Abstract
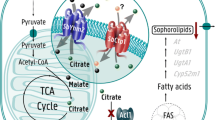
The mitochondrial citrate transport system, composed of citrate and malate transporters (MTs), can regulate the citrate efflux from mitochondria to cytosol, and then citrate is cleaved into OAA and acetyl-CoA which can be used for fatty acid (FA) biosynthesis. However, in the fungus Mucor circinelloides the molecular mechanism of citrate efflux from the mitochondria by this system and its role in FA synthesis is unclear. In the present study, we have analyzed the genome of high lipid-producing strain WJ11 and the low lipid-producing strain CBS 277.49 to find the potential genes involving in this system. Five potential genes are present in the genome of WJ11. These genes encode one citrate transport protein (CT), one tricarboxylate carrier (TCT), one MT, and two 2-oxoglutarate:malate antiporters (SoDIT-a and SoDIT-b). However, the genome of CBS 277.49 contains the same set of genes, except for the presence of just one SoDIT. The proteins from WJ11 had similar properties as their counterparts in CBS 277.49. Moreover, phylogenetic analyses revealed the evolutionary relationship of these proteins and illuminated their typical motifs related to potential functions. Additionally, the expression of these genes was analyzed to predict the possible functions in lipid metabolism in M. circinelloides. This is the first study to report the in silico analysis of structures and functions of the mitochondrial citrate transport system in M. circinelloides. This work showed a new strategy for research for the selection of candidate genes for further detailed functional investigation of the mitochondrial citrate transport system in lipid accumulation.




Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this article (and its Additional file).
References
Thorpe R, Ratledge C (1972) Fatty acid distribution in triglycerides of yeasts grown on glucose or n-alkanes. J Gen Microbiol 72:151–163
Chutrakul C, Jeennor S, Panchanawaporn S, Cheawchanlertfa P, Suttiwattanakul S, Veerana M, Laoteng K (2016) Metabolic engineering of long chain-polyunsaturated fatty acid biosynthetic pathway in oleaginous fungus for dihomo-gamma linolenic acid production. J Biotechnol 218:85–93. https://doi.org/10.1016/j.jbiotec.2015.12.003
Zhang Y, Luan X, Zhang H, Garre V, Song Y, Ratledge C (2017) Improved gamma-linolenic acid production in Mucor circinelloides by homologous overexpressing of delta-12 and delta-6 desaturases. Microb Cell Factories 16(1):113. https://doi.org/10.1186/s12934-017-0723-8
Khan M, Yang J, Hussain S, Zhang H, Garre V, Song Y (2019) Genetic Modification of Mucor circinelloides to construct stearidonic acid producing cell factory. Int J Mol Sci 20(7):1683. https://doi.org/10.3390/ijms20071683
Khan MAK, Yang J, Hussain SA, Zhang H, Liang L, Garre V, Song Y (2019) Construction of DGLA producing cell factory by genetic modification of Mucor circinelloides. Microb Cell Factories 18(1):64. https://doi.org/10.1186/s12934-019-1110-4
Ratledge C (2014) The role of malic enzyme as the provider of NADPH in oleaginous microorganisms: a reappraisal and unsolved problems. Biotechnol Lett 36(8):1557–1568. https://doi.org/10.1007/s10529-014-1532-3
Guay C, Madiraju SR, Aumais A, Joly E, Prentki M (2007) A role for ATP-citrate lyase, malic enzyme, and pyruvate/citrate cycling in glucose-induced insulin secretion. J Biol Chem 282(49):35657–35665. https://doi.org/10.1074/jbc.M707294200
Heart E, Cline GW, Collis LP, Pongratz RL, Gray JP, Smith PJ (2009) Role for malic enzyme, pyruvate carboxylation, and mitochondrial malate import in glucose-stimulated insulin secretion. Am J Physiol Endocrinol Metab 296(6):E1354–E1362. https://doi.org/10.1152/ajpendo.90836.2008
Zhao L, Tang X, Luan X, Chen H, Chen YQ, Chen W, Song Y, Ratledge C (2015) Role of pentose phosphate pathway in lipid accumulation of oleaginous fungus Mucor circinelloides. RSC Adv 5(118):97658–97664. https://doi.org/10.1039/c5ra20364c
Zhang H, Zhang L, Chen H, Chen YQ, Chen W, Song Y, Ratledge C (2014) Enhanced lipid accumulation in the yeast Yarrowia lipolytica by over-expression of ATP:citrate lyase from Mus musculus. J Biotechnol 192(Pt A):78–84. https://doi.org/10.1016/j.jbiotec.2014.10.004
Castegna A, Scarcia P, Agrimi G, Palmieri L, Rottensteiner H, Spera I, Germinario L, Palmieri F (2010) Identification and functional characterization of a novel mitochondrial carrier for citrate and oxoglutarate in Saccharomyces cerevisiae. J Biol Chem 285(23):17359–17370. https://doi.org/10.1074/jbc.M109.097188
Dolce V, Cappello AR, Capobianco L (2014) Mitochondrial tricarboxylate and dicarboxylate–tricarboxylate carriers: from animals to plants. IUBMB Life 66(7):462–471. https://doi.org/10.1002/iub.1290
Klingenberg M (1972) Kinetic study of the tricarboxylate carrier in rat liver mitochondria. Eur J Biochem 26(4):587–594
Spagnoletta A, De Santis A, Tampieri E, Baraldi E, Bachi A, Genchi G (2006) Identification and kinetic characterization of HtDTC, the mitochondrial dicarboxylate–tricarboxylate carrier of Jerusalem artichoke tubers. J Bioenerg Biomembr 38(1):57–65. https://doi.org/10.1007/s10863-006-9006-5
Shug AL, Shrago E (1973) Inhibition of phosphoenolpyruvate transport via the tricarboxylate and adenine nucleotide carrier systems of rat liver mitochondria. Biochem Biophys Res Commun 53(2):659–665
De Palma A, Prezioso G, Scalera V (2005) Kinetic evidence for the uniport mechanism hypothesis in the mitochondrial tricarboxylate transport system. J Bioenerg Biomembr 37(5):279–287. https://doi.org/10.1007/s10863-005-8639-0
Bisaccia F, De Palma A, Palmieri F (1990) Purification and reconstitution of the tricarboxylate carrier from rat liver mitochondria. Ital J Biochem 39(3):167A–169A
Zara V, Palmieri L, Franco MR, Perrone M, Gnoni GV, Palmieri F (1998) Kinetics of the reconstituted tricarboxylate carrier from eel liver mitochondria. J Bioenerg Biomembr 30(6):555–563
Gnoni GV, Giudetti AM, Mercuri E, Damiano F, Stanca E, Priore P, Siculella L (2010) Reduced activity and expression of mitochondrial citrate carrier in streptozotocin-induced diabetic rats. Endocrinology 151(4):1551–1559. https://doi.org/10.1210/en.2009-1352
Mancusso R, Gregorio GG, Liu Q, Wang DN (2012) Structure and mechanism of a bacterial sodium-dependent dicarboxylate transporter. Nature 491(7425):622–626. https://doi.org/10.1038/nature11542
Halperin ML, Schiller CM, Fritz IB (1971) The inhibition by methylmalonic acid of malate transport by the dicarboxylate carrier in rat liver mitochondria. A possible explantation for hypoglycemia in methylmalonic aciduria. J Clin Investig 50(11):2276–2282. https://doi.org/10.1172/JCI106725
Johnson RN, Chappell JB (1973) The transport of inorganic phosphate by the mitochondrial dicarboxylate carrier. Biochem J 134(3):769–774
Nozawa A, Fujimoto R, Matsuoka H, Tsuboi T, Tozawa Y (2011) Cell-free synthesis, reconstitution, and characterization of a mitochondrial dicarboxylate–tricarboxylate carrier of Plasmodium falciparum. Biochem Biophys Res Commun 414(3):612–617. https://doi.org/10.1016/j.bbrc.2011.09.130
Huypens P, Pillai R, Sheinin T, Schaefer S, Huang M, Odegaard ML, Ronnebaum SM, Wettig SD, Joseph JW (2011) The dicarboxylate carrier plays a role in mitochondrial malate transport and in the regulation of glucose-stimulated insulin secretion from rat pancreatic beta cells. Diabetologia 54(1):135–145. https://doi.org/10.1007/s00125-010-1923-5
Zhao L, Canovas-Marquez JT, Tang X, Chen H, Chen YQ, Chen W, Garre V, Song Y, Ratledge C (2016) Role of malate transporter in lipid accumulation of oleaginous fungus Mucor circinelloides. Appl Microbiol Biotechnol 100(3):1297–1305. https://doi.org/10.1007/s00253-015-7079-y
Palmieri EM, Spera I, Menga A, Infantino V, Porcelli V, Iacobazzi V, Pierri CL, Hooper DC, Palmieri F (1847) Castegna A (2015) Acetylation of human mitochondrial citrate carrier modulates mitochondrial citrate/malate exchange activity to sustain NADPH production during macrophage activation. Biochim Biophys Acta 8:729–738. https://doi.org/10.1016/j.bbabio.2015.04.009
Camarasa C, Bidard F, Bony M, Barre P, Dequin S (2001) Characterization of Schizosaccharomyces pombe malate permease by expression in Saccharomyces cerevisiae. Appl Environ Microbiol 67(9):4144–4151. https://doi.org/10.1128/aem.67.9.4144-4151.2001
Thammarongtham C, Nookaew I, Vorapreeda T, Srisuk T, Land ML, Jeennor S, Laoteng K (2017) Genome characterization of oleaginous Aspergillus oryzae BCC7051: a potential fungal-based platform for lipid production. Curr Microbiol. https://doi.org/10.1007/s00284-017-1350-7
Vongsangnak W, Ruenwai R, Tang X, Hu X, Zhang H, Shen B, Song Y, Laoteng K (2013) Genome-scale analysis of the metabolic networks of oleaginous Zygomycete fungi. Gene 521(1):180–190. https://doi.org/10.1016/j.gene.2013.03.012
Ratledge C, Wynn JP (2002) The biochemistry and molecular biology of lipid accumulation in oleaginous microorganisms. Adv Appl Microbiol 51:1–51
Tang X, Zhao L, Chen H, Chen YQ, Chen W, Song Y, Ratledge C (2015) Complete genome sequence of a high lipid-producing strain of Mucor circinelloides WJ11 and comparative genome analysis with a low lipid-producing strain CBS 277.49. PLoS ONE 10(9):e0137543. https://doi.org/10.1371/journal.pone.0137543
Yang J, Li S, Kabir Khan MA, Garre V, Vongsangnak W, Song Y (2019) Increased lipid accumulation in Mucor circinelloides by overexpression of mitochondrial citrate transporter genes. Ind Eng Chem Res 58(6):2125–2134. https://doi.org/10.1021/acs.iecr.8b05564
Saier MH Jr, Reddy VS, Tsu BV, Ahmed MS, Li C, Moreno-Hagelsieb G (2016) The Transporter Classification Database (TCDB): recent advances. Nucleic Acids Res 44(D1):D372–D379. https://doi.org/10.1093/nar/gkv1103
Kendrick A, Ratledge C (1992) Desaturation of polyunsaturated fatty acids in Mucor circinelloides and the involvement of a novel membrane-bound malic enzyme. Eur J Biochem 209(2):667–673
Zan X, Tang X, Chu L, Zhao L, Chen H, Chen YQ, Chen W, Song Y (2016) Lipase genes in Mucor circinelloides: identification, sub-cellular location, phylogenetic analysis and expression profiling during growth and lipid accumulation. J Ind Microbiol Biotechnol 43(10):1467–1480. https://doi.org/10.1007/s10295-016-1820-0
Palmieri F (2013) The mitochondrial transporter family SLC25: identification, properties and physiopathology. Mol Asp Med 34(2–3):465–484. https://doi.org/10.1016/j.mam.2012.05.005
Xu Y, Kakhniashvili DA, Gremse DA, Wood DO, Mayor JA, Walters DE, Kaplan RS (2000) The yeast mitochondrial citrate transport protein. Probing the roles of cysteines, Arg(181), and Arg(189) in transporter function. J Biol Chem 275(10):7117–7124
Ma C, Remani S, Sun J, Kotaria R, Mayor JA, Walters DE, Kaplan RS (2007) Identification of the substrate binding sites within the yeast mitochondrial citrate transport protein. J Biol Chem 282(23):17210–17220. https://doi.org/10.1074/jbc.M611268200
Majd H, King MS, Smith AC (1859) Kunji ERS (2018) Pathogenic mutations of the human mitochondrial citrate carrier SLC25A1 lead to impaired citrate export required for lipid, dolichol, ubiquinone and sterol synthesis. Biochim Biophys Acta 1:1–7. https://doi.org/10.1016/j.bbabio.2017.10.002
Azzi A, Glerum M, Koller R, Mertens W, Spycher S (1993) The mitochondrial tricarboxylate carrier. J Bioenerg Biomembr 25(5):515–524
Capobianco L, Ferramosca A, Zara V (2002) The mitochondrial tricarboxylate carrier of silver eel: dimeric structure and cytosolic exposure of both N- and C-termini. J Protein Chem 21(8):515–521
Siculella L, Sabetta S, Damiano F, Giudetti AM, Gnoni GV (2004) Different dietary fatty acids have dissimilar effects on activity and gene expression of mitochondrial tricarboxylate carrier in rat liver. FEBS Lett 578(3):280–284. https://doi.org/10.1016/j.febslet.2004.11.014
Miyake S, Yamashita T, Taniguchi M, Tamatani M, Sato K, Tohyama M (2002) Identification and characterization of a novel mitochondrial tricarboxylate carrier. Biochem Biophys Res Commun 295(2):463–468
Marchler-Bauer A, Bo Y, Han L, He J, Lanczycki CJ, Lu S, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Geer LY, Bryant SH (2017) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45(D1):D200–D203. https://doi.org/10.1093/nar/gkw1129
Bergeron MJ, Clemencon B, Hediger MA, Markovich D (2013) SLC13 family of Na(+)-coupled di- and tri-carboxylate/sulfate transporters. Mol Asp Med 34(2–3):299–312. https://doi.org/10.1016/j.mam.2012.12.001
Pajor AM (1999) Sodium-coupled transporters for Krebs cycle intermediates. Annu Rev Physiol 61:663–682. https://doi.org/10.1146/annurev.physiol.61.1.663
Liu R, Li B, Qin G, Zhang Z, Tian S (2017) Identification and functional characterization of a tonoplast dicarboxylate transporter in tomato (Solanum lycopersicum). Front Plant Sci 8:186. https://doi.org/10.3389/fpls.2017.00186
Medeiros DB, Barros KA, Barros JAS, Omena-Garcia RP, Arrivault S, Sanglard L, Detmann KC, Silva WB, Daloso DM, DaMatta FM, Nunes-Nesi A, Fernie AR, Araujo WL (2017) Impaired malate and fumarate accumulation due to the mutation of the tonoplast dicarboxylate transporter has little effects on stomatal behavior. Plant Physiol 175(3):1068–1081. https://doi.org/10.1104/pp.17.00971
Birkenfeld AL, Lee HY, Guebre-Egziabher F, Alves TC, Jurczak MJ, Jornayvaz FR, Zhang D, Hsiao JJ, Martin-Montalvo A, Fischer-Rosinsky A, Spranger J, Pfeiffer AF, Jordan J, Fromm MF, Konig J, Lieske S, Carmean CM, Frederick DW, Weismann D, Knauf F, Irusta PM, De Cabo R, Helfand SL, Samuel VT, Shulman GI (2011) Deletion of the mammalian INDY homolog mimics aspects of dietary restriction and protects against adiposity and insulin resistance in mice. Cell Metab 14(2):184–195. https://doi.org/10.1016/j.cmet.2011.06.009
Tang X, Chen H, Chen YQ, Chen W, Garre V, Song Y, Ratledge C (2015) Comparison of biochemical activities between high and low lipid-producing strains of Mucor circinelloides: an explanation for the high oleaginicity of strain WJ11. PLoS ONE 10(6):e0128396. https://doi.org/10.1371/journal.pone.0128396
Iacobazzi V, Infantino V (2014) Citrate—new functions for an old metabolite. Biol Chem 395(4):387–399. https://doi.org/10.1515/hsz-2013-0271
Kirimura K, Kobayashi K, Ueda Y, Hattori T (2016) Phenotypes of gene disruptants in relation to a putative mitochondrial malate–citrate shuttle protein in citric acid-producing Aspergillus niger. Biosci Biotechnol Biochem 80(9):1737–1746. https://doi.org/10.1080/09168451.2016.1164583
Bellou S, Triantaphyllidou IE, Mizerakis P, Aggelis G (2016) High lipid accumulation in Yarrowia lipolytica cultivated under double limitation of nitrogen and magnesium. J Biotechnol 234:116–126. https://doi.org/10.1016/j.jbiotec.2016.08.001
Broach JR (2012) Nutritional control of growth and development in yeast. Genetics 192(1):73–105. https://doi.org/10.1534/genetics.111.135731
Kurihara Y, Kanki T, Aoki Y, Hirota Y, Saigusa T, Uchiumi T, Kang D (2012) Mitophagy plays an essential role in reducing mitochondrial production of reactive oxygen species and mutation of mitochondrial DNA by maintaining mitochondrial quantity and quality in yeast. J Biol Chem 287(5):3265–3272. https://doi.org/10.1074/jbc.M111.280156
Acknowledgements
We would like to thank Professor Wanwipa Vongsangnak in Computational Biomodelling Laboratory for Agricultural Science and Technology (CBLAST), Faculty of Science, Kasetsart University, Thailand, for computational facilities and Yao Zhang for data analysis work supported by the Key Research and Development Project of Shandong Province (2018GSF121013).
Funding
This work was supported by National Natural Science Foundation of China (Grant No. 31670064), TaiShan Industrial Experts Programme (tscy 20160101), the Key Research and Development Project of Shandong Province (2018GNC110039), Starting Grant from Shandong University of Technology and the Plan to Introduce High-Quality Foreign Experts in Zibo City.
Author information
Authors and Affiliations
Contributions
Junhuan Yang and Md. Ahsanul Kabir Khan performed the experimental design, computational analysis, fermentation testing, manuscript writing, and figures and tables arrangement. Huaiyuan Zhang and Yao Zhang were involved in the experimental design. Victoriano Garre carried out results interpretation and reviewed of final draft. Yuanda Song proposed the project and was involved in data analysis, result interpretation, manuscript writing, and review of the final draft.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
284_2019_1822_MOESM1_ESM.docx
Supplementary file1 (DOCX 137 kb) S1. List of 51 candidates of mitochondrial transporter genes with predicted functions from TCDB. S2. Cell growth and lipid accumulation in M. circinelloides WJ11 and CBS 277.49. S3. Primers used in this study.
Rights and permissions
About this article
Cite this article
Yang, J., Khan, M.A.K., Zhang, H. et al. Mitochondrial Citrate Transport System in the Fungus Mucor circinelloides: Identification, Phylogenetic Analysis, and Expression Profiling During Growth and Lipid Accumulation. Curr Microbiol 77, 220–231 (2020). https://doi.org/10.1007/s00284-019-01822-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01822-5