Abstract
Pancreatic ductal adenocarcinoma (PDA) is a lethal and metastatic malignancy resistant to therapy. Elucidating how pancreatic tumor-specific T cells differentiate and are maintained in vivo could inform novel therapeutic avenues to promote T cell antitumor activity. Here, we show that the spleen is a critical site harboring tumor-specific CD8 T cells that functionally segregate based on differential Cxcr3 and Klrg1 expression. Cxcr3+ Klrg1- T cells express the memory stem cell marker Tcf1, whereas Cxcr3-Klrg1 + T cells express GzmB consistent with terminal differentiation. We identify a Cxcr3+ Klrg1+ intermediate T cell subpopulation in the spleen that is highly enriched for tumor specificity. However, tumor-specific T cells infiltrating primary tumors progressively downregulate both Cxcr3 and Klrg1 while upregulating exhaustion markers PD-1 and Lag-3. We show that antigen-specific T cell infiltration into PDA is Cxcr3 independent. Further, Cxcr3-deficiency results in enhanced antigen-specific T cell IFNγ production in primary tumors, suggesting that Cxcr3 promotes loss of effector function. Ultimately, however, Cxcr3 was critical for mitigating cancer cell dissemination following immunotherapy with CD40 agonist + anti-PD-L1 or T cell receptor engineered T cell therapy targeting mesothelin. In the absence of Cxcr3, splenic Klrg1 + GzmB + antitumor T cells wain while pancreatic cancer disseminates suggesting a role for these cells in eliminating circulating metastatic tumor cells. Intratumoral myeloid cells are poised to produce Cxcl10, whereas splenic DC subsets produce Cxcl9 following immunotherapy supporting differential roles for these chemokines on T cell differentiation. Together, our study supports that Cxcr3 mitigates tumor cell dissemination by impacting peripheral T cell fate rather than intratumoral T cell trafficking.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The incidence and mortality of pancreatic ductal adenocarcinoma (PDA) is on the rise [1] and predicted to become the 2nd leading cause of cancer-related deaths by 2030 [2]. While immunotherapies elicit clinical responses in many malignancies [3, 4], PDA is often resistant [5]. The robust fibroinflammatory tumor microenvironment (TME) characteristic of PDA is believed to limit T cell access, functionality and immunotherapy response [6]. However, our studies in faithful PDA animal models support that antigen-specific T cells preferentially accumulate in primary tumors and metastasis [7, 8]. Thus, understanding how T cells can access malignant sites and the factors that contribute to maintenance of functional antitumor T cells will inform effective immunotherapy design.
Chemokines are small, secreted proteins that instruct cellular migration through venules into and out of tissues. Cxcr3 is a chemokine receptor expressed by effector and memory T cells and binds interferon-inducible chemokines including Cxcl9 and Cxcl10 [9, 10]. Cxcr3 promotes T cell migration [11, 12] and PD-1 blockade efficacy [13, 14] in subcutaneous implantable tumor animal models. Cxcr3 and its ligands are associated with a T cell inflamed signature in many cancers [12] and plasma levels of Cxcl9 and Cxcl10 are associated with prolonged survival in PDA patients treated with chemotherapy [15]. Cxcr3 promotes effector T cell (TEFF) fate at the expense of central memory T cells (TCM) following a primary immune response to infection [16,17,18,19]. After viral infection, Cxcr3 guides CD8 T cells to the lymph node periphery promoting effector T cell differentiation [19]. During chronic T. gondii infection in mice, Cxcr3 regulates the differentiation of antigen-specific Cxcr3 + Klrg1- memory T cells into Cxcr3+ Klrg1+ intermediate effector/memory T cells in the spleen [20]. However, the role of Cxcr3 on a tumor-specific T cell response in PDA is unknown.
In contrast to the potential antitumor roles for Cxcr3, intratumoral Cxcr3 ligands are associated with a worse PDA patient prognosis [21]. Cxcr3 is expressed on a subset of Foxp3 + regulatory T cells (Treg) [22] and Cxcl10 expression by cancer-associated fibroblasts can recruit suppressive Treg [23]. Pancreatic tumor cells can also express Cxcr3, and Cxcr3:Cxcl10 interactions facilitate tumor cell migration to sensory neurons and contribute to cancer-related pain [24]. Thus, Cxcr3 has the potential to promote or mitigate PDA pathogenesis.
We previously showed that conventional type 1 dendritic cells (cDC1s) are critical for maintaining both spleen-residing and intratumoral tumor-specific T cells in PDA [25]. As cDC1s have been previously shown to produce Cxcr3 ligands in subcutaneous tumor models [13], we posited that the Cxcr3 pathway may play a role in tumor-specific T cell fate during pancreatic cancer growth and immunotherapy. We identify that both endogenous and adoptively transferred TCR engineered antigen-specific T cell infiltration into PDA is Cxcr3 independent. However, our study supports that Cxcr3 is critical for immunotherapy durability through maintaining peripheral GzmB + Klrg1 + effector T cells deemed critical for mitigating cancer cell dissemination. In primary tumors, Cxcr3 instead promotes T cell exhaustion. Thus, our study reveals a link between peripheral effector T cell differentiation and exhaustion in the tumor bed and tissue-specific roles for Cxcr3 on T cell differentiation and antitumor immunity.
Results
Cxcr3 and Klrg1 delineate endogenous tumor-specific T cell subpopulations during PDA progression
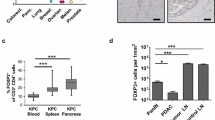
Elucidating how pancreatic tumor-specific T cells differentiate and are maintained in vivo could inform novel therapeutic avenues to promote their antitumor activity. Indeed, the phenotypic traits and qualities of tumor-specific T cells in secondary lymphoid organs as well as intratumorally in PDA are largely unclear. To investigate tumor-specific T cell differentiation during PDA progression, we employed the orthotopic KPC2a model in which tumor cells express click-beetle red luciferase (CB) that serves for tumor imaging [26, 27] and as a model neoantigen [8, 25, 28]. Tumor (CB)-specific T cells can be tracked longitudinally in vivo with a fluorescent CB101-109:H-2Db tetramer (Fig. 1A) [8, 25, 28]. KPC cells that express ovalbumin are rejected in syngeneic immunocompetent mice after orthotopic implantation [8] rendering the CB + tumor model a valuable alternative to study tumor-specific T cells. As T cell phenotype can identify similar differentiation states in diverse biological contexts, we assessed Cxcr3 and Klrg1 as landmark markers because functionally distinct pathogen-specific CD8 T cells can be distinguished based upon Cxcr3 and Klrg1 during chronic infection [20, 29]. Specifically, Cxcr3+ Klrg1- T cells exhibit sustained proliferative capacity and differentiate into Cxcr3+ Klrg1+ T cells, which give rise to terminally differentiated Cxcr3-Klrg1 + T cells during chronic infection [20, 29]. Therefore, we posited a similar differentiation pattern may be maintaining tumor-specific T cells. We identified that 60–80% of CB101-109:H-2Db tetramer + T cells expressed Cxcr3 in both spleen and tumor on day 7 posttumor (Fig. 1B). While 50–60% of splenic tetramer + T cells were Cxcr3+ until day 21, intratumoral tetramer + T cells progressively downregulated Cxcr3 (Fig. 1A-B). Klrg1+ tetramer + T cell frequency progressively increased in spleen and decreased in tumor (Fig. 1B). Cxcr3+ Klrg1-tetramer + T cells frequency progressively decreased in spleen and tumor over time (Fig. 1C). Within spleens, this corresponded to an increase in putatively more differentiated Cxcr3+ Klrg1 + and Cxcr3-Klrg1 + subpopulations (Fig. 1C). In contrast, the progressive loss of Cxcr3+ tetramer + T cells in tumor corresponded to an accumulation of Cxcr3-Klrg1- cells (Fig. 1C), a subset not previously identified in the chronic infection model [20, 29]. To identify the differentiation state of Cxcr3-Klrg1-tetramer + T cells in PDA, we performed ViSNE analysis on CD45 + cells. Intratumoral Cxcr3-Klrg1-tetramer + T cells exhibited elevated PD-1 and Lag-3 (Fig. 1D-F), markers of PDA-specific T cells defective in IFNγ and GzmB [8, 25, 28] and consistent with an acquisition of an exhausted T (TEX) cell state. ViSNE analysis showed Cxcr3 was mostly confined to T cells in murine PDA including a fraction of CD4 + Foxp3 + (Treg) and CD4 + Foxp3- T cells (Fig. 1D). In non-tumor-bearing mice, Cxcr3 was expressed by memory and effector T cells but not naïve T cells (Supplementary Fig. 1A-C), as expected [9]. Cxcr3 was also expressed by antigen experienced CD44 + CD4 + T cells (Supplementary Fig. 1D), maintained in spleen and progressively decreased on intratumoral Treg and CD4 + Foxp3- T cells (Supplementary Fig. 1E).
Cxcr3 and Klrg1 delineate endogenous tumor-specific T cell subpopulations during PDA progression. A Representative tetramer staining and gating strategy (left) and Cxcr3 and Klrg1 expression (right, gated on CD8 + tetramer + T cells) on day 21 post orthotopic KPC2a tumor implantation. B Frequency of Cxcr3+ or Klrg1+ tetramer + T cells gated on total CD8 + tetramer + T cells following KPC2a orthotopic tumor implantation. Data are mean ± S.E.M. n = 4–5 mice per timepoint. C Frequency of the indicated subsets gated on CD8 + tetramer + T cells. Data are mean ± S.E.M. n = 4–5 mice per timepoint. D ViSNE analysis of n = 3–4 concatenated samples per timepoint from KPC2a-bearing mice gated on live CD45 + cells. Red circle, CD8 + tetramer + cells. Orange circle, CD4 + T cells. E Representative plots of Cxcr3, PD1 and Lag3 on CD8 + tetramer + T cells on day 21 posttumor. F Frequency of tetramer + T cells that express PD-1 or Lag3 in KPC2a tumor-bearing mice. Data are mean ± S.E.M. n = 4–5 mice per timepoint. G Representative gating strategy for analyzing Cxcr3 and Klrg1 proportions among CD44hi CD8+ T cells in the KPC GEMM or the KPC2a orthotopic model. n = 3 mice per group. H Mean proportion of each subset among the indicated T cell population and model. n = 3 mice per group. I Gating strategy for assessing Cxcr3 and Klrg1 as markers to identify tetramer-binding T cells at 2 weeks post KPC2a orthotopic tumor implantation. Data are quantified (right) and are mean ± S.E.M. n = 3 mice per group. J Frequency of antigen-experienced (CD8 + CD44high) T cells or CD8 + tetramer + that co-express PD-1 and Lag-3 from tumors in the KPC GEMM or the KPC2a orthotopic model
The data above suggest that Cxcr3 and Klrg1 may define distinct T cell differentiation states during invasive tumor growth. However, one limitation of the above approach was the highly aggressive orthotopic model of invasive PDA fails to recapitulate the progression of PDA through a preinvasive state nor the robust fibroinflammatory response characteristic of the human disease. Therefore, we next investigated if phenotypically similar subsets were present in the autochthonous KPC genetically engineered mouse model (GEMM) of PDA. Since there are no known immunogenic epitopes in the KPC GEMM, we could not distinguish tumor-specific from non-specific T cells. Therefore, we gated on total CD8 + CD44hi antigen-experienced T cells in spleen and tumor from the KPC GEMM and the KPC2a model (Fig. 1G), with the understanding that this approach includes non-specific T cells. Despite this caveat, both Cxcr3+ Klrg1- and Cxcr3-Klrg1+ subsets were clearly present and at similar proportions in the two models (Fig. 1G-H). In contrast, in the spleen, the Cxcr3+ Klrg1+ intermediate population was quite rare (Fig. 1G-H), especially when comparing to the proportion of Cxcr3+ Klrg1+ tumor-specific (tetramer-binding) T cells (Fig. 1H). These data suggest that the Cxcr3 + Klrg1 + intermediate subpopulation in the spleen may be particularly enriched for tumor specificality. To test this hypothesis, we quantified the percentage of each Cxcr3/Klrg1 subset that bound tetramer in KPC2a tumor-bearing mice. Indeed, the Klrg1 + Cxcr3 + subset in the spleen was the most enriched for tetramer + T cells, with approximately 60% binding the CB101-109:H-2Db tetramer (Fig. 1I). As expected, there were very few PD-1 + Lag3 + T cells infiltrating tumors in the KPC GEMM (Fig. 1J), consistent with a lack of tumor antigen specificity and a distinction from the KPC2a model (Fig. 1J).
We next evaluated CXCR3 cellular distribution in human tissues. As expected, CXCR3 and KLRG1 were decreased in tumor vs. spleen (Supplementary Fig. 1F). Analysis of 6 merged human PDAs using a public scRNAseq dataset [30] showed that CXCR3 was mostly confined to T cells (Supplementary Fig. 1G), similar to our results in mice. A minor LILRA4 + population (cluster 21), which are likely plasmacytoid dendritic cells (DCs) [31], were also CXCR3+ in human PDA (Supplementary Fig. 1G). To identify T cell differentiation states in human PDA, we re-clustered just CD3E + cells (Supplementary Fig. 1H). Cluster 0 contained CD8A + CXCR3+ KLRG1- T cells; Cluster 1 contained CD4 + FOXP3 + TREG; Cluster 2 contained CD8A + CXCR3+ KLRG1High T cells; Cluster 3 contained CD4 + CCR7 + naïve or memory T cells; Cluster 4 contained NCR1 + NKG7 + NK cells; and Cluster 5, which was rare, contained CD8A + cells that were CXCR3+KLRG1Int. PDCD1 and LAG3 appeared enriched in CD8 T cell clusters 2 and 5 potentially reflective of TEX (Supplementary Fig. 1I). Cluster 2 that was highest for KLRG1 (Supplementary Fig. 1 J) was enriched for GZMA and GZMK suggesting enhanced cytotoxicity and an effector T cell differentiation state. Thus, the above markers may also be useful for delineating human T cell differentiation states in cancer.
Cxcr3 is dispensable for intratumoral antigen-specific T cell accumulation
As Cxcr3 is regulated on antigen-specific T cells during PDA growth (Fig. 1), we next tested if Cxcr3 deficiency impacted tumor growth and T cell infiltration. While spleens were significantly larger in Cxcr3−/− vs. Cxcr3+/+ mice on day 21 post orthotopic tumor implantation, primary tumor weights were similar in Cxcr3+/+ and Cxcr3−/− mice (Fig. 2A). CD8 T cell frequency was increased in Cxcr3−/− spleens on day 21 (Fig. 2B), which correlated with increased spleen size (Fig. 2A). CD8 T cell frequency and number were similar in tumors from Cxcr3−/− vs. Cxcr3+/+ mice at both timepoints (Fig. 2B). We next gated on tumor-specific T cells using the CB101-109:H-2Db tetramer (Fig. 2C). In the spleen, we detected a trend for increased frequency of tumor-specific T cells in Cxcr3−/− mice at day 21 (Fig. 2D), suggesting greater T cell retention, survival and/or proliferation. In PDA, tumor-specific T cell frequency and number were not significantly different among Cxcr3+/+ vs. Cxcr3−/− mice at both timepoints (Fig. 2C-D). Immunofluorescent (IF) staining showed that CD8 T cells were infiltrating the tumor parenchyma in both Cxcr3+/+ and Cxcr3−/− mice (Fig. 2E). Thus, Cxcr3 is dispensable for CD8 T cell infiltration into orthotopic PDA and had negligible impact on overall tumor control.
Cxcr3 is dispensable for intratumoral antigen-specific T cell accumulation. A Orthotopic KPC2a tumor and spleen weights on 14 or 21 days posttumor in Cxcr3+/+ and Cxcr3−/− mice. Each dot is an independent animal. Data are mean ± S.E.M. *p < 0.05, **p < 0.005, ANOVA with a Tukey’s posttest. B Frequency and number of CD8 + T cells per gram spleen or tumor from KPC2a-bearing Cxcr3+/+ and Cxcr3−/− mice on day 14 (d14) or day 21 (d21) posttumor. Each dot is an independent animal. Data are mean ± S.E.M. n = 3–4 mice per group. *p < 0.05, **p < 0.005, ANOVA with a Tukey’s posttest. C Representative H-2Db:CB101-109 tetramer staining gated on live CD8 + T cells infiltrating tumors from KPC2a-bearing Cxcr3+/+ and Cxcr3−/− mice on day 14 or day 21 posttumor. D Frequency of tetramer + T cells of CD8 + T cells (left) and number of CD8 + tetramer + T cells per gram tissue. E Immunofluorescent detection of CD8 T cells infiltrating orthotopic tumors isolated from Cxcr3+/+ and Cxcr3−/− mice on day 14. Scale bar, 50 μm. n = 3 mice per group. F Spleen and tumor weight from mice bearing orthotopic (Orth) or subcutaneous (S.C.) KPC2a tumors on day 14 posttumor. Each dot is an independent animal. Data are mean ± S.E.M. **p < 0.005, ***p < 0.0005, Student’s T test. G Representative IF for Cxcl9/Cxcl10 from normal mouse pancreas, KPC2 (CB-) parental tumors, KPC2a (CB +) tumors, or KPC2a tumors from mice treated with αPD-L1 on day 14. H Intensity units of Cxcl9/Cxcl10 + staining from samples in G. Each dot is an independent mouse. NP, normal pancreas. I Number of CD8 + T cells per field of view (FOV) from samples in G. J Correlation between CD8 + T cells and Cxcl9/Cxcl10 intensity from mice bearing KPC2a orthotopic tumors from G-I
A role for Cxcr3 in T cell accumulation in tumors has been described in subcutaneous (s.c.) tumor implantation models [11,12,13,14]. Mechanisms governing T cell migration into skin tumors may be distinct from tumors that originate in internal organs and could explain why our results contrast with prior studies. KPC2a orthotopic tumors were five–tenfold larger than s.c. tumors in syngeneic C57Bl/6 J mice on day 14 (Fig. 2F), indicating that the tissue is a major determinant of pancreatic tumor growth. Spleens were also significantly larger in mice bearing orthotopic tumors vs. s.c. tumors (Fig. 2F), which likely reflects increased tumor burden as tumor cells produce factors that promote splenomegaly [32]. To test the hypothesis that Cxcr3 is critical for T cell migration into s.c. tumors, we implanted KPC2a tumor cells s.c. into Cxcr3+/+ or Cxcr3−/− mice (Supplementary Fig. 2A). Tumor growth rate was similar early on, but by day 14, tumor volumes were modestly yet significantly reduced in Cxcr3+/+ mice compared to Cxcr3−/− mice (Supplementary Fig. 2B-C). Tumor ulceration was delayed in Cxcr3−/− mice (Supplementary Fig. 2D), which is consistent with a delay in tumor growth. Like our results in the orthotopic tumor model, the frequency and number of intratumoral CD8 T cells (Supplementary Fig. 2E) and tumor-specific CD8 T cells (Supplementary Fig. 2F) were similar in s.c. tumors from Cxcr3+/+ and Cxcr3−/− mice on day 14. Thus, despite the accelerated antitumor response in Cxcr3+/+ mice bearing s.c. tumors, Cxcr3 was ultimately dispensable for tumor-specific T cell intratumoral accumulation in tumors growing within the skin. Together, the studies indicate tumor-specific T cells infiltrate PDA in a Cxcr3-independent manner.
Cxcr3 may not be required for antigen-specific T cell infiltration into PDA because ligands for Cxcr3, including Cxcl9 and Cxcl10 are absent in PDA. Therefore, we stained tumor sections for Cxcl9 and Cxcl10 from untreated KPC2a-bearing mice and mice treated with αPD-L1 or CD40 agonist as such therapies may enhance IFNγ [8, 28], which induces Cxcl9/Cxcl10. As expected, Cxcl9/Cxcl10 were mostly undetectable in normal mouse pancreas or parental KPC2 (CB-negative) orthotopic tumors on day 14 posttumor (Fig. 2G). In contrast, Cxcl9/Cxcl10 were modestly yet significantly increased in CB + KPC2a tumors and even further increased after αPD-L1 (Fig. 2G-H). CD8 T cells were increased in KPC2a tumors compared to normal pancreas and KPC2 tumors (Fig. 2I) and intratumoral CD8 T cell number correlated with Cxcl9/Cxcl10 in KPC2a tumors from control and αPD-L1-treated mice (Fig. 2J). CD40 agonist also increased Cxcl9/Cxcl10 in KPC2a tumors and Cxcl9/Cxcl10 colocalized with both tumor-associated macrophages (TAMs) and tumor cells (Supplementary Fig. 2G-H). Since we observed differences in Cxcl9/Cxcl10 staining in KPC2 vs. KPC2a tumors, we next tested if there were tumor cell intrinsic differences in chemokine expression by KPC2 and KPC2a cells. KPC2a cells expressed tenfold more Cxcl9 and tenfold less Cxcl10 as compared to the parental line in response to IFNγ (Supplementary Fig. 2I). However, IFNγ increased Cxcl9 4–5 logs and Cxcl10 ~ 1–2 logs in both cell lines. These data are consistent with Cxcl9 induction following IFNγ, whereas Cxcl10 may be more sensitive to innate stimuli such as type I IFNs [9, 33]. Together, our results suggest that both antigenicity and immunotherapy increase Cxcl9/Cxcl10 in PDA.
Cxcr3 alters T cell fate in a tissue-specific manner
Cxcr3 promotes effector T cell differentiation at the expense of memory T cells during acute infection in mouse models [16,17,18,19]. Therefore, we hypothesized that Cxcr3 may influence tumor-specific T cell fate. We orthotopically implanted KPC2a tumor cells into the pancreas of Cxcr3+/+ and Cxcr3−/− mice and analyzed CD8 + tetramer + T cell phenotype on day 14 posttumor. Using CD44 and CD62L markers to distinguish naïve, effector and memory T cells, most tetramer + T cells were a CD44 + CD62L- effector phenotype in both Cxcr3+/+ and Cxcr3−/− mice (Fig. 3A). Klrg1 is expressed by effector T cells, whereas the memory/stem cell transcription factor Tcf1 is expressed by naïve and memory T cells. In both spleen and tumors from Cxcr3−/− mice, tetramer + T cells had a higher frequency of Tcf1 + Klrg1- memory T cells and a lower frequency of Tcf1-Klrg1 + effector T cells (Fig. 3A-B). We previously showed that Klrg1 + Lag3- identifies functional effector T cells and Lag3 + Klrg1- identifies TEX in KPC2a PDA [8, 25, 28]. In both Cxcr3+/+ and Cxcr3−/− mice, tumor-infiltrating T cells upregulated PD-1 and Lag3 and lost Klrg1 as compared to splenic tetramer + T cells (Fig. 3C). The frequency of PD-1 + and/or Lag3 + tetramer + (Fig. 3D) and tetramer- (Fig. 3E) CD8 + T cells were significantly decreased in tumors from Cxcr3−/− mice, suggesting Cxcr3 drives TEX. Supporting this interpretation, intratumoral Cxcr3−/− tetramer + T cells exhibited significantly increased IFNγ production in response to CB101-109 peptide ex vivo (Fig. 3E). Thus, while Cxcr3 promoted effector T cell differentiation in the spleen, it paradoxically promoted TEX differentiation in tumors.
Cxcr3 alters T cell fate in a tissue-specific manner A Phenotype of CD8 + tetramer + T cells on day 14 post KPC2a orthotopic implantation into Cxcr3+/+ and Cxcr3−/− mice. Flow cytometry plots show CD8 + tetramer + gates (left) for analyzing CD44, CD62L, Klrg1 and Tcf1. B Frequency of the indicated subpopulations from spleen and tumor gated on CD8 + tetramer + T cells 14 days following orthotopic KPC2a implantation. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, **p < 0.005. One-way ANOVA with a Tukey’s posttest. C Representative flow cytometric plots from spleen and tumor 14 days following orthotopic KPC2a implantation. Samples are gated on total CD8 + T cells (top row) to assess tetramer + cells and CD8 + tetramer + T cells (middle and bottom rows) to assess activation and exhaustion markers. D Proportion of CD8 + tetramer + T cells expressing the indicated receptors 14 days following orthotopic KPC2a implantation. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, **p < 0.05, ***p < 0.0005. ANOVA with a Tukey’s posttest. E Proportion of CD8 + tetramer- T cells expressing the indicated receptors 14 days following orthotopic KPC2a implantation. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, **p < 0.05, ***p < 0.0005. ANOVA with a Tukey’s posttest. F Proportion of total CD8 + T cells producing IFNγ and TNFα following a 5 h restimulation with CB101-109. Data are mean ± S.E.M. Each dot is an independent animal. **p < 0.005, Student’s T test
Cxcr3 sustains splenic Gzmb + antitumor T cells and mitigates cancer cell dissemination
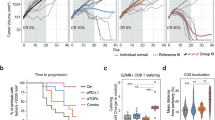
We previously showed that αPD-L1 + CD40 agonist significantly prolonged survival of KPC2a-bearing mice [28]. In cured animals, tumor-specific T cells persist for over 100 days and reject orthotopic tumor rechallenge indicative of memory T cell formation [25, 26]. In non-cured mice, bioluminescent CB + tumors emerge indicative of relapse (Fig. 4A). Tumor-specific T cells persist indefinitely (up to at least ~ 125 days posttumor) and a fraction express Cxcr3 (Supplementary Fig. 3A). As Cxcr3 altered antitumor T cell fate (Fig. 3), we next tested if Cxcr3 played a role in the long-term durability of αPD-L1 + CD40 agonist therapy. By IVIS imaging of tumor growth of mice for up to 91 days posttumor, Cxcr3 loss impaired the long-term antitumor activity of αPD-L1 + CD40 agonist (Fig. 4B). We next compared tumor burden and metastasis in Cxcr3+/+ and Cxcr3−/− mice that relapsed between 35–91 days posttumor. We observed increased ascites and/or metastasis in Cxcr3−/− vs. Cxcr3+/+ relapsed mice (p = 0.0155, Fig. 4C). Primary tumor weights were smaller in Cxcr3−/− vs. Cxcr3+/+ relapsed mice (Fig. 4D), indicating increased tumor dissemination in Cxcr3−/− mice was not merely due to larger primary tumors and consistent with increased T cell IFNγ production at the earlier timepoint (Fig. 3F). To understand the basis for increased tumor cell dissemination in Cxcr3−/− mice, we quantified tumor-specific T cells in relapsed Cxcr3+/+ and Cxcr3−/− mice by tetramer staining. The frequency and number of tetramer + T cells (Fig. 4E-G) were similar between Cxcr3+/+ and Cxcr3−/− mice, suggesting that increased tumor cell dissemination was not due to a deficit in maintaining tumor-specific T cell quantity. Overall CD8 + T cell frequency was also similar in multiple tissues from Cxcr3+/+ and Cxcr3−/− relapsed mice (Supplementary Fig. 3B).We next evaluated effector T cell phenotype at these later timepoints, and included a positive control cohort of animals that were vaccinated with CD40 agonist + PolyIC + CB101-109 peptide [34]. The phenotype of intratumoral tetramer + T cells, including GzmB and T-bet, was similar in Cxcr3−/− vs. Cxcr3+/+ mice (Supplementary Fig. 3C-D). In contrast, circulating and splenic tetramer + T cells from Cxcr3−/− mice had decreased Klrg1 compared to tetramer + T cells from Cxcr3+/+ mice (Fig. 4I). Circulating and splenic tetramer + T cells from Cxcr3−/− mice also exhibited decreased GzmB yet similar T-bet (Fig. 4J-K). These data suggest that the defect in Gzmb by Cxcr3−/− T cells is not merely due to a global defect in the induction of the T-bet transcription factor.
Cxcr3 sustains Klrg1 + Gzmb + antitumor T cells and mitigates cancer cell dissemination. A Simplified schematic for analysis of T cell persistence in mice that are cured or relapsed following immunotherapy. B Fraction of wild type and Cxcr3−/− animals bearing KPC2a tumors that are cured following αPD-L1 + CD40 agonist by day 70 posttumor. Fisher’s exact test. Data are pooled from n = 2–4 independent experiments. C Proportion of relapsed mice that have metastasis and/or ascites at required euthanasia due to tumor burden 35–91 days post αPD-L1 + CD40 agonist. Data are pooled from n = 2–4 independent experiments. D Primary tumor weight (grams) at endpoint from relapsed mice in C. Each dot is an independent animal, *p < 0.05, Student’s T test. E Representative plots gated on CD8 + CD3 + T cells 35–91 days post αPD-L1 + CD40 agonist from relapsed Cxcr3+/+ or Cxcr3−/− mice. Numbers in the plots are the percent tetramer + . F Frequency of tetramer + T cells of total CD8 + T cells. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, Student’s T test. G Number of CD8 + tetramer + T cells per gram of tissue from relapsed mice in D. Data are mean ± S.E.M. Each dot is an independent animal. H Representative histograms gated on splenic CD8 + tetramer + T cells from relapsed Cxcr3+/+ and Cxcr3−/− mice. Splenic CD8 + tetramer + T cells from Cxcr3+/+ mice immunized with CD40 agonist + Poly:IC + CB101-109 peptide (Trivax) and assayed on day 7 post vaccination were included as a positive control. I Frequency of Klrg1 + tetramer CD8 + T cells in tissues from relapsed Cxcr3+/+ and Cxcr3−/− mice. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, Student’s T test. J Representative histograms of intracellular GzmB and T-bet gated on splenic CD8 + tetramer + T cells from relapsed Cxcr3+/+ and Cxcr3−/− mice. CD8 + tetramer + T cells without the intracellular staining (no ICS) served as a negative control. Splenic CD8 + tetramer + T cells from Cxcr3+/+ mice immunized with TriVax (CD40 agonist + Poly:IC + CB101-109 peptide) and assayed on day 7 post vaccination served as a positive control. K Frequency of GzmB + cells gated on tetramer + T cells in tissues from relapsed Cxcr3+/+ and Cxcr3−/− mice. Data are mean ± S.E.M. Each dot is an independent animal. *p < 0.05, Student’s T test
CD40 agonist promotes splenic Cxcl9 + cDC1s and Cxcl9 + Cxcl10 + cDC2s
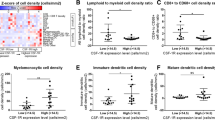
A prior study suggested that conventional type 2 dendritic cells (cDC2s) in the spleen produce Cxcl9/cxcl10 in mice response to T. gondii chronic infection and may interact with Cxcr3-Klrg1 + T cells in the red pulp of the spleen [20]. Our prior study showed that conventional type 1 dendritic cells (cDC1s) were critical for both priming and maintenance of transferred effector or memory tumor-specific T cells in both the spleen and tumor [25]. To investigate the cellular source(s) of chemokine ligands in that may contribute to differentiation and/or maintenance of Klrg1 + GzmB + antitumor T cells in the spleen, we obtained the reporter of Cxcr3 ligands (REX3) mouse strain in which red fluorescent protein (RFP) reports Cxcl9 and blue fluorescent protein (BFP) reports Cxcl10 [19, 35]. We orthotopically implanted KPC2a tumor cells in REX3 mice and on day 7, administered αPD-L1, CD40 agonist or the combination (Fig. 5A) [28]. αPD-L1, CD40 agonist or the combination decreased tumor size and CD40 agonist increased spleen size in REX3 mice on day 14 posttumor (Supplementary Fig. 4A), similar to our prior study [28]. The proportion of immune subsets expressing Cxcl9 and/or Cxcl10 in the spleen, as well as the pancreatic draining lymph node (dLN) and tumor were assessed using the gating strategy in Fig. 5B–C. In non-tumor-bearing mice, 20–50% of cDC1s were producing Cxcl9 in the dLN, spleen and pancreas (Fig. 5D–E, Supplementary Fig. 4B). CD40 agonist significantly increased Cxcl9 + splenic cDC1s, whereas Cxcl10 was largely unchanged by this subset in the spleen (Fig. 5D–E, Supplementary Fig. 4B). In contrast, CD40 agonist promoted Cxcl10 by cDC1s in tumors (Fig. 5D–E, Supplementary Fig. 4B). Tumorigenesis decreased Cxcl9 + cDC1 frequency in the dLN and CD40 agonist reverted this phenotype (Fig. 5D–E, Supplementary Fig. 4B). Tumorigenesis increased the frequency of cDC2s that were producing Cxcl10 or Cxcl9 and Cxcl10 in the dLN and tumor, yet not the spleen (Fig. 5F–G, Supplementary Fig. 4B). However, CD40 agonist or the combination therapy significantly increased the frequency of Cxcl9 + Cxcl10 + cDC2s in the spleen (Fig. 5F–G, Supplementary Fig. 4B). Thus, these data reveal that both tumorigenesis and CD40 agonist alters Cxcl9/Cxcl10 production by DCs in a tissue-specific manner. IF staining confirmed an abundance of Cxcl10 + and Cxcl9 + cells primarily located in the spleen red pulp following immunotherapy (Fig. 5H). We noted an expansion of CD8 + T cells in the red pulp in the spleen following αPD-L1 or αPD-L1 + CD40 agonist, which is consistent with an expansion of splenic CD8 + tetramer + T cells following PD-L1 blockade [8, 28]. In contrast, tumor tissue staining by IF showed predominantly Cxcl10 + cells (Supplementary Fig. 4C). As we showed previously [25], tumors are densely populated by macrophages and granulocytes and these cells are producing Cxcl10 rather than Cxcl9 (Supplementary Fig. 4D-F). The differential expression of Cxcl10 and Cxcl9 in tumor vs. spleen likely reflects different composition of myeloid cells and raises the possibility that the chemokines may play a differential role in the TEFF vs. TEX fate choice. However, further studies will be necessary to specifically identify the distinct contribution of DC subset-derived chemokines on antitumor T cell differentiation.
CD40 agonist increases Cxcl9 and Cxcl10 in a cell- and tissue-specific manner. A Schematic of immunotherapy in REX3 orthotopic KPC2a tumor-bearing mice. On day 7 post orthotopic tumor implantation, cohorts received isotype (-), αCD40 (100 μg), αPD-L1 (200 μg), or the combination as we described (28). αPD-L1 (200 μg) was also administered on day 10 and 12 posttumor for a total of 3 doses. B Flow cytometric gating strategy for analysis of indicated immune cells in spleen and tumor of mice depicted in (A). C Flow cytometric gating strategy for detecting Cxcl9 and/or Cxcl10 in non-tumor cells of REX3 mice depicted in (A). D Representative flow cytometry plots of cDC1s isolated from pancreatic draining lymph node (LN), spleen and tumor (or normal pancreas, e.g., no tumor) on day 14 posttumor. E Mean frequency of cDC1s expressing Cxcl9 and/or Cxcl10. F Representative flow cytometry plots of cDC2s isolated from pancreatic draining LN, spleen, and tumor (or normal pancreas, e.g., no tumor) on day 14 posttumor. G Mean frequency of cDC2s expressing Cxcl9 and/or Cxcl10. H Representative IF staining of CD8, B cells (B220) and Cxcl9 and Cxcl10 reporter expression in REX3 spleen and tumor from mice treated with the indicated therapies as in (A). Scale bar, 50 µm
Subcellular Cxcr3 ligand distribution is altered by immunotherapy and Cxcr3 promotes GzmB + engineered T cells that mitigate metastasis
CB is a model antigen and the CB101-109 binding affinity to H-2Db is predicted to be quite high [8], thereby modeling a particularly avid antitumor T cell response. Therefore, we next asked if Cxcr3 played a role in limiting metastasis during adoptive T cell therapy using mesothelin (Msln406-414:H-2Db)-specific 1045 TCR engineered T cells which we previously showed have therapeutic efficacy in the autochthonous KPC GEMM [7]. We first analyzed Cxcl9/Cxcl10 distribution in tumors from unmanipulated KPC GEMM. Unexpectedly, Cxcl9/Cxcl10 was largely confined to the apical surface in tumor cells from untreated KPC mice (Fig. 6A). The frequency of tumor cells expressing apical Cxcl9/Cxcl10 was significantly decreased in KPC recipients of 1045 T cells or 1045 T cells + CD40 agonist (Fig. 6B). Instead, engineered T cells + CD40 agonist, which we previously showed to promote the persistence of engineered T cells in autochthonous PDA [36], promoted tumor cytoplasmic Cxcl9/Cxcl10 (Fig. 6C-D, Supplementary Fig. 5A), indicating that immunotherapy alters the subcellular localization of Cxcr3 ligands in tumor cells. These data suggest that immunotherapy may result in tumor cell chemokine secretion into the parenchyma rather than into the ductal space, thereby orchestrating the intratumoral migration patterns of Cxcr3 + cells within the stroma. Since 1045 TCR engineered T cells express Cxcr3 prior to transfer (Supplementary Fig. 5B), we next quantified the frequency of Thy1.1 + 1045 T cells adjacent to Cxcl9/Cxcl10 + cells. We observed substantial variability in the frequency of Thy1.1 + T cells adjacent to Cxcl9/Cxcl10 + cells in PDA (Fig. 6E), suggesting that Cxcr3-independent factors influence effector T cell migration within the stroma. In stromal-rich regions that lacked CK + tumor cells, 1045 T cells + CD40 agonist increased Cxcl9/Cxcl10 expression by macrophage and 1045 T cells appeared to colocalize with the Cxcl9 + macrophage in these stromal regions (Supplementary Fig. 5C), similar to our results in resected human PDA [37]. Together, these data support that engineered T cell therapy alters Cxcl9/Cxcl10 spatial, cellular and subcellular localization in autochthonous PDA.
Subcellular Cxcr3 ligand distribution is altered by immunotherapy and Cxcr3 promotes GzmB + engineered T cells that mitigate metastasis. A Apical Cxcl9/10 by cytokeratin + (CK) ductal tumor cells from KPC GEMM tumors from untreated mice. Scale bar, 50 μm. B Percentage of CK + tumor cells that express apical Cxcl9/Cxc10. *p<0.05, **p<0.005, ***p<0.0005, one-way ANOVA with a Tukey's post test. C Number of tumor cells that express cytoplasmic Cxcl9/Cxc10 following immunotherapy. **p<0.005, one-way ANOVA with a Tukey's post test. D Cxcl9/Cxcl10 in KPC GEMM tumors on day 8 post Thy1.1 + 1045 T cells. Scale bar, 50 μm. D Representative IF of tumor sections from KPC mice treated with 1045 T cell therapy (1045 T cells) at 8 days post treatment. IF overlays of Cxcl9/Cxcl10, Thy1.1 expressed on engineered T cells, macrophage marker F4/80, tumor cell marker cytokeratin (CK) and DAPI nuclear stain. E Frequency of Thy1.1 + engineered T cells adjacent to Cxcl9/Cxc10 + cells in PDA from tissues depicted in D. F Experimental schematic for mice receiving orthotopic KPC2 (CB-) PDA cells, adoptive transfer of 1045 Thy1.1 + T cells and anti-Cxcr3 treatment days 6, 9 and 12, or untreated controls. G Representative histograms of Cxcr3 staining by adoptively transferred 1045 T cells in untreated (-) and anti-Cxcr3 treated mice compared to non-T cells (top panel) in mice depicted in F. H Proportion of splenic and intratumoral transferred Thy1.1 + CD8 + T cells in untreated (-) and anti-Cxcr3 treated mice depicted in F. I Proportion of total CD8 + T cells and total Thy1.1 + cell numbers per gram tissue of mice depicted in F. J Proportion of mice depicted in F which developed metastasis, n = 3 mice per group. K Representative flow cytometric plots of Klrg1 and Granzyme B (GzmB) staining on splenic and intratumoral Thy1.1 + CD8 + adoptively transferred 1045 CD8 T cells isolated from untreated (-) or anti-Cxcr3 treated mice 2 weeks following orthotopic tumor implantation. L Proportion of splenic and intratumoral Granzyme B + adoptively transferred Thy1.1 + CD8 T cells 2 weeks following orthotopic tumor implantation. n = 3 mice per group, significance determined by Student’s T test. *, p<0.05. M Simplified model depicting the role for Cxcr3 in promoting effector T cell differentiation in the spleen and exhausted T cell differentiation in the tumor microenvironment
To assess the requirement for Cxcr3 on 1045 T cell infiltration into PDA, 1045 T cells were transferred into mice bearing established and poorly immunogenic (CB-) KPC2 orthotopic tumors [8] using a similar therapy protocol that has efficacy in autochthonous KPC PDA [7] with or without anti-Cxcr3 (Fig. 6F). As expected, anti-Cxcr3 interfered with Cxcr3 on engineered T cells (Fig. 6G). Consistent with our studies of endogenous CB-specific T cells, Cxcr3 blockade did not impact 1045 T cell accumulation in PDA (Fig. 6H-I). However, Cxcr3 blockade resulted in more mice with detectable macrometastasis following engineered T cell therapy on day 14 (p = 0.0533, Fig. 6J). Notably, Cxcr3 blockade interfered with the frequency of splenic engineered T cells that expressed Granzyme B (GzmB) (Fig. 6K-L), paralleling the defect in endogenous GzmB + tumor-specific T cells in Cxcr3−/− mice (Fig. 4K). Together, our results support that Cxcr3, potentially by interacting with Cxcl9/Cxc10 + cDC1s in the spleen, promotes GzmB + cytotoxic T cells that may mitigate pancreatic cancer metastasis (Fig. 6M). Since Cxcr3 also promotes TEX in primary tumors (Figs. 3D-F and 6M) the data support a tissue-specific role for Cxcr3 on T cell fate during malignancy.
Discussion
Here, we identify paradoxical roles for Cxcr3 on tumor antigen-specific T cell fate and antitumor functionality during PDA progression and immunotherapy response. Cxcr3 sustained splenic Klrg1 + Gzmb + antitumor T cells that may aid in the control of disseminated disease. Thus, Cxcr3, potentially through positioning of T cells in secondary lymphoid organs [19], promotes cytotoxic T cells likely capable of targeting metastatic tumor cells. Unexpectedly, we identify that Cxcr3 was dispensable for intratumoral T cell accumulation and instead promoted TEX differentiation in primary tumors. Therefore, tissue-specific modulation of Cxcr3 may provide a novel therapeutic axis to uncouple TEX from TEFF differentiation.
Prior studies in other chronic antigen settings have identified rare PD-1 + Tcf1 + progenitor T cells that mediate response to PD-L1 blockade [38,39,40]. Since PD-1 is not clearly expressed on most splenic tumor-specific T cells (Fig. 1) [8, 25, 28], and migration of T cells from periphery into tumors is required for αPD-L1 transient antitumor activity [8], we sought other markers to identify peripheral tumor-specific T cell subsets that mediate immunotherapy response. During persistent T. gondii infection in mice, three subsets of splenic antigen-specific T cells were identified based on Cxcr3 and/or Klrg1 [29]. The Cxcr3 + Klrg1- subset exhibited the greatest proliferative capacity and seeded an intermediate Cxcr3+ Klrg1 + subset with memory and effector cell traits that gave rise to the cytotoxic terminally differentiated Cxcr3-Klrg1 + subset [29]. Here, we show in the spleen of tumor-bearing mice early on during tumor establishment, most tetramer + T cells expressed Cxcr3 and/or Klrg1 and the intermediate Cxcr3+ Klrg1 + subpopulation in the spleen is particularly enriched for tumor antigen specificity. Over time, Cxcr3 and Klrg1 were progressively decreased on tumor-specific T cells infiltrating PDA and these Cxcr3-Klrg1- T cells co-expressed PD-1 and Lag-3, markers that identify TEX cells defective in IFNγ production [8]. Cxcr3 and Klrg1 are also downregulated by TEX during chronic viral infection [41] and Cxcr3 is downregulated on T cells after multiple rounds of antigen exposure in vivo [42], suggesting that Cxcr3 loss in PDA is due to persistent TCR signaling. At least at the gene level, three CD8 + T cell clusters infiltrating human PDA expressed CXCR3 and variable levels of KLRG1, suggesting a similar pattern in human disease. However, additional studies are needed to determine the extent such markers can be utilized to map human CD8 T cell differentiation states. In human PDA, PDCD1 + LAG3 + were among KLRG1 + CD8 T cell clusters consistent with a connection between exhausted and effector T cell states [43].
We previously showed that Batf3 + Xcr1 + cDC1s are required to not only prime tumor-specific CD8 T cells in the spleen, but also maintain adoptively transferred Klrg1 + effector T cells in both spleen and tumor [25]. Our prior study also indicated that cDC1s promote Lag3 + TEX in the PDA [25], paralleling our findings here that show Cxcr3 can promote Klrg1 + T cells in the spleen and Lag3 + TEX in primary tumors. As we find that cDC1s uniquely express Cxcl9 without Cxcl10, the Cxcl9-Cxcr3 axis may be responsible for the tissue-specific outcomes on cytotoxic T cell fate. This chemokine axis may promote T cell:cDC1 interactions that impact T cell differentiation in a tissue-specific manner. In a prior study, Cxcl9 but not Cxcl10 expression by cDC1s was critical for αPD-L1 efficacy which was attributed to intratumoral T cell trafficking toward cDC1s in the tumor bed in s.c. implantable tumor models [13]. We find that cDC1s are poised to produce Cxcl9, whereas more abundant intratumoral myeloid cells including macrophages, cDC2s and granulocytes [25] are biased toward Cxcl10 production. A prior study suggested that cDC2s, which reside in the bridging channels in the spleen, sustains Klrg1 + pathogen-specific T cells during chronic infection [20]. As cDC1s and cDC2s contribute to antitumor immunity in part through induction of tumor-reactive CD8 + and CD4 + T cells, respectively, further investigation into the specific contribution of these cells on T cell differentiation during tumor growth and immunotherapy is of interest [44]. Additional factors beyond antigen presentation and Cxcl9/Cxcl10 that DCs are known to express including IL-15 [45] or IL-12 [46] may also be involved in impacting T cell differentiation and maintenance.
We show here that Cxcr3 is not a principal mechanism governing T cell migration into PDA and raise the possibility that other chemokine receptors may play a role in guiding antitumor T cells into the tumor bed. Alternatively, cognate antigen presentation by APCs or graft endothelial cells was sufficient for antigen-specific T cell migration in a heart allograft model [47], whereas the migration of non-specific bystander T cells remained G-protein signaling dependent [47]. Cxcr3 was shown to promote CD8 T cell migration in B16 melanoma [11, 48] and other s.c tumor models [49]. Since we observed a subtle delay in antitumor effects in Cxcr3−/− mice bearing s.c. tumors, our results suggest a potential role for Cxcr3 in the initial wave of T cell migration into the skin.
Cxcr3 promoted the differentiation of Klrg1 + PDA-specific T cells in the spleen of tumor-bearing mice, consistent with Cxcr3 promoting short-lived effector T cells in acute infection models [16,17,18,19]. Cxcr3 guides T cells to LN periphery to support effector T cell differentiation during infection [19]. During persistent T. gondii infection, Cxcr3 loss led to an expanded Ki67 + Klrg1- memory T cells defective in differentiating into Klrg1 + T cells [20]. Klrg1 can mark effector T cells that localize in the vasculature, including the red pulp of the spleen, which promotes enhanced capacity to clear systemic infections [50, 51]. Our results raise the possibility that this localization, in addition to cytotoxic functionality, may poise such T cells to rapidly respond to circulating metastatic cancer cells. In sum, our results support a role for Cxcr3 in the spleen to promote the differentiation of potent Gzmb + antitumor T cells that may aid in the control of tumor cell dissemination. As metastasis is a leading cause of cancer-related mortality, further investigation into how to generate and maintain such cells could inform superior T cell-based immunotherapies for cancer patient treatment.
Materials and methods
Animals
KrasLSLG12D/+;Trp53LSLR172H/+;p48Cre (KPC) mice previously speed-backcrossed to > 99.6% genetic similarity to C57BL/6 J mice were used [7]. 6- to 12-wk-old female and male C57BL/6 J mice were purchased from The Jackson Laboratory (000,664). C57BL/6 J backcrossed Cxcr3−/− mice were generously provided by Dr. Sara Hamilton (University of Minnesota). REX3 transgenic mice [35] were kindly provided by Dr. Andrew Luster (Massachusetts General Hospital) and obtained from Dr. Marc Jenkins (University of Minnesota). University of Minnesota Institutional Animal Care and Use Committee approved all animal studies.
Tumor cell lines
We previously described the KPC2 parental and the KPC2a cell line retrovirally transduced to express click-beetle red luciferase linked to eGFP (CB-eGFP) [8]. Tumor cells were cultured in Basic media: DMEM (Life Technologies) + 10% FBS (Life Technologies) + 2.5 mg/ml amphotericin B (Life Technologies) + 100 mg/ml penicillin/streptomycin (Life Technologies) + 2.5 mg dextrose (Fisher Chemical) at 37 C + 5% CO2. Media was sterile filtered and stored in the dark at 4 C. Cell lines used for experiments were maintained below passage 15. 0.25% trypsin–EDTA (Thermo Fisher) was used for serial passages.
Tumor cell implantation
For orthotopic tumor implantation, mice received slow-release buprenorphine injected subcutaneously prior to surgery for analgesia according to approved protocols. Mice were anesthetized using continuous flow of 2–5% isoflurane. Hair was removed using clippers followed by Nair. The abdomen was sterilized using a series of EtOH and Betadine washes. Once mice reached surgical plane anesthesia, a small incision was made in the abdomen followed by a small incision in the peritoneum to access the pancreas. 1 × 105 KPC2a cells in 20 µl of 60% Matrigel were injected into the pancreas using an insulin syringe (Covidien) as we described [8, 28]. Separate sets of sutures were used to close the peritoneum and skin (Ethicon). For subcutaneous (s.c.) tumor implantation, 50 µl containing 1 × 106 KPC2a cells in of 60% Matrigel (Discovery Labware) was injected above the hind flank. Tumor volume was measured using calipers.
In vivo antibody treatments
Seven days post orthotopic tumor implantation, a single 100 µg dose of agonistic αCD40 (FGK45, BioXcell) was administered alone or in combination with 200 µg αPD-L1 (10F.9G2, BioXcell) on days 7, 10, and 12 i.p. [28]. 200 µg of αCxcr3 (CXCR-173, BioXcell) was administered on days 6, 9 and 12.
Bioluminescent tumor imaging in vivo
Abdominal hair was removed with Nair. Mice were injected i.p. with 100 μg of D-Luciferin (Promega). Six–10 min after D-Luciferin injection, images were acquired after 0.5 s exposure time with a binning of 8 as we described [8, 28]. As luminescence saturation may occur, additional images with a binning of 2 and/or auto exposure setting were acquired. Tumor radiance was quantified in photons per second using IVIS 100 and Living Image software (Xenogen).
Preparation of mononuclear cells from tissues
Spleens were mechanically dissociated to single cells followed by red blood cell (RBC) lysis in 1 ml of Tris-ammonium chloride (ACK) lysis buffer (Life Technologies) for 2 min at rt. RBC lysis was quenched by addition of 9 ml of T cell media. Splenocytes were spun at 1400 rpm for 5 min and kept on ice. Tumors were mechanically digested to single-cell suspensions, collagenase digested and washed 2X to remove cell debris and pancreatic enzymes. Peripheral blood was collected in Eppendorf tubes containing 20 μL of 20 mM EDTA. PBMCs were spun for 8 min at 10,000 rpm at 4 °C and plasma was removed. RBCs were lysed with 1 ml of ACK Lysis buffer (GIBCO) for 8 min at rt. If incomplete lysis occurred, cells were spun down for a second time at 13,000 rpm for 30 s and the ACK lysis step was repeated.
Real-time PCR
KPC2 and KPC2a cells were cultured in vitro ± recombinant murine IFNγ (100 ng/mL, R&D Systems). After 24 h, RNA was extracted using the RNeasy Mini Kit (Qiagen) and concentration and purity of RNA was determined by Nanodrop. cDNA was generated using RT Buffer Mix and RT Enzyme Mix (Thermo Fisher). Real-time PCR was performed in triplicate on a BioRad CFX96 Touch Real-Time PCR Detection System by measuring SYBR Green fluorescence in triplicate for 40 cycles. ATP5b was the housekeeping gene. Relative quantification was determined using the delta-delta Ct method, also known as the 2–∆∆Ct method [52]. Fold gene expression was calculated using 2^-(∆∆Ct) where ∆Ct = Ct (gene of interest) – Ct (housekeeping gene) and ∆∆Ct = ∆Ct (Sample) – ∆Ct (control average).
Cell surface staining
Cells were stained in the presence of 1:500 Fc block (αCD16/32, Tonbo). CB101-109:H-2Db-BV421 tetramer was generated in laboratory [8, 28] and diluted 1:100 in FACs buffer (PBS + 2.5% FBS). For analysis of immune cells, fluorescently conjugated monoclonal antibodies were diluted 1:100 in FACs buffer and include CD45 (30F-11, Biolegend), CD3e (17A2, Biolegend), CD8 α(53–6.7, Tonbo), CD44 (IM7, BD), CD4 (RM4.5, Tonbo), Klrg1 (2F1, Biolegend), Cxcr3 (Cxcr3-173, Biolegend), PD-1 (J43, Invitrogen), Lag-3 (C9B7W, Biolegend), CD69 (H1.2F3, BD), CD62L (Mel14, Biolegend), CD11b (M1/70, Tonbo), CD19 (1D3, BD), NK1.1 (PK136, eBioscience), Ly6G (1A8, eBioscience), F4/80 (T45-2342, BD), CD11c (N418, BD), I-Ab (M5/114 15.2, Invitrogen), Ly6C (HK1.4, eBioscience) and Xcr1 (ZET, Biolegend). Cells were stained for 30 min at 4 °C in the dark and washed 2X with FACs buffer. Viability dyes BV510 or APCe780 (Tonbo, 1:500) were used to exclude dead cells and cell counting beads (Sigma-Aldrich) were added prior to analysis to calculate cell number per tissue gram. Cells were fixed in 2% PFA or fixation buffer (Tonbo) for 10–15 min at 4 °C in the dark prior to data acquisition. For detecting REX3 reporters, cells were not fixed and acquired immediately after staining. Cell analysis was performed using FlowJo software (version 10). Cells were acquired within 24 h using a Fortessa 1770 and data analyzed using FACS Diva software (BD).
Intracellular staining
The Foxp3 intracellular staining kit (Tonbo) was used for detecting intracellular proteins. Following cell surface staining, cells were washed 2X in FACs buffer, fixed for 30 min at 4 °C, washed 2X in permeabilization buffer and stained with Gzmb (NGZB, eBiosciences), Tcf1 (S33-966, BD), Foxp3 (FJK-16 s, eBioscience) and Helios (22F6, BD). Antibodies were diluted 1:100 in permeabilization buffer and cells were stained 1–2 h in the dark at 4 °C. Cells were washed 2X in permeabilization buffer and acquired within 24 h on a Fortessa 1770 flow cytometer following addition of cell counting beads (Sigma). Cell analysis was performed using FlowJo software (version 10).
Engineered T cell therapy
DMEM (Life Technologies) + 10% FBS (Life Technologies), 100 μg/ml penicillin/streptomycin (Life Technologies), 20 mM l-glutamine (Life Technologies), 1X NEAA (Life Technologies) and 50 μM 2-betamercaptoethanal (Sigma-Aldrich) were used to culture T cells in vitro. Mononuclear cells from spleens of P14 Thy1.1 + mice were stimulated in vitro with 1 mg/mL of αCD3 (145-2C11, BD) + 1 mg/mL αCD28 (37.51, BD) in 10 mL of T cell media containing human recombinant IL-2 (rIL2, 50 U/mL, Peprotech) in T25 flasks at 37 °C and 5% CO2 as described [7, 36]. On days 1 and 2 after stimulation, activated T cells were transduced with retroviral supernatant containing the 1045 TCR [7] by spinfection in 12-well plates containing polybrene (10 mg/mL) and rIL2 (50 U/mL) for 90 min at 1,000 × g at 32 °C. On day 7, T cells were restimulated with Msln406-414 (GQKMNAQAI) pulsed irradiated splenocytes and rIL2. Five days post the 2nd stim, T cells were injected i.p. into mice as we previously described [7, 36]. At the time of T cell transfer, recipients either received 100 µg i.p. of isotype or agonistic αCD40 (FGK45, BioXcell) as described [36]. For cell therapy experiments in Fig. 6, B6 mice were orthotopically implanted with 1 × 105 parental CB-negative KPC2 tumor cells [8]. Donor Thy1.1 + T cells expressing the high affinity Msln406-414:H-2Db-restricted TCR (clone 1045) were isolated from the spleens of 1045 TCR Trac knockin mice (Rollins et. al., in revision) for studies in Fig. 6. T cells were transferred 6 h post Cytoxan (180 mg/kg, Amneal Pharmaceuticals) on day 6 posttumor ± 200 μg anti-Cxcr3 (Cxcr3-173, BioXcell) i.p. on days 6, 9 and 12. Recipients received rhIL-2 104 U i.p. on day 6, 8, 10 and Msln406-414 pulsed irradiated splenocytes on day 6 to support donor T cell expansion similar to as described [7, 36].
Immunofluorescence
Tissues were embedded in OCT (Tissue-Tek) and stored at − 80 °C. 7 μm sections were cut using a Cryostat and fixed in acetone at − 20 °C for 10 min. Sections were rehydrated with PBS + 1% bovine serum albumin (BSA) and incubated for 1 h at room temperature (rt) with 1:200 αCD8α-PerCp-Cy5.5 (53–6.7, BD), 1:500 αThy1.1-BV510 (OX-7, Biolegend), 1:200 αF4/80-PE (BM8, Biolegend), 1:200 αpanCK-FITC (F3418, Sigma-Aldrich), 1:100 Cxcl9-EF660 (MIG-2F5.5, eBioscience) and 1:100 αCxcl10 (goat polyclonal, R&D Systems) diluted in PBS + 1% BSA. Slides were washed 3X in PBS + 1% BSA and incubated with anti-goat 1:500 AF647 (Jackson ImmunoResearch, for detecting αCXCL10) for 1 h at rt in the dark. Cxcl9 EF660 and Cxcl10 AF647 were detected in the APC channel. Tissues were washed 3X with PBS + 1% BSA, washed 3X with PBS, and mounted in DAPI Prolong Gold (Life Technologies). Images were acquired on a Leica DM6000 epifluorescent microscope at the University of Minnesota Center for Immunology using Imaris 9.1.0 (Bitplane). To determine CD8 T cell number, individual cells from n = 3 mice per group and a minimum of 3–10 fields per section were manually counted and recorded by an investigator blinded to the experimental conditions using Cell Counter in Fiji2.0. Cxcl9/10 staining intensity was measured by pixel intensity from 3 mice per group and 3–8 fields per section using Fiji2.0. In tissues from REX3 mice, Cxcl9 (PE) and Cxcl10 (BFP) reporters were detected ex vivo in 7 μm OCT-embedded sections. These sections were also stained with αCD8 (53–6.7, BD) and αB220 (RA3-6B2, Biolegend) diluted 1:100 in PBS + 1%BSA for 1 h in the dark, followed by washing 3X as above. These images were acquired on a Leica DM4 B microscope. Images were analyzed using Fiji2.0.
scRNAseq analysis
Filtered count matrices for 9 human samples from Elyada et al. study [30] were obtained after NIH approval at dbGaP (accession number phs001840.v1.p1) and were used as input data. Barcodes that were considered to represent noise and poor-quality cells were removed using the knee-inflection strategy implemented in DropletUtils package (version 1.10.3). For downstream analysis, Seurat package (version 3.1.0) was used. Genes expressed in fewer than 3 cells were filtered from the expression matrices as well as cells that expressed fewer than 200 genes. Finally, cells with a mitochondrial percentage more than the highest confidence interval for scaled mitochondrial content were filtered out. Each sample was normalized using the SCTransform function with mitochondrial content as a variable to regress out in a second non-regularized linear regression. Before integration, highly variable features across the samples were found by SelectIntegrationFeatures function with the number of genes equal to 2000, then the object was prepared for integration (PrepSCTIntegration function), the anchors were found (FindIntegrationAnchors function), and the samples were integrated into the whole object (IntegrateData function). Principal component analysis was applied for dimensionality reduction, and the first 20 principal components were used further to generate uniform manifold approximation and projection (UMAP) dimensionality reduction. The clustering procedure was performed by FindNeighbors and FindClusters (resolution 0.8). After that, cluster 7 enriched by low-quality cells was removed, and only 6 tumor samples (SRR9274536, SRR9274537, SRR9274538, SRR9274539, SRR9274542, SRR9274544) were preserved. Remaining cells were fully renormalized, reintegrated and re-clustered.
Statistical analysis
Statistical analyses were performed using GraphPad software (version 9.0). Mouse experiments include = 3–23 mice per group. Unpaired, two-tailed Student’s T test was used to compare two-group data. One-way ANOVA and Tukey posttest were used for comparing > 2-group data. Log-rank (Mantel–Cox) test was used to test for statistically significant differences in survival. Data are presented as mean ± standard error of the mean (SEM), and p < 0.05 was considered significant. *p < 0.05, **p < 0.005 and ***p < 0.0005.
References
Siegel RL, Miller KD, Jemal A (2020) Cancer statistics. CA Cancer J Clin 70(3):145–164
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM (2014) Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res [Internet] 74(11):2913–2921. https://doi.org/10.1158/0008-5472.CAN-14-0155
Ribas A, Hamid O, Daud A, Hodi FS, Wolchok JD, Kefford R et al (2016) Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. JAMA - J Am Med Assoc. 315(15):1600–1609
Lim WA, June CH (2017) The principles of engineering immune cells to treat cancer. Cell 168(4):724–740
Foley K, Kim V, Jaffee E, Zheng L (2016) Current progress in immunotherapy for pancreatic cancer. Cancer Lett 381(1):244–251
Stromnes IMIM, DelGiorno KEKE, Greenberg PDPD, Hingorani SRSR (2014) Stromal re-engineering to treat pancreas cancer. Carcinogenesis 35(7):1451–1460
Stromnes IM, Schmitt TM, Hulbert A, Brockenbrough JS, Nguyen HN, Cuevas C, et al. (2015) T cells engineered against a native antigen can surmount immunologic and physical barriers to treat pancreatic ductal adenocarcinoma. Cancer Cell [Internet]. 28(5):638–52. Available from: https://linkinghub.elsevier.com/retrieve/pii/S1535610815003773
Burrack AL, Spartz EJ, Raynor JF, Wang I, Olson M, Stromnes IM (2019) Combination PD-1 and PD-l1 blockade promotes durable neoantigen-specific t cell-mediated immunity in pancreatic ductal adenocarcinoma. Cell Rep [Internet]. 28(8):2140–2155.e6. Available from: https://linkinghub.elsevier.com/retrieve/pii/S2211124719309635
Groom JR, Luster AD (2011) CXCR3 ligands: redundant, collaborative and antagonistic functions. Immunol Cell Biol 89:207–215
Nagarsheth N, Wicha MS, Zou W (2017) Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat Rev Immunol 17:559–572
Mikucki ME, Fisher DT, Matsuzaki J, Skitzki JJ, Gaulin NB, Muhitch JB et al (2015) Non-redundant requirement for CXCR3 signalling during tumoricidal T-cell trafficking across tumour vascular checkpoints. Nat Commun 6:1–4
Dangaj D, Bruand M, Grimm AJ, Ronet C, Barras D, Duttagupta PA et al (2019) Cooperation between constitutive and inducible chemokines enables T Cell engraftment and immune attack in solid tumors. Cancer Cell 35(6):885–900
Chow MT, Ozga AJ, Servis RL, Frederick DT, Lo JA, Fisher DE et al (2019) Intratumoral activity of the CXCR3 chemokine system is required for the efficacy of anti-PD-1 therapy. Immunity 50(6):1498–1512
Han X, Wang Y, Sun J, Tan T, Cai X, Lin P et al (2019) Role of CXCR3 signaling in response to anti-PD-1 therapy. EBioMedicine 48:169–177
Qian L, Yu S, Yin C, Zhu B, Chen Z, Meng Z et al (2019) Plasma IFN-γ-inducible chemokines CXCL9 and CXCL10 correlate with survival and chemotherapeutic efficacy in advanced pancreatic ductal adenocarcinoma. Pancreatology 19(2):340–345
Kurachi M, Kurachi J, Suenaga F, Tsukui T, Abe J, Ueha S et al (2011) Chemokine receptor CXCR3 facilitates CD8+ T cell differentiation into short-lived effector cells leading to memory degeneration. J Exp Med 208(18):1605–1620
Hu JK, Kagari T, Clingan JM, Matloubian M (2011) Expression of chemokine receptor CXCR3 on T cells affects the balance between effector and memory CD8 T-cell generation. Proc Natl Acad Sci USA 108(21):E118–E127
Kohlmeier JE, Reiley WW, Perona-Wright G, Freeman ML, Yager EJ, Connor LM et al (2011) Inflammatory chemokine receptors regulate CD8+ T cell contraction and memory generation following infection. J Exp Med 208(18):1621–1634
Duckworth BC, Lafouresse F, Wimmer VC, Broomfield BJ, Dalit L, Alexandre YO et al (2021) Effector and stem-like memory cell fates are imprinted in distinct lymph node niches directed by CXCR3 ligands. Nat Immunol 22(4):434–448
Bangs DJ, Tsitsiklis A, Steier Z, Chan SW, Kaminski J, Streets A et al (2022) CXCR3 regulates stem and proliferative CD8+ T cells during chronic infection by promoting interactions with DCs in splenic bridging channels. Cell Rep 38:110266
Cannon A, Thompson CM, Carlo Maurer H, Atri P, Bhatia R, West S et al (2020) CXCR3 and cognate ligands are associated with immune cell alteration and aggressiveness of pancreatic ductal adenocarcinoma. Clin Cancer Res 26(22):6051–6063
Koch MA, Tucker-Heard G, Perdue NR, Killebrew JR, Urdahl KB, Campbell DJ (2009) The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat Immunol 10(6):595–602
Lunardi S, Jamieson NB, Lim SY, Griffiths KL, Carvalho-Gaspar M, Al-Assar O et al (2014) IP-10/CXCL10 induction in human pancreatic cancer stroma influences lymphocytes recruitment and correlates with poor survival. Oncotarget 5(22):11064
Hirth M, Gandla J, Höper C, Gaida MM, Agarwal N, Simonetti M et al (2020) CXCL10 and CCL21 promote migration of pancreatic cancer cells toward sensory neurons and neural remodeling in tumors in mice Associated With Pain in Patients. Gastroenterology 159(2):665–681
Burrack AL, Schmiechen ZC, Patterson MT, Miller EA, Spartz EJ, Rollins MR, Raynor JR, Mitchell Kaisho JT, Fife B, Stromnes I (2022) Distinct myeloid antigen presenting cells dictate differential fates of tumor specific CD8 T cells in pancreatic cancer. JCI Insights. 7(7):e151593. https://doi.org/10.1172/jci.insight.151593
Branchini BR, Southworth TL, Fontaine DM, Kohrt D, Welcome FS, Florentine CM et al (2017) Red-emitting chimeric firefly luciferase for in vivo imaging in low ATP cellular environments. Anal Biochem 534:36–39
Hall MP, Woodroofe CC, Wood MG, Que I, van’t Root M, Ridwan Y, et al. (2018) Click beetle luciferase mutant and near infrared naphthyl-luciferins for improved bioluminescence imaging. Nat Commun [Internet] 9(1):132. Available from: http://www.nature.com/articles/s41467-017-02542-9
Burrack AL, Rollins MR, Spartz EJ, Mesojednik TD, Schmiechen ZC, Raynor JF et al (2021) CD40 Agonist overcomes T cell exhaustion induced by chronic myeloid cell IL-27 production in a pancreatic cancer preclinical model. J Immunol [Internet] 206(6):1372–1384. https://doi.org/10.4049/jimmunol.2000765
Chu HH, Chan SW, Gosling JP, Blanchard N, Tsitsiklis A, Lythe G et al (2016) Continuous effector CD8+ T cell production in a controlled persistent infection is sustained by a proliferative intermediate population. Immunity 45(1):159–171
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA et al (2019) Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov 8:1102–1123
Cao W, Bover L (2010) Signaling and ligand interaction of ILT7: receptor-mediated regulatory mechanisms for plasmacytoid dendritic cells. Immunol Rev 234:163–176
Stromnes IM, Brockenbrough JS, Izeradjene K, Carlson MA, Cuevas C, Simmons RM et al (2014) Targeted depletion of an MDSC subset unmasks pancreatic ductal adenocarcinoma to adaptive immunity. Gut [Internet] 63(11):1769–1781. https://doi.org/10.1136/gutjnl-2013-306271
Sung JH, Zhang H, Ashley Moseman E, Alvarez D, Iannacone M, Henrickson SE et al (2012) Chemokine guidance of central memory T cells is critical for antiviral recall responses in lymph nodes. Cell 150(6):1249–1263
Il CH, Celis E (2009) Optimized peptide vaccines eliciting extensive CD8 T-cell responses with therapeutic antitumor effects. Cancer Res 69(23):9012–9019
Groom JR, Richmond J, Murooka TT, Sorensen EW, Sung JH, Bankert K et al (2012) CXCR3 chemokine receptor-ligand interactions in the lymph node optimize CD4+ T helper 1 cell differentiation. Immunity 37(6):1091–1103
Stromnes IM, Burrack AL, Hulbert A, Bonson P, Black C, Brockenbrough JS et al (2019) Differential effects of depleting versus programming tumor-associated macrophages on engineered T Cells in pancreatic ductal adenocarcinoma. Cancer Immunol Res [Internet] 7(6):977–989. https://doi.org/10.1158/2326-6066.CIR-18-0448
Stromnes IM, Hulbert A, Pierce RH, Greenberg PD, Hingorani SR (2017) T-cell Localization, activation, and clonal expansion in human pancreatic ductal adenocarcinoma. Cancer Immunol Res [Internet] 5(11):978–991. https://doi.org/10.1158/2326-6066.CIR-16-0322
Paley MA, Kroy DC, Odorizzi PM, Johnnidis JB, Dolfi DV, Barnett BE et al (2012) Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 80:1220–1225
Utzschneider DT, Charmoy M, Chennupati V, Pousse L, Ferreira DP, Calderon-Copete S et al (2016) T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45(2):415–427
Blank CU, Haining WN, Held W, Hogan PG, Kallies A, Lugli E et al (2019) Defining T cell exhaustion. Nat Rev Immunol 19:665–674
Wherry EJ, Ha SJ, Kaech SM, Haining WN, Sarkar S, Kalia V et al (2007) Molecular signature of CD8+ T cell Exhaustion during chronic viral infection. Immunity 27:670–684
Wirth TC, Xue HH, Rai D, Sabel JT, Bair T, Harty JT et al (2010) Repetitive antigen stimulation induces stepwise transcriptome diversification but preserves a core signature of memory CD8+ T cell differentiation. Immunity 33(1):128–140
Wherry EJ, Kurachi M (2015) Molecular and cellular insights into T cell exhaustion. Nat Rev Immunol 15:486–499
Noubade R, Majri-Morrison S, Tarbell KV (2019) Beyond CDC1: emerging roles of DC crosstalk in cancer immunity. Front Immunol 10:1014. https://doi.org/10.3389/fimmu.2019.01014
McGill J, Van Rooijen N, Legge KL (2010) IL-15 trans-presentation by pulmonary dendritic cells promotes effector CD8 T cell survival during influenza virus infection. J Exp Med 207(3):521–534
Böttcher JP, Reise SC (2018) The role of type 1 conventional dendritic cells in cancer immunity. Trends Cancer 4:784–792
Walch JM, Zeng Q, Li Q, Oberbarnscheidt MH, Hoffman RA, Williams AL et al (2013) Cognate antigen directs CD8+ T cell migration to vascularized transplants. J Clin Invest 123(6):2663–2671
Chheda ZS, Sharma RK, Jala VR, Luster AD, Haribabu B (2016) Chemoattractant receptors BLT1 and CXCR3 regulate antitumor immunity by facilitating CD8 + T cell migration into tumors. J Immunol 197(5):2016–2026
House IG, Savas P, Lai J, Chen AXY, Oliver AJ, Teo ZL et al (2020) Macrophage-derived CXCL9 and CXCL10 are required for antitumor immune responses following immune checkpoint blockade. Clin Cancer Res 26(2):487–504
Olson JA, McDonald-Hyman C, Jameson SC, Hamilton SE (2013) Effector-like CD8+ T cells in the memory population mediate potent protective immunity. Immunity 6:1250–1260
Renkema KR, Huggins MA, Borges da Silva H, Knutson TP, Henzler CM, Hamilton SE (2020) KLRG1 + Memory CD8 T cells combine properties of short-lived effectors and long-lived memory. J Immunol 205(4):1059–1069
Rao X, Huang X, Zhou Z, Lin X (2013) An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat Bioinforma Biomath 3(3):71–85
Acknowledgements
We acknowledge the University of Minnesota Flow Cytometry Resource for technical assistance and the University of Minnesota Research Animal Resource (RAR) staff for animal husbandry and veterinary services. We acknowledge the University of Minnesota Imaging Core for assistance with IVIS imaging and quantification. A.L.B was supported by a computational training award from the American Association of Immunologists. E.J.S. was funded through Carolyn L. Kuckein Student Research Fellowship and NIH T35 AI118620. M.R.R. is supported by National Institutes of Health (NIH) T32 AI 007313 and a Dennis Watson Fellowship (University of Minnesota). S.S. is supported by UL1 TR002494. M.F. and K.Z. were supported by Priority 2030 Federal Academic Leadership Program. I.M.S. is supported by NIH R01 CA249393, R01 CA255039 and P01CA254849, Department of Defense #PA200286, an American Association for Cancer Research (AACR) Pancreatic Cancer Action Network Career Development Award (17-20-25-STRO), an AACR Pancreatic Cancer Action Network Catalyst Award (19-35-STRO), American Cancer Society Institutional Research Grant (124166-IRG-58-001-55-IRG65), Randy Shaver Cancer and Community Fund and has pilot awards from the Masonic Cancer Center and Cancer Research Training Initiative (University of Minnesota Medical School).
Funding
NIH T35, AI118620, Carolyn L. Kuckein Student Research Fellowship, NIH T32,T32 AI 007313, Dennis Watson Fellowship, NIH, UL1 TR002494, Priority 2030 Federal Academic Leadership Program, NIH, NCI, R01 CA249393, Department of Defense, #PA200286, American Association for Cancer Research (AACR) Pancreatic Cancer Action Network Career Development Award, 17-20-25-STRO, AACR Pancreatic Cancer Action Network Catalyst Award, 19-35-STRO, American Cancer Society Institutional Research Grant, 124166-IRG-58-001-55-IRG65
Author information
Authors and Affiliations
Contributions
ALB, EJS and IMS conceptualized the study, performed experiments, analyzed data and wrote the manuscript. MRR, EAM, EC and IXW conducted experiments. EAM, EC, HN, SS, MF and KZ acquired and/or analyzed data. M.F.G. provided reagents and expertise. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burrack, A.L., Spartz, E.J., Rollins, M.R. et al. Cxcr3 constrains pancreatic cancer dissemination through instructing T cell fate. Cancer Immunol Immunother 72, 1461–1478 (2023). https://doi.org/10.1007/s00262-022-03338-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-022-03338-7