Abstract
A novel facultative anaerobic and Gram-stain-positive coccus, designated strain ChDC F135T, was isolated from human subgingival dental plaque of periodontitis lesion and was characterized by polyphasic taxonomic analysis. The 16S rRNA gene (16S rDNA) sequence of strain ChDC F135T was closest to that of Streptococcus sinensis HKU4T (98.2%), followed by Streptococcus intermedia SK54T (97.0%), Streptococcus constellatus NCTC11325T (96.0%), and Streptococcus anginosus NCTC 10713T (95.7%). In contrast, phylogenetic analysis based on the superoxide dismutase gene (sodA) and the RNA polymerase beta-subunit gene (rpoB) showed that the nucleotide sequence similarities of strain ChDC F135T were highly similar to the corresponding genes of S. anginosus NCTC 10713T (99.2% and 97.6%, respectively), S. constellatus NCTC11325T (87.8% and 91.4%, respectively), and S. intermedia SK54T (85.8% and 91.2%, respectively) rather than those of S. sinensis HKU4T (80.5% and 82.6%). The complete genome of strain ChDC F135T consisted of 1,901,251 bp and the G+C content was 38.9 mol %. Average nucleotide identity value between strain ChDC F135T and S. sinensis HKU4T or S. anginosus NCTC 10713T were 75.7% and 95.6%, respectively. The C14:0 composition of the cellular fatty acids of strain ChDC F135T (32.8%) was different from that of S. intermedia (6–8%), S. constellatus (6–13%), and S. anginosus (13–20%). Based on the results of phylogenetic and phenotypic analysis, strain ChDC F135T (= KCOM 2412T = JCM 33300T) was classified as a type strain of a novel species of the genus Streptococcus, for which we proposed the name Streptococcus periodonticum sp. nov.

Similar content being viewed by others
References
Allen BL, Katz B, Höök M (2002) Streptococcus anginosus adheres to vascular endothelium basement membrane and purified extracellular matrix proteins. Microb Pathog 32:191–204. https://doi.org/10.1006/mpat.2002.0496
Basaranoglu ST, Ozsurekci Y, Aykac K, Aycan AE, Bıcakcigil A, Altun B, Sancak B, Cengiz AB, Kara A, Ceyhan M (2018) Streptococcus mitis/oralis causing blood stream infections in pediatric patients. Jpn J Infect Dis. https://doi.org/10.7883/yoken.jjid.2018.074
Chatelain S, Lombardi T, Scolozzi P (2018) Streptococcus anginosus dental implant-related ssteomyelitis of the jaws: an insidious and calamitous entity. J Oral Maxillofac 76:1187–1193. https://doi.org/10.1016/j.joms.2018.01.010
Cho E, Park SN, Lim YK, Shin Y, Paek J, Hwang CH, Chang YH, Kook JK (2015) Fusobacterium hwasookii sp. nov., isolated from a human periodontitis lesion. Curr Microbiol 70:169–175. https://doi.org/10.1007/s00284-014-0692-7
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Clarridge JE III, Osting C, Jalali M, Osborne J, Waddington M (1999) Genotypic and phenotypic characterization of “Streptococcus milleri” group isolates from a Veterans Administration hospital population. J Clin Microbiol 37:3681–3687. https://doi.org/10.1099/00221287-135-4-831
Doern CD, Burnham CA (2010) It’s not easy being green: the viridans group streptococci, with a focus on pediatric clinical manifestations. J Clin Microbiol 48:3829–3835. https://doi.org/10.1128/JCM.01563-10
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gao XY, Zhi XY, Li HW, Klenk HP, Li WJ (2014) Comparative genomics of the bacterial genus Streptococcus illuminates evolutionary implications of species groups. PLoS ONE 30:e101229. https://doi.org/10.1371/journal.pone.0101229
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-01
Jensen A, Scholz CF, Kilian M (2016) Re-evaluation of the taxonomy of the Mitis group of the genus Streptococcus based on whole genome phylogenetic analyses, and proposed reclassification of Streptococcus dentisani as Streptococcus oralis subsp. dentisani comb. nov., Streptococcus tigurinus as Streptococcus oralis subsp. tigurinus comb. nov., and Streptococcus oligofermentans as a later synonym of Streptococcus cristatus. Int J Syst Evol Microbiol 66:4803–4820. https://doi.org/10.1099/ijsem.0.001433
Jo E, Park SN, Lim YK, Paek J, Shin Y, Kim H, Kim SH, Shin JH, Chang YH, Kook JK (2018) Capnocytophaga endodontalis sp. nov., isolated from a human refractory periapical abscess. Curr Microbiol 75:420–425. https://doi.org/10.1007/s00284-017-1397-5
Kawamura Y, Hou XG, Sultana F, Miura H, Ezaki T (1995) Determination of 16S rRNA sequences of Streptococcus mitis and Streptococcus gordonii and phylogenetic relationships among members of the genus Streptococcus. Int J Syst Bacteriol 45:406–408 (Erratum in: Int J Syst Bacteriol (1995) 45:882)
Kim HS, Lee DS, Chang YH, Kim MJ, Koh S, Kim J, Seong JH, Song SK, Shin HS, Son JB, Jung MY, Park SN, Yoo SY, Cho KW, Kim DK, Moon S, Kim D, Choi Y, Kim BO, Jang HS, Kim CS, Kim C, Choe SJ, Kook JK (2010) Application of rpoB and zinc protease gene for use in molecular discrimination of Fusobacterium nucleatum subspecies. J Clin Microbiol 48:545–553. https://doi.org/10.1128/JCM.01631-09
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0 (Erratum in: Int J Syst Evol Microbiol 2014;64:1825)
Kim SL, Gordon SM, Shrestha NK (2018) Distribution of streptococcal groups causing infective endocarditis: a descriptive study. Diagn Microbiol Infect Dis 91:269–272. https://doi.org/10.1016/j.diagmicrobio.2018.02.015
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60. https://doi.org/10.1186/1471-2105-14-60
Mohanty S, Panigrahi MK, Turuk J, Dhal S (2018) Liver abscess due to Streptococcus constellatus in an immunocompetent adult: a less known entity. J Natl Med Assoc 110:591–595. https://doi.org/10.1016/j.jnma.2018.03.006
Morita E, Narikiyo M, Yano A, Nishimura E, Igaki H, Sasaki H, Terada M, Hanada N, Kawabe R (2003) Different frequencies of Streptococcus anginosus infection in oral cancer and esophageal cancer. Cancer Sci 94:492–496. https://doi.org/10.1111/j.1349-7006.2003.tb01471.x
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Seta V, Teicher E, Fortineau N, Ladouceur M, Lambotte O (2015) Infective endocarditis caused by Streptococcus sinensis. Med Mal Infect 45:56–57. https://doi.org/10.1016/j.medmal.2014.11.001
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849. https://doi.org/10.1099/00207713-44-4-846
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis Version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Wagner KW, Schön R, Schumacher M, Schmelzeisen R, Schulze D (2006) Case report: brain and liver abscesses caused by oral infection with Streptococcus intermedius. Oral Surg Oral Med Oral Pathol Oral Radiol Endod 102:e21–e23. https://doi.org/10.1016/j.tripleo.2006.02.010
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) Report of the Ad Hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464. https://doi.org/10.1099/00207713-37-4-463
Whiley RA, Beighton D, Winstanley TG, Fraser HY, Hardie JM (1992) Streptococcus intermedius, Streptococcus constellatus, and Streptococcus anginosus (the Streptococcus milleri group): association with different body sites and clinical infections. J Clin Microbiol 30:243–244
Woo PC, Tam DM, Leung KW, Lau SK, Teng JL, Wong MK, Yuen KY (2002) Streptococcus sinensis sp. nov., a novel species isolated from a patient with infective endocarditis. J Clin Microbiol 40:805–810. https://doi.org/10.1128/JCM.40.3.805-810.2002
Acknowledgments
This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science and ICT (Grant No. 2017M3A9B8065844) and in part by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (Grant No. 2018R1A2B5002239).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
DPD number: TA00863. GenBank accession number of 16S rRNA gene for strain ChDC F135 T: MK452275. GenBank accession number of genome for strain ChDC F135T: CP034543.
Electronic supplementary material
Below is the link to the electronic supplementary material.
284_2019_1695_MOESM1_ESM.pptx
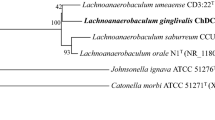
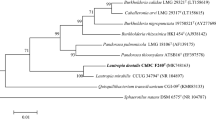
Supplementary material 1—Supplementary Fig S1. Maximum likelihood (A) and the minimum evolution (B) phylogenetic tree based on 16S rDNA sequences of strain ChDC F135T and type strains of related species. Stability of phylogenetic tree was assessed by bootstrap analysis of 1,000 replicates with MEGA version 6.06 [27]. Bars indicate 0.01 (A) or 0.005 (B) changes per nucleotide position (PPTX 90 kb)
Rights and permissions
About this article
Cite this article
Lim, Y.K., Park, SN., Shin, J.H. et al. Streptococcus periodonticum sp. nov., Isolated from Human Subgingival Dental Plaque of Periodontitis Lesion. Curr Microbiol 76, 835–841 (2019). https://doi.org/10.1007/s00284-019-01695-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01695-8