Abstract
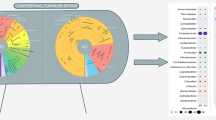
The contents of fed-batch composting (FBC) reactors often aggregate after prolonged operation. This process leads to irreversible breakdown of the decomposition reaction and possible alteration of the bacterial communities. We compared the structures of bacterial communities in reactors under aggregate and optimal conditions. The results of 16S rRNA gene clone analysis showed that populations of the family Bacillaceae (such as Bacillus spp., Cerasibacillus spp., Gracilibacillus spp.), which dominate (98%) under optimal condition, were significantly decreased under aggregate condition. In contrast, populations of the family Staphylococcaceae considerably increased after aggregation and accounted for 53% of the total. Phylogenetic analysis also showed that anaerobes or facultative anaerobes related to Tetragenococcus halophilus, Atopostipes suicloacalis, Jeotgalicoccus pinnipedialis, and Staphylococcus spp. were dominant in the aggregates. These results suggested that aerobic Gram-positive bacteria mainly contributed to organic degradation and that aggregation created some anaerobic environment, which promoted the growth of bacterial communities usually not found in well-functioning FBC reactors.





Similar content being viewed by others
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Ashelford KE, Chuzhanova NA, Fry JC et al (2005) At least 1 in 20 16S rRNA sequence records currently held in public repositories is estimated to contain substantial anomalies. Appl Environ Microbiol 71:7724–7736
Collins MD, Williams AM, Wallbanks S (1990) The phylogeny of Aerococcus and Pediococcus as determined by 16S rRNA sequence analysis: Description of Tetragenococcus gen. nov. FEMS Microbiol Lett 70:255–262
Collins MD, Wiernik A, Falsen E et al (2005) Atopococcus tabaci gen. nov., sp. nov., a novel Gram-positive, catalase-negative, coccus-shaped bacterium isolated from tobacco. Int J Syst Evol Microbiol 55:1693–1696
Cotta MA, Whitehead TR, Collins MD et al (2004) Atopostipes suicloacale gen. nov., sp. nov., isolated from an underground swine manure storage pit. Anaerobe 10:191–195
Dees P, Ghiorse W (2001) Microbial diversity in hot synthetic compost as revealed by PCR-amplified rRNA sequences from cultivated isolates and extracted DNA. FEMS Microbiol Ecol 35:207–216
Demharter W, Hensel R (1989) Bacillus thermocloacae sp. nov., a new thermophilic species from sewage sludge. Syst Appl Microbiol 11:272–276
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) version 3.5c. Distributed by the author, Department of Genetics, University of Washington
Haruta S, Kondo M, Nakamura K et al (2002) Microbial community changes during organic solid waste treatment analyzed by double gradient-denaturing gradient gel electrophoresis and fluorescence in situ hybridization. Appl Microbiol Biotechnol 60:224–231
Haruta S, Kondo M, Nakamura K et al (2004) Succession of a microbial community during stable operation of semi-continuous garbage-decomposing system. J Biosci Bioeng 98:20–27
Hoyles M, Collins MD, Foster G et al (2004) Jeotogalicoccus pinnipedialis sp. nov., from a southern elephant seal (Mirounga leonina). Int J Syst Evol Microbiol 54:745–748
Kurisu F, Satoh S, Mino T et al (2002) Microbial community analysis of thermophilic contact oxidation process by using ribosomal RNA approaches and the quinone profile method. Water Res 36:429–438
Kurosawa N, Itoh YH, Itoh T (2005) Thermus kawarayensis sp. nov., a new member of the genus Thermus, isolated from Japanese hot springs. Extremophiles 9:81–84
Madigan MT, Martinko JM, Parker J et al (2000) Brock biology of microorganisms, 9th ed. New Jersey, Prentice-Hall
Maidak BL, Cole JR, Lilburn TG et al (2001) The RDP-II (Ribosomal Database Project). Nucleic Acids Res 29:173–174
Nakamura K, Haruta S, Ueno S et al (2004) Cerasibacillus quisquiliarum gen. nov., sp. nov., isolated from a semi-continuous decomposing system of kitchen refuse. Int J Syst Evol Microbiol 54:1063–1069
Nakamura K, Haruta S, Nguyen HL et al (2004) Enzyme production-based approach for determining the functions of microorganisms within a community. Appl Environ Microbiol 70:3329–3337
Nagao N, Matsuyama T, Yamamoto H et al (2003) A novel hybrid system of solid state and submerged fermentation with recycle for organic solid waste treatment. Process Biochem 39:37–43
Nagao N, Osa S, Matsuyama T et al (2005) Optimization of washing rate in a hybrid recycling system of solid state and submerged fermentation. Process Biochem 40:3321–3326
Narihiro T, Yamanaka Y, Hiraishi A (2003) High culturability of bacteria in commercially available personal composters for fed-batch treatment of household biowaste. Microbes Environ 18:94–99
Narihiro T, Takebayashi S, Hiraishi A (2004) Activity and phylogenetic composition of proteolytic bacteria in mesophilic fed-batch garbage composters. Microbes Environ 19:292–300
Narihiro T, Abe T, Yamanaka Y et al (2004) Microbial population dynamics during fed-batch operation of commercially available garbage composters. Appl Microbial Biotechnol 65:488–495
Narihiro T, Hiraishi A (2005) Microbiology of fed-batch composting. Microbes Environ 20:1–13
Pedro MS, Haruta S, Nakamura K et al (2003) Isolation and characterization of predominant microorganisms during decomposition of waste materials in a field-scale composter. J Biosci Bioeng 95:368–373
Pikuta E, Lysenko A, Chuvilskaya N et al (2000) Anoxybacillus pushchinensis gen. nov., sp. nov., a novel aerobic, alkaliphilic, moderately thermophilic bacterium from manure, and description of Anoxybacillus flavithermus comb. nov. Int J Syst Evol Microbiol 50:2109–2117
Ryckeboer J, Mergaert J, Coosemans J et al (2003) Microbiological aspects of biowaste during composting in a monitored compost bin. J Appl Microbiol 94:127–137
Saitou N, Nei M. (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shibata A, Goto Y, Nakajima R (2006) Use of SYBR gold stain for enumerating bacteria and viruses in seawater samples [in Japanese]. Bull Plankton Soc Japan 53:64–68
Singleton DR, Furlong MA, Rathbun SL et al (2001) Quantitative comparisons of 16S rRNA gene sequence libraries from environmental samples. Appl Environ Microbiol 67:4374–4376
Strom PF (1985) Identification of thermophilic bacteria in solid-waste composting. Appl Environ Microbiol 50:906–913
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Waino M, Tindall BJ, Schumann P et al (1999) Gracilibacillus gen. nov., with description of Gracilibacillus halotolerans gen. nov., sp. nov.; transfer of Bacillus dipsosauri to Gracilibacillus dipsosauri comb. nov., and Bacillus salexigens to the genus Salibacillus gen. nov., as Salibacillus salexigens comb. nov. Int J Syst Bacteriol 49:821–831
Wang Q, Garrity GM, Tiedje JM et al (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Acknowledgments
This work was supported by University-Industry Joint Research Project for Private Universities and a matching fund subsidy from the Ministry of Education, Culture, Sports, Science and Technology-Japan, 2004-2008. We thank S. Osa for technical assistance, L. Y. Giak, and T. Yoshida for correcting English in the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Watanabe, K., Nagao, N., Toda, T. et al. Changes in Bacterial Communities Accompanied by Aggregation in a Fed-Batch Composting Reactor. Curr Microbiol 56, 458–467 (2008). https://doi.org/10.1007/s00284-008-9107-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-008-9107-y