Abstract
Purpose
The relentless rise in antimicrobial resistance is a major societal challenge and requires, as part of its solution, a better understanding of bacterial colonization and infection. To facilitate this, we developed a highly efficient no-wash red optical molecular imaging agent that enables the rapid, selective, and specific visualization of Gram-positive bacteria through a bespoke optical fiber–based delivery/imaging endoscopic device.
Methods
We rationally designed a no-wash, red, Gram-positive-specific molecular imaging agent (Merocy-Van) based on vancomycin and an environmental merocyanine dye. We demonstrated the specificity and utility of the imaging agent in escalating in vitro and ex vivo whole human lung models (n = 3), utilizing a bespoke fiber–based delivery and imaging device, coupled to a wide-field, two-color endomicroscopy system.
Results
The imaging agent (Merocy-Van) was specific to Gram-positive bacteria and enabled no-wash imaging of S. aureus within the alveolar space of whole ex vivo human lungs within 60 s of delivery into the field-of-view, using the novel imaging/delivery endomicroscopy device.
Conclusion
This platform enables the rapid and specific detection of Gram-positive bacteria in the human lung.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gram-positive bacteria are responsible for a vast range of serious clinical pathologies, including infections of wounds, colonization of indwelling medical devices including catheters and prosthetic joints, and ventilator-associated pneumonia (VAP) [1,2,3]. Such infections result in significant morbidity, mortality and healthcare costs meaning that accurate and rapid diagnosis of these infections is critical for the initiation of appropriate treatment [4]. Traditional diagnostic practices for infection rely on bacterial culture, which may take several days to yield a result or techniques such as the polymerase chain reaction (PCR), which have a tendency to be overly sensitive [5, 6].
Developments in optical technologies [7, 8] have opened new avenues for point-of-care diagnosis of disease; however, in the area of bacterial infection, success has been limited by a lack of targeted imaging agents with the required sensitivity, specificity, and compatibility [9, 10]. Gram-positive species such as Staphylococcus aureus and Streptococcus spp. are responsible for approximately one-third of all VAP in the intensive care unit. Current diagnostic strategies for VAP rely on the interpretation of non-specific clinical symptoms, meaning that half of patients receive an incorrect diagnosis and one-third of cases are missed [3, 11]. Thus, a VAP diagnostic strategy could benefit from technological advances offered by optical endomicroscopy and suitable molecular imaging agents [12, 13].
Here we describe the rational development and characterization of a highly specific, selective, rapidly reporting fluorescent glycopeptide-based imaging agent that has no requirement for a wash or processing step, and demonstrate its utility in detecting Gram-positive bacteria within seconds in a whole explanted human lung. This was achieved using a novel disposable endoscopic delivery/imaging device comprising two delivery capillaries and an optical fiber imaging bundle, allowing the imaging agent to be delivered (< 100 μg) directly into the field-of-view within the alveolar space.
The Gram-positive optical molecular imaging agent was rationally designed based on the clinically utilized glycopeptide antibiotics vancomycin and oritavancin—with the knowledge that oritavancin is produced by reductive amination (with hydrophobic groups) of the natural product chloroeremomycin (an analogue of vancomycin). This gives analogues with far greater activity than chloroeremomycin or vancomycin themselves due to the duality of binding modes—the hydrophobic anchor binding into and disrupting the cell membrane and the classic glycopeptide inhibition of transpeptidation, whilst retaining specificity towards Gram-positive bacteria [14,15,16,17].
Thus, our rationale was that the incorporation of an environmentally sensitive hydrophobic dye (merocyanine) by similar chemistry would give highly selective imaging probes (similar in function to oritavancin). Additionally, this chemistry would allow not only selective attachment at a known single site on vancomycin, but it would also leave behind an amino group, thus enhancing solubility and specificity. The merocyanine core was modified with a benzaldehyde type moiety to further mimic the chemistry of chloroeremomycin conversion to oritavancin, which uses an arylaldehyde.
The environmentally sensitive dye utilized was based on merocyanine, which comprises electron donor (D) and acceptor (A) components connected by double bonds (the so-called D-π-A system). This makes the dye very sensitive to changes in polarity and viscosity, while also displaying a high extinction coefficient and quantum yield; thus, it fluoresces brightly in hydrophobic environments (including the bacterial membrane) [18,19,20,21].
Merocy-Van is the first example of a red-shifted, environmentally activated bacterial imaging agent, and it demonstrates superior specificity and binding kinetics to Gram-positive bacteria compared with previously reported targeting agents [10, 22]. Moreover, these compounds also had “always-on” fluorophores, precluding them from topical application and rapid bedside imaging. Furthermore, the excitation and emission in the “red” shifts the dye away from the classical green autofluorescence of tissue, making it a promising candidate for imaging applications.
The imaging fiber was fabricated from optical fiber preforms [23] and was subsequently co-packaged with two glass capillaries into a Pebax® outer layer and a polyimide inner layer with a steel braid reinforcement. The distal end was terminated with a stainless steel end cap, and a fiber optic connector at the proximal end for connectivity to the imaging system [24].
In this study, the topical application of Merocy-Van via the capillary channels of our endoscopy device provided a novel delivery approach into ex vivo human lungs and was performed simultaneously with two-color endomicroscopy imaging. The imaging agent delivery and two-color imaging capability enabled both structural and functional information to be collected and exemplifies significant advances in bacterial optical imaging agent design, and in optical endomicroscopy.
Results
Synthesis of a Gram-positive specific imaging agent
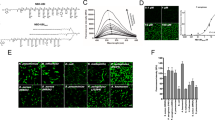
A benzaldehyde-modified merocyanine was synthesized in three steps (Supporting Information Scheme S1) starting from commercially available 2,3,3-trimethyl-3H-indolenine, which was alkylated with 4-(chloromethyl)benzyl alcohol. Microwave-assisted condensation [20] of the acceptor (indolenine) with the benzothiophene donor gave the benzyl alcohol–modified merocyanine, which was then oxidized (using Dess-Martin periodinane) to give the merocyanine-aldehyde (Merocy-Ald). The imaging agent (Merocy-Van) was generated via reductive amination chemistry [17] of Merocy-Ald via the vancosamine side chain of vancomycin, and purified by reverse phase HPLC (Fig. 1). It proved, as designed, to be environmentally sensitive with an exmax and emmax of 590 nm and 620 nm (Fig. 1d).
Synthesis and characterization of the environmentally responsive imaging agent. a Synthesis of Merocy-Van by reductive amination of vancomycin with the aldehyde dye. b HPLC trace of the purified imaging agent (detection at 600 nm). c HRMS spectra of Merocy-Van (experimental trace (top) vs theoretical (bottom)). d Absorbance (dashed trace) and emission spectra of the imaging agent with increasing levels of DMSO (DMSO/saline, vol/vol)
Merocy-Van demonstrated selectivity for Gram-positive bacteria
Selectivity for Gram-positive bacteria was assessed against a panel of clinically relevant Gram-positive and Gram-negative bacteria, and freshly isolated human peripheral blood granulocytes and mononuclear cells. Merocy-Van demonstrated rapid bacterial staining of encapsulated (S. pneumoniae) and non-encapsulated Gram-positive bacteria with high signal-to-noise in a concentration-dependent manner (50% max intensity at 0.1 μM) without the need to remove unbound excess probe or requiring a washing procedure (Fig. 2 and Fig. S1). Importantly, Merocy-Van was also able to label MRSA within biofilms of depths of > 10 μM (Fig. S2). Merocy-Van labelling of Gram-positive bacteria was fully orthogonal to the green Gram-negative imaging agent NBD-PMX (PMX (polymyxin)) [13] (Fig. 2c).
Merocy-Van selectively labels Gram-positive bacteria. a S. aureus, MRSA, S. epidermidis, S. pneumoniae, E. coli, P. aeruginosa, and S. aureus/mononuclear cell (PBMC) or granulocyte (PMN) co-cultures were imaged by confocal microscopy following the addition of 5 μM probe (magenta). Image inserts show counterstained bacteria/cells (green). Representative images from nine fields-of-view from three independent repeats. Scale bar = 5 μm. b Quantification of images from (a) (relative fluorescence intensity from n = 90 bacteria, n = 20 mononuclear cell, and n = 20 granulocyte per condition, one-way ANOVA ****P < 0.0001). Dashed line shows background fluorescence. c Wash-free confocal imaging of Gram-negative (green) and Gram-positive (magenta) bacteria with NBD-PMX and Merocy-Van. Primary isolated human mononuclear cells were pre-labeled with Hoechst prior to adding to the confocal chamber. Scale bar = 5 μm, n = 3
Merocy-Van retained Gram-positive toxicity but was non-cytotoxic
The probe, as expected for oritavancin, caused complete growth inhibition of S. aureus and S. pneumoniae at 1 μM while the growth rate of Escherichia coli remained unaffected (Fig. S3). Merocy-Van was non-toxic and non-hemolytic to eukaryotic cells (Fig. S4), an important pre-requisite for clinical translation.
Fiber development and performance
The three-in-one device was fabricated using an imaging fiber (8100 cores with an imaging square diagonal of 450 μm and a center-to-center core separation of 3.5 μm) and two fluid delivery channels formed from a fused silica glass tube drawn down to a 410-μm capillary (inner diameter of 326 μm). The imaging fiber was fabricated from optical fiber preforms produced for the telecommunications industry (Draka/Prysmian OM1 multi-mode preform) and showed improved performance in terms of imaging contrast with no compromise in resolution over conventional imaging fibers formed from significantly more expensive starting glasses [23]. Both the imaging fiber and the capillary fibers were coated, as they were drawn, with a UV curable acrylate polymer to protect the outer surfaces from damage. Biocompatible tubing (Pebax® outer layer and a polyimide inner layer with a steel braid reinforcement) was used to co-package the three individual fibers (2 × capillaries, 1 × imaging fiber), and the three fibers were fixed in place at the distal end inside a 5-mm long stainless steel end cap with an epoxy (Epotek 301). An epoxy fiber splitter was formed at the proximal end (Epotek 301). After the splitter, the three fibers were individually sheathed in the Pebax® tubing; the two fluid channels were terminated with Luer lock fittings and the imaging fiber was terminated with a large bore fiber optic connector (Fig. 3). The delivery system is intended to deliver small boluses to alveolar regions of less than 0.5 cm3 for point imaging/sampling. This was achieved, with the fluid channels able to readily deliver 0.5 mL of fluid from a 2-mL syringe in under 20 s in a continuous flow.
Merocy-Van selectively labels S. aureus in an ex vivo human lung model. a The endomicroscopy fluorescence imaging system used. b The packaged three-in-one fiber-based endomicroscopy device. c End of view of the distal tip of the fiber-based endomicroscopy device. The outer diameter of the device is 1.4 mm, and the optical imaging bundle field-of-view is 450 μm (right of the image) and the internal diameter of the glass capillaries is 326 μm. d Image of the imaging fiber (8100 cores). e Diagram of a human lung and infection/imaging locations used. f Image of an ex vivo human lung undergoing ventilation with the imaging device inserted within the distal lung (blue light within the white circle). Insert shows blow-up of device distal end location (blue light). g Representative images of real-time Merocy-Van (1 μM) delivery into human ex vivo lung (baseline and following 0.4 nmol delivery shown) within S. aureus (top) and control (bottom) lung lobes. Green: capturing lung autofluorescence. Red: capturing activated Merocy-Van. Each large panel shows two-color overlay; insert shows red channel only. Scale bar = 50 μm. h Average red fluorescence intensity per frame during real-time delivery of 0.4 nmol Merocy-Van (1 μM) into S. aureus and control lung lobes (data shown from lung 1, see Fig. S5 for lungs 2 and 3)
Delivery of Merocy-Van enabled rapid visualization of Gram-positive bacterial in an ex vivo human lung model
Merocy-Van and the optical fiber–based endoscope were evaluated using an explanted ventilated human lung model. S. aureus was instilled into the right lower lobe (a common area of the lung for VAP to occur [25]), and Pseudomonas aeruginosa was instilled into anatomically distinct lobes as a negative control (Fig. 3e and f). The bacterial inoculum instilled correlated to a bronchoalveolar lavage (BAL) retrieval of ~ 104 colony forming units (CFU) mL−1, in line with clinical thresholds for pneumonia diagnosis [12, 26]. Optical endomicroscopy imaging was performed with a wide-field imaging system [24] with appropriate LED and filter sets for the excitation/detection of tissue autofluorescence (green (488 nm, 500–550 nm)) and activated Merocy-Van (red (590 nm, 615–675 nm)) (Fig. 3a). Imaging was simultaneously captured in the two-color channels, with 500 μL of Merocy-Van (1 μM) delivered down the glass capillaries of the optical fiber–based endoscope into the imaging field-of-view. Videos were captured throughout the Merocy-Van instillation process to enable real-time, in situ visualization of probe activation.
In all three individual ex vivo lung experiments, a clear increase red fluorescence was observed for S. aureus lung segments following dosing of Merocy-Van (average 20-fold increase within 60 s compared with control lobes) (Fig. 3g and h, and Fig. S5 and S6, and Supporting Information Movies S1, S2, S3). The relative intensity collected from the green channel remained constant throughout. Following the imaging procedure, a BAL was performed and the retrieved sample plated for CFU determination. S. aureus was only retrieved from the S. aureus lobes, and for each of the three lungs, 104–105 CFU mL−1 was recovered.
(MP4 5438 kb).
(MP4 7132 kb).
(MP4 8652 kb).
Discussion
We have demonstrated the rational design of a red-shifted, environmental, highly selective optical molecular imaging agent to enable real-time detection of Gram-positive bacteria using a dual-color molecular alveoscopy approach. The probe Merocy-Van displayed Gram-positive specificity, no detectable cytotoxicity, and was able to generate high signal-to-noise within seconds of administration into the human distal lung with clinically relevant levels of Gram-positive bacteria. Merocy-Van was dosed directly into the imaging field-of-view without requiring a wash-step, enabling both structural (alveolar location) and functional (Gram-positive bacteria) information to be discerned in real time. Merocy-Van showed superior specificity and binding kinetics to Gram-positive bacteria compared with any other previously reported agents [10, 22]. The choice of fluorophore was critical—many fluorophores carry a surfeit of negative charge or are always “on”—which can inhibit the binding ligand from interacting with fidelity to its target, or generate a high background signal [10]. Our approach to the development of Merocy-Van demonstrates a significant advancement in this area and means that the probe could be utilized in a range of other diagnostic applications.
We evaluated Merocy-Van in a human lung model. The three-in-one optical fiber–based endoscope allows for optical imaging to be proximal to chemical probe delivery as well as providing a capillary for sample removal. Our approach showed it was sensitive and robust, with the alveoscopy procedure performed by three independent clinicians within this study, and offers advantages over conventional lavage processing because the detection and results are presented directly in real time at the site of infection.
Whilst the lung is a large organ and it is not feasible to sample each lobe or sub-section with our fiber-based technique, the clinician can be guided to the bronchopulmonary region of interest based on x-ray or CT scans for fiber-based interrogation and probe delivery, or select the posterior segment of the right lower lobe if changes were widespread, as has previously been demonstrated [13]. The initial clinical application is intended for the bedside evaluation of respiratory deterioration and new pulmonary infiltrates in intubated and ventilated patients in critical care. In these patients, with suspected VAP which has a high morbidity and mortality, antibiotic stewardship is important, yet often broad spectrum antibiotics are prescribed without microbial confirmation of distal lung infection. The intended clinical utility is to provide a high negative predictive value to excluding distal alveolar infection due to Gram-positive bacteria with a threshold to delineate low-grade colonization from de novo infection. The imaging will be performed in a standardized manner, and the technology now requires clinical validation in prospective studies to determine performance and health economic benefit as an adjunct to existing care pathways.
Beyond these initial clinical indications, our approach also allows access to the distal lung with targeted sampling of discrete regions; this has the potential to aid in microbiome research as traditional processing of lavage or tracheal aspirates are not necessarily indicative of the area of interest due to cross-contamination [27, 28]; however, these applications are beyond the scope of this study.
Conclusion
In conclusion, we have presented a strategy for the in situ detection of Gram-positive bacteria in a whole human ex vivo ventilated lung, paving the way for simultaneous, multiplexed, multicolor imaging in vivo in real time. Combining optical molecular imaging agents and our three-in-one optical fiber–based endoscope constructs could aid point-of-care diagnostic decisions and provide a platform to interrogate dynamic biological processes in situ.
Methods
Chemistry and biology reagents and methods
Chemistry and biology reagents and methods are described in detail in the Supporting Information.
Ethical approval
All experiments using human samples were performed following approval of the appropriate regional ethics committee (REC), NHS Lothian, and the South East Scotland Research Ethics Committee 02 (references 15/HV/013 and 11/SS/0103), and with informed consent. Experiments were conducted within three explanted human lungs (Fig. 3 and Fig. S5 and S6, and Supporting Information Movies S1, S2, S3).
Optical imaging agent characterization
All imaging agents were solubilized in 0.9% NaCl (used at concentrations of 1 μM or 5 μM). Absorbance and emission spectra were determined on a microplate reader (BioTek Synergy H1 multi-mode reader). Data were collected in duplicate from three independent repeats.
Human explanted lung model
Human lungs were obtained from solid organ donors after being declined by all UK transplant centers as being unsuitable for transplantation. They were flushed in situ and retrieved as per standard clinical practice for organ retrieval and transported on ice. Lungs were split and single lung ventilation (Drager Savina 300) performed using PEEP 5, tidal volume of 3.5 mL kg−1 and a respiratory rate of 12. Following recruitment of the lungs, the bronchoscope was wedged proximally in each lobe, and 2 mL of 108 CFU mL−1 S. aureus was instilled into the lower lobe, and 2 mL of 108 CFU mL−1 P. aeruginosa or 2 mL 0.9% NaCl were instilled into the upper or middle lobes as negative controls via a flexible catheter (1.5-mm-diameter APC catheter, Erbe). Following a 60-min incubation, the endoscopic device was passed down the working channel of a bronchoscope to the lobe where saline or bacteria had been instilled, and into the alveolar space. This was confirmed by imaging in the green channel. Five hundred microliters of 1 μM Merocy-Van was instilled into the lung through the capillaries of the endoscopic device within the field-of-view, in 5 × 100 μL (5 × 0.1 nmol) installments. Dual-color videos were captured throughout each instillation of Merocy-Van. Images were captured as previously described [24] with 50-ms exposure and a frame rate of 8 frames per second. The endoscopic device (including internally within the delivery capillaries) was cleaned with 8% H2O2 between imaging each lobe. Following all imaging, each lobe underwent BAL with 20 mL 0.9% NaCl (Baxter), with a retrieval rate of 40–50%. BAL samples were serially diluted and plated for CFU counting to confirm bacterial presence from each lobe. The study was completed using lungs from three individual donors.
Human explanted lung model image analysis
Bespoke image analysis tools were developed in-house. To extract fluorescence intensity levels, MATLAB scripts were written that detect and mark the area of the fiber facet as the region of interest (ROI) of an image. The script subsequently removes the background signal (acquired in a calibration step during every experiment) and, in a second step, computes the mean fluorescence intensity levels of the ROI of every image of a given set. Off target frames were removed from analysis.
For display purposes, the image processing we implemented was a method similar to one previously reported [29]. Our implementation first subtracts fiber bundle autofluorescence, then removes high spatial frequency noise from the image, and then corrects for fiber core transmission heterogeneity through normalization. Each video file was brightness- and contrast-enhanced (maintaining the same settings per experiment) and exported to create real-time imaging videos. Individual representative images from each video file were saved as TIFs.
Data availability
Data used within this publication can be accessed at https://doi.org/10.7488/ds/2813.
References
Bessa LJ, Fazii P, Di Giulio M, Cellini L. Bacterial isolates from infected wounds and their antibiotic susceptibility pattern: some remarks about wound infection. Int Wound J. 2015;12:47–52. https://doi.org/10.1111/iwj.12049.
Huotari K, Peltola M, Jamsen E. The incidence of late prosthetic joint infections: a registry-based study of 112,708 primary hip and knee replacements. Acta Orthop. 2015;86:321–5. https://doi.org/10.3109/17453674.2015.1035173.
Kalanuria AA, Zai W, Mirski M. Ventilator-associated pneumonia in the ICU. Crit Care. 2014;18:208. https://doi.org/10.1186/cc13775.
Jasovský D, Littmann J, Zorzet A, Cars O. Antimicrobial resistance—a threat to the world’s sustainable development. Ups J Med Sci. 2016;121:159–64. https://doi.org/10.1080/03009734.2016.1195900.
Bousbia S, Papazian L, Saux P, Forel JM, Auffray J-P, Martin C, et al. Repertoire of intensive care unit pneumonia microbiota. PLoS One. 2012;7:e32486. https://doi.org/10.1371/journal.pone.0032486.
Gadsby NJ, McHugh MP, Forbes C, MacKenzie L, Hamilton SKD, Griffith DM, et al. Comparison of Unyvero P55 Pneumonia Cartridge, in-house PCR and culture for the identification of respiratory pathogens and antibiotic resistance in bronchoalveolar lavage fluids in the critical care setting. Eur J Clin Microbiol Infect Dis. 2019;38:1171–8. https://doi.org/10.1007/s10096-019-03526-x.
Joshi BP, Wang TD. Gastrointestinal imaging in 2015: emerging trends in endoscopic imaging. Nat Rev Gastroenterol Hepatol. 2016;13:72–3. https://doi.org/10.1038/nrgastro.2015.214.
Yun SH, Kwok SJJ. Light in diagnosis, therapy and surgery. Nat Biomed Eng. 2017;1. https://doi.org/10.1038/s41551-016-0008.
van Oosten M, Hahn M, Crane LMA, Pleijhuis RG, Francis KP, van Dijl JM, et al. Targeted imaging of bacterial infections: advances, hurdles and hopes. FEMS Microbiol Rev. 2015;39:892–916. https://doi.org/10.1093/femsre/fuv029.
Mills B, Bradley M, Dhaliwal K. Optical imaging of bacterial infections. Clin Transl Imaging. 2016;4:163–74. https://doi.org/10.1007/s40336-016-0180-0.
Rubinstein E, Kollef MH, Nathwani D. Pneumonia caused by methicillin-resistant Staphylococcus aureus. Clin Infect Dis. 2008;46:S378–S85. https://doi.org/10.1086/533594.
Akram AR, Avlonitis N, Lilienkampf A, Perez-Lopez AM, McDonald N, Chankeshwara SV, et al. A labelled-ubiquicidin antimicrobial peptide for immediate in situ optical detection of live bacteria in human alveolar lung tissue. Chem Sci. 2015;6:6971–9. https://doi.org/10.1039/c5sc00960j.
Akram AR, Chankeshwara SV, Scholefield E, Aslam T, McDonald N, Megia-Fernandez A, et al. In situ identification of Gram-negative bacteria in human lungs using a topical fluorescent peptide targeting lipid A. Sci Transl Med. 2018;10:eaal0033. https://doi.org/10.1126/scitranslmed.aal0033.
Cooper RD, Snyder NJ, Zweifel MJ, Staszak MA, Wilkie SC, Nicas TI, et al. Reductive alkylation of glycopeptide antibiotics: synthesis and antibacterial activity. J Antibiot (Tokyo). 1996;49:575–81.
Kim SJ, Tanaka KS, Dietrich E, Rafai Far A, Schaefer J. Locations of the hydrophobic side chains of lipoglycopeptides bound to the peptidoglycan of Staphylococcus aureus. Biochemistry. 2013;52:3405–14. https://doi.org/10.1021/bi400054p.
Butler MS, Hansford KA, Blaskovich MAT, Halai R, Cooper MA. Glycopeptide antibiotics: back to the future. J Antibiot. 2014;67:631–44. https://doi.org/10.1038/ja.2014.111.
Guan D, Chen F, Xiong L, Tang F, Faridoon, Qiu Y, et al. Extra sugar on vancomycin: new analogues for combating multidrug-resistant Staphylococcus aureus and vancomycin-resistant Enterococci. J Med Chem. 2018;61:286–304. https://doi.org/10.1021/acs.jmedchem.7b01345.
Toutchkine A, Kraynov V, Hahn K. Solvent-sensitive dyes to report protein conformational changes in living cells. J Am Chem Soc. 2003;125:4132–45. https://doi.org/10.1021/ja0290882.
Kulinich AV, Ishchenko AA. Merocyanine dyes: synthesis, structure, properties and applications. Russ Chem Rev. 2009;78:141–64. https://doi.org/10.1070/RC2009v078n02ABEH003900.
MacNevin CJ, Gremyachinskiy D, Hsu CW, Li L, Rougie M, Davis TT, et al. Environment-sensing merocyanine dyes for live cell imaging applications. Bioconjug Chem. 2013;24:215–23. https://doi.org/10.1021/bc3005073.
Escobedo JO, Rusin O, Lim S, Strongin RM. NIR dyes for bioimaging applications. Curr Opin Chem Biol. 2010;14:64–70. https://doi.org/10.1016/j.cbpa.2009.10.022.
van Oosten M, Schafer T, Gazendam JA, Ohlsen K, Tsompanidou E, de Goffau MC, et al. Real-time in vivo imaging of invasive- and biomaterial-associated bacterial infections using fluorescently labelled vancomycin. Nat Commun. 2013;4:2584. https://doi.org/10.1038/ncomms3584.
Stone JM, Wood HAC, Harrington K, Birks TA. Low index contrast imaging fibers. Opt Lett. 2017;42:1484–7. https://doi.org/10.1364/OL.42.001484.
Krstajić N, Mills B, Murray I, Marshall A, Norberg D, Craven TH, et al. Low-cost high sensitivity pulsed endomicroscopy to visualize tricolor optical signatures. J Biomed Opt. 2018;23:1–12.
Koenig SM, Truwit JD. Ventilator-associated pneumonia: diagnosis, treatment, and prevention. Clin Microbiol Rev. 2006;19:637–57. https://doi.org/10.1128/CMR.00051-05.
Miller PR, Meredith JW, Chang MC. Optimal threshold for diagnosis of ventilator-associated pneumonia using bronchoalveolar lavage. J Trauma. 2003;55:263–7; discussion 7-8. https://doi.org/10.1097/01.TA.0000075786.19301.91.
Cabrera-Rubio R, Garcia-Núñez M, Setó L, Antó JM, Moya A, Monsó E, et al. Microbiome diversity in the bronchial tracts of patients with chronic obstructive pulmonary disease. J Clin Microbiol. 2012;50:3562–8. https://doi.org/10.1128/jcm.00767-12.
Moffatt MF, Cookson WO. The lung microbiome in health and disease. Clin Med. 2017;17:525–9. https://doi.org/10.7861/clinmedicine.17-6-525.
Vercauteren T, Ayache N, Savoire N, Malandain G, Perchant A. Processing of in vivo fibered confocal microscopy video sequences. In: Rittscher J, Machiraju R, Wong STC, editors. Microscopic image analysis for life science applications: Artech House; 2008. p. 441–63.
Acknowledgments
We would like to thank the CALM imaging facilities at the Queens Medical Research Institute, University of Edinburgh, Warren’s Wish (Registered Scottish Charity: SC 045290) and the donor’s and their families.
Funding
Open Access funding provided by University of Edinburgh. We would like to thank Engineering and Physical Sciences Research Council (EPSRC, United Kingdom) Interdisciplinary Research Collaboration grant EP/K03197X/1 and EP/R005257/1.
Author information
Authors and Affiliations
Contributions
JMS, MU, and BM prepared the manuscript. MU and AMF designed, prepared, and characterized the imaging agent. BM, SD, AM, AA, TQ, and IR designed and performed all biological experiments. ES and AMB developed the tissue collection pathway. DN developed the real-time image processing tools to the endomicroscopy device. JMS, TC, HACW, KH, MGT, HP, TAB, and JCK were responsible for the device development and testing. GW and NK built and modified the imaging device. MB and KD supervised the project. All authors contributed to manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Ethics approval
All experiments using human samples were performed following approval of the appropriate regional ethics committee (REC), NHS Lothian, and the South East Scotland Research Ethics Committee 02 (references 15/HV/013 and 11/SS/0103), and with informed consent.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Infection and inflammation
Electronic supplementary material
ESM 1
(DOCX 10.2 mb).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mills, B., Megia-Fernandez, A., Norberg, D. et al. Molecular detection of Gram-positive bacteria in the human lung through an optical fiber–based endoscope. Eur J Nucl Med Mol Imaging 48, 800–807 (2021). https://doi.org/10.1007/s00259-020-05021-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-020-05021-4