, Zhang T, Tang Y, Zhang X, Qiu R (2013) Silane-based coatings on the pyrite for remediation of acid mine drainage. Water Res 47(13):4391–4402. https://doi.org/10.1016/j.watres.2013.05.006
Dev S, Roy S, Bhattacharya J (2017) Optimization of the operation of packed bed bioreactor to improve the sulfate and metal removal from acid mine drainage. J Environ Manag 200:135–144. https://doi.org/10.1016/j.jenvman.2017.04.102
Sheoran AS, Sheoran V, Choudhary RP (2010) Bioremediation of acid-rock drainage by sulphate-reducing prokaryotes: a review. Miner Eng 23(14):1073–1100. https://doi.org/10.1016/j.mineng.2010.07.001
van den Brand TPH, Roest K, Chen GH, Brdjanovic D, van Loosdrecht MCM (2015) Potential for beneficial application of sulfate reducing bacteria in sulfate containing domestic wastewater treatment. World J Microbiol Biotechnol 31(11):1675–1681. https://doi.org/10.1007/s11274-015-1935-x
Knoblauch C, Jorgensen BB (1999) Effect of temperature on sulphate reduction, growth rate and growth yield in five psychrophilic sulphate-reducing bacteria from Arctic sediments. Environ Microbiol 1(5):457–467. https://doi.org/10.1046/j.1462-2920.1999.00061.x
Utgikar VP, Harmon SM, Chaudhary N, Tabak HH, Govind R, Haines JR (2002) Inhibition of sulfate-reducing bacteria by metal sulfide formation in bioremediation of acid mine drainage. Environ Toxicol 17(1):40–48. https://doi.org/10.1002/tox.10031
Reis MAM, Almeida JS, Lemos PC, Carrondo MJT (1992) Effect of hydrogen sulfide on growth of sulfate reducing bacteria. Biotechnol Bioeng 40(5):593–600. https://doi.org/10.1002/bit.260400506
Costa V, Angelini C, De Feis I, Ciccodicola A (2010) Uncovering the complexity of transcriptomes with RNA-Seq. J Biomed Biotechnol 2010:1–19. https://doi.org/10.1155/2010/853916
Plugge CM, Scholten JCM, Culley DE, Nie L, Brockman FJ, Zhang W (2010) Global transcriptomics analysis of the Desulfovibrio vulgaris change from syntrophic growth with Methanosarcina barkeri to sulfidogenic metabolism. Microbiology 156(9):2746–2756. https://doi.org/10.1099/mic.0.038539-0
He Z, Zhou A, Baidoo E, He Q, Joachimiak MP, Benke P, Phan R, Mukhopadhyay A, Hemme CL, Huang K, Alm EJ, Fields MW, Wall J, Stahl D, Hazen TC, Keasling JD, Arkin AP, Zhou J (2010) Global transcriptional, physiological, and metabolite analyses of the responses of Desulfovibrio vulgaris Hildenborough to salt adaptation. Appl Environ Microbiol 76(5):1574–1586. https://doi.org/10.1128/aem.02141-09
Zhang W, Culley DE, Hogan M, Vitiritti L, Brockman FJ (2006) Oxidative stress and heat-shock responses in Desulfovibrio vulgaris by genome-wide transcriptomic analysis. Antonie Van Leeuwenhoek 90(1):41–55. https://doi.org/10.1007/s10482-006-9059-9
Li J, Qi L, Guo Y, Yue L, Li Y, Ge W, Wu J, Shi W, Dong X (2015) Global mapping transcriptional start sites revealed both transcriptional and post-transcriptional regulation of cold adaptation in the methanogenic archaeon Methanolobus psychrophilus. Sci Rep. https://doi.org/10.1038/srep09209
Czechowski MH, Rossmoore HW (1990) Purification and partial characterization of ad(−)-lactate dehydrogenase from Desulfovibrio desulfuricans (ATCC 7757). J Ind Microbiol 6(2):117–122. https://doi.org/10.1007/bf01576430
Xu D, Li Y, Gu T (2014) d-Methionine as a biofilm dispersal signaling molecule enhanced tetrakis hydroxymethyl phosphonium sulfate mitigation of Desulfovibrio vulgaris biofilm and biocorrosion pitting. Mater Corros 65(8):837–845. https://doi.org/10.1002/maco.201206894
Wen J, Zhao K, Gu T, Raad I (2009) Chelators enhanced biocide inhibition of planktonic sulfate-reducing bacterial growth. World J Microbiol Biotechnol 26(6):1053–1057. https://doi.org/10.1007/s11274-009-0269-y
Vincent KA, Parkin A, Lenz O, Albracht SPJ, Fontecilla-Camps JC, Cammack R, Friedrich B, Armstrong FA (2005) Electrochemical definitions of O-2 sensitivity and oxidative inactivation in hydrogenases. J Am Chem Soc 127(51):18179–18189. https://doi.org/10.1021/ja055160v
Hatchikian EC, Magro V, Forget N, Nicolet Y, Fontecilla-Camps JC (1999) Carboxy-terminal processing of the large subunit of [Fe] hydrogenase from Desulfovibrio desulfuricans ATCC 7757. J Bacteriol 181(9):2947–2952. https://doi.org/10.1128/jb.181.9.2947-2952.1999
Cock PJA, Fields CJ, Goto N, Heuer ML, Rice PM (2010) The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res 38(6):1767–1771. https://doi.org/10.1093/nar/gkp1137
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2007) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36(Database):D480–D484. https://doi.org/10.1093/nar/gkm882
Morgan JF (2007) p Value fetishism and use of the Bonferroni adjustment. Evid Based Ment Health 10(2):34–35. https://doi.org/10.1136/ebmh.10.2.34
de Hoon MJL, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20(9):1453–1454. https://doi.org/10.1093/bioinformatics/bth078
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci 95(25):14863–14868. https://doi.org/10.1073/pnas.95.25.14863
Saldanha AJ (2004) Java Treeview—extensible visualization of microarray data. Bioinformatics 20(17):3246–3248. https://doi.org/10.1093/bioinformatics/bth349
Broadbent JR, Larsen RL, Deibel V, Steele JL (2010) Physiological and transcriptional response of Lactobacillus casei ATCC 334 to acid stress. J Bacteriol 192(9):2445–2458. https://doi.org/10.1128/jb.01618-09
Boyd DA, Cvitkovitch DG, Bleiweis AS, Kiriukhin MY, Debabov DV, Neuhaus FC, Hamilton IR (2000) Defects in d-Alanyl-lipoteichoic acid synthesis in Streptococcus mutans results in acid sensitivity. J Bacteriol 182(21):6055–6065. https://doi.org/10.1128/jb.182.21.6055-6065.2000
Azcarate-Peril MA, McAuliffe O, Altermann E, Lick S, Russell WM, Klaenhammer TR (2005) Microarray analysis of a two-component regulatory system involved in acid resistance and proteolytic activity in Lactobacillus acidophilus. Appl Environ Microbiol 71(10):5794–5804. https://doi.org/10.1128/aem.71.10.5794-5804.2005
Mortensen HD, Gori K, Siegumfeldt H, Nissen P, Jespersen L, Arneborg N (2005) Intracellular pH homeostasis plays a role in the NaCl tolerance of Debaryomyces hansenii strains. Appl Microbiol Biotechnol 71(5):713–719. https://doi.org/10.1007/s00253-005-0196-2
Abbott DA, Suir E, Duong G-H, de Hulster E, Pronk JT, van Maris AJA (2009) Catalase overexpression reduces lactic acid-induced oxidative stress in Saccharomyces cerevisiae. Appl Environ Microbiol 75(8):2320–2325. https://doi.org/10.1128/aem.00009-09
Heidelberg JF, Seshadri R, Haveman SA, Hemme CL, Paulsen IT, Kolonay JF, Eisen JA, Ward N, Methe B, Brinkac LM, Daugherty SC, Deboy RT, Dodson RJ, Durkin AS, Madupu R, Nelson WC, Sullivan SA, Fouts D, Haft DH, Selengut J, Peterson JD, Davidsen TM, Zafar N, Zhou L, Radune D, Dimitrov G, Hance M, Tran K, Khouri H, Gill J, Utterback TR, Feldblyum TV, Wall JD, Voordouw G, Fraser CM (2004) The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat Biotechnol 22(5):554–559. https://doi.org/10.1038/nbt959
Sachar DB (1995) Maintenance therapy in ulcerative colitis and Crohnʼs disease. J Clin Gastroenterol 20(2):117–122. https://doi.org/10.1097/00004836-199503000-00009
Chhabra SR, He Q, Huang KH, Gaucher SP, Alm EJ, He Z, Hadi MZ, Hazen TC, Wall JD, Zhou J, Arkin AP, Singh AK (2006) Global analysis of heat shock response in Desulfovibrio vulgaris Hildenborough. J Bacteriol 188(5):1817–1828. https://doi.org/10.1128/jb.188.5.1817-1828.2006
Clark ME, He Q, He Z, Huang KH, Alm EJ, Wan XF, Hazen TC, Arkin AP, Wall JD, Zhou JZ, Fields MW (2006) Temporal transcriptomic analysis as Desulfovibrio vulgaris Hildenborough transitions into stationary phase during electron donor depletion. Appl Environ Microbiol 72(8):5578–5588. https://doi.org/10.1128/aem.00284-06
He Q, Huang KH, He Z, Alm EJ, Fields MW, Hazen TC, Arkin AP, Wall JD, Zhou J (2006) Energetic consequences of nitrite stress in Desulfovibrio vulgaris Hildenborough, inferred from global transcriptional analysis. Appl Environ Microbiol 72(6):4370–4381. https://doi.org/10.1128/aem.02609-05
Stolyar S, He Q, Joachimiak MP, He Z, Yang ZK, Borglin SE, Joyner DC, Huang K, Alm E, Hazen TC, Zhou J, Wall JD, Arkin AP, Stahl DA (2007) Response of Desulfovibrio vulgaris to alkaline stress. J Bacteriol 189(24):8944–8952. https://doi.org/10.1128/jb.00284-07
Zhang W, Culley DE, Scholten JCM, Hogan M, Vitiritti L, Brockman FJ (2006) Global transcriptomic analysis of Desulfovibrio vulgaris on different electron donors. Antonie Van Leeuwenhoek 89(2):221–237. https://doi.org/10.1007/s10482-005-9024-z
Zhou A, Baidoo E, He Z, Mukhopadhyay A, Baumohl JK, Benke P, Joachimiak MP, Xie M, Song R, Arkin AP, Hazen TC, Keasling JD, Wall JD, Stahl DA, Zhou J (2013) Characterization of NaCl tolerance in Desulfovibrio vulgaris Hildenborough through experimental evolution. ISME J 7(9):1790–1802. https://doi.org/10.1038/ismej.2013.60
Hao X, Chen B, An T (2015) Pathway modification of industrial microorganisms to improve acid-stress tolerance. Sheng Wu Gong Cheng Xue Bao 31(8):1151–1161. https://doi.org/10.13345/j.cjb.140496
Abdullah Al M, Sugimoto S, Higashi C, Matsumoto S, Sonomoto K (2010) Improvement of multiple-stress tolerance and lactic acid production in Lactococcus lactis NZ9000 under conditions of thermal stress by heterologous expression of Escherichia coli dnaK. Appl Environ Microbiol 76(13):4277–4285. https://doi.org/10.1128/aem.02878-09
Zhang J, Fu RY, Hugenholtz J, Li Y, Chen J (2007) Glutathione protects Lactococcus lactis against acid stress. Appl Environ Microbiol 73(16):5268–5275. https://doi.org/10.1128/AEM.02787-06
Xue C, Zhao J, Chen L, Yang ST, Bai F (2017) Recent advances and state-of-the-art strategies in strain and process engineering for biobutanol production by Clostridium acetobutylicum. Biotechnol Adv 35(2):310–322. https://doi.org/10.1016/j.biotechadv.2017.01.007
Zuberi A, Misba L, Khan AU (2017) CRISPR Interference (CRISPRi) Inhibition of luxS gene expression in E. coli: an approach to inhibit biofilm. Front Cell Infect Microbiol 7:214. https://doi.org/10.3389/fcimb.2017.00214
Acknowledgements
We thank the research group of Mingding Li from Zhejiang University. This study was supported by the National Natural Science Foundation of China (Nos. 41330639 and 41720104004) and the Guangdong Natural Science Foundation (No. 2018A030313918).
Author information
Authors and Affiliations
Contributions
Conceived and designed the study: FL, ZD and CG. Collected and analyzed the data: HY, YL, XY, CH, ZJ and YO. Wrote and edited the manuscript: HY, YL and FL.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, H., Jiang, Z., Lu, Y. et al. Transcriptome Analysis of the Acid Stress Response of Desulfovibrio vulgaris ATCC 7757. Curr Microbiol 77, 2702–2712 (2020). https://doi.org/10.1007/s00284-020-02051-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02051-x