Abstract
Pancreatic ductal adenocarcinoma is characterized by a strong immunosuppressive network with a dense infiltration of myeloid cells including myeloid-derived suppressor cells (MDSC). Two distinct populations of MDSC have been defined: polymorphonuclear MDSC (PMN-MDSC) and monocytic MDSC (M-MDSC). Several factors influence the development and function of MDSC including the transcription factor interferon regulatory factor 4 (IRF4). Here, we show that IRF4 deficiency accelerates tumor growth and reduces survival, accompanied with a dense tumor infiltration with PMN-MDSC and reduced numbers of CD8+ T cells. As IRF4 has been described to modulate myeloid cell development and function, particularly of PMN-MDSC, we analyzed its role using MDSC-specific IRF4 knockout mice with the Ly6G or LysM knock-in allele expressing Cre recombinase and Irf4flox. In GM-CSF-driven bone marrow cultures, IRF4 deficiency increased the frequency of MDSC-like cells with a strong T cell suppressive capacity. Myeloid (LysM)-specific depletion of IRF4 led to increased tumor weight and a moderate splenic M-MDSC expansion in tumor-bearing mice. PMN cell (Ly6G)-specific depletion of IRF4, however, did not influence tumor progression or MDSC accumulation in vivo in accordance with our finding that IRF4 is not expressed in PMN-MDSC. This study demonstrates a critical role of IRF4 in the generation of an immunosuppressive tumor microenvironment in pancreatic cancer, which is independent of IRF4 expression in PMN-MDSC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The tumor microenvironment (TME) contains a variety of immune cells with opposing functions. While anti-tumoral cells such as cytotoxic T cells, T helper cells and natural killer cells are important in the immune surveillance and protection against tumors, certain immune modulatory cells including myeloid subsets shape the TME during immune equilibrium and escape phase [1] and enhance tumor progression by dampening the effector cells [2]. Three main subsets within the myeloid compartment are characterized by a strong T cell suppressive capacity. Depending on the cell of origin and marker expression, they can be classified in (1) monocytic myeloid-derived suppressor cells (M-MDSC), (2) polymorphonuclear myeloid-derived suppressor cells (PMN-MDSC) and (3) tumor-associated macrophages (TAM) [3, 4]. Under pathological activation such as chronic inflammation or cancer MDSC are expanded and gain a suppressive phenotype [5]. Increased numbers of MDSC are a negative prognostic factor for the survival of patients with pancreatic cancer (PDAC) [6, 7]. In addition, a MDSC-enriched TME is a negative predictive marker for the response to immunotherapy in a mouse model of PDAC [8]. Despite their high clinical relevance, MDSC-targeted treatment approaches are still insufficient [9] and there is great need for a better understanding of MDSC development and activation in cancer [10].
Both MDSC subsets develop from the granulocyte macrophage precursor (GMP) in the bone marrow, and several transcription factors are associated with development and function of MDSC [11]. Signal transducer and activator of transcription 3 (STAT3) has been shown to regulate MDSC expansion by mediating myeloid-specific growth factor signaling [12] as well as the suppressive activity by inducing Arginase 1 (ARG1) expression, which suppresses T cell activity by degrading the essential amino acid arginine [13]. Expression of arginase is one key suppressive mechanism of MDSC.[3]. CCAAT/enhancer-binding protein beta (C/EBPβ) regulates the development of polymorphonuclear cells under inflammatory conditions and has been also associated with PMN-MDSC development [14]. The transcription factor interferon regulatory factor 8 (IRF8) plays a central role in the development of monocytic and polymorphonuclear cells from GMP [15]. IRF8-deficient mice show leukemic-like symptoms with a dense accumulation of polymorphonuclear cells [16]. In the context of cancer, strong accumulation of PMN-MDSC has been reported for IRF8-deficient mice [17].
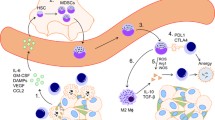
Another transcription factor of the IRF family, IRF4 is known for its function in lymphoid cells. There, IRF4 regulates antibody class switching in B cells [18, 19] and facilitates the differentiation of naïve CD4+ T cells into T helper cell subsets such as Th2, Th9 or Th17 [20]; in CD8+ T cells, IRF4 plays a crucial role in maintaining effector function [21, 22]. Besides its importance in the lymphoid lineage, IRF4 expression has been shown to regulate similar biological functions as observed for IRF8 in myeloid cells. IRF4 is expressed in myeloid-precursor cells and shifts the myeloid development from neutrophil to monocyte development, but to a lesser extent than IRF8 [23]. In contrast to early myeloid cell development, IRF4 and IRF8 play opposing roles in the development of conventional dendritic cells (cDC) that develop from GMP via pre-cDC precursor cells. While IRF4 is one of the lineage-defining transcription factors in the differentiation of cDC2, IRF8 is pivotal for the differentiation of cDC1 [24, 25]. Recently, it has been shown that differentiation of Ly6C+ monocytes to monocytic DC (moDC) requires IRF4 [26]. Moreover, IRF4 expression is induced by cytokines mediating M2 alternatively activated macrophage polarization such as IL-4 and IL-13 [27], and itself regulates M2 polarization by inducing M2 effector genes like Arg1 [28]. A recent publication indicates that IRF4 may regulate the differentiation and suppressive function of MDSC [29].
In this study, we aim at understanding the role of IRF4 in MDSC development and function in a murine model of PDAC which is characterized by strong expansion of MDSC [30, 31], by using mice with either global, myeloid cell- or granulocyte-specific Irf4 deletion.
Results
Systemic IRF4 deficiency accelerates PDAC tumor growth and expands MDSC in vivo
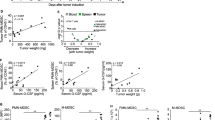
To study the role of IRF4 in pancreatic cancer we made use of a transplantable, orthotopic tumor model using the tumor cell line T110299, which originate from a genetically engineered, spontaneous mouse model with PDAC-characteristic driver mutations (KrasG12D Tp53R172H). We recently reported that this model is particularly enriched for myeloid cells such as MDSC [31]. Three weeks after orthotopic tumor induction, tumor weight of Irf4−/− mice was significantly increased compared to IRF4 sufficient Irf4flox/flox controls (Fig. 1a, b). Moreover, the survival of tumor-bearing Irf4−/− mice was significantly reduced (Fig. 1c). We noted no significant differences in histologic tumor morphology between both genotypes (Fig. S1). T110299 tumors from both control and Irf4−/− mice showed a moderately differentiated tubular to tubulo-papillary adenocarcinoma pattern with similar amounts of desmoplastic stroma and moderate chronic inflammatory infiltrates, resembling human pancreatic ductal adenocarcinoma. To study the immunological consequences of IRF4 deficiency, immune cell frequency of T110299 tumor-bearing mice was analyzed three weeks after tumor induction by flow cytometry. In the spleen of Irf4−/− mice, PMN-MDSC, M-MDSC and CD4+ T cell frequencies were significantly increased, whereas the frequency of CD8+ T cells was reduced (Fig. 1d). In the TME, PMN-MDSC frequency was significantly increased in Irf4−/− mice, whereas the frequency of CD8+ T cells was dramatically reduced, indicating a profound immunosuppressive TME in mice deficient for IRF4 (Fig. 1e) that may account for the more rapid tumor progression in these mice.
IRF4 deficiency accelerates PDAC tumor growth and expansion of MDSC populations in blood and spleen. a–e T110299 tumors were induced orthotopically in Irf4−/− and wild-type (WT) mice. After 21 days, tumor weight was measured (b). Kaplan–Meier curve depicts survival of orthotopic PDAC-bearing mice (c). Immune cell frequencies in spleen (d) and tumor (e) were analyzed by flow cytometry. Differences between genotypes were statically analyzed using the Mann–Whitney U test (b, d, e) or Log rank test (c). a–e Pooled data from 2–3 independent experiments are shown, error bars represent mean ± SEM (n = 3–8 mice/group). e IRF4 expression level of metastasized PDAC patients and Kaplan–Meier survival curve of patients from the upper and lower quartile of IRF4 expression is displayed for 36 months. Median survival of both groups in months is indicated with dotted lines (f). Contingency table of patients based on their survival status after one year and IRF4 expression level (g). Asterisks indicate *p < 0.05, **p < 0.01, ***p < 0.001
Low IRF4 expression in PDAC patients is associated with reduced one-year survival
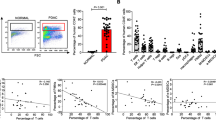
To confirm these findings in the human disease, the effect of IRF4 expression on the survival of metastasized PDAC patients was analyzed using the Cancer Genome Atlas (TCGA) data set. Metastasized PDAC patients were categorized based on their IRF4 expression level in two groups (upper and lower quartile) and survival of these selected groups was analyzed. The median survival of patients with low IRF4 expression was 15.9 months versus 18.7 months of patients with high expression levels. There was no significant difference in the overall survival in regard to the IRF4 expression (Fig. 1f). However, χ2-test of survival after 1 year revealed a significantly reduced survival of patients with lower IRF4 expression (χ2 = 5.33, p = 0.02) (Fig. 1g).
IRF4 is expressed in M-MDSC, but not PMN-MDSC
To better understand the role of IRF4 in the initiation of the immunosuppressive and myeloid cell-enriched TME, IRF4 expression of both monocytic and polymorphonuclear cells in tumor-free and T110299 tumor-bearing mice was analyzed by flow cytometry. IRF4 was expressed homogenously in monocytic cells in blood, spleen as well as tumor, and was independent of the tumor status. In contrast, IRF4 expression in polymorphonuclear cells in blood, spleen and tumor was absent (Fig. 2a, b).
IRF4 is expressed in monocytic but not polymorphonuclear cells. a–d KPC-derived T110299 tumors were induced orthotopically. a, b IRF4 expression of MDSC subsets from tumor-free and PDAC-bearing wild-type mice was analyzed by flow cytometry. a A representative histogram of anti-IRF4 staining as well as isotype control staining from a PDAC-bearing mouse is depicted. After 21 days, GFP expression of MDSC (c) and GMP (d) subsets of Ly6GCreIrf4fl/fl (green), LysMCreIrf4fl/fl (blue) and Irf4fl/fl (black) mice was determined by FACS analysis. e IRF4 expression of myeloid cells from bone marrow cultures were analyzed by flow cytometry in the absence and presence of IL-4 and IL-13. The difference between stimulated and unstimulated cells was statistically analyzed using unpaired student’s t-test. Error bars represent mean ± SEM (n = 2–5 mice/group), asterisks indicate *p < 0.05, **p < 0.01
As this is in contrast to published data [29], we further validated this finding by alternative approaches and used conditional myeloid-specific IRF4 knockout mouse models. Irf4flox mice have been designed to express green fluorescent protein (GFP) instead of IRF4 upon successful Cre-mediated recombination under the same physiological control as IRF4 [18]. Therefore, the conditional IRF4 knockout mice can also act as reporter mice. LysMCreIrf4fl/fl mice have an IRF4 deletion in polymorphonuclear cells, monocytes, macrophages and partly dendritic cells [32], whereas Ly6GCreIrf4fl/fl mice have the IRF4 deletion in polymorphonuclear cells only [33].
In line with our previous observation, M-MDSC of the LysMCreIrf4fl/fl mice were positive for GFP in the blood, spleen and tumor. No GFP expression was detectable in PMN-MDSC or M-MDSC of the Ly6GCreIrf4fl/fl mice. In contrast, in LysMCreIrf4fl/fl mice, a small fraction of approximately 15% of GFP+ PMN-MDSC was detectable in blood, spleen and tumor (Fig. 2c). As the GFP expression in those PMN-MDSC may derive from its precursor cell expressing LysM, we characterized GFP expression of GMP in LysMCreIrf4fl/fl mice. Indeed, 12% of GMP expressed GFP indicating that IRF4 is expressed early in the myeloid cell development (Fig. 2d). To further characterize the cell-specific expression pattern of IRF4 in myeloid cells, bone marrow cells were stimulated overnight with the known IRF4 inducers IL-4 and IL-13. As expected, CD11b+ CD11c+ MHC-II+ bone marrow-derived dendritic cells (BMDC) responded to cytokine stimulation by upregulating IRF4 expression. IRF4 expression was only inducible in CD11b+Ly6ChighLy6G− M-MDSC-like cells, but not in CD11b+Ly6CintLy6G+ PMN-MDSC-like cells (Fig. 2e). Taken together, the data demonstrate that IRF4 is expressed in M-MDSC and myeloid precursor cells but is absent in mature Ly6G-expressing PMN-MDSC.
IRF4 deficiency expands suppressive MDSC-like cells in vitro
GM-CSF-driven bone marrow cultures are an established model to study MDSC in vitro [14]. To evaluate the role of IRF4 in the development and function of MDSC, bone marrow cells of wild-type and Irf4−/− mice were cultured for seven days in the presence of GM-CSF, and cell composition was analyzed. The frequency of MDSC-like (Gr1+MHC-IIlow) cells was significantly increased in bone marrow cultures from Irf4−/− mice, while both MHC-IIint and MHC-IIhigh BMDC were significantly reduced (Fig. 3a, b). Next, T cell suppressive capacity of bone marrow-derived MDSC was measured in T cell co-cultures in the presence of anti-CD3/anti-CD28 beads. T cell proliferation was measured and revealed that IRF4-deficient bone marrow culture cells suppressed the proliferation of both CD4+ and CD8+ T cells more potently, as compared to wild-type controls (Fig. 3c). However, as the difference in MDSC composition may influence the overall suppressive activity, FACS-sorted Gr1highMHC-IIlow MDSC-like cells from wild-type and Irf4−/− mice were individually analyzed in a T cell suppression assay. Sorted Gr1+ MDSC-like cells of wild-type and Irf4−/− mice exhibited similar T cell suppressive capacities, arguing against a direct role of IRF4 in controlling the suppressive capacity of MDSC-like cells (Fig. 3d).
IRF4 deficiency leads to MDSC-like cell expansion in bone marrow cultures in vitro. a–d Bone marrow cells from WT and Irf4−/− mice were cultured for seven days in the presence of GM-CSF. a, b Cell composition of bone marrow cultures was measured. c Bone marrow cells from WT and IRF4-deficient mice were co-cultured with CFSE-labeled T cells and proportion of proliferated CFSElow CD8+ and CD4+ T cells was analyzed by flow cytometry. d Gr1high MHC-IIlow cell population of WT and IRF4-deficient bone marrow cultures were FACSorted, co-cultured with CFSE-labeled T cells and the proportion of proliferated CFSElow CD8+ and CD4+ was determined. The difference between genotypes was statistically analyzed using Mann–Whitney U test, error bars represent mean ± SEM (n = 3), asterisks indicate *p < 0.05 and ***p < 0.001
Myeloid-specific deletion of IRF4 accelerates PDAC growth and expands M-MDSC numbers
The two conditional IRF4 knockout mouse models described above were used to study the intrinsic role of IRF4 in myeloid cells in vivo. KPC-derived T110299 tumors were orthotopically induced in Ly6GCreIrf4fl/fl mice, LysMCreIrf4fl/fl mice as well as Irf4fl/fl control mice (Fig. 4a). While there was no significant difference in tumor growth between Ly6GCreIrf4fl/fl and control mice, tumor weight was significantly increased in LysMCreIrf4fl/fl mice, as compared to Irf4fl/fl control mice (Fig. 4b). Flow cytometric analysis of MDSC frequencies three weeks after tumor induction showed no difference in PMN-MDSC frequency in blood, spleen and tumor of LysMCreIrf4fl/fl mice and Ly6GCreIrf4fl/fl mice compared to Irf4fl/fl controls; however, M-MDSC frequency was moderately increased in the spleen of LysMCreIrf4fl/fl mice (Fig. 4c). Compared to Irf4fl/fl controls, there was no significant difference in CD4+ or CD8+ T cell frequencies in LysMCreIrf4fl/fl mice or Ly6GCreIrf4fl/fl mice (Fig. 4d). Of note, both myeloid-specific IRF4-deficiencies had no influence on the survival of PDAC-bearing mice (Fig. 4e).
Targeted deletion of IRF4 in LysMCre but not Ly6GCre mice increases tumor growth without influencing survival. a–d KPC-derived T110299 tumors were induced orthotopically in Ly6GCreIrf4fl/fl (green), LysMCreIrf4fl/fl (blue) and Irf4fl/fl (black) mice. After 21 days, a, b tumor weight was measured and c MDSC as well as d T cell frequency was analyzed by flow cytometry. e Survival of PDAC-bearing mice was monitored. Statistical analysis was performed using Kruskal Wallis test, followed by Dunn’s multiple comparison test between controls and conditional knockout mice. Pooled data from three independent experiments are depicted, error bars represent mean ± SEM (n = 8–14 mice/group), p value of Dunn’s test are shown in the graph, asterisks indicate *p < 0.05 and **p < 0.01
In conclusion, this study demonstrates that only early myeloid precursor cells and M-MDSC express IRF4, whereas mature PMN-MDSC lack IRF4 expression. In our hands, specific deletion of IRF4 in PMN-MDSC has no influence on PMN-MDSC expansion, their T cell suppressive capacity, tumor growth or survival. In contrast, myeloid cell-specific deletion of IRF4 in LysMCreIrf4fl/fl mice slightly accelerates tumor growth, which was accompanied by increased M-MDSC frequency in the spleen. No impact on PMN-MDSC numbers or survival is seen in these mice. These findings argue against a pro-tumoral effect of IRF4 in PMN-MDSC. Hence, as observed in mice with global IRF4 deficiency, the IRF4-mediated effects in other immune cells likely account for the immunosuppressive TME with its dense MDSC infiltration.
Discussion
In this study, we show that global IRF4 deficiency accelerates tumor growth, increases frequencies of tumor-infiltrating PMN-MDSC and reduces survival in a murine PDAC model. This is in line with human TCGA cohort data showing that the one-year survival of metastasized PDAC patients with low IRF4 expression is significantly reduced. Reporter mice for studying IRF4 expression in myeloid cells in vivo revealed the induction of IRF4 in myeloid precursor cells and M-MDSC, but not in PMN-MDSC. Accordingly, IRF4 deletion in Ly6G+ cells had no influence on MDSC frequency, the suppressive function, tumor growth or survival. In contrast, IRF4 deletion in LysM+ cells increased tumor weight and led to moderately expanded M-MDSC population in the spleen; yet there was no impact on survival.
A recent study reported that IRF4 is expressed by PMN-MDSC as well as M-MDSC and that 4T1 breast tumors reduced the expression of IRF4 in both cell types. Furthermore, the study demonstrated by knock-down and overexpression experiments in bone marrow cultures showed that lower IRF4 expression increased the frequency of PMN-MDSC and was associated with higher suppressive activity of MDSC. A myeloid cell-specific knockout of IRF4 increased the tumor weight in B16F10 tumor bearing mice [29]. In contrast to this report, we clearly demonstrate by the use of reporter mice as well as antibody staining that in our PDAC model IRF4 expression is limited to M-MDSC, whereas PMN-MDSC lack IRF4 expression [29]. As the 4T1 cell line was generated from a BALB/c mouse and the PDAC model we used originated from a C57BL/6 mouse, it remains open whether this discrepancy is due to different tumor models or mouse strains used.
A fraction of GMP was found to express IRF4, which is why they likely contain precursors for both M-MDSC and PMN-MDSC. In line with this finding, a small proportion of PMN-MDSC exhibited a weak GFP expression in LysMCreIrf4flox reporter mice. Taking together, the findings argue that IRF4 is expressed in some precursors of granulocytes, but is lost during PMN-MDSC differentiation. Considering our data and the current literature of IRF4 and IRF8 data on early myeloid cell development, IRF4 and IRF8 have seemingly overlapping functions in myeloid cell development. The data also suggest that IRF8 may compensate for IRF4 function in IRF4-deficient animals [23]; however, the exact molecular mechanism of how IRF4-regulates cell fate decision remains elusive.
The suppressive function of MDSC has been linked to their immature state [34]. The GM-CSF-driven bone marrow culture system is frequently used to study dendritic cells and MDSC in vitro [14, 29]. IRF4-deficiency shifts the bone marrow culture from mature dendritic cells towards immature MDSC-like cells. This shift in the population frequency questions again earlier findings showing that induction of IRF4 decreases and inhibition of IRF4 increases the suppressive capacity of MDSC intrinsically [29]. Using sorted MDSC-like cells from these cultures as well as genetic deletion instead of siRNA knock-down, we demonstrate that the intrinsic suppressive activity is not directly influenced by IRF4.
Given the result that IRF4 is not expressed in polymorphonuclear cells, it was not surprising that the deletion of IRF4 in Ly6G+ cells in vivo did not influence tumor growth, overall survival or MDSC cell frequency. In line with the previous report [29], myeloid-specific deletion of IRF4 using the LysMCreIrf4fl/fl mice, accelerated tumor growth, but had no influence on the overall survival. The frequency of M-MDSC was moderately elevated in the spleen, but not in tumors, suggesting that IRF4 rather impacts monocyte differentiation or proliferation.
Besides the minor effects of myeloid-specific deletion of IRF4, the accelerated tumor growth in global IRF4-deficient mice demonstrates a central role of IRF4 in shaping the TME and anti-tumor immunity. IRF4 has been described to be pivotal for efficient antigen cross-presentation of moDC [26] and to be required for sustained CD8+ T cell activation [21, 22]. Given the substantial reduction of intratumoral CD8+ T cells, it seems more likely that this is due to an impaired sustained activation of antitumoral T cells than due to the amplification and action of MDSC in IRF4-deficent mice. We recently demonstrated that in the PDAC model used the PMN-MDSC frequency correlates with tumor weight [31]. One can therefore argue that the elevated levels of PMN-MDSC in the global IRF4 deficient mouse could be a secondary effect of the increased tumor size.
In summary, we demonstrate that IRF4 plays an important role in shaping the immune cell composition in the TME of murine pancreatic cancer. Due to the increased PMN- and M-MDSC frequency in IRF4 knockout mice, we hypothesized that a MDSC-intrinsic role of IRF4 might explain the effect on tumor progression. However, polymorphonuclear cells do not express IRF4 in tumor bearing mice and in line with that, the deletion of IRF4 in Ly6G+ cells did not alter tumor development. Despite some effects of IRF4 deletion in LysM+ cells on tumor growth, again, no effect on survival or PMN-MDSC accumulation was observed. Our results, therefore, suggest that the observed in vivo effects in globally IRF4-deficient mice are secondary and due to globally imbalanced immune regulation, but not due to an IRF4-intrinsic effect in MDSC.
Material and methods
Mice
C57BL/6 wild-type mice were purchased from Janvier, France. Irf4flox mice (B6.129S1-Irf4tm1Rdf/J) were a kind gift from Prof. Bopp (Institute of Immunology, Universitätsmedizin Mainz), Ly6GCre mice (C57BL/6-Ly6g(tm2621(Cre−tdTomato)Arte)) were a kind gift from Prof. Gunzer (Institute for Experimental Immunology and Imaging, University of Duisburg-Essen), LysMCre mice (B6.129P2-Lyz2tm1(cre)Ifo/J) were a kind gift from PD Dr. Lech (Institute of Clinical Biochemistry, Klinikum der Universität München). FLP1 recombinase expressing FLPe mice (B6;SJL-Tg(ACTFLPe)9205Dym)/J) were purchased from Jackson Laboratory (Sulzfeld, Germany). All mice were kept on C57BL/6 background with a 12-h light/dark cycle, water ad lib. and regular chow diet (sniff, Soest, Germany) at the Klinikum der Universität München, Munich, Germany. Experiments were performed according to national ethical guidelines approved by the local government (Regierung von Oberbayern, Munich, Germany; file number 55.2-1-54-2532-175-12). LysMCre were cross-bred with Irf4flox mice to obtain LysMCreIrf4flox, and Ly6GCre were cross-bred with Irf4flox mice to obtain Ly6GCreIrf4flox. Both mouse strains were kept on homozygous Irf4fl/fl background. Exons 1 and 2 of Irf4 in B6.129S1-Irf4tm1Rdf/J mice are flanked by two FRT sites. To generate global IRF4-deficient mice, B6;SJL-Tg(ACTFLPe)9205Dym/J mice were cross-bred with B6.129S1-Irf4tm1Rdf/J mice and, as described before, Irf4−/− mice were obtained [18]. IRF4 sufficient mice originating from those breedings were used as littermate controls. Genotypes of all mice were routinely analyzed by PCR.
DNA isolation and genotyping
DNA from ear or tail biopsies was extracted and analyzed as described before [35]. Briefly, biopsies were incubated in 75 µl alkaline lysis buffer (25 mM NaOH, 0.2 mM EDTA in H2O) for 30 min at 95 °C and reaction was stopped by adding 75 µl neutralization buffer (40 mM Tris-HCl in H2O). Supernatant containing genomic DNA was subsequently used for genotyping with locus specific primer pairs listed in supplementary table 1 by using genotype-specific cycling programs, as summarized in supplementary table 2.
Cell culture
Primary cells were cultured in RPMI-1640 medium (Sigma-Aldrich, Taufkirchen, Germany), supplemented with 10% fetal calf serum (FCS), 2 mM l-glutamine, 100 U/l penicillin, 0.1 mg/ml streptomycin, 100 mM non-essential amino acids (all gibco®, Thermo Fisher Scientific, Karlsruhe, Germany), 1 mM sodium pyruvate and 50 mM 2-mercaptoethanol (both Sigma Aldrich). T110299 tumor cells had been isolated from tumors of genetically-engineered Ptf1a-Cre KrasG12D p53fl/R172H (KPC) mice and kindly provided by Prof. Siveke (West German Cancer Center (WTZ), University Hospital Essen). T110299 cells were cultured in DMEM high glucose media (Sigma-Aldrich), supplemented with 10% FCS, 2 mM l -glutamine, 100 U/l penicillin and 0.1 mg/ml streptomycin (all gibco®). All cells were kept in a humidified incubator at 37 °C and 5% CO2 and were regularly tested for mycoplasma contamination.
Bone marrow culture
Bone marrow cells were isolated by flushing femur and tibia. 2 × 106 cells per 10 ml were seeded in a 10 cm cell culture round plate in primary cell medium supplemented with 20 ng/ml GM-CSF (Peprotech, London, United Kingdom). After three days, 10 ml primary cell medium supplemented with 20 ng/ml GM-CSF was added. Five days after cell isolation, 66% of the medium containing 20 ng/ml GM-CSF was exchanged. If indicated, cells were stimulated overnight with 10 ng/ml IL-4 and 10 ng/ml IL-13 (both Peprotech, London, United Kingdom).
Orthotopic tumor induction
Orthotopic tumors were induced by surgical implantation, as described before [36]. Briefly, 6–12 weeks old mice were anesthetized, and by surgical incision of the skin and peritoneum, the pancreas was carefully mobilized. After the injection of 2 × 105 T110299 cells in 25 µl PBS, the pancreas was relocated, and the incision was closed by surgical suture. Mice were monitored daily and distressed mice were sacrificed. Tumor weight of sacrificed animals was measured and normalized to average tumor weight of WT animals in the respective experiment.
Cell isolation
Spleens and tumors were isolated from the mice and blood was drawn. Spleens were processed through a 70 µm cell strainer. Erythrocytes from spleen and blood were removed using the red blood cell lysis buffer (BD Biosciences, Heidelberg, Germany). Tumor tissue was minced into pieces and mechanically dissociated using the mouse Tumor Dissociation Kit with the gentleMACS™ Dissociator application (both Miltenyi Biotech, Bergisch Gladbach, Germany), according to the manufacturer´s instructions. The cell suspension was separated from tissue debris by sequentially using 100 µm and 70 µm cell strainers. For functional assays, untouched T cells were isolated using the Pan T cell isolation Kit II, and for MDSC isolation the Myeloid-Derived Suppressor Cell Kit was used (both Miltenyi Biotec). In brief, in a two-step separation process, PMN-MDSC (CD11b+ Ly6Cint Ly6G+) were isolated with anti-Ly6G beads followed by M-MDSC (CD11b+ Ly6Chigh Ly6G−) isolation using anti-Gr1 beads. The purity of isolated cells was > 95% for T cells and between 75 and 90% for MDSC.
Cells from bone morrow cultures were isolated by FACSorting. Cells were stained as described for FACS analysis. FVDnegGr1highMHC-IIlow and FVDnegGr1lowMHC-IIhigh were sorted on a BD Aria III system (BD Bioscience, Heidelberg, Germany).
FACS analysis
Prior to fluorochrome staining of single cell suspensions, FcR III/II blocking was performed using the anti-CD16/CD32 TrueStain fcX™ antibody (BioLegend, London, UK). Dead cells were stained for exclusion with fixable viability dye (FVD) (Thermo Fisher Scientific, Karlsruhe, Germany). For cell-specific surface staining, cells were labeled with CD4 (clone GK1.5), CD8 (clone 53–6.7), CD11b (clone M1/70), CD11c (clone N418), CD45 (clone 30-F11), Gr1 (clone RB6-8C5), Ly6C (clone HK1.4), Ly6G (clone 1A8), MHC-II (clone AF6-120.1; all BioLegend, London, UK). IRF4 (clone IRF4.3E4; BioLegend, London UK) or isotype control (clone RTK2071; both BioLegend, London, UK) were stained intracellularly using the one-step intracellular staining protocol of the eBioscience™ FoxP3/Transcription Factor Staining Buffer Kit (Thermo Fisher Scientific, Karlsruhe, Germany). Samples were acquired on a BD LSRFortessa system (BD Bioscience, Heidelberg, Germany), and data were analyzed with FlowJo X software (FLOWJO LLC, Ashland, OR, USA).
T cell suppression assay
Isolated T cells were stained with 2.5 µM Carboxyfluorescein succinimidyl ester (CFSE; Thermo Fisher Scientific) in PBS for 4 min at room temperature and reaction was stopped with FCS. For the assessment of MDSC suppressive capacity, MDSC were co-cultured with anti-CD3/anti-CD28 stimulated CFSE-labeled T cells. For this, 5 × 104 T cells (per well) were seeded into 96-well plates and cocultured with 0.31 × 104, 0.63 × 104, 1.25 × 104, 2.5 × 104 or 5 × 104 MDSC. Each well was supplemented with 1 µl beads (Dynabeads™ Mouse T-Activator CD3/CD28, gibco®, Thermo Fisher Scientific, Karlsruhe, Germany). After 72 h, CFSE dilution of CD4+ and CD8+ T cells was analyzed by flow cytometry (BD Canto II system, BD Bioscience, Heidelberg, Germany). Unstimulated CFSE-labeled T cells only were used to set the threshold of proliferated T cells (CFSElow).
Human dataset analysis
Survival data, IRF4 expression level as well as information on metastasis status of PDAC patients from the TCGA cohort were retrieved via the Xena browser [37]. Two groups of patients with metastasized PDAC were analyzed: Patients with low IRF4 expression level (≤ the lower quartile) and patients with high IRF4 expression level (≥ the upper quartile). The survival of these two groups was analyzed in a Kaplan–Meier analysis and compared with a Wilcoxon test. The survival status after one year was displayed in a contingency table. As all expected cell frequencies were above five, χ2 test was used to compare one-year survival of the two groups. All analysis on the TCGA data set was performed using IBM® SPSS® Statistics 25. Graphs were generated in Graphpad Prism 8.3.
Histology
Tumors were embedded in Tissue-Tek® O.C.T.™ (Sakura Finetek GmbH, Staufen, Germany), rapidly frozen in liquid nitrogen and stored at − 80 °C. Prior to analysis, samples were thawed, once washed in PBS, fixed in ROTI®Histofix 4% paraformaldehyde (Roth, Karlsruhe, Germany) and subjected to automated routine histological tissue processing. After paraffin embedding, four µm thick whole tissue sections were stained using haematoxylin-eosine (HE) staining (haematoxylin: Waldeck, Münster, Germany; eosine: Sigma) in an automated tissue stainer (Tissue Tek Prisma, Sakura Finetek).
Statistical analysis
Data represent individual mice and are displayed as mean with standard error of the mean (SEM). To test for statistically significant differences between two groups, student’s t test (for expression data only) or Mann–Whitney U test was used. To compare more than two groups, we applied Kruskal–Wallis test followed by Dunn’s multiple comparison post hoc tests between selected samples. To analyze differences in the survival, Mantel–Cox Logrank test was conducted. For the statistical analysis of suppression assays, only the difference between the conditions with the highest concentration of MDSC was analyzed using a Mann–Whitney U test. Statistical analysis of murine data was performed in Graphpad Prism 8.3.
References
Schreiber RD, Old LJ, Smyth MJ (2011) Cancer immunoediting: integrating immunity's roles in cancer suppression and promotion. Science 331(6024):1565–1570. https://doi.org/10.1126/science.1203486
Kleeff J, Beckhove P, Esposito I, Herzig S, Huber PE, Lohr JM, Friess H (2007) Pancreatic cancer microenvironment. Int J Cancer 121(4):699–705. https://doi.org/10.1002/ijc.22871
Bronte V, Brandau S, Chen SH, Colombo MP, Frey AB, Greten TF, Mandruzzato S, Murray PJ, Ochoa A, Ostrand-Rosenberg S, Rodriguez PC, Sica A, Umansky V, Vonderheide RH, Gabrilovich DI (2016) Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat Commun 7:12150. https://doi.org/10.1038/ncomms12150
Si Y, Merz SF, Jansen P, Wang B, Bruderek K, Altenhoff P, Mattheis S, Lang S, Gunzer M, Klode J, Squire A, Brandau S (2019) Multidimensional imaging provides evidence for down-regulation of T cell effector function by MDSC in human cancer tissue. Sci Immunol 4(40):eaaw9159. https://doi.org/10.1126/sciimmunol.aaw9159
Veglia F, Perego M, Gabrilovich D (2018) Myeloid-derived suppressor cells coming of age. Nat Immunol 19(2):108–119. https://doi.org/10.1038/s41590-017-0022-x
Gabitass RF, Annels NE, Stocken DD, Pandha HA, Middleton GWJCI (2011) Elevated myeloid-derived suppressor cells in pancreatic, esophageal and gastric cancer are an independent prognostic factor and are associated with significant elevation of the Th2 cytokine interleukin-13. Cancer Immunol Immunother 60(10):1419. https://doi.org/10.1007/s00262-011-1028-0
Mowbray NG, Griffith D, Hammoda M, Shingler G, Kambal A, Al-Sarireh B (2018) A meta-analysis of the utility of the neutrophil-to-lymphocyte ratio in predicting survival after pancreatic cancer resection. HPB Off J Int Hepato Pancreato Biliary Assoc 20(5):379–384. https://doi.org/10.1016/j.hpb.2017.12.009
Li J, Byrne KT, Yan F, Yamazoe T, Chen Z, Baslan T, Richman LP, Lin JH, Sun YH, Rech AJ, Balli D, Hay CA, Sela Y, Merrell AJ, Liudahl SM, Gordon N, Norgard RJ, Yuan S, Yu S, Chao T, Ye S, Eisinger-Mathason TSK, Faryabi RB, Tobias JW, Lowe SW, Coussens LM, Wherry EJ, Vonderheide RH, Stanger BZ (2018) Tumor cell-intrinsic factors underlie heterogeneity of immune cell infiltration and response to immunotherapy. Immunity 49(1):178–193.e177. https://doi.org/10.1016/j.immuni.2018.06.006
Sanford DE, Porembka MR, Panni RZ, Mitchem JB, Belt BA, Plambeck-Suess SM, Lin G, Denardo DG, Fields RC, Hawkins WG, Strasberg SM, Lockhart AC, Wang-Gillam A, Goedegebuure SP, Linehan DC (2013) A study of zoledronic acid as neo-adjuvant, perioperative therapy in patients with resectable pancreatic ductal adenocarcinoma. J Cancer Ther 4(3):797–803. https://doi.org/10.4236/jct.2013.43096
Morrison AH, Byrne KT, Vonderheide RH (2018) Immunotherapy and prevention of pancreatic cancer. Trends Cancer 4(6):418–428. https://doi.org/10.1016/j.trecan.2018.04.001
Condamine T, Mastio J, Gabrilovich DI (2015) Transcriptional regulation of myeloid-derived suppressor cells. J Leukocyte Biol 98(6):913–922. https://doi.org/10.1189/jlb.4ri0515-204r
Thorn M, Guha P, Cunetta M, Espat NJ, Miller G, Junghans RP, Katz SC (2016) Tumor-associated GM-CSF overexpression induces immunoinhibitory molecules via STAT3 in myeloid-suppressor cells infiltrating liver metastases. Cancer Gene Ther 23(6):188–198. https://doi.org/10.1038/cgt.2016.19
Vasquez-Dunddel D, Pan F, Zeng Q, Gorbounov M, Albesiano E, Fu J, Blosser RL, Tam AJ, Bruno T, Zhang H, Pardoll D, Kim Y (2013) STAT3 regulates arginase-I in myeloid-derived suppressor cells from cancer patients. J Clin Investig 123(4):1580–1589. https://doi.org/10.1172/JCI60083
Marigo I, Bosio E, Solito S, Mesa C, Fernandez A, Dolcetti L, Ugel S, Sonda N, Bicciato S, Falisi E, Calabrese F, Basso G, Zanovello P, Cozzi E, Mandruzzato S, Bronte V (2010) Tumor-induced tolerance and immune suppression depend on the C/EBPbeta transcription factor. Immunity 32(6):790–802. https://doi.org/10.1016/j.immuni.2010.05.010
Yanez A, Ng MY, Hassanzadeh-Kiabi N, Goodridge HS (2015) IRF8 acts in lineage-committed rather than oligopotent progenitors to control neutrophil vs monocyte production. Blood 125(9):1452–1459. https://doi.org/10.1182/blood-2014-09-600833
Holtschke T, Lohler J, Kanno Y, Fehr T, Giese N, Rosenbauer F, Lou J, Knobeloch KP, Gabriele L, Waring JF, Bachmann MF, Zinkernagel RM, Morse HC 3rd, Ozato K, Horak I (1996) Immunodeficiency and chronic myelogenous leukemia-like syndrome in mice with a targeted mutation of the ICSBP gene. Cell 87(2):307–317
Waight JD, Netherby C, Hensen ML, Miller A, Hu Q, Liu S, Bogner PN, Farren MR, Lee KP, Liu K, Abrams SI (2013) Myeloid-derived suppressor cell development is regulated by a STAT/IRF-8 axis. J Clin Invest 123(10):4464–4478. https://doi.org/10.1172/jci68189
Klein U, Casola S, Cattoretti G, Shen Q, Lia M, Mo T, Ludwig T, Rajewsky K, Dalla-Favera R (2006) Transcription factor IRF4 controls plasma cell differentiation and class-switch recombination. Nat Immunol 7(7):773–782. https://doi.org/10.1038/ni1357
Sciammas R, Shaffer AL, Schatz JH, Zhao H, Staudt LM, Singh H (2006) Graded expression of interferon regulatory factor-4 coordinates isotype switching with plasma cell differentiation. Immunity 25(2):225–236. https://doi.org/10.1016/j.immuni.2006.07.009
Huber M, Lohoff M (2014) IRF4 at the crossroads of effector T-cell fate decision. Eur J Immunol 44(7):1886–1895. https://doi.org/10.1002/eji.201344279
Yao S, Buzo BF, Pham D, Jiang L, Taparowsky EJ, Kaplan MH, Sun J (2013) Interferon regulatory factor 4 sustains CD8(+) T cell expansion and effector differentiation. Immunity 39(5):833–845. https://doi.org/10.1016/j.immuni.2013.10.007
Nayar R, Schutten E, Bautista B, Daniels K, Prince AL, Enos M, Brehm MA, Swain SL, Welsh RM, Berg LJ (2014) Graded levels of IRF4 regulate CD8+ T cell differentiation and expansion, but not attrition, in response to acute virus infection (Baltimore, Md: 1950). J Immunol 192(12):5881–5893. https://doi.org/10.4049/jimmunol.1303187
Yamamoto M, Kato T, Hotta C, Nishiyama A, Kurotaki D, Yoshinari M, Takami M, Ichino M, Nakazawa M, Matsuyama T, Kamijo R, Kitagawa S, Ozato K, Tamura T (2011) Shared and distinct functions of the transcription factors IRF4 and IRF8 in myeloid cell development. PLoS ONE 6(10):e25812. https://doi.org/10.1371/journal.pone.0025812
Guilliams M, Dutertre CA, Scott CL, McGovern N, Sichien D, Chakarov S, Van Gassen S, Chen J, Poidinger M, De Prijck S, Tavernier SJ, Low I, Irac SE, Mattar CN, Sumatoh HR, Low GHL, Chung TJK, Chan DKH, Tan KK, Hon TLK, Fossum E, Bogen B, Choolani M, Chan JKY, Larbi A, Luche H, Henri S, Saeys Y, Newell EW, Lambrecht BN, Malissen B, Ginhoux F (2016) Unsupervised high-dimensional analysis aligns dendritic cells across tissues and species. Immunity 45(3):669–684. https://doi.org/10.1016/j.immuni.2016.08.015
Sichien D, Scott Charlotte L, Martens L, Vanderkerken M, Van Gassen S, Plantinga M, Joeris T, De Prijck S, Vanhoutte L, Vanheerswynghels M, Van Isterdael G, Toussaint W, Madeira Filipe B, Vergote K, Agace William W, Clausen Björn E, Hammad H, Dalod M, Saeys Y, Lambrecht Bart N, Guilliams M (2016) IRF8 transcription factor controls survival and function of terminally differentiated conventional and plasmacytoid dendritic cells. Respect Immun 45(3):626–640. https://doi.org/10.1016/j.immuni.2016.08.013
Briseno CG, Haldar M, Kretzer NM, Wu X, Theisen DJ, Kc W, Durai V, Grajales-Reyes GE, Iwata A, Bagadia P, Murphy TL, Murphy KM (2016) Distinct transcriptional programs control cross-priming in classical and monocyte-derived dendritic cells. Cell Rep 15(11):2462–2474. https://doi.org/10.1016/j.celrep.2016.05.025
El Chartouni C, Schwarzfischer L, Rehli M (2010) Interleukin-4 induced interferon regulatory factor (Irf) 4 participates in the regulation of alternative macrophage priming. Immunobiology 215(9):821–825. https://doi.org/10.1016/j.imbio.2010.05.031
Satoh T, Takeuchi O, Vandenbon A, Yasuda K, Tanaka Y, Kumagai Y, Miyake T, Matsushita K, Okazaki T, Saitoh T, Honma K, Matsuyama T, Yui K, Tsujimura T, Standley DM, Nakanishi K, Nakai K, Akira S (2010) The Jmjd3-Irf4 axis regulates M2 macrophage polarization and host responses against helminth infection. Nat Immunol 11(10):936–944. https://doi.org/10.1038/ni.1920
Nam S, Kang K, Cha JS, Kim JW, Lee HG, Kim Y, Yang Y, Lee MS, Lim JS (2016) Interferon regulatory factor 4 (IRF4) controls myeloid-derived suppressor cell (MDSC) differentiation and function. J Leukocyte Biol. https://doi.org/10.1189/jlb.1a0215-068rr
Clark CE, Hingorani SR, Mick R, Combs C, Tuveson DA, Vonderheide RH (2007) Dynamics of the immune reaction to pancreatic cancer from inception to invasion. Cancer Res 67(19):9518–9527. https://doi.org/10.1158/0008-5472.Can-07-0175
Metzger P, Kirchleitner SV, Kluge M, Koenig LM, Hörth C, Rambuscheck CA, Böhmer D, Ahlfeld J, Kobold S, Friedel CC, Endres S, Schnurr M, Duewell P (2019) Immunostimulatory RNA leads to functional reprogramming of myeloid-derived suppressor cells in pancreatic cancer. J Immuno Therapy Cancer 7(1):288. https://doi.org/10.1186/s40425-019-0778-7
Abram CL, Roberge GL, Hu Y, Lowell CA (2014) Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J Immunol Methods 408:89–100. https://doi.org/10.1016/j.jim.2014.05.009
Hasenberg A, Hasenberg M, Mann L, Neumann F, Borkenstein L, Stecher M, Kraus A, Engel DR, Klingberg A, Seddigh P, Abdullah Z, Klebow S, Engelmann S, Reinhold A, Brandau S, Seeling M, Waisman A, Schraven B, Gothert JR, Nimmerjahn F, Gunzer M (2015) Catchup: a mouse model for imaging-based tracking and modulation of neutrophil granulocytes. Nat Methods 12(5):445–452. https://doi.org/10.1038/nmeth.3322
Gabrilovich DI, Ostrand-Rosenberg S, Bronte V (2012) Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol 12:253. https://doi.org/10.1038/nri3175
Truett GE, Heeger P, Mynatt RL, Truett AA, Walker JA, Warman ML (2000) Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT). BioTechniques 29(1):52–54. https://doi.org/10.2144/00291bm09
Ellermeier J, Wei J, Duewell P, Hoves S, Stieg MR, Adunka T, Noerenberg D, Anders HJ, Mayr D, Poeck H, Hartmann G, Endres S, Schnurr M (2013) Therapeutic efficacy of bifunctional siRNA combining TGF-beta1 silencing with RIG-I activation in pancreatic cancer. Cancer Res 73(6):1709–1720. https://doi.org/10.1158/0008-5472.CAN-11-3850
Goldman M, Craft B, Hastie M, Repečka K, McDade F, Kamath A, Banerjee A, Luo Y, Rogers D, Brooks AN, Zhu J, Haussler D (2019) The UCSC Xena platform for public and private cancer genomics data visualization and interpretation. BioRxiv. https://doi.org/10.1101/326470
Acknowledgements
Open Access funding provided by Projekt DEAL. We thank PD Dr. Michael Hristov (Institute for Cardiovascular Prevention (IPEK), University Hospital, LMU Munich) for performing the FACSorting of MDSC and we acknowledge the iFlow Core Facility of the University Hospital, LMU Munich for assistance with the generation of flow cytometry data.
Funding
This work was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Project Number 315563603 to M.S. and P.D., Project Number 329628492 – SFB 1321 – to K.L. and M.S., project number 238187445 – SFB1123/2 – to P.D., Project Number 385829722 to M.L. and project number 405344257 – KFO 337 – to M.G., the international doctoral program “i-Target: Immunotargeting of cancer” funded by the Elite Network of Bavaria to M.S. and L.M.K. as well as the Marie-Sklodowska-Curie Training Network for the Immunotherapy of Cancer (IMMUTRAIN, Grant Number 641549) funded by the H2020 program of the European Union to S.E., M.S. and L.M.K. and the National Science Center, Poland (Grant no 2016/23/B/NZ6/00086) to M.L. as well as the Mildred-Scheel- Doktorandenprogramm funded by the Deutsche Krebshilfe to S.V.K. and M.S. and the Förderprogramm für Forschung und Lehre (FöFoLe) funded by the Ludwig-Maximilians Universität München to P.D. and M.S.
Author information
Authors and Affiliations
Contributions
Conceptualization: PM, PD, MS, LMK, Formal Analysis: PM, AE, Funding acquisition: ML, KL, SE, PD, MS, LMK, Investigation: PM, SVK, DB, CH, SO, Resources: ML, MG, Supervision: ML, KL, SE, PD, MS, LMK, Writing-original draft: PM, Writing-review and editing: PM, MS, LMK, All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
This work has been performed for the doctoral thesis of P.M. at the Ludwig-Maximilians-Universität München.
Ethical approval
Experiments were performed according to national ethical guidelines approved by the local government (Regierung von Oberbayern, Munich, Germany; file number 55.2-1-54-2532-175-12).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
262_2020_2605_MOESM1_ESM.pdf
Supplementary Figure S1:Histology of T110299 tumors from wild-type and Irf4-/- mice displays no apparent morphologic differences. Representative sections of orthotopic T110299 tumors (two tumors per genotype) stained with haematoxylin and eosin (H&E) are depicted. Scale bars indicates 100 μm. (PDF 56572 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Metzger, P., Kirchleitner, S.V., Boehmer, D.F.R. et al. Systemic but not MDSC-specific IRF4 deficiency promotes an immunosuppressed tumor microenvironment in a murine pancreatic cancer model. Cancer Immunol Immunother 69, 2101–2112 (2020). https://doi.org/10.1007/s00262-020-02605-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02605-9