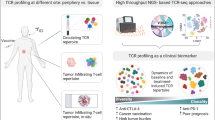
Abstract
Despite the conventional view that a truly random V(D)J recombination process should generate a highly diverse immune repertoire, emerging reports suggest that there is a certain bias toward the generation of shared/public immune receptor chains. These studies were performed in viral diseases where public T cell receptors (TCR) appear to confer better protective responses. Selective pressures generating common TCR clonotypes are currently not well understood, but it is believed that they confer a growth advantage. As very little is known about public TCR clonotypes in cancer, here we set out to determine the extent of shared TCR clonotypes in the intra-tumor microenvironments of virus- and non-virus-driven head and neck cancers using TCR sequencing. We report that tumor-infiltrating T cell clonotypes were indeed shared across individuals with the same cancer type, where the majority of shared sequences were specific to the cancer type (i.e., viral versus non-viral). These shared clonotypes were not particularly enriched in EBV-associated nasopharynx cancer but, in both cancers, exhibited distinct characteristics, namely shorter CDR3 lengths, restricted V- and J-gene usages, and also demonstrated convergent V(D)J recombination. Many of these shared TCRs were expressed in patients with a shared HLA background. Pattern recognition of CDR3 amino acid sequences revealed strong convergence to specific pattern motifs, and these motifs were uniquely found to each cancer type. This suggests that they may be enriched for specificity to common antigens found in the tumor microenvironment of different cancers. The identification of shared TCRs in infiltrating tumor T cells not only adds to our understanding of the tumor-adaptive immune recognition but could also serve as disease-specific biomarkers and guide the development of future immunotherapies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
The collective expression of different T cell receptors (TCR) in an individual, known as the TCR repertoire, is central to each person’s ability to recognize a vast range of pathogens and to initiate specific adaptive immune response. Each TCR consists of a heterodimer of two chains (either α + β or γ + δ), each generated through random rearrangement of germline variable (V), diversity (D) and joining (J) segments and the non-templated insertion and deletion of nucleotides at the V(D)J junctions. Although the theoretical estimate of the diversity of the TCR is more than 1 × 1015 [1], only about 106–107 unique TCRβ chains are expressed in approximately 1012 circulating T cells from healthy adults [2, 3]. T cells are activated and driven to clonally expand upon antigenic stimulation, with some formation of memory subsets that persist for long periods. Hence, the TCR repertoire not only reflects the current state of the adaptive immune system, but also provides an immunological footprint of its past history.
Despite the low expected likelihood of having two individuals that share the same TCR sequences, there are consistent observations that a small fraction of TCR chains, also known as public TCRs, are shared between different individuals, both in mice [4, 5] and humans [4, 6, 7]. This phenomenon is detectable in circulating blood of healthy adults [6, 7] and also encompasses public T cell responses to viruses [8, 9] and auto-immune diseases [10,11,12]. The selection of public TCRs appears to confer improved T cell survival in certain circumstances. For example, in human immunodeficiency virus (HIV), public TCRs are found in a population of HIV-responsive CD8 and CD4 T cells [9]. HIV-associated public TCRs are endowed with features of high avidity and cross-reactivity, both major determinants of antiviral efficacy. Similarly, the T cell response to other human viral diseases such as Epstein–Barr virus (EBV), cytomegalovirus (CMV) and influenza virus is peculiarly limited among individuals with the same MHC haplotypes [13]. There is clearly a need to fully understand the processes that bias toward the production of public TCRs, whose functional consequences may have important translational applications.
In the domain of tumor immunology, there were some early reports to indicate sharing of TCR clonotypes in tumors using earlier technologies [14,15,16], but a comprehensive analysis of TCR sharing across tumor types and examination of their characteristics has been limited. Undoubtedly, the identification of these common sequences would have a number of important consequences. First, these could be used to trace common immune histories across patients with the same cancer types in order to understand disease biology. Second, targeting universally shared TCR clonotypes could be an important approach in future TCR-based gene or cellular immunotherapy. Finally, disease-specific public TCRs could lead to the development of biomarkers specific to disease states.
In this study, we sought to determine the extent of shared TCR clonotypes in the intra-tumor microenvironment of two different head and neck cancer types: EBV-linked nasopharyngeal carcinoma (NPC) and non-virally linked head and neck squamous carcinomas (HNSCC). We hypothesized that public clonotypes may be more abundant in the virus-driven cancers like NPC compared to non-virus-driven cancers, as a result of selection for TCR clones consequent to persistent viral insults to the tumor environment.
Results
Targeted T cell receptor sequencing for the TCRβ chain was performed on RNA samples from 19 NPC patients and 10 HNSCC patients; the former cohort were all confirmed to be positive for EBV, while the latter were negative for EBV or HPV. The clinical characteristics of these patients are shown in Supplementary Table 1. The quality, filtering and analysis of the TCR sequencing data are described in Supplementary Table 2 and Methods. We obtained on average 330,578 productive TCR reads per sample (range 6716–1,150,255), after correcting for duplication and sequencing errors, and using only unique barcoded reads aligned against the TCRβ sequences from the ImMunoGeneTics (IMGT) database. Of these productive reads, an average of 32,599 unique CDR3 clonotypes per sample were obtained per sample (range 1446–164,463). TCR clonotypes were defined here as TCR sequences sharing the same CDR3 amino acid sequence and the same V- and J-gene region. Whilst we did not exhaustively capture the full TCR repertoire from these patients, this level of TCR sequencing depth can allow for detailed analysis of public clonotypes (Supplementary Figure 1).
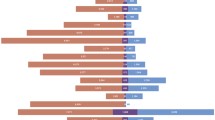
We next determined whether we could identify unique, tumor-specific TCRs within the microenvironment of each cancer type. We calculated the extent of clonal overlap between any two samples using Jaccard similarity index: the majority of each TCR repertoire was unique to its sample (average Jaccard index: 0.0007). However, a subset of TCRs were present in multiple samples (Fig. 1a). The extent of TCR sharing in each cancer type was determined by examining the number of private and shared CDR3s found across different patients within that type. There were 5878 and 1060 TCRs found in at least two NPC and two HNSCC patients, respectively (Fig. 1b). Subsequently, in our study, we defined as shared TCR clonotypes those clonotypes with the same CDR3 amino acid sequence, V-gene and J-gene, and present in at least 30% of the patients in each cancer cohort. Each sample had to have at least two uniquely barcoded TCR reads for each TCR clonotype to be confidently deemed present. There were a total of 49 shared TCR clonotypes found in our NPC cohort and 67 shared TCR clonotypes in the HNSCC samples. Out of these, only 3 TCRs were found shared in both cancer types. The baseline frequency of shared TCRs was calculated from the number of shared TCRs each patient expressed as a percentage of the total number of unique clonotypes found in each patient. We found that there was no statistical difference between the two cancer types with an average of 0.002% for virally driven NPC compared to 0.004% for HNSCC (p = 0.16, Fig. 1c). The median baseline frequency of shared TCRs that were found common in both cancers was 0.001% across all samples. These results suggest that contrary to our hypothesis, the presence of persistent viral stimulation does not contribute to the selection of more public TCR clonotypes.
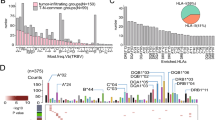
Sharing of TCR clonotypes is common across patients in two different cancers. a Jaccard similarity matrix of TCR clonal overlap between samples. Boxes shaded in gray are similarities between self. b Number of shared TCR clonotypes found between number of patients. x-axis: Log2(counts + 1) of shared TCRs found in any number of patients (y-axis). c Baseline frequency of shared TCR clonotypes (found in each cancer type or in both cancer types) over total number of unique clonotypes per sample. d Frequency of shared TCR clonotypes per patient. Colors represent the number of patients that share the same TCR clonotype within the cancer cohort. Shapes represent TCR clonotypes found uniquely within a single cancer type, in both cancers or found in an independent cohort of healthy individuals. e Proportion of shared TCRs that are found in healthy donors, single cancer type or both cancer types. f Frequency of high-frequency clonally expanded TCRs per patient. Legend same as 1D. g Proportion of private, less shared, shared TCR clonotypes in the high-frequency clonally expanded TCRs. h CDR3 length distribution of private (green), less shared (red) and shared (blue) TCRs. i V- and J-gene usage heatmap of shared TCR clonotypes. Fold-change in V + J use between shared and private TCRs. j Number of nucleotides coding for each shared TCR clonotype. Each dot represents a single shared TCR. X-axis: number of patients each shared TCR found in. y-axis: number of nucleotides found in each cohort that codes for same shared TCR
To determine whether these shared TCR clonotypes could result from recent exposures to antigenic stimulation, the clonal frequency of shared TCRs in individual samples was plotted to determine their clonal sizes (Fig. 1d). Of the total number of different shared TCRs each patient expressed from both cancer cohorts, 83.7% of shared TCRs were small clones with frequency of less than 0.1% (Supplementary Table 3). However, there were 10 HNSCC TCR clones and 12 NPC TCR clones that were dominantly expanded in a few samples, with clonal frequencies of more than 5%. We next asked if these shared CDR3s were cancer-specific and searched for these clonotypes in an independent, large cohort of TCR sequences obtained from the blood of normal healthy adults [6, 17] and found that only 22 of all the shared TCR clonotypes in both cancers (n = 116) were found in the blood of healthy donors. Remarkably, the majority of the shared TCRs could only be found within each cancer type (69.4% NPC-only and 80.6% HNSCC-only, Fig. 1e, Supplementary Table 3) and 50 of these shared TCRs were significantly enriched in each cancer group (p value < 0.05) (Supplementary Table 4), suggesting that these tumor-specific shared TCRs may be driven by common antigens found uniquely within each cancer type. Conversely, high-frequency, clonally expanded TCRs (Supplementary Figure 2A, 2B, prevalence more than 1% clonal frequency in each tumor), tend to be private and less shared (84.7% in NPC and 65.8% in HNSCC) (Fig. 1f, g), suggesting that the high-frequency TCR clones are predominantly tumor- and individual-specific and may represent a more recent patient-tumor-specific antigenic event.
Certain TCR traits such as CDR3 length, V- and J-gene usages are distinctive in some infectious or auto-immune diseases. For example, type 1 diabetic patients were found to have shorter CDR3 regions [18], and restrictions in V- and J-gene usages were found common in public TCR clonotypes to viral antigens [19]. We therefore queried if the public TCR clonotypes identified here presented similar features. For this and subsequent analyses, we removed shared TCR clonotypes that were also found in the cohort of healthy donors and only analyzed cancer patient-specific clonotypes. Shared TCRs in both cancer cohorts had shorter CDR3 sequence lengths compared to private TCRs (13 versus 15 amino acids, respectively) (Fig. 1h). Moreover, there were significantly different V- and J-gene usages between the two groups. In particular, TRBV13-TRBJ2-6 was used 21 times more commonly in shared HNSCC TCRs vs private HNSCC TCRs, and TRBV13-TRBJ1-1/-4/-5 were used 7 times more commonly in shared NPC TCRs vs private NPC TCRs (Fig. 1i). Both cancers also exhibited preferential V- and J-gene usages within each cancer type. Lastly, we asked whether shared TCRs display a higher level of convergent recombination, as seen in other reports [5]. Increased sharing was associated with an increasing number of nucleotides that encode the same CDR3 sequence (Fig. 1j). This was seen in both cancers, demonstrating a high level of convergent recombination driving the selection of these public TCR clonotypes.
Given the obvious association between antigen presentation and HLA subtypes, we next set out to examine the relationship (if any) between HLA types and the public intra-tumoral TCR clonotypes identified here. HLA typing for each patient is shown in Supplementary Table 5. To determine the association, we first calculate the percentage of patients with the same HLA alleles over the total number of patients expressing that shared TCR, for each public TCR clonotype at each HLA allele. We then counted the number of TCRs that were shared and expressed in patients with different percentages of HLA sharing (Fig. 2a). For example, there were 24 TCR clonotypes in HNSCC patients that had at least 75% patients with the same HLA-DRB1*12:02 background (Fig. 2b). Notably, there were a total of 36 public TCR clonotypes that were shared in more than 75% of HNSCC patients with the same HLA background, while there was only 1 such public TCR clonotype in the NPC cohort (Supplementary Table 6a, 6b).
Public TCR clonotypes converge on motif signatures that are unique to cancers. a Number of shared TCRs in patients expressing same HLA alleles. X-axis: percentage of patients with same HLA = Patients with same HLA type and expressing same shared TCR/total number of patients expressing same shared TCR. Y-axis: number of shared TCRs. Color represents cancer cohort. b Inset diagram of shared TCRs that are shared in more than 75% of patients with the same HLA-types. c Representative convergent motifs found in shared TCR clonotypes that are specific to each cancer type. Black box: HNN Red box: NPC. Blue number on top left of box reveals total number of patients who have convergent TCRs. Red number on bottom right of box shows total number of different TCRs that contain that motif. d Heatmap of motif convergence to TCR sequences from public databases. Y-axis is individual motif patterns. X-axis is source of database. Red denotes convergence, while blue denotes non-convergence
Much of the data thus far support the notion that intra-tumor shared TCR clonotypes converge on shared tumor-specific antigens. To test this hypothesis further, we extended the actual amino sequences to motif signatures for each of the TCR clonotypes using GLIPH2 (grouping of lymphocyte interaction by paratope hotspots version 2) [20]. Interestingly, we identified 8 and 6 shared motifs in HNSCC and NPC clonotypes, respectively, all of which were tumor-type-specific (representative motifs shown in Fig. 2c; detailed motif search shown in Supplementary Table 7a, 7b).
Finally, the intra-tumoral public TCR clonotypes identified here were compared with TCR sequences against known antigens. To do this, we obtained TCRβ sequences from public databases, VDJDB [21] and McPAS [22], which contain TCRs associated with different viruses, auto-immune diseases and tumors. We found that only 6 TCR clonotypes had the exact CDR3, V-, J-gene and HLA match to those in the databases, and these were associated with common viruses like CMV, EBV and influenza (Supplementary Table 8), while one had a match to an antigen epitope from colorectal cancer. We next searched for common motifs between the shared TCR clonotypes with those from the public databases and found 92 and 83 common motifs in the HNSCC and NPC cohorts that converged with TCRβ sequences associated predominantly with viruses like CMV, EBV, HCV, HIV-1 and Influenza (Fig. 2d, Supplementary Table 9a, 9b). The heatmap shows that there is a NPC-specific cluster (Clusters 2a and 2b) that overlaps with the part of the viral cohort, while a separate and distinct HNSCC cluster (Clusters 3a and 3b) demonstrates another group of motifs, which overlaps a different set of the viral cohort. Interestingly, these also overlap with common motifs found in the general tumor neoantigens and highly antigenic melanoma cohorts, but not with specific cancer types like leukemia and lymphoma. Separately, the identification of common motifs across TCR sequences from different viruses, self-antigens and tumor (Cluster 1) may support future work on understanding the roles of shared TCR clonotypes in the concept of TCR cross-reactivity.
Discussion
In this study in two cancer types with different etiologies, we found that each tumor-type harbors a set of cancer-specific public TCR clonotypes that contain distinct features from private TCRs. Previous studies have identified the existence of shared TCR clones in the intra-tumoral microenvironment [14,15,16], yet the characteristics of these remain unclear. We show that TCR clonotypes linked to the same tumor microenvironment have distinctive characteristics: shorter CDR3 lengths and restricted V- and J-gene usages, display convergent recombination and demonstrate a high level of converging motif signatures. These are surprisingly overlapping with previously published studies on public TCRs observed from circulating blood of healthy humans [6, 23, 24], infants [25] and mice [5, 26] and suggest a common underlying mechanism and function for public TCR convergence, yet the motifs and sequences identified here also show cancer-type specificity.
The phenomenon of immune convergence is not only seen in the T cell-mediated response, but also frequently extends to B cells. These are well-described in infectious diseases, auto-immune diseases but less in cancer. For example, converging TCR sequences were identified in healthy donors exposed to cytomegalovirus (CMV), Epstein–Barr virus (EBV), mycobacterium tuberculosis (TB), influenza and the yellow fever virus YV-17D vaccine [27,28,29,30]. In HIV, TCR repertoire analysis revealed that rare patients who controlled their HIV well showed a highly skewed TCR repertoire that was characterized by a predominance of TRAV24 and TRBV2 variable genes, shared CDR3 motifs, and a high frequency of public clonotypes [9]. The most prevalent public clonotypes generated TCRs with high binding affinities and are associated with superior functions that control HIV well. Recently, analysis of the B-cell receptor repertoire in convalescent blood of recovered COVID-19 patients revealed expanded clones of receptor-binding domain (RBD)-specific memory B cells (toward SARS-CoV-2 Spike protein (S)) expressing closely related antibodies in different individuals [31]. There was high neutralization efficacy of these antibodies to three distinct epitopes on RBD. A separate study also showed a high preference for the IGHV3-53 gene as the most frequently used IGHV gene for targeting the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein [32]. Several mechanisms including structural bias or convergent recombination have been proposed to explain the public immune responses. Regardless of the evolutionary driver, these studies consistently signify a selective advantage for the convergence of the T or B cell repertoire across individuals, one that is geared toward a more effective immune response. Future studies investigating public TCRs shared across cancers may provide insights into the clonotypes that may mediate desirable immune responses toward malignancy.
Our data also show many of the motifs we identified were common between public tumor TCR clonotypes and those with viral and neoantigen-associated TCR sequences from public databases. Combined with observations that shorter CDR3s were also found to be highly enriched in antigen-experienced memory T cells [24, 33], we postulate that the public TCR clonotypes observed in the tumor microenvironment could be memory T cells that were either selected for after past encounters with viruses or tumor neoantigen within the tumor environment, or are virus-specific memory T cells extending their surveillance in the tumors. Nevertheless, this could present a window of opportunity to re-activate known antiviral T cells in the tumor and a unique therapeutic approach for cancer immunotherapy. Rosato et al. have concluded a proof-of-concept study where they reactivated surveillant antiviral T cells by injecting adjuvant-free non-replicating viral peptides into tumors explanted in mice [34]. They showed that viral peptide treatment mimics a viral reinfection event to memory CD8 + T cells and arrest the growth of checkpoint blockade-resistant and poorly immunogenic tumors. Similarly, delineating the antigen repertoire recognized by intra-tumoral bystander T cells through TCR sequencing could pave the way to re-direct these to target the tumor instead.
Interestingly, majority of the shared TCR clonotypes in each cancer was specific to the cancer type and converged to similar motifs that were unique to each cancer. This suggests a shared immune background specific to different cancers and may reveal an antigenic footprint of common etiology unique to each. In addition, this presents a unique circumstance to identify cancer-specific TCR sequences, which can be further developed into TCR-based diagnostics that can be used to track and monitor diseases. Viral-specific TCR sequences have been discovered [17, 35], and future work can be extended into cancer studies.
We acknowledge that future studies can examine the TCRα chain sequence pairing to fully understand the antigen-binding specificity of each shared TCR. Public TCRα sequences have also been found at high frequencies in multiple individuals [36, 37]. However, only single-cell sequencing is able to fully recapitulate the extent of public paired TCRαβ chain, which is an expensive endeavor. In addition, the unsaturated sequencing depth achieved in this study limits the discovery of more low-frequency shared TCR clonotypes and the baseline frequency of shared TCRs reported here could be an under-representation of the true extent of TCR sharing in tumors. In spite of these limitations, we present this study to first highlight the extent of public TCR sharing in the tumor microenvironment and their distinct characteristics, to show that selective pressures act in the tumor to drive the TCR repertoire to convergent signatures, which may be beneficial for future analysis and development of diagnostic and therapeutic applications.
Conclusions
In this study, we discovered that TCRs are shared in the tumor microenvironment across multiple patients within two different head and neck cancer subtypes: EBV-linked nasopharyngeal carcinoma (NPC) and non-virally linked head and neck squamous carcinomas (HNSCC). These shared TCRs consistently display features of shorter CDR3 lengths, restricted V- and J-gene usages and demonstrate convergent recombination. They were also expressed in patients with a common HLA background. Most strikingly, these shared tumor TCRs were unique to each cancer-type and revealed specific cancer-type motif signatures. This study provides a useful resource for the future development of TCR-based cancer diagnostics or therapeutics.
Materials and methods
Patient samples
Tumor tissues were obtained from patients with head and neck squamous cancers (HNSCC) or nasopharyngeal cancers (NPC) undergoing resection surgery at National Cancer Centre Singapore (NCCS) after obtaining informed consent. All protocols were reviewed and approved by the Institutional Review Board (IRB) at NCCS. Samples were collected from 10 HNN patients and 19 NPC patients spanning stage II–IV disease. Clinical information of all patients is annotated in Supplementary Table 1.
TCR sequencing
RNA was extracted from tissue biopsies of all patient samples using Qiagen AllPrep DNA/RNA Mini Kit (Qiagen). The TCR libraries were prepared according to previous published method [38]. MiSeq libraries were prepared using Illumina protocols and sequenced using 300-bp paired-end MiSeq kits (Illumina).
TCR repertoire analysis
Raw MiSeq forward and reverse reads were merged using published Paired-End reAd mergeR (PEAR) tool. Universal barcoded regions were identified in each read using the following sequence: TNNNNTNNNNTNNNNT. Reads with identical universal barcode regions were condensed into a single read, and primers and constant regions were trimmed from all reads. Sequences were annotated using the reference TRBV, D- and J-genes from the IMGT database with IMGT/HighV-QUEST tool [39]. Non-TCR reads and non-productive TCR rearrangements were removed for further analysis.
Clonal overlap analysis
Clonal overlap analysis was performed by calculating the Jaccard similarity index between two samples, where a minimum of two unique barcoded reads must be present to be determined as present. The Jaccard index is calculated as:
where J is the Jaccard similarity index, X, Y = number of TCR clonotypes present in dataset X, Y and \(\left| {X \cap Y} \right|\) represents the number of TCR clonotypes found in both X, Y datasets. Dendrogram clustering was performed using hclust method in R, with ‘complete’ linkage method.
HLA typing
HLA genotypes of all NPC samples and HNSCC samples, except for HNN220 and HNN228, were obtained from available aligned reads from RNAseq data. ArcasHLA tool [40] was then used to extract and define HLA genotypes, where IMGT-HLA database version 3.39 was used as the reference database. Whole exome sequencing data (WES) but not RNAseq data were available for HNN220 and HNN228 and were used to extract HLA reads using HLA-HD [REF]. There were no available WES/RNAseq/HLA data for NCC010, NCC014, NCC022 and NCC028. Only HLA-A, HLA-B, HLA-C, HLA-DPA1, HLA-DPB1, HLA-DQA1, HLA-DQB1 and HLA-DRB1 were analyzed in this study.
Motifs searching
To look for common motif signatures, we used the published method, Grouping of Lymphocyte interactions by paratope hotspots version 2-Gliph2 [20], through the web service: http://50.255.35.37:8080. The CD4/8 reference option was used, together with all other default options.
Curation of public databases
TCRβ sequences were extracted from two public databases, VDJDB [21] and McPAS [22]. We selected all sequences associated with human species and specific to viruses, allergy and cancer.
Statistical tests
All statistical analyses were performed using R version 3.6.3 and associated packages. Kolmogorov–Smirnov test was performed to determine the significance of the distribution of CDR3 lengths between the private, less shared and shared TCRs. Wilcoxon test was used to compare statistical significance between two groups. Rarefaction analysis of estimated species richness was performed using the R package, rtk [41]. Fisher’s exact test was used to examine the association of specific shared TCRs within cancer groups.
Availability of data and materials
Sequence data have been deposited at the European Genome-Phenome Archive (EGA), which is hosted by the EBI and the CRG, under accession study number EGAS00001005480 (to be released upon acceptance). Further information about EGA can be found on https://ega-archive.org and “The European Genome-Phenome Archive of human data consented for biomedical research” (http://www.nature.com/ng/journal/v47/n7/full/ng.3312.html). Custom scripts to run the analyses described in the manuscript are available at https://github.com/yhfteng/Shared-TCRs-in-cancers.
Change history
22 October 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00262-021-03086-0
References
Davis MM, Bjorkman PJ (1988) T-cell antigen receptor genes and T-cell recognition. Nature 334:395–402
Robins HS, Campregher PV, Srivastava SK, Wacher A, Turtle CJ, Kahsai O, Riddell SR, Warren EH, Carlson CS (2009) Comprehensive assessment of T-cell receptor beta-chain diversity in alphabeta T cells. Blood 114:4099–4107
Arstila TP, Casrouge A, Baron V, Even J, Kanellopoulos J, Kourilsky P (1999) A direct estimate of the human alphabeta T cell receptor diversity. Science 286:958–961
Madi A, Poran A, Shifrut E, Reich-Zeliger S, Greenstein E, Zaretsky I, Arnon T, Laethem FV, Singer A, Lu J et al (2017) T cell receptor repertoires of mice and humans are clustered in similarity networks around conserved public CDR3 sequences. Elife 6:e22057
Madi A, Shifrut E, Reich-Zeliger S, Gal H, Best K, Ndifon W, Chain B, Cohen IR, Friedman N (2014) T-cell receptor repertoires share a restricted set of public and abundant CDR3 sequences that are associated with self-related immunity. Genome Res 24:1603–1612
Chu ND, Bi HS, Emerson RO, Sherwood AM, Birnbaum ME, Robins HS, Alm EJ (2019) Longitudinal immunosequencing in healthy people reveals persistent T cell receptors rich in highly public receptors. BMC Immunol 20:19
Soto C, Bombardi RG, Kozhevnikov M, Sinkovits RS, Chen EC, Branchizio A, Kose N, Day SB, Pilkinton M, Gujral M et al (2020) High frequency of shared clonotypes in human T cell receptor repertoires. Cell Rep 32:107882
Price DA, Asher TE, Wilson NA, Nason MC, Brenchley JM, Metzler IS, Venturi V, Gostick E, Chattopadhyay PK, Roederer M et al (2009) Public clonotype usage identifies protective Gag-specific CD8+ T cell responses in SIV infection. J Exp Med 206:923–936
Benati D, Galperin M, Lambotte O, Gras S, Lim A, Mukhopadhyay M, Nouel A, Campbell KA, Lemercier B, Claireaux M et al (2016) Public T cell receptors confer high-avidity CD4 responses to HIV controllers. J Clin Investig 126:2093–2108
Marrero I, Aguilera C, Hamm DE, Quinn A, Kumar V (2016) High-throughput sequencing reveals restricted TCR Vbeta usage and public TCRbeta clonotypes among pancreatic lymph node memory CD4(+) T cells and their involvement in autoimmune diabetes. Mol Immunol 74:82–95
Zhao Y, Nguyen P, Ma J, Wu T, Jones LL, Pei D, Cheng C, Geiger TL (2016) Preferential use of public TCR during autoimmune encephalomyelitis. J Immunol 196:4905–4914
Zhao Y, Nguyen P, Vogel P, Li B, Jones LL, Geiger TL (2016) Autoimmune susceptibility imposed by public TCRbeta chains. Sci Rep 6:37543
Miles JJ, Douek DC, Price DA (2011) Bias in the alphabeta T-cell repertoire: implications for disease pathogenesis and vaccination. Immunol Cell Biol 89:375–387
Chung YL, Wu ML (2018) Spatiotemporal homogeneity and distinctness of the T-cell receptor beta-chain repertoires in Epstein-Barr virus-associated primary and metastatic nasopharyngeal carcinomas. Int J Cancer 143:610–620
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA et al (2018) The immune landscape of cancer. Immunity 48:812-830-e814
Munson DJ, Egelston CA, Chiotti KE, Parra ZE, Bruno TC, Moore BL, Nakano TA, Simons DL, Jimenez G, Yim JH et al (2016) Identification of shared TCR sequences from T cells in human breast cancer using emulsion RT-PCR. Proc Natl Acad Sci USA 113:8272–8277
Emerson RO, DeWitt WS, Vignali M, Gravley J, Hu JK, Osborne EJ, Desmarais C, Klinger M, Carlson CS, Hansen JA et al (2017) Immunosequencing identifies signatures of cytomegalovirus exposure history and HLA-mediated effects on the T cell repertoire. Nat Genet 49:659–665
Gomez-Tourino I, Kamra Y, Baptista R, Lorenc A, Peakman M (2017) T cell receptor beta-chains display abnormal shortening and repertoire sharing in type 1 diabetes. Nat Commun 8:1792
Chen G, Yang X, Ko A, Sun X, Gao M, Zhang Y, Shi A, Mariuzza RA, Weng NP (2017) Sequence and structural analyses reveal distinct and highly diverse human CD8(+) TCR repertoires to immunodominant viral antigens. Cell Rep 19:569–583
Huang H, Wang C, Rubelt F, Scriba TJ, Davis MM (2020) Analyzing the Mycobacterium tuberculosis immune response by T-cell receptor clustering with GLIPH2 and genome-wide antigen screening. Nat Biotechnol 38:1194–1202
Bagaev DV, Vroomans RMA, Samir J, Stervbo U, Rius C, Dolton G, Greenshields-Watson A, Attaf M, Egorov ES, Zvyagin IV et al (2020) VDJdb in 2019: database extension, new analysis infrastructure and a T-cell receptor motif compendium. Nucleic Acids Res 48:D1057–D1062
Tickotsky N, Sagiv T, Prilusky J, Shifrut E, Friedman N (2017) McPAS-TCR: a manually curated catalogue of pathology-associated T cell receptor sequences. Bioinformatics 33:2924–2929
Hou X, Wang M, Lu C, Xie Q, Cui G, Chen J, Du Y, Dai Y, Diao H (2016) Analysis of the repertoire features of TCR beta chain CDR3 in human by high-throughput sequencing. Cell Physiol Biochem 39:651–667
Hou X, Zeng P, Zhang X, Chen J, Liang Y, Yang J, Yang Y, Liu X, Diao H (2019) Shorter TCR beta-chains are highly enriched during thymic selection and antigen-driven selection. Front Immunol 10:299
Carey AJ, Hope JL, Mueller YM, Fike AJ, Kumova OK, van Zessen DBH, Steegers EAP, van der Burg M, Katsikis PD (1859) Public clonotypes and convergent recombination characterize the naive CD8(+) T-cell receptor repertoire of extremely preterm neonates. Front Immunol 2017:8
Venturi V, Kedzierska K, Price DA, Doherty PC, Douek DC, Turner SJ, Davenport MP (2006) Sharing of T cell receptors in antigen-specific responses is driven by convergent recombination. Proc Natl Acad Sci USA 103:18691–18696
Pogorelyy MV, Minervina AA, Touzel MP, Sycheva AL, Komech EA, Kovalenko EI, Karganova GG, Egorov ES, Komkov AY, Chudakov DM et al (2018) Precise tracking of vaccine-responding T cell clones reveals convergent and personalized response in identical twins. Proc Natl Acad Sci USA 115:12704–12709
Glanville J, Huang H, Nau A, Hatton O, Wagar LE, Rubelt F, Ji X, Han A, Krams SM, Pettus C et al (2017) Identifying specificity groups in the T cell receptor repertoire. Nature 547:94–98
Dash P, Fiore-Gartland AJ, Hertz T, Wang GC, Sharma S, Souquette A, Crawford JC, Clemens EB, Nguyen THO, Kedzierska K et al (2017) Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547:89–93
Fink K (2019) Can we improve vaccine efficacy by targeting T and B cell repertoire convergence? Front Immunol 10:110
Robbiani DF, Gaebler C, Muecksch F, Lorenzi JCC, Wang Z, Cho A, Agudelo M, Barnes CO, Gazumyan A, Finkin S et al (2020) Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature 584:437–442
Yuan M, Liu H, Wu NC, Lee CD, Zhu X, Zhao F, Huang D, Yu W, Hua Y, Tien H et al (2020) Structural basis of a shared antibody response to SARS-CoV-2. Science 369:1119–1123
McHeyzer-Williams MG, Davis MM (1995) Antigen-specific development of primary and memory T cells in vivo. Science 268:106–111
Rosato PC, Wijeyesinghe S, Stolley JM, Nelson CE, Davis RL, Manlove LS, Pennell CA, Blazar BR, Chen CC, Geller MA et al (2019) Virus-specific memory T cells populate tumors and can be repurposed for tumor immunotherapy. Nat Commun 10:567
Wolf K, Hether T, Gilchuk P, Kumar A, Rajeh A, Schiebout C, Maybruck J, Buller RM, Ahn TH, Joyce S, DiPaolo RJ (2018) Identifying and tracking low-frequency virus-specific TCR clonotypes using high-throughput sequencing. Cell Rep 25:2369-2378-e2364
Hamrouni A, Aublin A, Guillaume P, Maryanski JL (2003) T cell receptor gene rearrangement lineage analysis reveals clues for the origin of highly restricted antigen-specific repertoires. J Exp Med 197:601–614
Tynan FE, Borg NA, Miles JJ, Beddoe T, El-Hassen D, Silins SL, van Zuylen WJ, Purcell AW, Kjer-Nielsen L, McCluskey J et al (2005) High resolution structures of highly bulged viral epitopes bound to major histocompatibility complex class I. Implications for T-cell receptor engagement and T-cell immunodominance. J Biol Chem 280:23900–23909
De Mattos-Arruda L, Sammut SJ, Ross EM, Bashford-Rogers R, Greenstein E, Markus H, Morganella S, Teng Y, Maruvka Y, Pereira B et al (2019) The genomic and immune landscapes of lethal metastatic breast cancer. Cell Rep 27:2690-2708-e2610
Li S, Lefranc MP, Miles JJ, Alamyar E, Giudicelli V, Duroux P, Freeman JD, Corbin VD, Scheerlinck JP, Frohman MA et al (2013) IMGT/HighV QUEST paradigm for T cell receptor IMGT clonotype diversity and next generation repertoire immunoprofiling. Nat Commun 4:2333
Orenbuch R, Filip I, Comito D, Shaman J, Pe’er I, Rabadan R (2020) arcasHLA: high-resolution HLA typing from RNAseq. Bioinformatics 36:33–40
Saary P, Forslund K, Bork P, Hildebrand F (2017) RTK: efficient rarefaction analysis of large datasets. Bioinformatics 33:2594–2595
Acknowledgements
The computational work for this article was fully performed on resources of the National Supercomputing Centre, Singapore (https://www.nscc.sg). We also want to thank staff from Laboratory of Cell Therapy and Cancer Vaccine, National Cancer Centre Singapore for facilitating our use of their equipment, Agilent Bioanalyzer. Many thanks to Chua Khi Pin, Sim Adelene, Wang Haitao for their technical assistance and valuable input.
Funding
This work was supported by National Medical Research Council (NMRC) Clinician Scientist Award—Senior Investigator (CSA-SI) MOH-000325 to N.G.I, NMRC CS-IRG MOH-000375 and NMRC/CSA-INV/0025/2017 to D.W-T.L, and National Cancer Centre Early Career Research Fund NCCRF-YR2019-JUL-ECCRS1 to Y.H-F.T.
Author information
Authors and Affiliations
Contributions
YHT and NGI conceived and designed the studies. DWL and NGI recruited patient samples. HSQ and LS prepared the samples. YHT performed the TCR sequencing. JMLD, AM, RJMBR and GSV led the TCR sequencing analysis pipeline. YHT and NGI analyzed the data. DWL, MLC, DST and NGI supervised the study. YHT and NGI prepared the manuscript with input from co-authors. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Accession study number was incorrect in the section “Availability of data and materials”.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Teng, Y.H.F., Quah, H.S., Suteja, L. et al. Analysis of T cell receptor clonotypes in tumor microenvironment identifies shared cancer-type-specific signatures. Cancer Immunol Immunother 71, 989–998 (2022). https://doi.org/10.1007/s00262-021-03047-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-021-03047-7