Abstract
Background
The non-invasive biomarkers for predicting immunotherapy response are urgently needed to prevent both premature cessation of treatment and ineffective extension. This study aimed to construct a non-invasive model for predicting immunotherapy response, based on the integration of deep learning and habitat radiomics in patients with advanced non-small cell lung cancer (NSCLC).
Methods
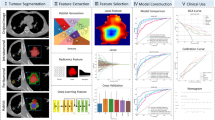
Independent patient cohorts from three medical centers were enrolled for training (n = 164) and test (n = 82). Habitat imaging radiomics features were derived from sub-regions clustered from individual’s tumor by K-means method. The deep learning features were extracted based on 3D ResNet algorithm. Pearson correlation coefficient, T test and least absolute shrinkage and selection operator regression were used to select features. Support vector machine was applied to implement deep learning and habitat radiomics, respectively. Then, a combination model was developed integrating both sources of data.
Results
The combination model obtained a strong well-performance, achieving area under receiver operating characteristics curve of 0.865 (95% CI 0.772–0.931). The model significantly discerned high and low-risk patients, and exhibited a significant benefit in the clinical use.
Conclusion
The integration of deep-leaning and habitat radiomics contributed to predicting response to immunotherapy in patients with NSCLC. The developed integration model may be used as potential tool for individual immunotherapy management.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung cancer (NSCLC) is one of the main cause of cancer-related mortality in global [1]. Recently, Immunotherapy, the immune checkpoint inhibitors such as PD-1 or PD-L1, has revolutionized the treatment paradigm for malignant cancer, establishing itself as the novel benchmark for managing locally advanced and metastatic non-small cell lung cancer (NSCLC) patients [2]. Numerous studies have demonstrated that the efficacy of immunotherapy on improving long-term survival rates when used as a monotherapy or in integration with other therapies [3,4,5,6]. Despite these advancements, only a part of patients, ranging from 20 to 50%, exhibits responsiveness to the therapy [7, 8]. The uncommon patterns of immunotherapy response, such as delayed response or pseudo-progression, makes traditional response prediction approaches obsolete. Additionally, there is a potential risk of immune-related adverse events which influences the patients' prognosis [9].
The identification of biomarkers with immunotherapy response power has become increasingly important for effective condition monitoring. Various biomarkers, including PD-L1 expression and tumor mutational burden, have raised researcher’s attention. And their relationship to immunotherapy response have been reported in previous studies, though conclusion differs [10, 11]. Moreover, the reliability of these biomarkers may be influenced by tumor heterogeneity, as they mostly depend on biopsy samples that do not reveal the full spectrum of the tumor microenvironment.
Due to the significant heterogeneity of malignancies, tumors present diverse microenvironments and microstructures [12,13,14,15]. Radiomics, an approach that extracts high throughput features to classify disease condition using machine learning techniques, holds promise for personalized medicine decision-making. However, conventional radiomic analysis generally focuses on the entire tumor, ignoring the sub-regional phenotypic variations within the tumor [16]. Recently, a novel method known as 'habitat' has emerged, which partitions tumors into sub-regions by identifying grayscale voxels with similar imaging characteristics [16]. This approach has demonstrated potential in enhancing the differentiation of tumor heterogeneity [17, 18]. Furthermore, recent developments in deep learning have revealed that neural networks can automatically extract radiomic features without the need for human intervention, leading to improvement of predictive performance [19, 20].
Additionally, the integration of multimodal information, such as deep learning features and habitat radiomic features, could offer tumor-specific multi dimension data for better immunotherapy management [21, 22].
The present study was conducted to explore the potential improvement in predictive accuracy through the integration of imaging data during the early stages of treatment. The support vector machine (SVM), was developed and subsequently validated on an external dataset to predict the possible benefit of immunotherapy in NSCLC patients at 6 months following treatment commencement.
Method
Patient selection
A total of 246 patients diagnosed with NSCLC treated with immune checkpoint inhibitor between January, 2018, and May, 2020 were enrolled at the Wenzhou Medical University First Affiliated hospital, Yueqing people’s hospital, Xiamen Second hospital in this study, as depicted Fig. 1. This study has been approved by their institutional reviews board. Written informed consent was waived because the data used in this manuscript were anonymized by removing personal information and its nature of retrospective design. All enrolled patients met the following inclusion criteria: (1) pathological diagnosis of NSCLC; (2) chest contrast CT examinations were available. Patients were excluded if: (1) Basic clinical data (such as age, sex) were missing; (2) preoperative treatment or the time between CT examination and subsequent surgery exceeded 1 month; (3) CT images were of low quality; and (4) molecular testing results were difficult to determine.
Finally, the patients from Wenzhou Medical University First Affiliated hospital were used as training set (n = 164), the remaining patients from two other institutions were utilized as test set (n = 82).
Clinical endpoint
The primary objective of this study was the durable clinical benefit, as defined by progression-free survival (PFS). This metric represents the interval from the start of the first immunotherapy cycle until the occurrence of death, disease progression, or the last follow-up. Disease progression was judged by the individual’s overall clinical condition and the response evaluation criteria in solid tumors [23, 24], derived from imaging evaluations. Patients who exhibited a durable clinical benefit with a PFS exceeding 6 (PFS6) months were classified as responders [25], whereas the remaining were labeled as non-responders. Patients with censored data 6 months post-treatment were omitted from the analysis.
The secondary clinical outcome, overall survival (OS), was defined as the duration in months from the commencement of immunotherapy to the occurrence of death or the last follow-up date.
Image acquisition and Tumor segmentation
All patients received a CT scan within a one-month period prior to the administration of immunotherapy. The contrast CT images were obtained after contrast injection during an inspiratory breath-hold, in accordance with the established protocol for contrast-enhanced CT of the chest.
Two experienced radiologists were tasked with delineating the border of tumors. The definitive VOIs were derived from their mutual consensus. In the event of a disagreement, a third radiologist was consulted to provide opinion. The delineation was processed using the ITK-SNAP (Version 4.0).
VOI delineation and sub-region clustering
Habitat analysis employs entropy and voxel metrics extracted from CT imaging to categorize volumes of interest (VOIs) into different sub-regions [26,27,28,29]. The count of voxels for each VOI were derived by a conventional approach. Entropy values are obtained from CT images.
The K-means clustering algorithm was utilized to label the VOI of each patient, leading to the formation of multiple habitats. Euclidean distance, based on voxel and entropy values, was used to identify the relationships among samples. We tested the number of habitats ranging from 2 to 10. To determine the optimal k value, we employed the Consensus Cluster Plus approach. The optimal k-value was crucial for determining the suitable number of clusters within the patients, as depicted in Fig. 2. Finally, our analysis determined the optimal k-value to be 5. Subsequently, we used the Python to extract each patient's radiomics information, and labeled each sub-regions named habitat 1, habitat 2, habitat 3, habitat4, habitat5.
Feature selection and model development
To tackle differences in imaging features due to changes in the reconstruction layer thickness and pixel size, we resampled the images to a dimension of 1 × 1 × 1 m^3, and normalized them to a grayscale range of 0–255. The PyRadiomics [30] Python package was applied to independently extract radiomics features from six distinct habitats: habitat 1, habitat 2, habitat 3, habitat 4, habitat 5, and the entire tumor. This Python package is in compliance with the Imaging Biomarker Standardization Initiative [31].
On the other hand, we also used a 3D ResNet model as the CT image feature extractor to obtain deep learning features. The deep learning feature extraction module processed the region of interest (ROI) as input. When compared with traditional 2D image analysis, the 3D ResNet takes 3D context into consideration, thereby capturing more comprehensive image information from different level of one solid tumor, and make better decision. Finally, we obtained deep learning features from the whole tumor.
As for the features selection, at first, the features with an intraclass correlation coefficient (ICC) below 0.75 were filtered out. Subsequently, the remaining imaging features were underwent Z-score standardization to normalize the data to a mean of zero and a variance of one. Then we assessed the inter-feature relationships using the Pearson correlation coefficient. In instances where the correlation over the threshold of 0.9, we only kept one feature from each pair of highly correlated features to avoid redundancy. Finally, we applied two-sample T test and the least absolute shrinkage and selection operator (LASSO) regression model to further refine the remaining features in the training dataset.
A support vector machine (SVM) classification model was constructed utilizing features from each single habitat, the entire tumor, and deep learning features with five-fold cross-validation. Then, also fused each habitat tumor radiomics features with deep learning features, and fused all habitat tumor radiomics features with deep learning features as a combination to predict the risk of clinical endpoint.
Model explanation
The SHAP (SHapley Additive exPlanations) algorithm, based on Cooperative Game Theory and local explanations, was utilized to elucidate the contribution of each feature to the final prediction output of the SVM model. By assigning an importance value to each feature for every prediction, SHAP visualized the contribution of each feature on the model's prediction, indicating how it either increased or decreased the likelihood of a particular outcome. SHAP analysis was performed using the KernelExplainer package (version 0.40.0) in Python (Version 3.7.0).
Statistical analyses
Statistical analyses in this study were conducted using R software (version 4.0.2). Univariate analyses, comprising chi-square tests, t tests, and Mann–Whitney U tests were performed according to the normality of variables. Intraclass correlation coefficient (ICC), least absolute shrinkage and selection operator (LASSO), receiver operating characteristic (ROC) analysis, survival analysis, and decision curve analysis (DCA) were carried out using R statistical software. A two-sided p value of less than 0.05 was deemed to indicate statistical significance.
Results
Baseline clinical characteristic
A total of 246 patients from three institutions with NSCLC had received immune checkpoint inhibitor treatment and met the other inclusion criteria were finally included in our study. The mean age was 66.4 ± 8.4 [standard deviation]. Men were predominant (n = 215, 87.4%). There were 16 patients were never smoker. Regarding our study, we found the following: 59.3% included patients responded to immune checkpoint inhibitor after 6 months therapy. The most histology type was the adenocarcinoma. And 93.5% enrolled patients (n = 230) were current or former smokers. The therapy included monotreatment (n = 134, 54.5%), treatment with chemotherapy (n = 26, 10.6%), treatment with radiotherapy (n = 13, 5.3%) and combined immunological agents (n = 41, 16.7%), the other detailed demographic and clinical characteristic are shown in Table 1. Patients were split into training and test set based on their source. One hundred and sixty-four patients were used for model training, and 82 patients were utilized for external validation. There were no significant differences between the training and test set in demographic and clinical characteristics variables.
Feature selection
A total of 372 radiomic features were extracted from the imaging data based on each single habitat and the entire tumor. And 512 deep learning features were derived from the entire tumor. To reduce feature dimension and simplify model, we adopted a series of screening approach. After the screening using the ICC, Pearson correlation coefficients, two-sample t test, and the least absolute shrinkage selection operator regression method as depicted in Fig. 2c, optimal radiomic features were obtained from habitat 1, habitat 2, habitat3, habitat4, habitat 5, and the entire tumor. And deep learning features were selected based on the whole tumor.
Performance evaluation of models
The SVM machine learning algorithm was developed mainly based on two groups: One consisted of each single habitat, the entire tumor radiomic features, and the whole tumor deep learning features. And the other one is the combination between deep learning features and each radiomic features from different sub-regions, and combination of deep learning features and all habitat radiomics features. The prediction efficiency of each model is summarized in Table 2. Table 2 described the prediction performance of models based on different sub-regions, the entire tumor radiomic and deep learning features. Then we fused the deep learning features with radiomic features to construct novel machine learning models. Among of different combination, the deep learning features plus habitat 4 performed best as depicted as Fig. 3d, with AUC 0.865 (95% CI 0.772–0.931), accuracy 0.82, precision score 0.89, recall score 0.68, F1-score 0.77 in the external test cohort. Figure 3a–h demonstrated ROC, decision curve analysis of the different combinations in the training and test set, respectively.
ROC curves of the SVM machine learning models in the training (a) and external test cohorts (b) based on single source features. The ROC curves of the SVM machine learning models in the training (c) and external test cohorts (d) based on combined features. The DCA curves of the SVM machine learning models in the training (e) and external test cohorts (f) based on single source features. The DCA curves of the SVM machine learning models in the training (g) and external test cohorts (h) based on combined features
Then, we investigated the performance of the combination of habitat 4 radiomic features and deep learning features in predicting PFS and OS. As depicted in Fig. 4a–b, included individuals were grouped into high-risk and low-risk group based on the optimum cutoff of value generated by ROC curve in the both of training and test set. Figure 4 shows the Kaplan–Meier survival curves for PFS and OS on the independent test set for the combination SVM models. The combination SVM could significantly stratify PFS and OS for the clinical endpoint (p value < 0.05) (Table 3).
Model explainability
The SHAP algorithm was utilized to illustrate the contribution of each feature to the model's final prediction. This algorithm was specifically applied to the model of the SVM, where a positive SHAP value indicated an elevated risk of disease progression for each prediction. As depicted in Fig. 5, the original_glszm_SmallAreaHighGrayLevelEmphasis and Feature 271 from deep learning extracted variable emerged as the most influential clinical variables.
a Variable importance of the SVM model in the training set, showing that the original_glszm_SmallAreaHighGrayLevelEmphasis was the most important feature. b SHAP of the SVM model. The closer the values of the features were to 1, the more likely patients were to progress to be non-responders to the immunotherapy
Discussion
Despite the immune checkpoint blockade has revolutionized treatment of NSCLC, a large number of patients remain non-responsive to the therapy [32]. Under these circumstances, there is an urgent need for a reliable signature to identify patients who may benefit from this advanced therapeutic approach. In this multi-institutional study, deep learning imaging features and radiomics based on habitat was employed to predict the clinical durable benefit at 6 months following the initiation of immune checkpoint inhibitor therapy in patients with NSCLC. The ensemble, which combines the deep learning features with the habitat radiomics, obtain the area under the curve (AUC) values of 0.943 (95% CI 0.895–0.973) training cohort and 0.865 (95% CI 0.772–0.931) for the external validation cohort.
CT is routinely applied in clinical settings to evaluate the immune therapy response to immune therapy in NSCLC patients [33]. Lesion diameters in the CT images could directly reflect tumor burden. There is growing evidence supporting the use of tumor diameters in CT scans to predict the responsivity to neoadjuvant chemotherapy and immune therapy [34, 35]. However, the immune therapy effect may precede any observable change of the tumor size [36]. Therefore, the reliance on mere superficial characteristics from CT image is insufficient for precise prediction of the therapy response. It necessitates the extraction of high-dimensional and high-throughput imaging features to accurately quantify the risk of prognosis in patients with NSCLC. This study underlined the need for more sophisticated imaging analytics in the evaluation of complex therapeutic outcomes.
When compared with conventional entire tumor radiomics, habitat radiomics have been considered as important for the prognosis [37, 38]. Moreover, the development of deep learning-based radiomics marks a significant advancement in the field, with its capacity to unearth deep-level imaging features not being recognized to the human eye [39], thereby offering novel perspectives on the prediction of the efficacy of immunotherapy. Deep learning's integration into conventional radiomics analysis of tumors is becoming increasingly important in the field of disease diagnosis, therapeutic strategy formulation, and prognostication [40,41,42].
To the best of our knowledge, no previous studies have revealed that if the integration of deep learning features and habitat radiomics features can stratify the patients at high risk of non-responsive to immune checkpoint inhibitor. Similarly, Vanguri et al. [21] previously reported that a multimodal model integrating baseline medical imaging, histopathological, and genomic features surpassed the performance of unimodal models achieving an AUC of 0.80 in predicting the response to immunotherapy.
Our study is subject to some limitations. First of all, the nature of retrospective and multicenter design of our study introduces heterogeneity within the cohort, particularly concerning treatment approaches and imaging protocols. Secondly, the interpretation of the deep features is often not straightforward since they are optimized to minimize the prediction error and are not designed to match human intuition or knowledge.
Conclusion
In conclusion, an ensemble of deep learning features and habitat radiomics has been employed to predict the durable benefit of immunotherapy at 6 months after treatment. This suggests its potential as a viable non-invasive biomarker for drug responsivity prediction. The model's robust performance has been substantiated by a stable AUC in an external test cohort.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Siegel RL, Miller KD, Wagle NS, Jemal A (2023) Cancer statistics. CA Cancer J Clin 73(1):17–48
Gridelli C, Peters S, Mok T, Forde PM, Reck M, Attili I, de Marinis F (2022) First-line immunotherapy in advanced non-small-cell lung cancer patients with ECOG performance status 2: results of an international expert panel meeting by the italian association of thoracic oncology. ESMO Open 7(1):100355
Doroshow DB, Sanmamed MF, Hastings K, Politi K, Rimm DL, Chen L, Melero I, Schalper KA, Herbst RS (2019) Immunotherapy in non-small cell lung cancer: facts and hopes. Clin Cancer Res 25(15):4592–4602
Patel SA, Weiss J (2020) Advances in the treatment of non-small cell lung cancer: immunotherapy. Clin Chest Med 41(2):237–247
Broderick SR (2020) Adjuvant and neoadjuvant immunotherapy in non-small cell lung cancer. Thorac Surg Clin 30(2):215–220
Paz-Ares L, Ciuleanu TE, Cobo M, Schenker M, Zurawski B, Menezes J, Richardet E, Bennouna J, Felip E, Juan-Vidal O et al (2021) First-line nivolumab plus ipilimumab combined with two cycles of chemotherapy in patients with non-small-cell lung cancer (CheckMate 9LA): an international, randomised, open-label, phase 3 trial. Lancet Oncol 22(2):198–211
Blons H, Garinet S, Laurent-Puig P, Oudart JB (2019) Molecular markers and prediction of response to immunotherapy in non-small cell lung cancer, an update. J Thorac Dis 11(Suppl 1):S25-s36
Ribas A, Wolchok JD (2018) Cancer immunotherapy using checkpoint blockade. Science 359(6382):1350–1355
Suresh K, Naidoo J, Lin CT, Danoff S (2018) Immune checkpoint immunotherapy for non-small cell lung cancer: benefits and pulmonary toxicities. Chest 154(6):1416–1423
Bai R, Lv Z, Xu D, Cui J (2020) Predictive biomarkers for cancer immunotherapy with immune checkpoint inhibitors. Biomark Res 8:34
Dong A, Zhao Y, Li Z, Hu H (2021) PD-L1 versus tumor mutation burden: Which is the better immunotherapy biomarker in advanced non-small cell lung cancer? J Gene Med 23(2):e3294
Junttila MR, de Sauvage FJ (2013) Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 501(7467):346–354
Gillies RJ, Brown JS, Anderson ARA, Gatenby RA (2018) Eco-evolutionary causes and consequences of temporal changes in intratumoural blood flow. Nat Rev Cancer 18(9):576–585
Janiszewska M (2020) The microcosmos of intratumor heterogeneity: the space-time of cancer evolution. Oncogene 39(10):2031–2039
Shi Z, Huang X, Cheng Z, Xu Z, Lin H, Liu C, Chen X, Liu C, Liang C, Lu C et al (2023) MRI-based quantification of intratumoral heterogeneity for predicting treatment response to neoadjuvant chemotherapy in breast cancer. Radiology 308(1):e222830
Gatenby RA, Grove O, Gillies RJ (2013) Quantitative imaging in cancer evolution and ecology. Radiology 269(1):8–15
Kim J, Ryu SY, Lee SH, Lee HY, Park H (2019) Clustering approach to identify intratumour heterogeneity combining FDG PET and diffusion-weighted MRI in lung adenocarcinoma. Eur Radiol 29(1):468–475
Park JE, Kim HS, Kim N, Park SY, Kim YH, Kim JH (2021) Spatiotemporal heterogeneity in multiparametric physiologic MRI is associated with patient outcomes in idh-wildtype glioblastoma. Clin Cancer Res 27(1):237–245
Mu W, Jiang L, Shi Y, Tunali I, Gray JE, Katsoulakis E, Tian J, Gillies RJ, Schabath MB (2021) Non-invasive measurement of PD-L1 status and prediction of immunotherapy response using deep learning of PET/CT images. J Immunother Cancer. https://doi.org/10.1136/jitc-2020-002118
Tian P, He B, Mu W, Liu K, Liu L, Zeng H, Liu Y, Jiang L, Zhou P, Huang Z et al (2021) Assessing PD-L1 expression in non-small cell lung cancer and predicting responses to immune checkpoint inhibitors using deep learning on computed tomography images. Theranostics 11(5):2098–2107
Vanguri RS, Luo J, Aukerman AT, Egger JV, Fong CJ, Horvat N, Pagano A, Araujo-Filho JAB, Geneslaw L, Rizvi H et al (2022) Multimodal integration of radiology, pathology and genomics for prediction of response to PD-(L)1 blockade in patients with non-small cell lung cancer. Nat Cancer 3(10):1151–1164
Farina B, Guerra ADR, Bermejo-Peláez D, Miras CP, Peral AA, Madueño GG, Jaime JC, Vilalta-Lacarra A, Pérez JR, Muñoz-Barrutia A et al (2023) Integration of longitudinal deep-radiomics and clinical data improves the prediction of durable benefits to anti-PD-1/PD-L1 immunotherapy in advanced NSCLC patients. J Transl Med 21(1):174
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M et al (2009) New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45(2):228–247
Seymour L, Bogaerts J, Perrone A, Ford R, Schwartz LH, Mandrekar S, Lin NU, Litière S, Dancey J, Chen A et al (2017) iRECIST: guidelines for response criteria for use in trials testing immunotherapeutics. Lancet Oncol 18(3):e143–e152
Dercle L, McGale J, Sun S, Marabelle A, Yeh R, Deutsch E, Mokrane FZ, Farwell M, Ammari S, Schoder H et al (2022) Artificial intelligence and radiomics: fundamentals, applications, and challenges in immunotherapy. J Immunother Cancer. https://doi.org/10.1136/jitc-2022-005292
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures. They Are Data Radiology 278(2):563–577
Xie C, Yang P, Zhang X, Xu L, Wang X, Li X, Zhang L, Xie R, Yang L, Jing Z et al (2019) Sub-region based radiomics analysis for survival prediction in oesophageal tumours treated by definitive concurrent chemoradiotherapy. EBioMedicine 44:289–297
Wang S, Liu X, Wu Y, Jiang C, Luo Y, Tang X, Wang R, Zhang X, Gong J (2023) Habitat-based radiomics enhances the ability to predict lymphovascular space invasion in cervical cancer: a multi-center study. Front Oncol 13:1252074
Chen L, Liu K, Zhao X, Shen H, Zhao K, Zhu W (2021) Habitat imaging-based (18)F-FDG PET/CT radiomics for the preoperative discrimination of non-small cell lung cancer and benign inflammatory diseases. Front Oncol 11:759897
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillion-Robin JC, Pieper S, Aerts H (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77(21):e104–e107
Zwanenburg A, Vallières M, Abdalah MA, Aerts H, Andrearczyk V, Apte A, Ashrafinia S, Bakas S, Beukinga RJ, Boellaard R et al (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2):328–338
Akinboro O, Larkins E, Pai-Scherf LH, Mathieu LN, Ren Y, Cheng J, Fiero MH, Fu W, Bi Y, Kalavar S et al (2022) FDA Approval summary: pembrolizumab, atezolizumab, and cemiplimab-rwlc as single agents for first-line treatment of advanced/metastatic PD-L1-High NSCLC. Clin Cancer Res 28(11):2221–2228
Liu C, Zhao W, Xie J, Lin H, Hu X, Li C, Shang Y, Wang Y, Jiang Y, Ding M et al (2023) Development and validation of a radiomics-based nomogram for predicting a major pathological response to neoadjuvant immunochemotherapy for patients with potentially resectable non-small cell lung cancer. Front Immunol 14:1115291
Trotman J, Barrington SF, Belada D, Meignan M, MacEwan R, Owen C, Ptáčník V, Rosta A, Fingerle-Rowson GR, Zhu J et al (2018) Prognostic value of end-of-induction PET response after first-line immunochemotherapy for follicular lymphoma (GALLIUM): secondary analysis of a randomised, phase 3 trial. Lancet Oncol 19(11):1530–1542
Shi W, Huang X, Wang Y, Wan X, He J, Xu Y, Zhang W, Chen R, Xu L, Zha X et al (2022) A novel nomogram containing efficacy indicators to predict axillary pathologic complete response after neoadjuvant systemic therapy in breast cancer. Front Endocrinol (Lausanne) 13:1042394
Chiou VL, Burotto M (2015) Pseudoprogression and immune-related response in solid tumors. J Clin Oncol 33(31):3541–3543
Cui Y, Tha KK, Terasaka S, Yamaguchi S, Wang J, Kudo K, Xing L, Shirato H, Li R (2016) Prognostic imaging biomarkers in glioblastoma: development and independent validation on the basis of multiregion and quantitative analysis of MR images. Radiology 278(2):546–553
Zhou M, Chaudhury B, Hall LO, Goldgof DB, Gillies RJ, Gatenby RA (2017) Identifying spatial imaging biomarkers of glioblastoma multiforme for survival group prediction. J Magn Reson Imaging 46(1):115–123
Parekh VS, Jacobs MA (2019) Deep learning and radiomics in precision medicine. Expert Rev Precis Med Drug Dev 4(2):59–72
Wang C, Ma J, Shao J, Zhang S, Li J, Yan J, Zhao Z, Bai C, Yu Y, Li W (2022) Non-invasive measurement using deep learning algorithm based on multi-source features fusion to predict PD-L1 expression and survival in NSCLC. Front Immunol 13:828560
Zhang L, Dong D, Zhang W, Hao X, Fang M, Wang S, Li W, Liu Z, Wang R, Zhou J et al (2020) A deep learning risk prediction model for overall survival in patients with gastric cancer: a multicenter study. Radiother Oncol 150:73–80
Song J, Shi J, Dong D, Fang M, Zhong W, Wang K, Wu N, Huang Y, Liu Z, Cheng Y et al (2018) A new approach to predict progression-free survival in stage IV EGFR-mutant NSCLC patients with EGFR-TKI therapy. Clin Cancer Res 24(15):3583–3592
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Contributions
Junkai Chen designed the study. Xiao Wu extracted, collected and analyzed data. Yongxian Chen prepared tables and figures. Weimin Caii, Yubo Shi, Kun Guo reviewed the results, interpreted data, and wrote the manuscript. All authors have made an intellectual contribution to the manuscript and approved the submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval and consent to participate
This research protocol was approved by the Medical Ethics Committee of the First affiliated hospital of Wenzhou medical university, Xiamen Second hospital, and Yueqing people’s hospital. It was conducted according to the guidelines of the Declaration of Helsinki. All data included in this research were anonymized; thus, the need for informed consent has been waived.
Consent for publication
All authors approved the final manuscript and the submission to this journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Caii, W., Wu, X., Guo, K. et al. Integration of deep learning and habitat radiomics for predicting the response to immunotherapy in NSCLC patients. Cancer Immunol Immunother 73, 153 (2024). https://doi.org/10.1007/s00262-024-03724-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00262-024-03724-3