Abstract
Objective
Most recurrent glioblastoma (rGBM) patients do not benefit from immune checkpoint inhibition, emphasizing the necessity for response biomarkers. This study evaluates whether tumor in situ fluid (TISF) circulating tumor DNA (ctDNA) could serve as a biomarker for response to low-dose bevacizumab (Bev) plus anti-PD-1 therapy in rGBM patients, aiming to enhance systemic responses to immunotherapy.
Methods
In this phase II trial, 32 GBM patients with first recurrence after standard therapy were enrolled and then received tislelizumab plus low-dose Bev each cycle. TISF samples were analyzed for ctDNA using a 551-gene panel before each treatment.
Results
The median progression-free survival (mPFS) and overall survival (mOS) were 8.2 months (95% CI, 5.2–11.1) and 14.3 months (95% CI, 6.5–22.1), respectively. The 12-month OS was 43.8%, and the objective response rate was 56.3%. Patients with more than 20% reduction in the mutant allele fraction and tumor mutational burden after treatment were significantly associated with better prognosis compared to baseline TISF-ctDNA. Among detectable gene mutations, patients with MUC16 mutation, EGFR mutation & amplification, SRSF2 amplification, and H3F3B amplification were significantly associated with worse prognosis.
Conclusions
Low-dose Bev plus anti-PD-1 therapy significantly improves OS in rGBM patients, offering guiding significance for future individualized treatment strategies. TISF-ctDNA can monitor rGBM patients' response to combination therapy and guide treatment.
Clinical trial registration
This trial is registered with ClinicalTrials.gov, NCT05540275.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma (GBM) is the most malignant primary brain tumor and is prone to recurrence. Despite multidisciplinary treatments including surgery, radiotherapy, chemotherapy, targeted therapy, and supportive care, the overall prognosis remains poor [1,2,3,4]. Updated guidelines for the management of gliomas still encourage clinical trials for rGBM due to the limited efficacy of available salvage therapies at the time of tumor recurrence, with a median survival of only 6–8 month [5]. The search for novel therapeutic options to improve the prognosis of rGBM patients is ongoing, with research focusing on combining antiangiogenic agents with immunotherapy to enhance antitumor immune responses [6, 7].
Preliminary results have shown that immune checkpoint inhibitors combined with anti-angiogenesis drugs have a good safety profile in treating rGBM [8, 9]. Non-clinical studies have demonstrated that bevacizumab, an antiangiogenic targeted agent, can inhibit vascular endothelial growth factor, promote tumor vascular normalization, increase T cell infiltration, and reduce immunosuppressive cell activity, thereby improving immunotherapy efficacy [10]. Although bevacizumab combined with immunotherapy has been feasible and safe in treating other solid tumors, it has not improved OS of rGBM patients [11]. A complete response to concurrent anti-PD-1 and low-dose anti-VEGF therapy was reported in one patient with rGBM [12]. Therefore, larger clinical trials are needed to investigate whether low-dose Bev can promote immunotherapy responses.
Biomarkers are critical to maximizing therapeutic efficacy and minimizing toxicity in rGBM treated with low-dose Bev plus anti-PD-1 therapy [13,14,15,16]. Analyzing circulating tumor DNA (ctDNA) as an emerging biomarker in solid tumors faces technical challenges due to the specificity of GBM's location [17, 18]. The collection of TISF for ctDNA analysis has been reported by our research group multiple times, yet there is limited literature on ctDNA changes in rGBM after immunotherapy combined with low-dose Bev treatment [17,18,19,20]. Thus, the feasibility of ctDNA as a biomarker in rGBM patients needs further investigation.
Combining tislelizumab with low-dose Bev in treating rGBM, we hypothesized that low-dose Bev treatment might improve the immunotherapy response. To assess ctDNA's efficacy for monitoring rGBM patients' response to combination therapy, we collected TISF samples at baseline and each subsequent immunotherapy cycle.
Methods
Study design and participants
This open-label phase 2 study (Clinical Trials ID: NCT 05540275) recruited rGBM patients at Zhengzhou University People's Hospital (Zhengzhou University). From March 28, 2022, patients received tislelizumab (200 mg) and bevacizumab (3 mg/kg) intravenously every 3 weeks until disease progression or intolerance. Magnetic resonance imaging (MRI) was performed at baseline and every 4–8 weeks thereafter. Tumor volume measurement and RANO 2.0 assessment were performed using 3D slicer software (National Institutes of Health, Bethesda, USA) [21].
Eligible patients were aged 18–75 years with confirmed rGBM, a Karnofsky Performance Status (KPS) ≥ 70, and had undergone ≥ 1 prior systemic GBM therapy. Exclusion criteria included systemic glucocorticoid or other immunosuppressive therapy within 7 days after enrollment, known or suspected active autoimmune disease, active hepatitis B or C, HIV infection, extracranial metastases, significant leptomeningeal disease, or tumors primarily in the brain stem or spinal cord.
Treatment Regimens
Primary GBM: Patients received concurrent chemoradiotherapy (TMZ 75 mg/m2/d for 42 days) 4 weeks after surgery, followed by TMZ (150 mg/m2/d every 4 weeks for 5 days, repeated every 28 days for 6 cycles).
Recurrent GBM: Surgery was recommended. Patients who refused surgery were given bevacizumab (5 mg/kg IV) combined with TMZ (150 mg/m2/d orally for 5 days, repeated every 21 days for 6 cycles), followed by bevacizumab (3 mg/kg) and tislelizumab (200 mg IV) every 21 days for six cycles.
Sample collection, DNA extraction, and library preparation
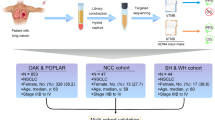
Tumor in situ fluid (TISF) samples were collected as previously described [18,19,20, 22]. A small amount of TISF (0.5–2 ml) was obtained by syringe from the implanted reservoir sac every 4 to 8 weeks (Fig. 1A). TISF is the fluid present in the local surgical cavity. ctDNA profiles from tumor tissue and TISF samples can be used to assess the dynamic evolution of the tumor in real time, while 5 ml of blood is collected as a germline DNA control.
Treatment schema and patient characteristics. A, Schematic showing the timing of treatment and tumor in situ fluid (TISF) collection. TISF was collected for ctDNA analysis pre-treatment and prior to each cycle of immunotherapy. B, The KPS scores of 32 patients with recurrent glioblastoma treated with anti-PD-1 antibody combined with low-dose bevacizumab were significantly higher than those before treatment
Genomic DNA (gDNA) and cell-free DNA (cfDNA) were extracted from fresh tissue, formalin-fixed, paraffin-embedded (FFPE) tissue, leukocytes, and TISF using kits (Kai Shuo, Thermo), according to the manufacturer's instructions. DNA was quantified using the Qubit dsDNA HS Assay Kit (Thermo, Fisher) and its quality assessed using the Agilent 4200 TapeStation (Agilent).
Library sequencing and bioinformatics analysis
Qualified DNA libraries were sequenced using the Illumina NovaSeq6000 platform (Illumina, San Diego, CA) to generate 150 bp paired-end reads. Adapter trimming and filtering of low-quality bases were performed using the software fastp (v.2.20.0). Reads were aligned to the reference genome (hg19, GRCh37 of UCSC) using BWA-MEM (v.0.7.17). Duplicate reads were removed using Dedup and Error Correct. SNVs/indels were called and annotated using VarDict (v.1.5.7) and InterVar, respectively, and screened for common SNPs from public databases (1000 Genome Project, ExAC). CNVs were analyzed using CNVkit (dx1.1) and fusion genes using factera (v1.4.4).
TMB calculation
To calculate the TMB using the 551-solid cancer-gene targeted next-generation sequencing (NGS) panel, all base substitutions and indels in the coding region of targeted genes were summed, excluding synonymous alterations, alterations with AF < 0.02, and alterations listed as known somatic alterations in COSMIC.
Statistical analysis
The primary outcome was overall survival (OS, defined as time from enrollment to death or last clinical follow-up). Secondary outcomes included OS rate at 12 months, progression-free survival (PFS, defined as time from treatment initiation to first disease progression, death, or last follow-up imaging), and the objective response rate (ORR, defined as complete response plus partial response). The mutant allele fraction (MAF) was defined as the sum of all mutations detectable in each sample. Exploratory endpoints included drug safety and toxicity (Common Terminology Criteria for Adverse Events, CTCAE 5.0). The functional status of tumor patients was assessed using the Karnofsky Performance Status (KPS) scoring criteria. PFS and OS were analyzed using the Kaplan–Meier method, and the stratified Cox proportional hazards model was employed to calculate the hazard ratio (HR) and 95% confidence interval (CI). Clinical response was assessed using RANO 2.0 criteria, classifying responses as complete response (CR), partial response (PR), stable disease (SD), or progressive disease (PD) [23]. The Wilcoxon rank-sum test compared continuous variables between two groups, while Spearman's rank correlation estimated the correlation between two continuous variables. P < 0.05 indicated statistical significance. Statistical analyses were conducted using Prism 9.5 or R, version 4.2.1.
Results
Patient characteristics
Between March 28, 2022, and December 31, 2023, 32 rGBM patients were enrolled (Fig. 1A). The median time from diagnosis to relapse was 5.5 months (range, 1.6–10.3 months), with a median age of 52.5 years (range, 49–65 years), and 53.1% (n = 17) were female (Table 1). KPS scores were significantly higher after treatment with tislelizumab and low-dose bevacizumab than before treatment (P = 0.038, Table 1, Fig. 1B).
At the data cutoff (December 31, 2023), with a median follow-up of 11.0 months (95% CI, 9.0–16.2), 22 patients (68.8%) had discontinued study treatment, primarily due to disease progression (n = 21, 95.5%) and study drug-related toxic effects (n = 1, 4.5%) (Table 1). All patients received at least one cycle of combination therapy, with a median of 4.5 cycles completed, allowing efficacy evaluation using RANO 2.0 criteria.
Circulating tumor DNA analysis
Despite challenges posed by the COVID-19 pandemic, at least one TISF or tissue sample from 31 patients was analyzed by ctDNA, with 19 samples containing before-and-after controls. High-throughput sequencing of TISF using a custom panel designed for solid tumors was performed (Fig. 2A). TERT emerged as the most prevalent genetic mutation, consistent with previous studies [24].
Patient treatment events and ctDNA outcomes. A, Oncoplot depicting the genomic alteration of 32 recurrent GBM patients at different time points. Plot of tumor variants identified from 551-panel sequencing and tracked using ctDNA analysis for each patient. The top panel shows the total number of single nucleotide variants (SNVs) and copy number alterations (CNAs) tracked, and the left panel shows the number of patients with mutations in each gene. Only the most frequently mutated genes are displayed. B, Event chart showing time points for low-dose Bev + anti-PD-1 treatment, treatment response assessed according to RANO2.0 criteria, and the results of ctDNA testing for each patient with at least one TISF sample or tissue-sample time point analyzed. C, Proportion of patients with ctDNA detected in at least one TISF sample time point. Treatment efficacy (PD progressive disease; PR partial response; SD stable disease)
To explore ctDNA levels and tumor mutational burden's prognostic predictive value, we calculated the mutant allele fraction (MAF) and tumor mutation burden (TMB) for all detectable mutations in each sample. Baseline ctDNA was detected in 78% of patients (n = 25), and ctDNA was detected at least once in 97% of patients (n = 31) (Fig. 2B and C). TISF or tissue samples were collected from 19 patients before and after treatment, with ctDNA levels elevated in 9 SD patients and decreased in 10 patients (1 PD patient, 9 PR patients) (Fig. 3A). TISF-ctDNA dynamic changes significantly correlated with treatment response (P = 0.0001, Fig. 3A, B). There was a significant correlation between baseline ctDNA levels and tumor volume burden measured on imaging (P = 0.03, Fig. 3C). The COX risk regression model showed that RANO 2.0 response assessment (PR vs. PD/SD) was significantly associated with PFS and OS (PFS: P < 0.0001, HR: 25.3; OS: P < 0.0001, HR: 14.4), while MAF and TMB of ctDNA at baseline and post-treatment did not significantly correlate with PFS and OS (Fig. 3D). Interestingly, 2 patients with ctDNA negative (ctDNA-) before or after treatment had better prognosis (P12: PFS 6.0 months, OS 14.3 months; P27: PFS 15.6 months, OS 16.5 months). Among the 12 patients with high baseline TISF-ctDNA levels (MAF > 5%), prognosis improved when gene mutations in TISF-ctDNA significantly changed post-combination therapy (PFS: P = 0.0002, HR: 0.12; OS: P = 0.0002, HR: 0.08, Fig. 3E).
Analysis of ctDNA in patients treated with low-dose Bev + anti-PD-1. A, Spider plot of ctDNA levels before and after treatment with low-dose Bev + anti-PD-1 treatment. Patients are colored by RANO2.0 response, and the ctDNA allele fraction at each time point was divided by the pre-treatment allele fraction. In the one patient with ctDNA not detected prior to treatment, the pre-treatment limit of detection was used for normalization based on the number of mutations tracked and average sequencing depth as described in the methods. B, Fisher precise test analysis showed that changes in ctDNA levels were significantly correlated with treatment response. C, Correlation between baseline tumor burden measured by 3D slicer and baseline ctDNA mutant allele fraction (MAF). D, Forest plot depicting progression-free survival (PFS) and overall survival (OS) improvements for each variable in patients treated with low-dose Bev + anti-PD-1 therapy. The HRs and statistical significance of the difference were computed using the Cox proportional hazards model and Wald test. E, Kaplan–Meier curves showed that patients in the TISF-ctDNA significant changes group had significantly improved PFS and OS after receiving low-dose Bev plus anti-PD-1 therapy. Spearman's correlation coefficient, 95% confidence interval, and P-value are displayed on the graph
We also analyzed whether MAF and TMB changes could predict prognosis. Patients were divided into ctDNA/TMB response and non-response groups based on whether post-treatment MAF and TMB decreased by 20% compared to baseline. Post-grouping analysis revealed that patients in the ctDNA/TMB response group (≥ 20%) had significantly better PFS and OS (ctDNA: PFS P = 0.0009, HR = 0.16; OS P = 0.008, HR = 0.10; TMB: PFS P = 0.0005, HR = 0.18, OS P = 0.008, HR = 0.17; Fig. 4A).
Oncogenic alterations correlated with fewer benefits from low-dose Bev + anti-PD-1 therapy. A, Kaplan–Meier curves showed that patients in the ctDNA response group had significantly improved PFS and OS after receiving low-dose Bev plus anti-PD-1 therapy. B, Kaplan–Meier curves depict PFS and OS improvements in patients with partial gene wild-type mutations on low-dose Bev + anti-PD-1 therapy. C, Copy number changes in H3F3B amplification in 8 rGBM patients throughout treatment
Oncogenic alterations correlated with fewer benefits from low-dose Bev + anti-PD-1 therapy
We applied COX regression models to evaluate whether gene mutations were associated with low-dose Bev + anti-PD-1 efficacy. Stratified analysis of baseline TISF-ctDNA revealed that MUC16 mutation (PFS: P = 0.03, HR = 2.90; OS: P = 0.004, HR = 4.20), H3F3B amplification (PFS: P = 0.025, HR = 3.38; OS: P = 0.038, HR = 2.87), and SRSF2 amplification (PFS: P = 0.18, HR = 2.25; OS: P = 0.043, HR = 3.37) were significantly associated with worse prognosis (Fig. 4B). Post-combination therapy TISF-ctDNA showed only EGFR mutations and amplification significantly associated with poorer OS and PFS (PFS: P < 0.0001, HR = 7.64; OS: P = 0.001, HR = 5.41; Fig. 4B). Interestingly, none of the eight samples with primary tumor tissue or pre-recurrent TISF had detectable H3F3B amplification (Fig. 4C), suggesting H3F3B amplification emerged during the period of standard therapy and was associated with resistance and relapse.
Two patients demonstrated H3F3B amplification's ability to track combination therapy response. Patient 21 had a near-complete imaging response at 6.4 months but progressed after being lost to follow-up for 3.1 months due to the COVID-19 pandemic (Fig. 5A). Patient 11 consistently had H3F3B amplification detected during follow-up and progressed 1.7 months after starting therapy (Fig. 5B). These results suggest H3F3B amplification may lead to drug resistance by altering the tumor immune microenvironment, and ctDNA has potential to monitor combination therapy response in rGBM patients.
Monitoring response to low-dose Bev + anti-PD-1 therapy using ctDNA analysis. A–B, Examples of longitudinal radiographic imaging and ctDNA monitoring in A a patient with progressive disease on first surveillance imaging and (B) a patient with sustained disease remission after starting treatment. Circulating tumor DNA allele fraction is shown in teal, and H3F3B copy numbers are shown in red. C, Bubble plots show pathway alterations by KEGG enrichment analysis at baseline and after low-dose Bev plus anti-PD-1 treatment. D, Bubble plots showing pathway alterations by KEGG enrichment analysis at baseline and after low-dose Bev plus anti-PD-1 treatment. There was a significant increase in mutated genes associated with cell cycle and transcription dysregulation pathways and a decrease in mutated genes associated with microRNAs in cancer pathways in TISF-ctDNA at relapse compared with baseline. E, The median PFS and OS of all patients were analyzed
Finally, 8 patients experienced a second relapse after combination therapy, all from the ctDNA/TMB non-response group. KEGG pathway enrichment analysis revealed significant increases in mutated genes associated with cell cycle and transcriptional misregulation pathways in ctDNA at second recurrence and significant decreases in genes associated with microRNAs in cancer pathways (Fig. 5C). More cohort studies are needed to verify these changes in detail, revealing the related mechanisms of low-dose Bev + anti-PD-1 therapy and acquired resistance.
Patient outcomes and safety
Among all patients, 18 (56.3%) had PR, 9 (28.1%) had SD, and 5 (15.6%) had PD, with an ORR of 56.3%. The 12-month OS was 43.8%. Patients who achieved PR had a median response duration of 13.4 months (95% CI, 7.0–19.9). Median PFS and OS were 8.2 months (95% CI, 5.2–11.1) and 14.3 months (95% CI, 6.5–22.1), respectively (Fig. 5D).
Observed toxicities included anemia (50.0%), fatigue (34.1%), hypokalemia (31.3%), increased alanine aminotransferase (31.3%), and decreased white blood cell count (25.0%). One patient experienced grade 4 acute pancreatitis, and another had tertiary toxicity with elevated ALT levels. No grade 5 adverse events occurred (Table 2).
Discussion
In this study, 32 rGBM patients received tislelizumab plus low-dose Bev, hypothesizing that low-dose bevacizumab would normalize vascular conditions and facilitate immunotherapy, using TISF-ctDNA as a biomarker to track treatment response and gene evolution [8, 9]. The observed ORR of 56.3% significantly benefited patients, exceeding the 7.8% ORR for GBM with nivolumab in the CheckMate 143 trial [13]. This study is the first to perform biomarker analysis in rGBM patients treated with this combination therapy.
Standard-dose bevacizumab combined with anti-PD-1 has been confirmed effective in other solid tumors [28,29,30,31], but efficacy in rGBM is poor [32]. Bevacizumab, a humanized monoclonal antibody inhibiting VEGF, enhances tumor-specific immune response by promoting immunosuppressive tumor microenvironment, normalizing vascular structure, increasing T cell infiltration, and activating local immune microenvironment [12, 33,34,35,36]. The 2021 ASCO Annual Meeting reported no benefit of low-dose Bev + anti-PD-1 compared with standard Bev for rGBM, and standard Bev can help older rather than younger patients [32]. Therefore, it is essential to find suitable biomarkers that guarantee to maximize the therapeutic effect [37]. However, TMB and PD-L1 expression has not predicted anti-PD-1 monotherapy efficacy in rGBM [38,39,40,41]. Although single time-point MAF and TMB expression did not correlate with response to low-dose Bev plus anti-PD-1, dynamic changes predicted response. Two patients with ctDNA negative (ctDNA-) before or after treatment had better prognosis, and patients with high baseline ctDNA levels (MAF > 5%) had better prognosis with significant post-treatment TISF-ctDNA gene mutation changes. This facilitates screening high-risk recurrence patients and timely treatment regimen adjustments.
CtDNA is a promising biomarker in solid tumors (lung, breast, prostate, colorectal, melanoma, glioma) [17, 42, 43], used for early cancer detection, treatment selection, MRD detection, recurrence surveillance, and treatment response monitoring [44], used for early cancer detection, treatment selection, MRD detection, recurrence surveillance, and treatment response monitoring [44, 45]. Based on this research, early treatment of high-risk postoperative recurrence GBM patients (i.e., ctDNA recurrence) is planned.
Specific oncogenic alterations can disrupt the cancer immune cycle and influence immunotherapy efficacy [46, 47]. We identified various oncogenic alterations posing higher risk and reducing low-dose Bev + anti-PD-1 therapy benefits, including MUC16 mutation, EGFR mutation, H3F3B amplification, and SRSF2 amplification. MUC16 mutations confer immune evasion and resistance to immunotherapy in tumors [25, 48, 49]. SRSF2 expression correlates with cancer progression in malignant ovarian tissues [27]. Histone H3.3 point mutations are frequently observed in pediatric high-grade glioma (pHGG) [50,51,52,53]. But associated copy number variation in glioblastomas has not been reported. Amplification of H3F3B associated with aortic dissection disease may explain resistance to low-dose bevacizumab + anti-PD-1 treatment [26, 54]. EGFR mutation and amplification are poor prognostic markers for glioma [4]. EGFR signaling pathway plays crucial roles in cancer immune evasion [55], with SEC61G as an EGFR-coamplified gene promoting GBM immune evasion [56]. Even with low-dose Bev combined with anti-PD-1, these oncogenic alterations hinder immunotherapy effectiveness. MUC16 mutations, EGFR mutation, SRSF2 amplification, and H3F3B amplification accounted for 24, 20, 12, and 40% of patients, respectively, providing practical value for patient selection.
This study's limitations include the limited data size and all participants were Chinese. Future genomic data from cohorts with low-dose bevacizumab plus anti-PD-1 therapy are needed to validate identified biomarkers. Validation using a combination therapy dataset could demonstrate intrinsic associations between biomarkers and antitumor immunity, affirming their predictive value for immunostimulatory chemotherapy and anti-PD-1 therapy benefits. Technological advancements are needed to reduce genome sequencing costs and ensure speedy analysis for clinical application.
By performing high-throughput sequencing on samples from 97% of patients, we identified four oncogenic risk alterations as reliable biomarkers for low-dose bevacizumab plus anti-PD-1 therapy outcomes in rGBM patients. These findings provide a basis for individualized treatment and future biological studies of its immuno-oncology characteristics, inspiring biomarker exploration of low-dose bevacizumab + anti-PD-1 in other cancer types.
Conclusions
Anti-PD-1 antibody combined with low-dose bevacizumab can significantly prolong PFS and OS in rGBM patients without significant adverse reactions, improving quality of life and providing a new effective treatment for rGBM. TISF-ctDNA dynamic changes can predict the treatment response, identify drug resistance mechanisms, monitor high-risk recurrence (ctDNA molecular recurrence) populations, and provide a basis for early intervention decision making. TISF-ctDNA characterizes in vivo gene evolution in rGBM patients treated with anti-PD-1 antibody combined with low-dose bevacizumab, providing molecular information for drug resistance mechanism studies in rGBM.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Weller M, Cloughesy T, Perry JR, Wick W (2013) Standards of care for treatment of recurrent glioblastoma—are we there yet? Neuro Oncol 15(1):4–27. https://doi.org/10.1093/neuonc/nos273
Tan AC, Ashley DM, López GY, Malinzak M, Friedman HS, Khasraw M (2020) Management of glioblastoma: state of the art and future directions. CA Cancer J Clin 70(4):299–312. https://doi.org/10.3322/caac.21613
Omuro A, DeAngelis LM (2013) Glioblastoma and other malignant gliomas: a clinical review. JAMA 310(17):1842–1850. https://doi.org/10.1001/jama.2013.280319
Weller M, van den Bent M, Preusser M et al (2021) EANO guidelines on the diagnosis and treatment of diffuse gliomas of adulthood. Nat Rev Clin Oncol 18(3):170–186. https://doi.org/10.1038/s41571-020-00447-z
Nassiri F, Patil V, Yefet LS et al (2023) Oncolytic DNX-2401 virotherapy plus pembrolizumab in recurrent glioblastoma: a phase 1/2 trial. Nat Med 29(6):1370–1378. https://doi.org/10.1038/s41591-023-02347-y
Lim M, Xia Y, Bettegowda C, Weller M (2018) Current state of immunotherapy for glioblastoma. Nat Rev Clin Oncol 15(7):422–442. https://doi.org/10.1038/s41571-018-0003-5
Reardon DA, Gokhale PC, Klein SR, Ligon KL, Rodig SJ, Ramkissoon SH, Jones KL, Conway AS, Liao X, Zhou J, Wen PY (2016) Glioblastoma eradication following immune checkpoint blockade in an orthotopic, immunocompetent model. Cancer Immunol Res 4(2):124–135. https://doi.org/10.1158/2326-6066.Cir-15-0151
Wen PY, Weller M, Lee EQ et al (2020) Glioblastoma in adults: a society for neuro-oncology (sno) and european society of neuro-oncology (eano) consensus review on current management and future directions. Neuro Oncol 22(8):1073–1113. https://doi.org/10.1093/neuonc/noaa106
Huang Y, Goel S, Duda DG, Fukumura D, Jain RK (2013) Vascular normalization as an emerging strategy to enhance cancer immunotherapy. Cancer Res 73(10):2943–2948. https://doi.org/10.1158/0008-5472.Can-12-4354
Seto T, Nosaki K, Shimokawa M, Toyozawa R, Sugawara S, Hayashi H, Murakami H, Kato T, Niho S, Saka H, Oki M (2022) Phase II study of atezolizumab with bevacizumab for non-squamous non-small cell lung cancer with high PD-L1 expression (@ Be Study). J Immunother Cancer 10(2):e004025. https://doi.org/10.1136/jitc-2021-004025
Reardon DA, Brandes AA, Omuro A et al (2020) Effect of nivolumab vs bevacizumab in patients with recurrent glioblastoma. JAMA Oncol 6(7):1003–1010. https://doi.org/10.1001/jamaoncol.2020.1024
Guo X, Wang S, Wang Y, Ma W (2021) Anti-PD-1 plus anti-VEGF therapy in multiple intracranial metastases of a hypermutated. IDH Wild-Type Glioblastoma Neuro-Oncol 23(4):699–701. https://doi.org/10.1093/neuonc/noab005
Reardon DA, Brandes AA, Omuro A et al (2020) Effect of nivolumab vs bevacizumab in patients with recurrent glioblastoma: the checkmate 143 phase 3 randomized clinical trial. JAMA Oncol 6(7):1003–1010. https://doi.org/10.1001/jamaoncol.2020.1024
Soffietti R, Bettegowda C, Mellinghoff IK et al (2022) Liquid biopsy in gliomas: a RANO review and proposals for clinical applications. Neuro Oncol 24(6):855–871. https://doi.org/10.1093/neuonc/noac004
Miller AM, Shah RH, Pentsova EI et al (2019) Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 565(7741):654–658. https://doi.org/10.1038/s41586-019-0882-3
Westphal M, Lamszus K (2015) Circulating biomarkers for gliomas. Nat Rev Neurol 11(10):556–566. https://doi.org/10.1038/nrneurol.2015.171
Xu S, Sheng Z, Yu J et al (2022) Real-time longitudinal analysis of human gliomas reveals in vivo genome evolution and therapeutic impact under standardized treatment. Clin Transl Med 12(7):e956. https://doi.org/10.1002/ctm2.956
Sheng Z, Yu J, Deng K et al (2021) Integrating real-time in vivo tumour genomes for longitudinal analysis and management of glioma recurrence. Clin Transl Med 11(11):e567. https://doi.org/10.1002/ctm2.567
Liu G, Bu C, Guo G et al (2023) Genomic alterations of oligodendrogliomas at distant recurrence. Cancer Med 12(16):17171–17183. https://doi.org/10.1002/cam4.6327
Sheng Z, Yu J, Deng K et al (2021) Characterizing the genomic landscape of brain glioma with circulating tumor DNA from tumor in situ fluid. Front Oncol 11:584988. https://doi.org/10.3389/fonc.2021.584988
Fedorov A, Beichel R, Kalpathy-Cramer J et al (2012) 3D Slicer as an image computing platform for the quantitative imaging network. Magn Reson Imaging 30(9):1323–1341. https://doi.org/10.1016/j.mri.2012.05.001
Sheng Z, Bu C, Mei J, Xu S, Zhang Z, Guo G, Gao Y, Xing L, Chen Z, Hernesniemi J, Zemmar A (2023) Tracking tumor evolution during the first-line treatment in brain glioma via serial profiling of cell-free tumor DNA from tumor in situ fluid. Front Oncol 13:1238607. https://doi.org/10.3389/fonc.2023.1238607
Youssef G, Rahman R, Bay C et al (2023) Evaluation of standard response assessment in neuro-oncology, modified response assessment in neuro-oncology, and immunotherapy response assessment in neuro-oncology in newly diagnosed and recurrent glioblastoma. J Clin Oncol 41(17):3160–3171. https://doi.org/10.1200/jco.22.01579
Draaisma K, Chatzipli A, Taphoorn M et al (2020) Molecular evolution of IDH wild-type glioblastomas treated with standard of care affects survival and design of precision medicine trials: a report from the EORTC 1542 study. J Clin Oncol 38(1):81–99. https://doi.org/10.1200/jco.19.00367
Gao R, Lou N, Han X, Shi Y (2022) MUC16: the novel target for tumor therapy. Chin J Lung Cancer 25(7):452–459. https://doi.org/10.3779/j.issn.1009-3419.2022.101.31
Zhang X, Che Y, Mao L, Li D, Deng J, Guo Y, Zhao Q, Zhang X, Wang L, Gao X, Chen Y (2023) H3.3B controls aortic dissection progression by regulating vascular smooth muscle cells phenotypic transition and vascular inflammation. Genomics 115(5):110685. https://doi.org/10.1016/j.ygeno.2023.110685
Fischer DC, Noack K, Runnebaum IB et al (2004) Expression of splicing factors in human ovarian cancer. Oncol Rep 11(5):1085–1090
Ren Z, Xu J, Bai Y et al (2021) Sintilimab plus a bevacizumab biosimilar (IBI305) versus sorafenib in unresectable hepatocellular carcinoma (ORIENT-32): a randomised, open-label, phase 2–3 study. Lancet Oncol 22(7):977–990. https://doi.org/10.1016/s1470-2045(21)00252-7
Rini BI, Powles T, Atkins MB et al (2019) Atezolizumab plus bevacizumab versus sunitinib in patients with previously untreated metastatic renal cell carcinoma (IMmotion151): a multicentre, open-label, phase 3, randomised controlled trial. Lancet 393(10189):2404–2415. https://doi.org/10.1016/s0140-6736(19)30723-8
Socinski MA, Jotte RM, Cappuzzo F et al (2018) Atezolizumab for first-line treatment of metastatic nonsquamous NSCLC. N Engl J Med 378(24):2288–2301. https://doi.org/10.1056/NEJMoa1716948
Zhu AX, Abbas AR, de Galarreta MR et al (2022) Molecular correlates of clinical response and resistance to atezolizumab in combination with bevacizumab in advanced hepatocellular carcinoma. Nat Med 28(8):1599–1611. https://doi.org/10.1038/s41591-022-01868-2
Ahluwalia MS, Rauf Y, Li H, Wen PY, Peereboom DM, Reardon DA (2021) Randomized phase 2 study of nivolumab (nivo) plus either standard or reduced dose bevacizumab (bev) in recurrent glioblastoma (rGBM). J Clin Oncol. https://doi.org/10.1200/JCO.2021.39.15_suppl.2015
Fukumura D, Kloepper J, Amoozgar Z, Duda DG, Jain RK (2018) Enhancing cancer immunotherapy using antiangiogenics: opportunities and challenges. Nat Rev Clin Oncol 15(5):325–340. https://doi.org/10.1038/nrclinonc.2018.29
Yi M, Jiao D, Qin S, Chu Q, Wu K, Li A (2019) Synergistic effect of immune checkpoint blockade and anti-angiogenesis in cancer treatment. Mol Cancer 18:1–2. https://doi.org/10.1186/s12943-019-0974-6
Jain RK (2014) Antiangiogenesis strategies revisited: from starving tumors to alleviating hypoxia. Cancer Cell 26(5):605–622. https://doi.org/10.1016/j.ccell.2014.10.006
Alfaro C, Suarez N, Gonzalez A et al (2009) Influence of bevacizumab, sunitinib and sorafenib as single agents or in combination on the inhibitory effects of VEGF on human dendritic cell differentiation from monocytes. Br J Cancer 100(7):1111–1119. https://doi.org/10.1038/sj.bjc.6604965
Tie J, Cohen JD, Lahouel K et al (2022) Circulating tumor dna analysis guiding adjuvant therapy in stage II colon cancer. N Engl J Med 386(24):2261–2272. https://doi.org/10.1056/NEJMoa2200075
Merchant M, Ranjan A, Pang Y et al (2021) Tumor mutational burden and immunotherapy in gliomas. Trends Cancer 7(12):1054–1058. https://doi.org/10.1016/j.trecan.2021.08.005
Brown MC, Ashley DM, Khasraw M (2022) Low tumor mutational burden and immunotherapy in gliomas. Trends Cancer 8(5):345–346. https://doi.org/10.1016/j.trecan.2022.01.006
McGrail DJ, Pilié PG, Rashid NU et al (2021) High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann Oncol 32(5):661–672. https://doi.org/10.1016/j.annonc.2021.02.006
Capper D, Reifenberger G, French PJ et al (2023) EANO guideline on rational molecular testing of gliomas, glioneuronal, and neuronal tumors in adults for targeted therapy selection. Neuro Oncol 25(5):813–826. https://doi.org/10.1093/neuonc/noad008
Cheng ML, Pectasides E, Hanna GJ, Parsons HA, Choudhury AD, Oxnard GR (2021) Circulating tumor DNA in advanced solid tumors: clinical relevance and future directions. CA Cancer J Clin 71(2):176–190. https://doi.org/10.3322/caac.21650
Liu G, Bu C, Guo G, Zhang Z, Sheng Z, Deng K, Wu S, Xu S, Bu Y, Gao Y, Wang M (2023) Molecular and clonal evolution in vivo reveal a common pathway of distant relapse gliomas. Iscience 26(9):107528. https://doi.org/10.1016/j.isci.2023.107528
Cohen SA, Liu MC, Aleshin A (2023) Practical recommendations for using ctDNA in clinical decision making. Nature 619(7969):259–268. https://doi.org/10.1038/s41586-023-06225-y
Tan AC, Lai GGY, Saw SPL et al (2024) Detection of circulating tumor DNA with ultradeep sequencing of plasma cell-free DNA for monitoring minimal residual disease and early detection of recurrence in early-stage lung cancer. Cancer 130(10):1758–1765. https://doi.org/10.1002/cncr.35263
Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A (2017) Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168(4):707–723. https://doi.org/10.1016/j.cell.2017.01.017
Keenan TE, Burke KP, Van Allen EM (2019) Genomic correlates of response to immune checkpoint blockade. Nat Med 25(3):389–402. https://doi.org/10.1038/s41591-019-0382-x
Thomas D, Sagar S, Liu X et al (2021) Isoforms of MUC16 activate oncogenic signaling through EGF receptors to enhance the progression of pancreatic cancer. Mol Ther 29(4):1557–1571. https://doi.org/10.1016/j.ymthe.2020.12.029
Sharma SK, Mack KN, Piersigilli A, Pourat J, Edwards KJ, Keinänen O, Jiao MS, Zhao H, White B, Brooks CL, de Stanchina E (2022) ImmunoPET of ovarian and pancreatic cancer with AR9.6, a novel MUC16-targeted therapeutic antibody. Clin Cancer Res 28(5):948–959. https://doi.org/10.1158/1078-0432.Ccr-21-1798
Bressan RB, Southgate B, Ferguson KM, Blin C, Grant V, Alfazema N, Wills JC, Marques-Torrejon MA, Morrison GM, Ashmore J, Robertson F (2021) Regional identity of human neural stem cells determines oncogenic responses to histone H3.3 mutants. Cell Stem Cell 28(5):877–893. https://doi.org/10.1016/j.stem.2021.01.016
Lewis PW, Müller MM, Koletsky MS et al (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340(6134):857–861. https://doi.org/10.1126/science.1232245
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K, Sturm D, Fontebasso AM, Quang DA, Tönjes M, Hovestadt V (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231. https://doi.org/10.1038/nature10833
Wu G, Broniscer A, McEachron TA et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44(3):251–253. https://doi.org/10.1038/ng.1102
Aldera AP, Govender D (2022) Gene of the month: H3F3A and H3F3B. J Clin Pathol 75(1):1–4. https://doi.org/10.1136/jclinpath-2021-207751
Li CW, Lim SO, Chung EM et al (2018) Eradication of triple-negative breast cancer cells by targeting glycosylated PD-L1. Cancer Cell 33(2):187-201.e10. https://doi.org/10.1016/j.ccell.2018.01.009
Zeng K, Zeng Y, Zhan H, Zhan Z, Wang L, Xie Y, Tang Y, Li C, Chen Y, Li S, Liu M (2023) SEC61G assists EGFR-amplified glioblastoma to evade immune elimination. Proceed Nat Acad Sci 120(32):e2303400120. https://doi.org/10.1073/pnas.2303400120
Acknowledgements
Thanks to all the contributors.
Funding
This work was supported by the Natural Science Foundation of Henan Province (232300421171) and the Research Foundation of the Health Commission of Henan Province (SBGJ202302007).
Author information
Authors and Affiliations
Contributions
GG and YB helped in study concept and design and statistical analysis. All the authors contributed to acquisition, analysis, or interpretation of data and administrative, technical, or material support. GG, SX, and YB contributed to drafting of the manuscript. GL and YB supervised the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest that could be perceived as prejudicing the impartiality of the research reported.
Ethical approval
The protocol was approved by the Institutional Review Board at Zhengzhou University People's Hospital, China (MR-41–23-037636).
Consent to participate
All studies were conducted in accordance with the Helsinki Declaration, and all patients obtained informed written consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, G., Zhang, Z., Zhang, J. et al. Predicting recurrent glioblastoma clinical outcome to immune checkpoint inhibition and low-dose bevacizumab with tumor in situ fluid circulating tumor DNA analysis. Cancer Immunol Immunother 73, 193 (2024). https://doi.org/10.1007/s00262-024-03774-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00262-024-03774-7