Abstract
Water management measures in the 1970s in the Netherlands have produced a large number of “resident” populations of three-spined sticklebacks that are no longer able to migrate to the sea. This may be viewed as a replicated field experiment, allowing us to study how the resident populations are coping with human-induced barriers to migration. We have previously shown that residents are smaller, bolder, more exploratory, more active, and more aggressive and exhibited lower shoaling and lower migratory tendencies compared to their ancestral “migrant” counterparts. However, it is not clear if these differences in wild-caught residents and migrants reflect genetic differentiation, rather than different developmental conditions. To investigate this, we raised offspring of four crosses (migrant ♂ × migrant ♀, resident ♂ × resident ♀, migrant ♂ × resident ♀, resident ♂ × migrant ♀) under similar controlled conditions and tested for differences in morphology and behavior as adults. We found that lab-raised resident sticklebacks exhibited lower shoaling and migratory tendencies as compared to lab-raised migrants, retaining the differences in their wild-caught parents. This indicates genetic differentiation of these traits. For all other traits, the lab-raised sticklebacks of the various crosses did not differ significantly, suggesting that the earlier-found contrast between wild-caught fish reflects differences in their environment. Our study shows that barriers to migration can lead to rapid differentiation in behavioral tendencies over contemporary timescales (~ 50 generations) and that part of these differences reflects genetic differentiation.
Significance statement
Many organisms face changes to their habitats due to human activities. Much research is therefore dedicated to the question whether and how organisms are able to adapt to novel conditions. We address this question in three-spined sticklebacks, where water management measures cut off some populations, prohibiting their seasonal migration to the North Sea. In a previous study, we showed that wild-caught “resident” fish exhibited markedly different behavior than migrants. To disentangle whether these differences reflect genetic differentiation or differences in the conditions under which the wild-caught fish grew up, we conducted crosses, raising the F1 offspring under identical conditions. As their wild-caught parents, the F1 of resident × resident crosses exhibited lower migratory and shoaling tendencies than the F1 of migrant × migrant crosses, while the F1 of hybrid crosses were intermediate. This suggests that ~ 50 years of isolation are sufficient to induce behaviorally relevant genetic differentiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Habitat fragmentation resulting from human activities is considered to be a major threat for many animal populations (Foley et al. 2005; Fischer and Lindenmayer 2007). Habitat fragmentation is characterized by a reduction in habitat size, habitat loss, and loss of habitat connectivity (Fahrig 2003). This poses a threat to animal populations, especially for migratory species which rely on connectivity between functional habitats for reproduction and survival (Legrand et al. 2017). Migratory species would thus need to respond via adaptive changes in life history and behavior to thrive in disconnected patches (Bohlin et al. 2001; Kraabøl et al. 2009; Junge et al. 2014). Therefore, understanding the underlying mechanisms of these responses is crucial as they directly affect the future adaptive potential and evolutionary trajectories of populations (Kawecki and Ebert 2004; Wang and Bradburd 2014) as well as conservation measures (Stockwell et al. 2003).
Individuals need to maintain a match between their phenotypes and the environment to enhance their local performance, thereby allowing populations to subsist or grow in an altered environment. Depending on the underlying mechanism involved, such adaptive responses may occur more or less rapidly and may influence population genetic structure (Hedrick et al. 1976, 2006; Nicolaus and Edelaar 2018). For example, phenotypic adjustment may result from natural selection favoring some phenotypes over others, potentially leading to population genetic differentiation across multiple generations when phenotypic variation has a genetic basis (Kawecki and Ebert 2004). Non-exclusively, individuals may match their phenotype to local conditions through plasticity, be it reversible plasticity (or phenotypic flexibility sensu Piersma and Drent 2003), developmental plasticity, or transgenerational plasticity (through parental and epigenetic effects). Plasticity, defined as the ability of a genotype to exhibit different phenotypes in response to the environment (Via et al. 1995; Pigliucci 2005), can thus provide a rapid mechanism to respond to environmental changes (Ghalambor et al. 2007). Importantly, selection may favor genotypes with varying levels of plasticity (Scheiner 1993; Nussey et al. 2007), implying that the mentioned mechanisms are intertwined (Edelaar et al. 2017) and that observed population divergence could reflect genetic differentiation and/or differences in the environments under which individuals grow up. In migratory species, migrants would have to exhibit phenotypic plasticity or bet-hedging strategies, as they are exposed to different environmental conditions (Botero et al. 2015). In the case where migrants are no longer able to migrate (forced “residents”), we expect selection to act on either the traits themselves or on the degree of plasticity.
In this study, we focus on behavior as it is the primary way through which animals interact with their environment and respond to changes (Wong and Candolin 2015). Behavior is often considered highly flexible and hence less prone to genetic divergence in response to environmental changes. However, plastic responses could evolve rapidly through genetic divergence compared to fixed traits (van Gestel and Weissing 2018). In addition, “animal personality” research points that behaviors are highly structured and form correlations over time (consistency) and over contexts (syndromes) (Réale et al. 2007; Stamps and Groothuis 2010; Wolf and Weissing 2012). Furthermore, individual differences within populations are often repeatable (Bakker 1986; Réale et al. 2007) and, to some extent, heritable (Bakker 1986; Dingemanse et al. 2009; Dochtermann et al. 2014). As a consequence, personality variation may retard or accelerate rates of microevolution and population divergences (Wagner and Altenberg 1996; Wolf and Weissing 2012; Dochtermann and Dingemanse 2013; van Gestel and Weissing 2018). Here, we aim to study whether genetic differentiation underlies the rapid behavioral differentiation following habitat fragmentation. We capitalize on an unintended field experiment in the north of the Netherlands, where the construction of pumping stations in the 1970s has led to the forced residency of replicate populations of anadromous three-spined sticklebacks (Gasterosteus aculeatus). A previous study in this system has revealed extensive phenotypic differentiation (morphology and behavior) between the ancestral “migrant” and its derived “resident” populations (Ramesh et al. 2021). Compared to migrants, wild-caught residents are smaller, more active and aggressive, more exploratory, and bolder and showed reduced shoaling and migratory tendencies (Ramesh et al. 2021; see also Bakker 1994). These differences parallel the behavioral divergence reported between freshwater and marine populations of sticklebacks over ~ 12,000 years (Di-Poi et al. 2014). However, it remains to be determined if similar behaviorally relevant genetic differentiation has evolved in our system over much shorter time scales (~ 50 years). This knowledge is important because conservation efforts are underway to reconnect the waterways, and therefore, we need to better understand the current state of fish populations in order to predict the eco-evolutionary consequences of barrier removal.
We conducted a common garden experiment to test whether genetic differentiation underlies the observed divergence in morphology and behavior. We raised F1 juveniles from four types of crosses (migrant parents (MM), resident parents (RR), hybrids with a migrant mother (RM), and hybrids with a resident mother (MR); Fig. 1a) under similar laboratory conditions and quantified variation in activity, exploration, shoaling, boldness, and migratory tendencies among these crosses. We expect that (1) if the behavioral differentiation is genetic, individuals of MM crosses will differ significantly from RR crosses (similar to their wild-caught parents, Fig. 1b); (2) if the behavioral differences between wild-caught residents and migrants are induced by differences in their environments, there will be no differences between the “common garden” crosses (Fig. 1b); and (3) if parental effects are involved, we will see asymmetric changes in the reciprocal hybrid crosses (Fig. 1b). Specifically, if behavioral variation is strongly influenced by maternal effects, the hybrids resulting from the MR cross will have a similar score as the RR cross and the hybrids resulting from the RM cross will have a similar score as the MM cross (Fig. 1b). A similar trend can be expected in the case of paternal effects, but we eliminated that possibility to a large extent by raising juveniles without paternal care (Giesing et al. 2011; McGhee and Bell 2014; Heckwolf et al. 2018).
a Schematic of breeding design. We obtained four F1 crosses—migrant male × migrant female (MM), resident male × resident female (RR), migrant male × resident female (MR), and resident male × migrant female (RM). b Expectations of mean behavioral scores (e.g., shoaling) if the underlying basis for behavioral differentiation in wild-caught parents is due to genetic differentiation, environmental experiences during development or through maternal effects (letters of migrant and resident female in the maternal effects prediction plot are colored according to the origin for ease of interpretation of patterns in hybrids, when they are under the control of maternal effects. The expected mean value of hybrids would correspond to the migrant or resident status of the female)
Methods
Study populations
The waterways in the Netherlands consist of rivers and canals that are open to the sea and of land-locked smaller ditches (< 1-m deep) located inside polders. We caught incoming migrants at two sea locks (“TER” (53°18′7.24″, 7°2′17.11″) and “NSTZ” (53°13′54.49″, 7°12′30.99″)), whereas residents were caught in two land-locked polders (“LL-A” (53°17′56.14″, 7°2′1.28″) and “LL-B” (53°17′16.52″, 7°2′26.46″)) (Ramesh et al. 2021). Sticklebacks were caught over a period of 4 weeks between March and April in 2019. All individuals were transported to the laboratory within 2 h of capture in aerated bags (5–6 fish/3-L bag). They were housed outdoors separated by their origin in groups of five fish in 50-L aerated tanks filled with freshwater, exposed to the natural day-light cycles and temperatures. They were fed brine ad libitum with brine shrimps and blood worms (3F Frozen Fish Food company). Males were separated once they reached breeding colors, and females were checked daily for signs of gravidity.
Lab-bred F1 juveniles
Lab-bred F1 juveniles of resident, migrant, and hybrid sticklebacks arose from a partial factorial breeding design (Fig. 1a) using three resident males, three resident females, three migrant males, and three migrant females (six migrants from “NSTZ,” five residents from “LL-A,” and one resident female from “LL-B”). Each family consisted of all combinations of crosses between a male and female migrant and male and female resident, leading to F1 offspring of different crosses: pure migrant (MM) or resident (RR) and hybrids with migrant father and resident mother (MR) and vice versa (RM). From the offspring pool, a total of 40 fish were used per cross for the experiment, with each cross containing at least five fish from each family.
For obtaining F1 juveniles, we followed a split-clutch in vitro fertilization protocol, where eggs of ripe females were stripped, then weighed and split into two halves for artificial insemination with sperm extracted from freshly euthanized migrant and resident fathers respectively (Barber and Arnott 2000). All offspring were raised without paternal care to prevent undesired long-lasting effects of father on offspring behavior (McGhee and Bell 2014). The larvae hatched 5 to 7 days after fertilization and started maintaining buoyancy and independent feeding one week after hatching. The fish larvae were fed a mixture of frozen cyclops, freshly hatched Artemia nauplii, and zebrafish diet (GEMMA Micro 75, Skretting, Tooele, Utah) daily. The densities never exceeded 40 fish larvae in 5 liter “home-tanks” (30 × 16 × 18 cm (L × W × H)). Once fish reached ~ 2 cm, they were isolated, assigning ten random individuals from the same family into separate home tanks. After this, the individuals were fed ad libitum with brine shrimps and blood worms (3F Frozen Fish Food company), and tanks were connected to the same water system at 16 °C. The photoperiod was set at 16:8 (L:D), mimicking summer conditions during juvenile growth. When the fish reached a length of ~ 4 cm, they received a unique identification (see below). We induced autumn conditions when the fish were ~ 12–13 months old, characterized by 12:12 (L:D) photoperiod and temperatures being lowered to 13–14 °C. All fish were in non-breeding conditions and kept in autumn conditions during the period of experimentation. Experimentation started when fish were ~ 15–16 months old.
Individual identification
When the juveniles reached 4 cm length (~ 12 months), we used clipped spines or injection of an 8-mm Passive Integrated Transponder (PIT tag; Trovan, Ltd., Santa Barbara, California) for unique individual identification. We used PIT tag injection only for half of the tested fish (20 fish × 4 crosses = 80 fish), while the rest were tagged using a combination of dorsal and pelvic spine clipping (20 fish × 4 crosses = 80 fish). This was because PIT tag retention was low in these fish (~ 15% loss in the first week after tagging), and we did not retag the fish to prevent excess handling. PIT tags were injected in the abdominal cavity and under anesthesia following the standard protocol (following Cousin et al. 2012). During tagging/clipping, we also measured weight and standard length (the length from the tip of the snout to the base of the tail) as a proxy for size. Lateral plates were not very clearly visible in juvenile fish and hence were not measured. After individual tagging, we mixed juveniles from different families to be housed together in groups of ten in their home tanks while keeping them together with the same cross (MM, RR, MR or RM).
Large-scale movement tendencies in mesocosm (migratory tendencies)
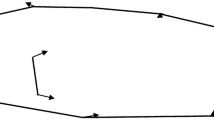
For the subset of PIT tagged fish, movement assays were performed in semi-natural mesocosms before subjecting them to the lab-based tests. The mesocosm system consisted of five connected outdoor ponds of diameter 1.6 m connected by four pipes of length ~ 1.5 m and diameter 11 cm, filled with water from a nearby freshwater ditch, with a linear flow in the system of connected ponds similar to those typically experienced in the canals and ditches (flow speed < 0.7 cm/s) (Fig. 2a). This was done to create a cue for migration-like movement. All connecting tubes were fitted with circular PIT antennas around the entrance and exit of each pond to record fish movement between ponds. The sticklebacks were tested in pond experiments after at least 1 week of recovery from tagging. A group of ten fish of one cross (MM, RR, MR, or RM) was introduced in the first pond and acclimatized for 5 h in the first morning, after which the connection to the rest of the ponds was opened. We then recorded the movement of fish as the number of crossings between ponds for the next 16 h (~ 4 pm–8 am). We attempted to have 20 tagged fish/cross and tested them in groups of ten each, making it two groups/cross. However, due to tag loss, we ended up with < 20 fish/cross. Instead of changing group size, which could have an effect on behavior, we decided to spread the final number of tagged fish between two groups and supplement the remaining with untagged fish from the same cross to make up to ten. In total, two groups, each from a randomly chosen cross, were tested, making a total of eight groups with 56 fish (NMM = 12, NMR = 17, NRM = 15, NRR = 12). In groups with less than ten tagged individuals, untagged fish from the same cross were added to maintain constant group size.
Schematic of behavioral assays. a Migration tendencies were tested in a linear setup of five connected pond mesocosm with groups of 10 fish. There is water flow (rate < 0.7 cm/s). PIT antennas are present at both ends of the corridors connecting the ponds. b Lab assays were performed in the following order: activity (day 1), exploration (day 1), shoaling (day 2), boldness (day 3)
Lab behavior assays
Three days before testing, fish were selected randomly and acclimatized in visually separated and isolated tanks, same as their home tanks, at an ambient temperature of 19 °C. We attempted to test 40 fish/cross, but some fish were lost due to mortality. Hence, in total, 154 fish (NMM = 40, NMR = 39, NRM = 35, NRR = 40) were randomly selected for testing and split into four batches. One round of testing consisted of four batches of approximately ten fish of each cross and lasted 1 week where we assayed activity, exploration, shoaling, and boldness in that order (Fig. 2b). Overall, roughly 40 fish were tested each week. The interval between the first and the second rounds of testing of each individual was thus at least 4 weeks. Fish were returned to their home tanks between the testing rounds. The sample sizes for the second round were lower (N = 151) due to mortality between the two rounds (NMM = 39, NMR = 38, NRM = 34, NRR = 40). All lab assays were filmed from the top using a Raspberry Pi camera (Raspberry Pi NoIR Camera Board V2 – 8MP, Raspberry Pi Foundation, UK) in tanks placed in illuminated wooden boxes to prevent external disturbance. Behavioral assays were conducted in fixed order as below, and videos were analyzed using EthovisionXT (Noldus Information Technology company). In all tests, observers were blind with respect to the cross to which the test fish belonged and further bias was reduced by analyzing the videos using automated video tracking techniques.
Activity (day 1)
Activity of the fish was measured as the total distance the fish swam in a tank identical to its home tank during a total of 20 min (with 5 min for acclimatization).
Exploration (day 1)
Just after activity was recorded, the fish was isolated to one corner of the tank using a sheet partition, and the setup in the tank was changed. Five stone pillars extending above the water’s surface were added in a specific position, forcing the fish to move around them. After 5 min, the sheet was removed remotely without opening the box, and the fish was recorded in this novel arena for 20 min. The total distance travelled by the fish in this novel environment was used as a proxy for exploratory tendency of fish as it highly correlates with space use (Ramesh et al. 2021).
Shoaling (day 2)
For the shoaling assay, a larger tank (60 × 30 × 30 cm) was filled with water up to 10-cm height. The tank was divided into three compartments: the central testing arena where the focal fish was released and two end compartments containing the stimulus shoal (N = 10 unfamiliar conspecifics of mixed crosses), and the distractor fish (N = 2 unfamiliar conspecifics) (adapted from Wark et al. 2011). The position of the distractor and shoal fish compartments was switched to prevent biases and replaced with new distractor and shoal fish every seven tests. At the start of the test, the focal fish was allowed to acclimatize for 5 min in the central arena without viewing the end compartments which were covered with opaque barriers. Then, the opaque barriers were lifted remotely from outside the box, and the response of the focal fish was recorded for the next 20 min. The water was refreshed after testing seven fish in the arena. In total, we had four groups of shoal fish and five pairs of distractor fish, which were randomly used to avoid biases. The proportion of time the focal fish spent within one-fish distance (6 cm) from the side containing the stimulus shoal was used as a proxy for shoaling.
Boldness (day 3)
In the boldness tests, we measured the responses of the focal fish toward visual cue of a European perch (Perca fluviatilis) (model with soft body, Kozak and Boughman 2012) and olfactory predation cues (50 mL of water from freshly dead sticklebacks mixed with 50 mL of water containing live perch scent, Sanogo et al. 2011). The focal fish was moved from its home tank into a bigger, novel tank (60 × 30 × 30 cm) with three compartments filled with 10 cm of water. The predator model was randomly presented in one of the end compartments, while the focal fish was acclimatized in the other end compartment (Kozak and Boughman 2012). After 5 min of acclimatization, the fish was released remotely into the arena with view of the predator model, and the assay lasted for 20 min. We changed the side of predator compartment systematically in order to avoid biases. Further, the water was refreshed and new predatory olfactory cues were added after testing seven fish in the arena. The proportion of time the focal fish spent within one-fish distance (6 cm) from the predator compartment was taken as a proxy for boldness.
Statistical analyses
Variation in size and behaviors (activity, exploration, shoaling, and boldness) was analyzed using linear mixed models (LMM) in which repeat (first vs. second round) and cross identity (MM, MR, RM or RR) were included as fixed factors. We also included the interactive effects (cross x round) to test for cross-specific habituation effects. Individual identity (Fish ID), mother identity (Mother ID), and father identity (Father ID) were included as random effects. For shoaling behavior, we added identity of the test shoal (Shoal ID) as an additional random effect. For migratory tendencies, only one round of tests was performed and we fitted a Poisson generalized linear mixed model with log-link function (GLMM), with number of pond crosses as the response variable and cross identity as fixed factor. As random effects, we included mother identity (Mother ID) and father identity (Father ID), and further, to prevent overestimation of predictive power caused due to overdispersion, we added observation level random effects (OLRE) (Harrison 2014).
All LMMs/GLMMs were constructed in R v. 3.6.1, R Core Team (2019) using the “lmer” function of the “lme4” package, package version 1.1-27.1 (Bates et al. 2015). The statistical significance of fixed effects was assessed based on the 95% confidence interval (CI): an effect was considered significant when its 95% CI did not include zero. In addition, Tukey’s HSD post hoc test was performed using the functions “emmeans” and “pairs” to give pairwise comparisons using the package “emmeans,” package version 1.6.1 (Lenth 2020). LMMs were used to decompose the phenotypic variance of behaviors into between-individual (VFish ID), between-mother (VMother ID), between-father (VFather ID), and within-individual (VResidual) variances that we subsequently used to calculate repeatabilities, i.e., the proportion of total phenotypic variation (Vp) attributable to differences between individuals (RFish ID), between mothers (RMother ID), and between father (RFather ID):
Raw (without fixed effects), adjusted repeatabilities (after accounting for fixed effects, cross x round), and their confidence intervals were calculated using “rpt” function with 1000 bootstraps in “rptR,” package version 0.9.22 (Stoffel et al. 2017).
Results
Our prime goal was to test if RR and MM crosses that were raised under similar conditions, exhibited similar behavioral differences as observed in their wild-caught population of origin and if these differences were consistent over time. We found that RR crosses were consistently less active than MM crosses in the two rounds (Fig. 3a; Table 1; overall effect of crosses on activity: χ2 = 17.35, df = 3, p < 0.01, Supp. Table 1). We further found that shoaling and migratory tendencies varied significantly and consistently between RR and MM crosses in the same direction, with RR crosses exhibiting lower shoaling and migratory tendencies than MM crosses (Fig. 3c, e; Table 1; overall effect of crosses on shoaling: χ2 = 17.91, df = 3, p < 0.01, on migratory tendency: χ2 = 14.37, df = 3, p < 0.01). MM but not RR crosses shoaled more than expected by chance (score > 0.5) (Table 1). RR and MM crosses did not differ consistently in levels of exploration and boldness (Fig. 3b, d; Table 1, Supp. Table 1). For boldness, RR cross differed from MM cross but only in round 2 (Fig. 3d, significant effects of round and round x cross RR, Table 1, Supp. Table 1), implying that the observed difference was not consistent over time (Fig. 3d). Crosses did not differ in body size (Fig. 3f, Table 1, Supp. Table 1).
Mean scores and standard errors for behaviors and size of F1 fish of different crosses. a “Activity,” total distance travelled in meters (m); b “exploration,” total distance travelled in a novel arena in m; c “shoaling,” proportion of time spent near shoal compartment; d “boldness,” proportion of time spent near predator. For lab-based behaviors, the mean behavioral scores for the two repeats are represented separately (sample sizes round 1, NMM = 40, NMR = 39, NRM = 35, NRR = 40; round 2, NMM = 39, NMR = 38, NRM = 34, NRR = 40); e “migratory tendency,” total number of pond crosses (NMM = 12, NMR = 17, NRM = 15, NRR = 12); f “size,” standard length in mm (NMM = 40, NMR = 39, NRM = 35, NRR = 40)
We did not find evidence for parental effects. For all traits investigated, we did not observe a clear directional asymmetry between the reciprocal hybrid crosses or trends in the distribution of individual behavior (Fig. 3, Supp. Fig. 2). Overall, only a small fraction of the variance in behaviors was attributable to differences between fathers and mothers (between 0 and 0.18, Table 2). In contrast, individual identity explained a significant part of the behavioral variation across the two rounds of measurement (adjusted Rind = 0.31 to 0.38; raw Rind = 0.14 to 0.43) (Supp. Fig. 1, Table 2), i.e., individual behavior is consistent (to a certain extent), despite potential effects of habituation or sensitization to handling (Fig. 3, Table 1).
Discussion
We aimed to study whether genetic differentiation underlies the behavioral differentiation following habitat fragmentation in sticklebacks. Using a common garden experiment, we showed that the differences between residents and migrants in shoaling and migration tendency (and to some extent also activity) have a genetic basis. In contrast, there were no clear patterns regarding differences in other behaviors or size between crosses. The earlier observed differences in these traits between wild-caught residents and migrants might therefore reflect differences in the respective developmental environments of the two ecotypes of fish. We discuss below the likely causes of divergence in our system and compare the patterns to those observed in post-glacial divergence of marine and freshwater sticklebacks. Then, we discuss the eco-evolutionary implications of our findings in link with conservation plans of our study area.
Our common garden experiment revealed that the divergence in at least two of the five behavioral traits studied have a genetic basis. This corroborates a previous study on sticklebacks showing that the expression of heritable variation, i.e., the fraction of phenotypic variance owing to additive effects of genes (Lynch and Walsh 1998), substantially varied depending on the personality trait considered and the evolutionary history of the populations (Dingemanse et al. 2009). An interesting future avenue will be to quantify population specific trait heritabilities and the relative contribution of genetic and non-genetic sources of variation in those behaviors. Furthermore, it remains to be tested if the genetic differences we uncovered reflect local adaptation as opposed to other processes such as genetic drift or founder effects. Shoaling and migration tendencies are very crucial for the ancestral migratory fish. Their migratory lifestyle involves group schooling tendencies and potentially higher shoaling tendencies due to increased predator pressure owing to “openness” of habitats in the sea. In residents, shoaling tendencies may be less strongly selected for, leading to the pattern of random association with the shoal that we have recovered in our experiments (Fig. 3c). Alternatively, lower shoaling tendencies may be selected for due to increased competition, for instance, in winter, when resources are scarce leading to a trade-off between intra-specific aggression and competition (Lacasse and Aubin-Horth 2014). Studies on marine-freshwater stickleback pairs have also revealed potential genetic underpinnings of shoaling via eda gene (freshwater sticklebacks shoaled less and schooled less efficiently than migrants; Wark et al. 2011; Di-Poi et al. 2014; Archambeault et al. 2020) and migratory tendencies via genetic divergence in thyroxine response mechanisms (Kitano et al. 2010). One next step will be to test whether the genetic differentiation of shoaling and migratory tendencies reflect local adaptation using either a genomic approach to detect signature of adaptive divergence (using, e.g., a whole genome and/or a candidate gene (eda allele) approach) or a transplant experiment where we would raise crosses in different environmental conditions (marine vs freshwater) to infer fitness.
We expected similar differentiation in other traits, as they were found to be different between wild-caught migrants and residents over two study years (Ramesh et al. 2021). For instance, studies have shown moderately heritable and additive genetic components in behaviors such as exploration and boldness in sticklebacks (Dingemanse et al. 2009). However, in our experiment, body size and behaviors such as exploration and boldness did not show differences between crosses. For body size, responses may be potentially plastically adjusted to the ecological conditions as seen in previous studies (e.g., predation pressure; Frommen et al. 2011; niche specialization, Day and McPhail 1996; Wund et al. 2008). Similar to body size, behaviors such as exploration and boldness may also be environmentally determined. Alternatively, these behaviors could also be state-dependent (state being size or mass in this case), owing to differences in resource availability during growth of migrants and residents (Luttbeg and Sih 2010; Wolf and Weissing 2010). It also remains possible that the differences in behavior in wild migrants are due to plastic responses of migrants in freshwater vs sea conditions, which have not been tested here.
In our current study, we found little evidence for maternal effects as maternal contribution to trait variation was small and not significant (19% for activity, 3% for exploration, and 3% for shoaling tendencies) and we did not find clear systematic differences between the reciprocal hybrid crosses (RM and MR). However, we raised the juveniles in the absence of paternal care. Hence, it remains possible that the behavioral differences observed between wild-caught migrants and residents (Ramesh et al. 2021) are related to differences in paternal care. This is an interesting avenue warranting further investigation because there is evidence for parental programming through maternal effects and paternal care in sticklebacks (Giesing et al. 2011; McGhee et al. 2012, 2015; McGhee and Bell 2014; Mommer and Bell 2014; Stein and Bell 2014).
Our studies revealing genetic differentiation between ancestral migrant and resident populations in behaviors related to migration and shoaling are timely and have important consequences for conservation efforts. Water authorities are currently implementing conservation measures which aim at restoring river connectivity via barrier removal or the construction of fishways. Reconnecting migratory and genetically differentiated land-locked populations can be viewed as a large scale eco-evolutionary experiment that raises exciting questions such as the following: will migratory and resident sticklebacks intermix and introgress in sympatry (Ravinet et al. 2021)?; Will hybrids be selected against?; or Will we have incomplete gene flow and partial migration occurring in these populations (Berner et al. 2011; Ingram et al. 2015; Hanson et al. 2016; Lackey and Boughman 2017)? From our studies, residents and hybrids show lowered migratory and shoaling tendencies. This could potentially drive divergent selection and lead to the genetic differentiation of sympatric populations with partial migration upon reconnection. Divergence may also be maintained or enhanced by size-assortative mating of migrants and residents as size difference at maturity has been detected in the wild (Ramesh et al. 2021) or by phenotype-dependent microhabitat choice (Maciejewski et al. 2020; Dean et al. 2021). Irrespective of the mechanisms involved in the observed phenotypic differentiation between migrants and residents, whether the migrant-resident ecotype divergence will persist in the absence of migration barriers, needs to be investigated.
Overall, using a common garden experiment, we found evidence for genetic differentiation in shoaling, migratory tendencies, and potentially activity. These results suggest that residents may have locally adapted to their novel environmental conditions in our system. Few imminent questions that follow this finding are whether our results can be generalized to other freshwater and migratory fish species that have undergone isolation and how conservation plans may be affected (Tuomainen and Candolin 2011; Franssen et al. 2013). Conservation methods indeed should not only aim at restoring the ecosystem to its original state because this may lead to unwanted consequences (Stockwell et al. 2003). For example, reversal of responses to restorations may not be possible if newly adapted populations or species lack genetic variation, leading to a rapid population decline after conservation measures are in place (Lahti et al. 2009; Mable 2019). Alternatively, newly adapted populations or species may, in fact, have selected for invasive phenotypes such as novel foraging tactics and increased aggression and boldness, leading to unwanted expansions causing unpredictable effects on other species and communities (Holway and Suarez 1999; Sol et al. 2002). Hence, conservation efforts should be aimed at implementing methods taking an informed approach of the current state of the system and assessing the evolutionary changes undergone in the species assemblages they are aimed at.
Data availability
The final processed data used for the figures and analyses of this study are made available as supplementary material.
References
Archambeault SL, Bärtschi LR, Merminod AD, Peichel CL (2020) Adaptation via pleiotropy and linkage: association mapping reveals a complex genetic architecture within the stickleback Eda locus. Evol Lett 4:282–301
Bakker TCM (1986) Aggressiveness in sticklebacks (Gasterosteus aculeatus L.): a behaviour-genetic study. Behaviour 98:1–144
Bakker TCM (1994) Evolution of aggressive behaviour in the threespine stickleback. In: Bell MA, Foster SA (eds) The Evolutionary Biology of the Threespine Stickleback. Oxford University Press, Oxford, pp 345–380
Barber I, Arnott SA (2000) Split-clutch IVF: A technique to examine indirect fitness consequences of mate preferences in sticklebacks. Behaviour 137:1129–1140
Bates D, Mächler M, Bolker BM, Walker SC (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48
Berner D, Kaeuffer R, Grandchamp AC, Raeymaekers JAM, Räsänen K, Hendry AP (2011) Quantitative genetic inheritance of morphological divergence in a lake-stream stickleback ecotype pair: implications for reproductive isolation. J Evol Biol 24:1975–1983
Bohlin T, Pettersson J, Degerman E (2001) Population density of migratory and resident brown trout (Salmo trutta) in relation to altitude: evidence for a migration cost. J Anim Ecol 70:112–121
Botero CA, Weissing FJ, Wright J, Rubenstein DR (2015) Evolutionary tipping points in the capacity to adapt to environmental change. Proc Natl Acad Sci USA 112:184–189
Cousin X, Daouk T, Péan S, Lyphout L, Schwartz M, Bégout M (2012) Electronic individual identification of zebrafish using radio frequency identification (RFID) microtags. J Exp Biol 215:2729–2734
Day T, McPhail JD (1996) The effect of behavioural and morphological plasticity on foraging efficiency in the threespine stickleback (Gasterosteus sp.). Oecologia 108:380–388
Dean LL, Dunstan HR, Reddish A, MacColl ADC (2021) Courtship behavior, nesting microhabitat, and assortative mating in sympatric stickleback species pairs. Ecol Evol 11:1741–1755
Dingemanse N, Van der Plas F, Wright J, Réale D, Schrama M, Roff DA, Van der Zee E, Barber I (2009) Individual experience and evolutionary history of predation affect expression of heritable variation in fish personality and morphology. Proc R Soc Lond B 276:1285–1293
Di-Poi C, Lacasse J, Rogers SM, Aubin-Horth N (2014) Extensive behavioural divergence following colonisation of the freshwater environment in threespine sticklebacks. PLoS One 9:98980
Dochtermann NA, Dingemanse NJ (2013) Behavioral syndromes as evolutionary constraints. Behav Ecol 24:806–811
Dochtermann NA, Schwab T, Sih A (2014) The contribution of additive genetic variation to personality variation: Heritability of personality. Proc R Soc B 282:20142201
Edelaar P, Jovani R, Gomez-Mestre I (2017) Should I change or should I go? Phenotypic plasticity and matching habitat choice in the adaptation to environmental heterogeneity. Am Nat 190:506–520
Fahrig L (2003) Effects of habitat fragmentation on biodiversity. Annu Rev Ecol Evol Syst 34:487–515
Fischer J, Lindenmayer DB (2007) Landscape modification and habitat fragmentation: a synthesis. Glob Ecol Biogeogr 16:265–280
Foley JA, DeFries R, Asner GP et al (2005) Global consequences of land use. Science 309:570–574
Franssen NR, Harris J, Clark SR, Schaefer JF, Stewart LK (2013) Shared and unique morphological responses of stream fishes to anthropogenic habitat alteration. Proc R Soc B 280:20122715
Frommen JG, Herder F, Engqvist L, Mehlis M, Bakker TCM, Schwarzer J, Thünken T (2011) Costly plastic morphological responses to predator specific odour cues in three-spined sticklebacks (Gasterosteus aculeatus). Evol Ecol 25:641–656
Ghalambor CK, McKay JK, Carroll SP, Reznick DN (2007) Adaptive versus non-adaptive phenotypic plasticity and the potential for contemporary adaptation in new environments. Funct Ecol 21:394–407
Giesing ER, Suski CD, Warner RE, Bell AM (2011) Female sticklebacks transfer information via eggs: Effects of maternal experience with predators on offspring. Proc R Soc Lond B 278:1753–1759
Hanson D, Moore JS, Taylor EB, Barrett RDH, Hendry AP (2016) Assessing reproductive isolation using a contact zone between parapatric lake-stream stickleback ecotypes. J Evol Biol 29:2491–2501
Harrison XA (2014) Using observation-level randomeffects to model overdispersion in count data in ecology and evolution. PeerJ 2014:e616
Heckwolf MJ, Meyer BS, Döring T, Eizaguirre C, Reusch TBH (2018) Transgenerational plasticity and selection shape the adaptive potential of sticklebacks to salinity change. Evol Appl 11:1873–1885
Hedrick PW (2006) Genetic polymorphism in heterogeneous environments: the age of genomics. Annu Rev Ecol Evol Syst 37:67–93
Hedrick PW, Ginevan ME, Ewing EP (1976) Genetic polymorphism in heterogeneous environments. Annu Rev Ecol Syst 7:1–32
Holway DA, Suarez AV (1999) Animal behavior: an essential component of invasion biology. Trends Ecol Evol 14:328–330
Ingram T, Jiang Y, Rangel R, Bolnick DI (2015) Widespread positive but weak assortative mating by diet within stickleback populations. Ecol Evol 5:3352–3363
Junge C, Museth J, Hindar K, Kraabøl M, Vøllestad LA (2014) Assessing the consequences of habitat fragmentation for two migratory salmonid fishes. Aquat Conserv Mar Freshw Ecosyst 24:297–311
Kawecki TJ, Ebert D (2004) Conceptual issues in local adaptation. Ecol Lett 7:1225–1241
Kitano J, Lema SC, Luckenbach JA, Mori S, Kawagishi Y, Kusakabe M, Swanson P, Peichel CL (2010) Adaptive divergence in the thyroid hormone signaling pathway in the stickleback radiation. Curr Biol 20:2124–2130
Kozak GM, Boughman JW (2012) Plastic responses to parents and predators lead to divergent shoaling behaviour in sticklebacks. J Evol Biol 25:759–769
Kraabøl M, Johnsen SI, Museth J, Sandlund OT (2009) Conserving iteroparous fish stocks in regulated rivers: the need for a broader perspective! Fish Manag Ecol 16:337–340
Lacasse J, Aubin-Horth N (2014) Population-dependent conflict between individual sociability and aggressiveness. Anim Behav 87:53–57
Lackey ACR, Boughman JW (2017) Evolution of reproductive isolation in stickleback fish. Evolution 71:357–372
Lahti DC, Johnson NA, Ajie BC, Otto SP, Hendry AP, Blumstein DT, Coss RG, Donohue K, Foster SA (2009) Relaxed selection in the wild. Trends Ecol Evol 24:487–496
Legrand D, Cote J, Fronhofer EA, Holt RD, Ronce O, Schtickzelle N, Travis JMJ, Colbert J (2017) Eco-evolutionary dynamics in fragmented landscapes. Ecography 40:9–25
Lenth R (2020) Emmeans: estimated marginal means, aka least-squares means. https://CRAN.R-project.org/package=emmeans. Accessed 1 Apr 2021
Luttbeg B, Sih A (2010) Risk, resources and state-dependent adaptive behavioural syndromes. Philos Trans R Soc B 365:3977–3990
Lynch M, Walsh B (1998) Genetics and Analysis of Quantitative Traits. Sinauer Associates, Sunderland, MA
Mable BK (2019) Conservation of adaptive potential and functional diversity: integrating old and new approaches. Conserv Genet 20:89–100
Maciejewski MF, Jiang C, Stuart YE, Bolnick DI (2020) Microhabitat contributes to microgeographic divergence in threespine stickleback. Evolution 74:749–763
McGhee KE, Bell AM (2014) Paternal care in a fish: Epigenetics and fitness enhancing effects on offspring anxiety. Proc R Soc B 281:20141146
McGhee KE, Pintor LM, Suhr EL, Bell AM (2012) Maternal exposure to predation risk decreases offspring antipredator behaviour and survival in threespined stickleback. Funct Ecol 26:932–940
McGhee KE, Feng S, Leasure S, Bell AM (2015) A female’s past experience with predators affects male courtship and the care her offspring will receive from their father. Proc R Soc B 282:20151840
Mommer BC, Bell AM (2014) Maternal experience with predation risk influences genome-wide embryonic gene expression in threespined sticklebacks (Gasterosteus aculeatus). PLoS One 9:e98564
Nicolaus M, Edelaar P (2018) Comparing the consequences of natural selection, adaptive phenotypic plasticity, and matching habitat choice for phenotype–environment matching, population genetic structure, and reproductive isolation in meta-populations. Ecol Evol 8:3815–3827
Nussey DH, Wilson AJ, Brommer JE (2007) The evolutionary ecology of individual phenotypic plasticity in wild populations. J Evol Biol 20:831–844
Piersma T, Drent J (2003) Phenotypic flexibility and the evolution of organismal design. Trends Ecol Evol 18:228–233
Pigliucci M (2005) Evolution of phenotypic plasticity: Where are we going now? Trends Ecol Evol 20:481–486
Ramesh A, Groothuis TGG, Weissing FJ, Nicolaus M (2021) Habitat fragmentation induces rapid divergence of migratory and isolated sticklebacks. Beh Ecol XX(XX):1–11. https://doi.org/10.1093/beheco/arab121
Ravinet M, Kume M, Ishikawa A, Kitano J (2021) Patterns of genomic divergence and introgression between Japanese stickleback species with overlapping breeding habitats. J Evol Biol 4:114–127
Réale D, Reader SM, Sol D, McDougall PT, Dingemanse NJ (2007) Integrating animal temperament within ecology and evolution. Biol Rev 82:291–318
Sanogo YO, Hankison S, Band M, Obregon A, Bell AM (2011) Brain transcriptomic response of threespine sticklebacks to cues of a predator. Brain Behav Evol 77:270–285
Scheiner SM (1993) Plasticity as a selectable trait: reply to Via. Am Nat 142:371–373
Sol D, Timmermans S, Lefebvre L (2002) Behavioural flexibility and invasion success in birds. Anim Behav 63:495–502
Stamps J, Groothuis TGG (2010) The development of animal personality: relevance, concepts and perspectives. Biol Rev 85:301–325
Stein LR, Bell AM (2014) Paternal programming in sticklebacks. Anim Behav 95:165–171
Stockwell CA, Hendry AP, Kinnison MT (2003) Contemporary evolution meets conservation biology. Trends Ecol Evol 18:94–101
Stoffel MA, Nakagawa S, Schielzeth H (2017) rptR: repeatability estimation and variance decomposition by generalized linear mixed-effects models. Methods Ecol Evol 8:1639–1644
Tuomainen U, Candolin U (2011) Behavioural responses to human-induced environmental change. Biol Rev 86:640–657
van Gestel J, Weissing FJ (2018) Is plasticity caused by single genes? Nature 555:E19–E20
Via S, Gomulkiewicz R, De Jong G, Scheiner SM, Schlichting CD, Van Tienderen PH (1995) Adaptive phenotypic plasticity: consensus and controversy. Trends Ecol Evol 10:212–217
Wagner GP, Altenberg L (1996) Perspective: complex adaptations and the evolution of evolvability. Evolution 50:967–976
Wang IJ, Bradburd GS (2014) Isolation by environment. Mol Ecol 23:5649–5662
Wark AR, Greenwood AK, Taylor EM, Yoshida K, Peichel CL (2011) Heritable differences in schooling behavior among threespine stickleback populations revealed by a novel assay. PLoS One 6:e18316
Wolf M, Weissing FJ (2010) An explanatory framework for adaptive personality differences. Philos Trans R Soc B 365:3959–3968
Wolf M, Weissing FJ (2012) Animal personalities: Consequences for ecology and evolution. Trends Ecol Evol 27:452–461
Wong BBM, Candolin U (2015) Behavioral responses to changing environments. Behav Ecol 26:665–673
Wund MA, Baker JA, Clancy B, Golub JL, Foster SA (2008) A test of the “flexible stem” model of evolution: ancestral plasticity, genetic accommodation, and morphological divergence in the threespine stickleback radiation. Am Nat 172:449–462
Acknowledgments
We thank Dennis de Worst and Willem Diederik for help with fish care and advice on experimental design. We thank Peter Paul Schollema, at the Water Authorities Hunze en Aa’s and Jeroen Huisman at van Hall Larenstein, University of Applied Sciences, for help with acquiring samples of sticklebacks. We thank Jakob Gismann, Jana Riederer, Raphaël Scherrer, Kevin Kort, and Albertas Janulevicius for their useful comments on the manuscript. We also thank the two reviewers, who provided excellent comments and suggestions that improved the manuscript.
Funding
This work is supported by PhD fellowship of the Adaptive Life program of the University of Groningen to AR, by funding from European Research Council to FJW (ERC Advanced Grant No. 789240) and from the Netherlands Organization for Scientific Research to FJW and MN (NWO-ALW; ALWOP.668). This work was also supported by grants from the Gratama Foundation to AR (2020GR040), the Dr. J.L. Dobberke Foundation (KNAWWF/3391/1911), and the Waddenfonds - Ruim Baan voor Vissen 2 (01755849/WF-2019/200914).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Sampling of wild animals and handling methods were done following a fishing permit from Rijksdienst voor Ondernemend Nederland (the Netherlands) and an angling permit from the Hengelsportfederatie Groningen-Drenthe. Animal housing and behavioral tests adhered to the project permit from the Centrale Commissie Dierproeven (the Netherlands) under the license number AVD1050020174084. All methods were carried out under the applicable international, national, and institutional guidelines for the use of animals (Art. 9, Wet op de Dierproeven & European directive 2010/63/EU).
Consent for publication
All authors have given their consent for publication.
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by J. Lindström.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is a contribution to the Topical Collection Using behavioral ecology to explore adaptive responses to anthropogenic change – Guest Editors: Jan Lindström, Constantino Macias Garcia, Caitlin Gabor
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ramesh, A., Domingues, M.M., Stamhuis, E.J. et al. Does genetic differentiation underlie behavioral divergence in response to migration barriers in sticklebacks? A common garden experiment. Behav Ecol Sociobiol 75, 161 (2021). https://doi.org/10.1007/s00265-021-03097-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00265-021-03097-y