Abstract
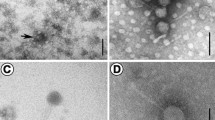
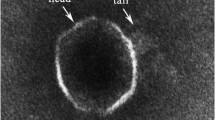
We isolated and purified a novel virulent Pseudoalteromonas bacteriophage PHq0 and the host bacterium Pseudoalteromonas BQ0 from seawater collected in a coastal area of the Yellow Sea of China. (36°06′N, 120°32′E). Transmission electron microscopy revealed that the phage had an icosahedral head of 50 nm in diameter with a long tail of 100 nm. The one-step growth curve showed the latent period of about 15 min, a rise period of 15 min, and a burst size of about 363 virions. The genome of phage PHq0 was found to consist of a linear, double-stranded 33,399-bp DNA molecule with a GC content of 40.29 % and 56 putative open reading frames (ORFs). Among these genes, 23 conserved domains were detected by BLASTP, 17 were functionally known, leaving 39 unknown putative genes, BLASTP results show that 57.14 % of the 56 predicted ORFs were not found to have any matches of putative functions or conserved domains in the BLASTP database which should be classified as a new member of the Siphoviridae family. The phage PHq0 genome adds a new Siphoviridae-family phage genome for marine bacteriophages which will provide useful basic information for further molecular research on interaction mechanism between bacteriophages and their hosts.




Similar content being viewed by others
References
Ahiwale S, Prakash D, Gajbhiye M et al (2012) BVPaP-3, a T7-like lytic phage of pseudomonas aeruginosa: its isolation and characterisation. Curr Microbiol 64:305–311
Bai Q, Zhang W, Yang YC et al (2012) Characterization and genome sequencing of a novel bacteriophage infecting Streptococcus agalactiae with high similarity to a phage from Streptococcus pyogenes. Arch Virol 158:1733–1741
Brown JM, LaBarre BA, Hewson I (2013) Characterization of trichodesmium-associated viral communities in the eastern Gulf of Mexico. FEMS Microbiol Ecol 84(3):603–613
Duhaime MB, Wichels A, Waldmann J et al (2011) Ecogenomics and genome landscapes of marine Pseudoalteromonas phage H105/1. ISME J 5(1):107–121
Duran AE, Muniesa M, Mendez X et al (2002) Removal and inactivation of indicator bacteriophages in fresh waters. J Appl Microbiol 92:338–347
Harald B (2009) The not so universal tree of life or the place of viruses in the living world. Phil Trans R Soc B 364:2263–2274
Hardies SC, Hwang YJ, Hwang CY et al (2013) Morphology, physiological characteristics, and complete sequence of marine bacteriophage ϕRIO-1 infecting pseudoalteromonas marina. J Virol 87(16):9189–9198
Huang G-T, Le S, Peng Y-Z et al (2013) Characterization and genome sequencing of phage Abp1, a new phiKMV-like virus infecting multidrug-resistant Acinetobacter baumannii. Curr Microbiol 66:535–543
Jamalludeen N, Johnson RP, Friendship R et al (2007) Isolation and characterization of nine bacteriophages that lyse O149 enterotoxigenic Escherichia coli. Vet Microbiol 124:47–57
Kropinski AM, Mazzocco A, Waddell TE et al (2009) Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol Biol 501:69–76
Lara E, Holmfeldt K, Solonenko N et al (2015) Life-style and genome structure of marine pseudoalteromonas siphovirus B8b isolated from the northwestern mediterranean sea. PLoS One 10:e0114829
Liao W, Song SY, Sun F et al (2008) Isolation, characterization and genome sequencing of phage MZTP02 from Bacillus thuringiensis MZ1. Arch Virol 153:1855–1865
Männistö RH, Kivelä HM, Paulin L et al (1999) The complete genome sequence of PM2, the first lipid-containing bacterial virus to be isolated. Virology 262(2):355–363
Mathias M, Chan Amy M, Bertelsen Sif K (2010) Isolation and life cycle characterization of lytic viruses infecting heterotrophic bacteria and cyanobacteria. Man Aquat Viral Ecol Mave Chap 13:118–133
Moebus K (1992) Further investigations on the concentration of marine bacteriophages in the water around Helgoland, with reference to the phage-host systems encountered. Helgoland Mar Res 46:275–292
Shin H, Lee JH, Ahn CS et al (2014) Complete genome sequence of marine bacterium Pseudoalteromonas phenolica bacteriophage TW1. Arch Virol 159(1):159–162
Sillankorva S, Neubauer P, Azeredo J (2008) Isolation and characterization of a T7-like lytic phage for Pseudomonas fluorescens. BMC Biotechnol 8:80
Thomas T, Evans FF, Schleheck D et al (2008) Analysis of the Pseudoalteromonas tunicata genome reveals properties of a surface-associated life style in the marine environment. PLoS One 3:e3252
Verheust C, Jensen G, Mahillon J (2003) pGIL01, a linear tectiviral plasmid prophage originating from B. Thuringiensis serovar israelensis. Microbiology 149:2083–2092
Weinbauer MG, Hofle MG (1998) Significance of viral lysis and flagellate grazing as factors controlling bacterioplankton production in a eutrophic lake. Appl Environ Microbiol 64:431–438
Zhang YY, Huang CX, Yang J et al (2011) Interactions between marine microorganisms and their phages. Chinese Sci Bull 56:1770–1777
Acknowledgments
We are thankful to Professor Zhang Xiao-Hua, Microbiology lab, College of Marine Life, Ocean University of China, for providing bacterial strains to carry out phage host-range experiments and suggestions during the preparation of this manuscript. We are grateful to the research vessel Dong Fang Hong 2, for providing the seawater samples. The research was funded by the National Natural Science Foundation of China (No. 41076088) and China Postdoctoral Science Foundation, No.2015M5Z0612.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Wang, Db., Li, Y., Sun, Mq. et al. Complete Genome of a Novel Pseudoalteromonas Phage PHq0. Curr Microbiol 72, 81–87 (2016). https://doi.org/10.1007/s00284-015-0919-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-015-0919-2