Abstract
Purpose
To understand the tumor immune microenvironment precisely, it is important to secure the quantified data of tumor-infiltrating immune cells, since the immune cells are true working unit. We analyzed unit immune cell number per unit volume of core tumor tissue of high-grade gliomas (HGG) to correlate their immune microenvironment characteristics with clinical prognosis and radiomic signatures.
Methods
The number of tumor-infiltrating immune cells from 64 HGG core tissue were analyzed using flow cytometry and standardized. After sorting out patient groups according to diverse immune characteristics, the groups were tested if they have any clinical prognostic relevance and specific radiomic signature relationships. Sparse partial least square with discriminant analysis using multimodal magnetic resonance images was employed for all radiomic classifications.
Results
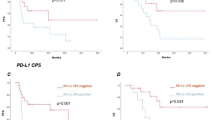
The median number of CD45 + cells per one gram of HGG core tissue counted 865,770 cells which was equivalent to 8.0% of total cells including tumor cells. There was heterogeneity in the distribution of immune cell subpopulations among patients. Overall survival was significantly better in T cell-deficient group than T cell-enriched group (p = 0.019), and T8 dominant group than T4 dominant group (p = 0.023). The number of tumor-associated macrophages (TAM) and M2-TAM was significantly decreased in isocitrate dehydrogenase mutated HGG. Radiomic signature classification showed good performance in predicting immune phenotypes especially with features extracted from apparent diffusion coefficient maps.
Conclusions
Absolute quantification of tumor-infiltrating immune cells confirmed the heterogeneity of immune microenvironment in HGG which harbors prognostic impact. This immune microenvironment could be predicted by radiomic signatures non-invasively.




Similar content being viewed by others
Data availability
All data are published within this paper and within accompanying supporting files (indicated in text) and accessed via weblink on the journal site.
References
Brooks WH, Markesbery WR, Gupta GD, Roszman TL (1978) Relationship of lymphocyte invasion and survival of brain tumor patients. Ann Neurol 4(3):219–224. https://doi.org/10.1002/ana.410040305
Louveau A, Smirnov I, Keyes TJ, Eccles JD, Rouhani SJ, Peske JD, Derecki NC, Castle D, Mandell JW, Lee KS, Harris TH, Kipnis J (2015) Structural and functional features of central nervous system lymphatic vessels. Nature 523(7560):337–341. https://doi.org/10.1038/nature14432
Hu X, Deng Q, Ma L, Li Q, Chen Y, Liao Y, Zhou F, Zhang C, Shao L, Feng J, He T, Ning W, Kong Y, Huo Y, He A, Liu B, Zhang J, Adams R, He Y, Tang F, Bian X, Luo J (2020) Meningeal lymphatic vessels regulate brain tumor drainage and immunity. Cell Res 30(3):229–243. https://doi.org/10.1038/s41422-020-0287-8
Orrego E, Castaneda CA, Castillo M, Bernabe LA, Casavilca S, Chakravarti A, Meng W, Garcia-Corrochano P, Villa-Robles MR, Zevallos R, Mejia O, Deza P, Belmar-Lopez C, Ojeda L (2018) Distribution of tumor-infiltrating immune cells in glioblastoma. CNS oncol 7(4):CNS21. https://doi.org/10.2217/cns-2017-0037
Han S, Zhang C, Li Q, Dong J, Liu Y, Huang Y, Jiang T, Wu A (2014) Tumour-infiltrating CD4(+) and CD8(+) lymphocytes as predictors of clinical outcome in glioma. Br J Cancer 110(10):2560–2568. https://doi.org/10.1038/bjc.2014.162
Hussain SF, Yang D, Suki D, Aldape K, Grimm E, Heimberger AB (2006) The role of human glioma-infiltrating microglia/macrophages in mediating antitumor immune responses. Neuro Oncol 8(3):261–279. https://doi.org/10.1215/15228517-2006-008
Palma L, Di Lorenzo N, Guidetti B (1978) Lymphocytic infiltrates in primary glioblastomas and recidivous gliomas. Incidence, fate, and relevance to prognosis in 228 operated cases. J Neurosurg 49(6):854–861. https://doi.org/10.3171/jns.1978.49.6.0854
Quail DF, Joyce JA (2017) The microenvironmental landscape of brain tumors. Cancer Cell 31(3):326–341. https://doi.org/10.1016/j.ccell.2017.02.009
Liu Z, Meng Q, Bartek J Jr, Poiret T, Persson O, Rane L, Rangelova E, Illies C, Peredo IH, Luo X, Rao MV, Robertson RA, Dodoo E, Maeurer M (2017) Tumor-infiltrating lymphocytes (TILs) from patients with glioma. Oncoimmunology 6(2):e1252894. https://doi.org/10.1080/2162402X.2016.1252894
Gabrusiewicz K, Rodriguez B, Wei J, Hashimoto Y, Healy LM, Maiti SN, Thomas G, Zhou S, Wang Q, Elakkad A, Liebelt BD, Yaghi NK, Ezhilarasan R, Huang N, Weinberg JS, Prabhu SS, Rao G, Sawaya R, Langford LA, Bruner JM, Fuller GN, Bar-Or A, Li W, Colen RR, Curran MA, Bhat KP, Antel JP, Cooper LJ, Sulman EP, Heimberger AB (2016) Glioblastoma-infiltrated innate immune cells resemble M0 macrophage phenotype. JCI Insight. https://doi.org/10.1172/jci.insight.85841
Brandenburg S, Turkowski K, Mueller A, Radev YT, Seidlitz S, Vajkoczy P (2017) Myeloid cells expressing high level of CD45 are associated with a distinct activated phenotype in glioma. Immunol Res 65(3):757–768. https://doi.org/10.1007/s12026-017-8915-1
Berghoff AS, Kiesel B, Widhalm G, Wilhelm D, Rajky O, Kurscheid S, Kresl P, Wohrer A, Marosi C, Hegi ME, Preusser M (2017) Correlation of immune phenotype with IDH mutation in diffuse glioma. Neuro Oncol 19(11):1460–1468. https://doi.org/10.1093/neuonc/nox054
Yu P, Fu YX (2006) Tumor-infiltrating T lymphocytes: friends or foes? Laboratory Invest 86(3):231–245. https://doi.org/10.1038/labinvest.3700389
Zhang J, Caruso FP, Sa JK, Justesen S, Nam DH, Sims P, Ceccarelli M, Lasorella A, Iavarone A (2019) The combination of neoantigen quality and T lymphocyte infiltrates identifies glioblastomas with the longest survival. Commun Biol 2:135. https://doi.org/10.1038/s42003-019-0369-7
Castaneda CA, Castillo M, Aliaga K, Bernabe LA, Casavilca S, Sanchez J, Torres-Cabala CA, Gomez HL, Mas L, Dunstan J, Cotrina JM, Abugattas J, Chavez I, Ruiz E, Montenegro P, Rojas V, Orrego E, Galvez-Nino M, Felix B, Landa-Baella MP, Vidaurre T, Villa MR, Zevallos R, Taxa L, Guerra H (2019) Level of tumor-infiltrating lymphocytes and density of infiltrating immune cells in different malignancies. Biomark Med 13(17):1481–1491. https://doi.org/10.2217/bmm-2019-0178
Martinez-Lage M, Lynch TM, Bi Y, Cocito C, Way GP, Pal S, Haller J, Yan RE, Ziober A, Nguyen A, Kandpal M, O’Rourke DM, Greenfield JP, Greene CS, Davuluri RV, Dahmane N (2019) Immune landscapes associated with different glioblastoma molecular subtypes. Acta Neuropathol Commun 7(1):203. https://doi.org/10.1186/s40478-019-0803-6
Zhang B, Chang K, Ramkissoon S, Tanguturi S, Bi WL, Reardon DA, Ligon KL, Alexander BM, Wen PY, Huang RY (2017) Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro Oncol 19(1):109–117
Choi KS, Choi SH, Jeong B (2019) Prediction of IDH genotype in gliomas with dynamic susceptibility contrast perfusion MR imaging using an explainable recurrent neural network. Neuro Oncol 21(9):1197–1209
Binnewies M, Roberts EW, Kersten K, Chan V, Fearon DF, Merad M, Coussens LM, Gabrilovich DI, Ostrand-Rosenberg S, Hedrick CC (2018) Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med 24(5):541–550
Wang G, Li W, Vercauteren T, Ourselin S (2019) Automatic brain tumor segmentation based on cascaded convolutional neural networks with uncertainty estimation. Front Comput Neurosci 13:56
Shinohara RT, Sweeney EM, Goldsmith J, Shiee N, Mateen FJ, Calabresi PA, Jarso S, Pham DL, Reich DS, Crainiceanu CM (2014) Statistical normalization techniques for magnetic resonance imaging. NeuroImage Clin 6:9–19
Van Griethuysen JJ, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RG, Fillion-Robin J-C, Pieper S, Aerts HJ (2017) Computational radiomics system to decode the radiographic phenotype. Can Res 77(21):e104–e107
Rohart F, Gautier B, Singh A, Lê Cao K-A (2017) mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol 13(11):e1005752
Le Cao KA, Boitard S, Besse P (2011) Sparse PLS discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinformatics 12:253. https://doi.org/10.1186/1471-2105-12-253
Barker M, Rayens W (2003) Partial least squares for discrimination. J Chemom 17(3):166–173
Hendry S, Salgado R, Gevaert T, Russell PA, John T, Thapa B, Christie M, van de Vijver K, Estrada MV, Gonzalez-Ericsson PI, Sanders M, Solomon B, Solinas C, Van den Eynden G, Allory Y, Preusser M, Hainfellner J, Pruneri G, Vingiani A, Demaria S, Symmans F, Nuciforo P, Comerma L, Thompson EA, Lakhani S, Kim SR, Schnitt S, Colpaert C, Sotiriou C, Scherer SJ, Ignatiadis M, Badve S, Pierce RH, Viale G, Sirtaine N, Penault-Llorca F, Sugie T, Fineberg S, Paik S, Srinivasan A, Richardson A, Wang Y, Chmielik E, Brock J, Johnson DB, Balko J, Wienert S, Bossuyt V, Michiels S, Ternes N, Burchardi N, Luen SJ, Savas P, Klauschen F, Watson PH, Nelson BH, Criscitiello C, O’Toole S, Larsimont D, de Wind R, Curigliano G, Andre F, Lacroix-Triki M, van de Vijver M, Rojo F, Floris G, Bedri S, Sparano J, Rimm D, Nielsen T, Kos Z, Hewitt S, Singh B, Farshid G, Loibl S, Allison KH, Tung N, Adams S, Willard-Gallo K, Horlings HM, Gandhi L, Moreira A, Hirsch F, Dieci MV, Urbanowicz M, Brcic I, Korski K, Gaire F, Koeppen H, Lo A, Giltnane J, Rebelatto MC, Steele KE, Zha J, Emancipator K, Juco JW, Denkert C, Reis-Filho J, Loi S, Fox SB (2017) Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the international immuno-oncology biomarkers working group: part 2: TILs in melanoma, gastrointestinal tract carcinomas, non-small cell lung carcinoma and mesothelioma, endometrial and ovarian carcinomas, squamous cell carcinoma of the head and neck, genitourinary carcinomas, and primary brain tumors. Adv Anat Pathol 24(6):311–335. https://doi.org/10.1097/PAP.0000000000000161
Fridman WH, Pages F, Sautes-Fridman C, Galon J (2012) The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 12(4):298–306. https://doi.org/10.1038/nrc3245
Kim YH, Jung TY, Jung S, Jang WY, Moon KS, Kim IY, Lee MC, Lee JJ (2012) Tumour-infiltrating T-cell subpopulations in glioblastomas. Br J Neurosurg 26(1):21–27. https://doi.org/10.3109/02688697.2011.584986
Yang I, Tihan T, Han SJ, Wrensch MR, Wiencke J, Sughrue ME, Parsa AT (2010) CD8+ T-cell infiltrate in newly diagnosed glioblastoma is associated with long-term survival. J Clin Neurosci 17(11):1381–1385. https://doi.org/10.1016/j.jocn.2010.03.031
Yue Q, Zhang X, Ye HX, Wang Y, Du ZG, Yao Y, Mao Y (2014) The prognostic value of Foxp3+ tumor-infiltrating lymphocytes in patients with glioblastoma. J Neurooncol 116(2):251–259. https://doi.org/10.1007/s11060-013-1314-0
Franklin RA, Liao W, Sarkar A, Kim MV, Bivona MR, Liu K, Pamer EG, Li MO (2014) The cellular and molecular origin of tumor-associated macrophages. Science 344(6186):921–925. https://doi.org/10.1126/science.1252510
Hambardzumyan D, Gutmann DH, Kettenmann H (2016) The role of microglia and macrophages in glioma maintenance and progression. Nat Neurosci 19(1):20–27. https://doi.org/10.1038/nn.4185
Tomaszewski W, Sanchez-Perez L, Gajewski TF, Sampson JH (2019) Brain tumor microenvironment and host state: implications for immunotherapy. Clin Cancer Res 25(14):4202–4210. https://doi.org/10.1158/1078-0432.CCR-18-1627
Prosniak M, Harshyne LA, Andrews DW, Kenyon LC, Bedelbaeva K, Apanasovich TV, Heber-Katz E, Curtis MT, Cotzia P, Hooper DC (2013) Glioma grade is associated with the accumulation and activity of cells bearing M2 monocyte markers. Clin Cancer Res 19(14):3776–3786. https://doi.org/10.1158/1078-0432.CCR-12-1940
Poon CC, Gordon PMK, Liu K, Yang R, Sarkar S, Mirzaei R, Ahmad ST, Hughes ML, Yong VW, Kelly JJP (2019) Differential microglia and macrophage profiles in human IDH-mutant and -wild type glioblastoma. Oncotarget 10(33):3129–3143. https://doi.org/10.18632/oncotarget.26863
Sun R, Limkin EJ, Vakalopoulou M, Dercle L, Champiat S, Han SR, Verlingue L, Brandao D, Lancia A, Ammari S (2018) A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol 19(9):1180–1191
Choi Y, Nam Y, Lee YS, Kim J, Ahn K-J, Jang J, Shin N-Y, Kim B-S, Jeon S-S (2020) IDH1 mutation prediction using MR-based radiomics in glioblastoma: comparison between manual and fully automated deep learning-based approach of tumor segmentation. Eur J Radiol. https://doi.org/10.1016/j.ejrad.2020.109031
Yu J, Shi Z, Lian Y, Li Z, Liu T, Gao Y, Wang Y, Chen L, Mao Y (2017) Noninvasive IDH1 mutation estimation based on a quantitative radiomics approach for grade II glioma. Eur Radiol 27(8):3509–3522
Patel SH, Poisson LM, Brat DJ, Zhou Y, Cooper L, Snuderl M, Thomas C, Franceschi AM, Griffith B, Flanders AE (2017) T2–FLAIR mismatch, an imaging biomarker for IDH and 1p/19q status in lower-grade gliomas: a TCGA/TCIA project. Clin Cancer Res 23(20):6078–6085
Acknowledgements
We thank Hye Jung Hong, R.N. for her support in collecting clinical data and laboratory management in contribution to this study.
Funding
This work was supported by the following funding sources: a National Research Foundation of Korea (NRF) grant funded by the Korea government Ministry of Science and ICT (MSIT) (NRF-2019R1A2C2005144 to C.-K.P.), the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2020R1A2C2008949 to S.H.C.), the Creative-Pioneering Researchers Program through Seoul National University (SNU; to S.H.C.), and the Institute for Basic Science (IBS-R006-A1 to S.H.C.)
Author information
Authors and Affiliations
Contributions
ARK, KSC, MSK, SHC, E-CS and C-K.P wrote the manuscript. C.-KP, E-CS and SHC contributed to the conception and design of the project and revised the manuscript. SK, TC, HJY, and CEL collected tissue samples and performed the experiments. ARK and E-CS performed flow cytometric analysis. KSC and SHC performed the radiomics analysis. K-MK, HK, JHL, S-TL, JWK, Y-HK, and TMK contributed to the collection and analysis of clinical data. JKW and S-HP. performed histological diagnosis and molecular/genetic examinations. All authors participated in drafting and revising the article for important intellectual content and approved the final version for publication.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This study involving human tissue and cells was approved by the Ethics Committee of the Seoul National University Hospital (IRB No. H-1902-062-1010).
Informed consent to participate
All tissue and data were anonymized. This study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, A.R., Choi, K.S., Kim, MS. et al. Absolute quantification of tumor-infiltrating immune cells in high-grade glioma identifies prognostic and radiomics values. Cancer Immunol Immunother 70, 1995–2008 (2021). https://doi.org/10.1007/s00262-020-02836-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02836-w



