Abstract
Glycoside Hydrolase family 65 (GH65) is a unique family of carbohydrate-active enzymes. It is the first protein family to bring together glycoside hydrolases, glycoside phosphorylases and glycosyltransferases, thereby spanning a broad range of reaction types. These enzymes catalyze the hydrolysis, reversible phosphorolysis or synthesis of various α-glucosides, typically α-glucobioses or their derivatives. In this review, we present a comprehensive overview of the diverse reaction types and substrate specificities found in family GH65. We describe the determinants that control this remarkable diversity, as well as the applications of GH65 enzymes for carbohydrate synthesis.
Key points
• GH65 is the first CAZy family to contain hydrolases, phosphorylases and transferases
• Distinct residues and loops are determinants of substrate specificity in family GH65
• GH65 enzymes hold strong potential for carbohydrate synthesis via coupled reactions
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
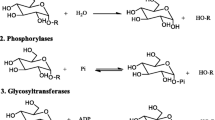
Glycoside Hydrolase family 65 (GH65) is an enzyme family in the carbohydrate-active enzymes database (CAZy, www.cazy.org) (Drula et al. 2022) and contains several α-glycoside hydrolases (GHs), phosphorylases (GPs) and α-glycosyltransferases (GTs) (Fig. 1a; Table 1) (De Beul et al. 2021; Sato et al. 2024). Currently, over 12,000 protein sequences are classified within family GH65. In vivo, GH65 enzymes are typically involved in breakdown reactions in the carbohydrate metabolism (D’Enfert and Fontaine 1997; Andersson et al. 2001; Pedreño et al. 2004; Mokhtari et al. 2013; Jung et al. 2014; Sánchez-Fresneda et al. 2014, 2015; Kabisch et al. 2014; Zilli et al. 2015; Andersen et al. 2020; Kido et al. 2021; Miyazaki et al. 2022; Nakamura et al. 2024), but they also play a role in antibiotic biosynthesis (Sato et al. 2024) or the modification of collagen glycosylation (Hamazaki and Hamazaki 2016).
a Reactions catalyzed by glycoside hydrolases (GHs), glycoside phosphorylases (GPs) and glycosyltransferases (GTs) in Glycoside Hydrolase family 65 (GH65). b Reaction scheme for the inverting single displacement mechanism of GH65 enzymes. For GHs: ROH = a carbohydrate derivative and R’OH = water; for GPs: ROH = a carbohydrate derivative and R’OH = inorganic phosphate; for GTs: ROH = an acceptor group, R’OH = a nucleoside diphosphate (NDP). βGlc1P β-glucose 1-phosphate, NDP-βGlc NDP-β-glucose, NDP-4a4d-βGlc NDP-4-amino-4-deoxy-β-glucose
All members of the GH65 family share great similarities in both their catalytic mechanism and structure. They all break or form α-glucosidic bonds through a single displacement mechanism which inverts the anomeric configuration of the substrate (Fig. 1b) (Sun and You 2021; Li et al. 2022). In a one-step reaction the glucosyl acceptor performs a direct nucleophilic attack on the anomeric carbon of the glucosyl donor (Nakai et al. 2010b; Nakamura et al. 2021). The transition state has an oxocarbenium ion character, with a partially positive charge at the anomeric carbon that is resonance-stabilized by the ring-oxygen (Desmet and Soetaert 2011). For many GPs in the GH65 family, it has been demonstrated that the reaction proceeds via a sequential bi-bi mechanism, which requires both substrates to bind before the reaction takes place and the products are released (Andersson et al. 2001; Nihira et al. 2012a, c, 2013, 2014b, c; Taguchi et al. 2017; Gao et al. 2019; Bi et al. 2022).
GH65 enzymes share a common structure that consists of four regions: an N-terminal β-sandwich domain, a helical linker region, an (α/α)6-barrel catalytic domain and a C-terminal β-sheet domain (Fig. 2) (Egloff et al. 2001; Okada et al. 2014; Touhara et al. 2014; Nakamura et al. 2021). Family GH65 is a member of clan GH-L, together with families GH15, GH125 and GH178 (Drula et al. 2022), and there are also structural similarities to enzymes from families GH94 (Hidaka et al. 2004) and GH95 (Curiel et al. 2021). Most described GH65 enzymes are present as dimers in solution (D’Enfert and Fontaine 1997; Hüwel et al. 1997; Chaen et al. 1999a; Egloff et al. 2001; Inoue et al. 2002a; Hidaka et al. 2005; Nakai et al. 2009, 2010b; Van Hoorebeke et al. 2010; Yamamoto et al. 2011; Van der Borght et al. 2011; Nihira et al. 2012a, 2013, 2014b, c; Okada et al. 2014; Touhara et al. 2014; Zilli et al. 2015; Taguchi et al. 2017; Gao et al. 2019). Two distinct types of dimers have been observed in crystal structures: a ‘head-to-head’ dimer formed by interactions in the catalytic domain (Egloff et al. 2001; Touhara et al. 2014) and a dimer stabilized by a disulfide bridge in the N-terminal domain (Okada et al. 2014). There are also two reports of GH65 enzymes that adopt a hexameric form (Chaen et al. 1999b; Nakamura et al. 2021).
Overall structure of GH65 enzymes illustrated by ribbon diagram of one monomer of the kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP) in complex with glucose (green) and phosphate (orange) (PDB ID: 3WIR). The four structural regions are: an N-terminal β-sandwich domain (residues 1-256, blue), a helical linker region (residues 257–294, cyan), an (α/α)6-barrel catalytic domain (residues 300–691, yellow) and a C-terminal β-sheet domain (residues 295–299 and 692–756, red)
Although all GH65 enzymes operate with a similar mechanism and share the same three-dimensional fold, family GH65 is also a very diverse enzyme family, especially with respect to the many enzymatic functions that it harbors. This review provides an overview of the remarkable functional diversity within family GH65, with a special focus on the determinants that dictate these differences and how this variety of activities can be exploited for glycoside synthesis. While some recent reviews about glycoside phosphorylases or trehalases discuss selected aspects of certain GH65 enzymes (Sakaguchi 2020; Awad 2021; Sun and You 2021; Li et al. 2022; Shrestha et al. 2023), this work presents the first comprehensive discussion of this intriguing protein family.
Diversity of reaction types in GH65
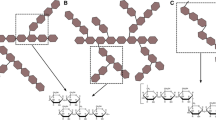
The most common reaction type found in family GH65 is the reversible phosphorolysis of glycosidic bonds, with eight different GP specificities known to date. However, family GH65 also contains various enzymes that catalyze the hydrolytic degradation of glycosidic bonds. It has previously been hypothesized that many of the GPs found in nature may have evolved from GHs, and the occurrence of both reaction types in this family reflects their close evolutionary relatedness (Fig. 3) (Franceus et al. 2021, 2022). For the substrates trehalose and kojibiose, family GH65 even contains a dedicated GH as well as a dedicated GP. Other examples of co-existing GHs and GPs that recognize the same substrate can be found in CAZy families GH3 (N-acetyl-β-glucosaminides) (Franceus et al. 2021), GH13 (sucrose) (Franceus and Desmet 2020; Miyazaki and Park 2020) and GH130 (β-1,2-mannosides) (Li et al. 2020). Together, these enzyme pairs are attractive case studies for future research to further elucidate the adaptations that are required to turn hydrolases into phosphorylases (Franceus et al. 2022).
Phylogenetic tree of GH65 enzymes with the characterized representatives indicated with colored circles. The legend specifies the colors used for each specificity. Glycoside phosphorylases (GPs) and glycoside hydrolases (GHs) are found in two separate clades of the tree, while the glycosyltransferase (GT) specificity is located within a branch of archaeal kojibiose phosphorylases. The tree was constructed by extracting all sequences from the CAZy database (10 June 2024) (Drula et al. 2022), clustering them at 90% sequence identity with CD-HIT (Huang et al. 2010) and removing fragments of less than 400 amino acids, followed by alignment with MAFFT (Katoh et al. 2019) and tree inference with FastTree (Price et al. 2010) using default parameters. Tree visualization with iTOL (Letunic and Bork 2021)
For a long time, all characterized GPs in GH65 originated from bacteria, whereas all eukaryotic family members were GHs. However, this changed when two bacterial representatives of a recently discovered GH specificity, an α-1,2-glucosidase, were simultaneously described by our group (De Beul et al. 2021) and by Nakamura et al. (2021). Later, an additional bacterial GH, active on α-1,2-glucosidic branch points in dextran, was identified (Miyazaki et al. 2022).
Very recently, a third reaction type was identified in family GH65. When reconstructing the biosynthetic pathway of apramycin in Streptoalloteichus tenebrarius, Sato et al. (2024) discovered that an enzyme from family GH65 acts as a GT that performs the final glycosylation step in the synthesis route of this antibiotic. It catalyzes the transfer of a glycosyl moiety from NDP-β-glucose or NDP-4-amino-4-deoxy-β-glucose to the precursor aprosamine 5-phosphate. This was a remarkable finding in two respects. Firstly, it is the first example of a natural glycosylation reaction that uses an NDP-β-d-hexose as glycosyl donor, in contrast to conventional Leloir-type GT reactions that exclusively use either NDP-α-d-hexoses or NDP-β-l-hexoses as glycosyl donor. Secondly, the discovery makes GH65 the first and only CAZy family known to harbor GHs, GPs and GTs. A similar evolutionary link between GH, GP and GT activities had already been described, albeit for enzymes that are more distantly related and belong to two different CAZy families: GH130 and GT108. GH130 contains GHs and GPs active on β-1,2-mannosides, while GT108 consists of strict GTs and dual-activity GP-GT enzymes for the same substrate. Despite their low sequence similarity, enzymes in both families share the same three-dimensional scaffold and active site architecture and are thought to have a common evolutionary origin (Sernee et al. 2019; Li et al. 2020). In analogy to a recent study of ancestral sequences in GT108 that showed how subtle distal mutations form the evolutionary link between GP and GT activity in GT108 (Franceus et al. 2024), it would be highly intriguing to uncover whether similar processes were involved in the emergence of the GH, GP and GT activities in GH65.
The diversity of reaction types in GH65 enzymes is also reflected in the slight variations in their reaction mechanism and active site architecture. GHs require two catalytic residues: a catalytic base that deprotonates a water molecule to facilitate a direct nucleophilic attack, and a catalytic acid that protonates the leaving group. Contrastingly, GPs and GTs operate with only one catalytic residue, since the phosphate or nucleotide group is already ionized under physiological conditions and the catalytic base is therefore no longer required (Desmet and Soetaert 2011). The catalytic acid (E483 in the kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP), Fig. 4; Table 2, Fig. S1) is conserved in all GH65 enzymes and was first identified by Egloff et al. (2001) by comparing the crystal structure of the maltose phosphorylase of Lactobacillus brevis (LbMP) with that of a related GH15 glucoamylase. For a long time, the identity of the catalytic base in GHs remained elusive, but Hamazaki and Hamazaki (2016) were the first to propose a candidate in human protein-α-glucosyl-1,2-β-galactosyl-l-hydroxylysine α-glucosidase based on mutational analysis of conserved carboxyl residues. Their hypothesis was later confirmed when the crystal structure of the α-1,2-glucosidase of Flavobacterium johnsoniae (FjKH) was solved and Glu616 was pinpointed as catalytic base (Fig. 4; Table 2, Fig. S1) (Nakamura et al. 2021). For several GH65 representatives it has been demonstrated experimentally that they become catalytically inactive when the catalytic acid or the catalytic base is mutated (Egloff et al. 2001; Hidaka et al. 2005; Nihira et al. 2014c; Hamazaki and Hamazaki 2016; Nakamura et al. 2021).
The active site of glycosides phosphorylases (GPs) and glycoside hydrolases (GHs) in GH65. The catalytic acid is conserved for all GH65 enzymes, but the catalytic base of GHs is replaced by a phosphate binding site in GPs. Stick representation of important residues in the kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP, teal, PDB ID: 3WIR) and the kojibiose hydrolase of Flavobacterium johnsoniae (FjKH, grey, PDB ID: 7FE4). Phosphate (orange) and glucose molecules in the − 1 and + 1 subsites (green) originate from the structure of CsKP (PDB ID: 3WIR)
In GPs the catalytic base is replaced by a highly conserved phosphate binding site formed by a lysine or arginine, two consecutive serines and a histidine residue (K330, S631, S632 and H675 in CsKP). Additionally, in the recently discovered GT, a tyrosine that is completely conserved in both GHs and GPs (Y337 in CsKP), is mutated to a histidine. In GPs and GHs, this tyrosine forms a hydrogen bond with the phosphate molecule or with the catalytic base, respectively (Fig. 4; Table 2, Fig. S1) (Egloff et al. 2001; Okada et al. 2014; Touhara et al. 2014; Nakamura et al. 2021). The tyrosine-to-histidine mutation in the GT is hypothesized to broaden the phosphate-binding site and allow recognition of a sugar nucleotide as glycosyl donor (Fig. S2) (Sato et al. 2024).
Diversity of substrate specificities in GH65
Overview of substrate specificities
GH65 is also a particularly diverse family on the level of substrate preference, with currently 13 characterized enzyme specificities (Fig. 3; Tables 1, S1, S2), of which eight are GPs, four are GHs and one is a GT. Especially the number of different phosphorylase specificities is remarkable. Of all ten GP-containing CAZy families, only family GH94 harbors more different GP specificities within one family than GH65 (Awad 2021; Li et al. 2022; Drula et al. 2022).
α-Glucobioses are the typical substrates of GH65 enzymes. The reversible phosphorolysis of trehalose (α-1,1-α-glucobiose), kojibiose (α-1,2-glucobiose), nigerose (α-1,3-glucobiose) and maltose (α-1,4-glucobiose) is catalyzed by trehalose phosphorylase (TP), kojibiose phosphorylase (KP), nigerose phosphorylase (NP) and maltose phosphorylase (MP), respectively. MPs, TPs and NPs have a strict preference for the phosphorolysis of the disaccharide (Table S1). On the other hand, all characterized KPs, except for the KP from Escherichia coli K-12, are also active on α-1,2-oligoglucans with a higher degree of polymerization (DP) (Table S1). The chain length preference of those KPs can vary between enzymes. For example, when comparing the synthetic capabilities of the KPs from Caldicellulosiruptor saccharolyticus (CsKP) and Thermoanaerobacter brockii (TbKP) in reverse phosphorolysis mode, the former had the tendency to produce kojioligosaccharides with a higher DP than the latter (Yamamoto et al. 2011).
Not all phosphorylases in the family are active on glucobioses. α-1,3-Oligoglucan phosphorylases (ONPs) phosphorolyze α-1,3-oligosaccharides starting from a DP of 3 and show no activity on the disaccharide nigerose. ONPs are often located up- or downstream of genes encoding an NP, suggesting that the ONP first breaks down α-1,3-glucosidic chains to β-glucose 1-phosphate (βGlc1P) and nigerose, which is subsequently used as substrate of the NP (Nihira et al. 2014b). Similar combinations of a strict disaccharide phosphorylase with a phosphorylase only active on longer chains are found in certain bacterial metabolic pathways for cellooligosaccharides (Liu et al. 2019), β-1,3-glucans (Kuhaudomlarp et al. 2019) and β-1,4-mannooligosaccharides (La Rosa et al. 2019).
Besides those α-glucobiose and α-oligoglucan phosphorylases, GPs active on the phosphate sugar trehalose 6-phosphate, the disaccharide 3-O-α-glucosyl-l-rhamnose and the osmolyte 2-O-α-glucosylglycerol are also classified within GH65. The latter was the first report of a GP that prefers a polyol instead of a carbohydrate as glycosyl acceptor in reverse phosphorolysis reactions (Nihira et al. 2014c). Remarkably, an enzyme from family GH13 was later found to also catalyze the phosphorolysis of 2-O-α-glucosylglycerol, but via a retaining mechanism, releasing αGlc1P instead of βGlc1P (Franceus et al. 2018). The inverting trehalose phosphorylase in GH65 also has such a retaining counterpart in family GT4. Together, 2-O-α-glucosylglycerol and trehalose are the only two substrates for which both an inverting and retaining phosphorylase have been described.
Next to this variety of GPs, also four GH specificities are found in GH65. Many fungal trehalases, which hydrolyze the α-1,1-α-bond in trehalose, have been described. To make the distinction with other trehalases classified in families GH15 and GH37, GH65 trehalases are sometimes also termed acid trehalases, due to their optimal activity under acidic conditions (Sakaguchi 2020). The other three GH specificities in GH65 are all active on α-1,2-glucosidic bonds, but prefer different substrates. Kojibiose hydrolases (KHs) perform the hydrolysis of kojibiose and longer linear kojioligosaccharides like kojitriose, kojitetraose and kojipentaose (De Beul et al. 2021; Nakamura et al. 2021). Nakamura et al. (2021) reported that the KH from Flavobacterium johnsoniae (FjKH) also has weak activity on α-1,2-branched dextran. Dextran α-1,2-debranching enzyme (DDE) has exactly the opposite substrate preference: its main activity is the release of glucose from α-1,2-glucosidic branchpoints in dextran, while it only shows minor activity on kojibiose and kojitriose (Miyazaki et al. 2022). The third α-1,2-active GH specificity in GH65 is the protein-α-glucosyl-1,2-β-galactosyl-l-hydroxylysine α-glucosidase. This enzyme hydrolyzes the disaccharide unit that decorates l-hydroxylysine residues of collagen and related proteins, releasing free glucose (Hamazaki and Hotta 1979; Hamazaki and Hamazaki 2016).
Lastly, GH65 also contains one GT specificity, the apramycin-5-phosphate synthase (A5PS) (Sato et al. 2024), which was discussed above.
Almost all GH65 enzymes strictly bind only their native substrate in the breakdown direction (Table S1). Promiscuous activity on the preferred substrates of other enzymes in the family is rarely observed. The only exception is the overlapping activity of KHs, KPs and NPs on nigerose and kojibiose. Both KHs and KPs have been reported to also hydrolyze or phosphorolyze nigerose (0.2–5.5% compared to their activity on kojibiose) (Yamamoto et al. 2011; Okada et al. 2014; Jung et al. 2014; De Beul et al. 2021; Nakamura et al. 2021), whereas NPs are also able to degrade kojibiose (0.5–8.4% compared to their activity on nigerose) (Nihira et al. 2012a; Bi et al. 2022). In contrast to the strict specificity for the glucosyl group in subsite − 1 and for the glycosidic linkage type of their native substrate, GH65 enzymes are more flexible when it comes to the identity of the substrate in subsite + 1. Most GPs tolerate a variety of alternative carbohydrates as glycosyl acceptor in the synthesis direction of their reversible phosphorolysis reaction (Table S2).
Determinants of substrate specificity
Although our understanding of what controls substrate specificity in family GH65 is not entirely complete, several important aspects of it have been described over the last two decades. Those insights can be grouped in three main topics, which will be discussed in more detail below: (1) specific substrate recognition motifs in the active site, (2) residues involved in a specificity-determining correlation network, and (3) distinctive larger structural elements.
Active site motifs for substrate recognition
Conservation patterns for residues in the active site of GH65 enzymes are strongly dependent on the subsite. While the − 1 subsite of all GH65 enzymes is largely conserved, the + 1 subsite is highly divergent between enzymes with different substrate preferences (subsite nomenclature according to Davies et al. (1997)). The − 1 subsite strictly binds a glucosyl group, except for the S. tenebrarius A5PS, which also binds 4-amino-4-deoxy-β-glucose. Fully conserved tryptophan, aspartate, lysine and glutamine residues (W343, D344, K596 and Q597 in CsKP) shape this binding pocket and form hydrogen bonds with all hydroxyl groups of the glucosyl moiety in subsite − 1 (Fig. 5; Table 2, Fig. S1) (Okada et al. 2014). In contrast, different GH65 enzymes have evolved to bind different substrate moieties in subsite + 1, and the residues in that subsite are adapted accordingly. Based on structural analyses and mutational studies, several authors have successfully identified such substrate binding motifs within a number of GH65 specificities.
Based on a homology model of the MP of Lactobacillus acidophilus (LaMP), Nakai et al. (2010b) identified two distinctive residues that form hydrogen bonds with the glucose moiety of maltose in the + 1 subsite of LaMP: His413 and Glu415 (Fig. 5a; Table 2, Fig. S1). Those amino acids are very conserved for all MPs, while other enzymes show different motifs at the corresponding positions. The importance of these residues for maltose binding was confirmed by replacing His413, Asn414 and Glu415 (HNE motif) by the corresponding residues in the KP (TPK) or TP (SAY) of Thermoanaerobacter brockii (TbKP or TbTP). The catalytic efficiency of those mutants for the phosphorolysis of maltose decreased dramatically by 3 to 4 orders of magnitude, caused by both a decrease of kcat and increase of Km. Those findings confirmed that the HNE motif is involved in key interactions with maltose. The mutants also obtained a low activity on kojibiose or trehalose (~ 3.5% compared to the activity of wild-type LaMP on maltose), depending on whether the KP or TP motif was introduced. This suggests that those residues participate at least to some extent in the accommodation of the glucose unit of kojibiose or trehalose in the + 1 subsite.
However, when Okada et al. (2014) solved the crystal structure of CsKP a few years later, they found that the side chains of the TPK motif point away from kojibiose, or are too small to interact with the substrate. Instead, the residues forming hydrogen bonds with kojibiose in subsite + 1 of CsKP were found to be Trp391, Glu392 and Thr417 (Fig. 5b; Table 2, Fig. S1). This WET motif is highly conserved for all KPs, and was later also shown to be present in KHs, which is the other GH65 specificity with activity on kojibiose (Nakamura et al. 2021). Other conserved amino acids are present in other enzyme specificities: MVI in MPs, WRA in TPs and WEF in NPs. Trp391 and Glu392, especially the latter, were shown to be crucial for kojibiose recognition in CsKP via mutational studies. Mutants with other amino acids at positions 391 and 392 had a severely impaired kojibiose-activity (< 2.1% of the activity of wild-type CsKP on kojibiose), whereas mutating Thr417 had a less dramatic impact (> 14% retained activity). While it is clear that the WET motif is important for kojibiose binding, it was not possible to induce MP, TP or NP activity by replacing it with the corresponding patterns of MPs, TPs or NPs (Okada et al. 2014). It is probably not a coincidence that NPs and DDEs, which are both promiscuous towards kojibiose, also possess the tryptophan and glutamate residues in their + 1 subsite (Miyazaki et al. 2022).
Remarkably, 2-O-α-glucosylglycerol phosphorylase (GGP) is the only GH65 specificity that strictly recognizes a polyol instead of a carbohydrate group in its + 1 subsite. Since the glycerol group is more flexible than a sugar ring, more extensive interactions are necessary in order to correctly position it in the active site (Fig. 5c; Table 2, Fig. S1). In the GGP of Bacillus selenitireducens (BsGGP), the hydrophobic atoms of the glycerol molecule interact with three phenylalanine residues (Phe395, Phe396 and Phe409). The O1 and O3 hydroxyl groups of glycerol form direct hydrogen bonds with Tyr327 and Trp381. On the other hand, the O2 atom of glycerol is kept in place by an extensive water-mediated hydrogen bond network formed by Tyr327, Tyr572 and Lys587. The authors postulate that this hydrogen-bonded water network can also position a water molecule instead of glycerol for nucleophilic attack, explaining the ability of BsGGP to hydrolyze βGlc1P. Tyr327 and Lys587 are conserved positions in GH65, but all other residues involved in glycerol recognition are specific for GGP. Mutating positions Tyr327, Trp381, Tyr572 or Lys587 removes or drastically decreases the reverse phosphorolysis activity of BsGGP (< 2% catalytic efficiency compared to wild-type BsGGP). Mutations at Tyr327, Tyr572 or Lys587 also largely abolish the hydrolysis of βGlc1P, while the mutant W381F still hydrolyzes βGlc1P, albeit with a 4 times lower affinity. Mutating Trp381 probably has less impact on βGlc1P hydrolysis because it is not involved in the network that keeps the water molecule in place (Touhara et al. 2014).
While the substrate recognition motifs of MPs, KPs, KHs and GGPs have been described in detail based on the available crystal structures of LbMP, CsKP, FjKH and BsGGP, the active site patterns responsible for binding the native substrates of other family members remain more elusive. Van der Borght et al. (2011) and Miyazaki et al. (2022) briefly mention amino acids involved in substrate binding in TPs and DDEs, based on a homology model or AlphaFold2 model, resp. (results summarized in Table 2, Fig. S1). More thorough investigation is required in the future to comprehensively map the determinants of specificity in family GH65.
Substrate recognition motifs in the active site of a the maltose phosphorylase of Lactobacillus acidophilus (LaMP), b the kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP) and c the 2-O-α-glucosylglycerol phosphorylase of Bacillus selenitireducens (BsGGP). Conserved amino acids that recognize the glucose unit in the − 1 subsite and the catalytic acid are shown in gray. Hydrogen bonds are shown as dashed lines. a The maltose-like moiety of acarbose (green) was positioned in the AlphaFold2 model of LaMP according to the method of Nakai et al. (2010b). The glucose unit in the + 1 subsite forms hydrogen bonds with His413 and Glu415 (magenta) of the HNE motif. b CsKP (PDB ID: 3WIQ) is shown in complex with kojibiose (green). The WET motif that recognizes the non-reducing end of the substrate is shown in teal. c BsGGP (PDB ID: 4KTR) is shown in complex with glycerol (green). The glucose unit in subsite − 1 (green) originates from another structure of the same enzyme (PDB ID: 4KTP). Residues that form (in)direct hydrogen bonds with glycerol are shown in orange. Three phenylalanine residues that form hydrophobic interactions with glycerol are shown as thin orange lines. Two water molecules involved in the water-mediated hydrogen bond network are shown as pale blue spheres
Correlated positions as specificity determinants
Studying the active site of enzymes via elucidation of their crystal structure and mutational experiments is a time-consuming and case-by-case approach. Moreover, non-obvious positions outside of the active site are usually overlooked, while in reality distant positions can actually have a significant impact on enzyme activity and specificity (Wilding et al. 2019; Osuna 2021; Gu et al. 2023; Franceus et al. 2024). In an attempt to study potential specificity determinants for the entire GH65 family at once in a more systematic manner, we recently used correlated mutations analysis (CMA) to uncover a specificity-based correlation network in GH65 (De Beul et al. 2021). Essentially, CMA detects co-evolving positions in a multiple sequence alignment (MSA). At those positions, only certain amino acid combinations are observed in the MSA, suggesting that other combinations are deleterious and thus purged by selective pressures. It is assumed that positions that are critical to a certain substrate specificity show this behavior. Indeed, such positions are typically highly conserved, but when they do mutate to accommodate a different substrate, they tend to mutate together. Those co-evolution patterns can be detected with CMA and visualized in a correlation network (Franceus et al. 2017).
In GH65, we unveiled a network of 24 correlated positions, of which positions 64, 392, 394, 402, 416 and 585 (CsKP numbering) showed the strongest evolutionary correlation (Fig. 6a, Fig. S1). Based on this analysis, 22 putative specificity subgroups could be defined. The amino acids at the correlated positions show distinct patterns within one subgroup but vary between subgroups (Fig. 6b). It has been demonstrated that those sequence patterns can be used to annotate the substrate specificity of unknown GH65 enzymes, leading to annotation of a large part of the phylogenetic tree. For example, subgroup 5 contains all MPs and has a highly conserved KV[MF]NES motif, while the EEAPxx motif is characteristic of KPs. However, some subgroups display motifs at the correlated positions that do not match those of already characterized representatives, suggesting that they may harbor enzymes with novel specificities. A targeted search within those clades previously led to the discovery of the KH specificity in subgroup 18 (De Beul et al. 2021).
Correlated positions as specificity determinants in family GH65. a Correlation network of the GH65 family alignment. Nodes represent the alignment positions, numbering according to the kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP). Node size indicates the number of edges. Edge thickness indicates the strength of pair-wise correlation. The six selected positions are highlighted in yellow. b Sequence logo and consensus motif at those six selected positions (left to right: positions 64, 392, 394, 402, 416 and 585; CsKP numbering) for each subgroup that contains characterized GH65 enzymes. A sequence logo for subgroup 7 is not shown, as this subgroup contained only one sequence. Adapted from De Beul et al. (2021)
While the amino acids at the correlated positions are undoubtedly somehow involved in specificity control, it remains unclear how they really affect substrate binding, especially since some residues are up to 27 Å away from the active site. In any case, one should be careful when substituting amino acids that are part of the correlation network, to avoid disrupting the intricate interactions between those residues. Introducing a mutation at one position might require compensatory mutations at other positions in the network to maintain enzyme activity.
Loops around the active site differ between substrate specificities
Larger structural elements can be indicative of the substrate preference of GH65 enzymes as well. Especially several loops around the active site were suggested by multiple authors to be related to specificity. Already in 2006b, Yamamoto et al. hypothesized the importance of the loop that connects α-helices α3 and α4 of the catalytic (α/α)6-barrel domain, termed loop 3. They constructed several chimeric enzymes of TbKP and TbTP. Surprisingly, a certain chimera of 785 amino acids contained only one segment of 125 residues originating from the KP (Met384–Thr512) but still exhibited KP activity. This region, ranging from α3 to α6 of the (α,α)6-barrel catalytic domain, was therefore suspected to contain crucial residues for substrate recognition. Via inspection of a multiple sequence alignment, they pinpointed loop 3 as a potential specificity determinant. This loop is highly divergent between different specificities, both in length and amino acid sequence (Yamamoto et al. 2006b; Nakai et al. 2010b).
This suggestion was further investigated by Nakai et al. (2010b) and Okada et al. (2014), who both identified a motif for substrate recognition within or close to this loop 3 in LaMP and CsKP, resp. (see above). Okada et al. (2014) also described a second loop that shows significant differences between specificities, namely a loop that connects two β-strands in the N-domain, termed the N-loop. In LbMP, this N-loop is long and helps shape the active site, while loop 3 is shorter. This is in contrast with the structure of CsKP, that has a shorter N-loop that is not involved in formation of the active site, and longer loop 3, which forms an anti-parallel β-sheet and covers the active site (Table 3; Figs. 7, S1). However, this longer loop 3 does not seem to be a strict requirement for KP activity, since exchanging the entire loop 3 of LaMP for its longer equivalent of TbKP resulted in a lower activity on kojibiose than when only the three residues of the HNE motif were swapped with the corresponding KP-conserved residues (TPK) (Nakai et al. 2010b).
Ribbon representation of four loops around the active site of GH65 enzymes: N-loop (green), loop 3 (magenta), loop 7 (blue), loop 11 (orange). a Maltose phosphorylase of Lactobacillus brevis (LbMP, PDB ID: 1H54) in complex with phosphate (orange). b Kojibiose phosphorylase of Caldicellulosiruptor saccharolyticus (CsKP, PDB ID: 3WIR) in complex with glucose (green) and phosphate (orange). c 2-O-α-Glucosylglycerol phosphorylase from Bacillus selenitireducens (BsGGP, PDB ID: 4KTR) in complex with the glucose analogue isofagomine and glycerol (green). d Kojibiose hydrolase from Flavobacterium johnsoniae (FjKH, PDB ID: 7FE4) in complex with glucose (green)
The crystal structure of BsGGP was determined shortly after that of CsKP. Again, the importance of certain loops around the active site was highlighted: loop 3, loop 7 (between α-helices α7 and α8) and loop 11 (between α-helices α11 and α12) (termed loop 1, 2 and 3 in the paper, resp.) completely cover the active site in the crystal structure of BsGGP. Therefore, one or more loops should flexibly move to allow the substrates and products to enter and leave the active site. As loop 11 has an average B-factor that is slightly higher than the overall average value and it is not located at the dimer interface, this loop is most likely the one that opens and allows the substrates to reach the active site. Again, these three loops vary in length and sequence between different specificities (Table 3; Figs. 7, S1). In LbMP, loops 3, 7 and 11 are short, which results in a relatively open active site, whereas a long loop 3 in CsKP partially covers the active site. Loop 7 is long in CsKP but is in an open configuration, while loop 11 is shorter than in BsGGP. Based on a multiple-sequence alignment, NPs have similar loop lengths than BsGGP, while trehalose-6-phosphate phosphorylases (T6PPs) have a long loop 3 and 7 and short loop 11 like CsKP. ONPs and TPs have three short loops like LbMP (Touhara et al. 2014).
In summary, several authors have already discussed the importance of four different loops in and around the active site for specificity: the N-loop (Thr59-Thr63 in CsKP), loop 3 (Glu398-Glu421 in CsKP), loop 7 (Met577-Asp587 in CsKP) and loop 11 (Gly668-Leu672 in CsKP). However, much more research is required to fully understand how they contribute to substrate specificity and how they may be engineered to change the substrate preference of these enzymes.
Finally, it should be noted that there is substantial overlap among the findings from efforts to identify specificity determinants through the analysis of active site residues, correlation networks and structural elements. Many of the mentioned active site positions or correlated positions are located in or near the N-loop or loop 3, 7 or 11. Moreover, three residues from the active site motifs are also part of the correlation network, i.e. Trp392 and Thr417 of the WET motif in CsKP, and Asn414 of the HNE motif in LaMP (corresponding to Pro402 in the TPK motif of CsKP).
Carbohydrate synthesis with GH65 phosphorylases
The broad portfolio of reactions catalyzed by GH65 enzymes can be exploited for carbohydrate synthesis. Especially GPs are interesting in this respect due to the reversible nature of the phosphorolysis reaction they catalyze. In the phosphorolysis direction, GPs can be applied to produce sugar 1-phosphates, while their reverse phosphorolysis reaction allows synthesis of valuable carbohydrates or glycosides.
Synthesis of sugar phosphates
The most straightforward use case of GH65 phosphorylases is the production of βGlc1P. This sugar phosphate is not only the preferred donor of all GPs in CAZy families GH3 (Franceus et al. 2021) and GH65 (Awad 2021; Li et al. 2022), but it can also serve as a chemical glycosylation agent (Plante et al. 1999; Luley-Goedl and Nidetzky 2010) or NDP-sugar precursor (Yang et al. 2004). Moreover, sugar phosphates can also have nutritional or pharmaceutical applications, for example as stimulant of intestinal active calcium transport (Fujinaka et al. 2007), phosphate source in parenteral nutrition (Ronchera-Oms et al. 1995), anti-inflammatory drug (Parish et al. 1992), immunosuppressant (Parish et al. 1992) or building block for a vitamin D-like medicine (Fujinaka et al. 2004).
In theory, all GPs in GH65 can be used to generate βGlc1P, but the most convenient ones are those that can start from a cheap substrate such as trehalose or maltose. Since industrial carbohydrate conversions are preferably carried out at high temperatures and no MPs are available that remain active after several hours at elevated temperatures, Van der Borght et al. (2010) produced βGlc1P from trehalose with the thermostable TbTP. However, βGlc1P is an activated sugar with a high energy content, whereas the substrate trehalose is very stable and low-energetic. Therefore, the unfavorable thermodynamic equilibrium of the reaction limits the theoretical maximum yield to 26%. Despite this limitation, βGlc1P was successfully produced from 200 mM trehalose in 200 mM phosphate buffer on 2-L scale, after which two different purification methods were compared. In the first method, anion exchange chromatography was used to separate βGlc1P from the remaining substrates and from glucose, which is a by-product of the reaction. Alternatively, the combined action of baker’s yeast (Saccharomyces cerevisiae) and a trehalase treatment removed all unwanted sugars, whereas phosphate was precipitated as struvite. Although the product could be recovered almost completely (> 99%) and with high purity (> 99%) via both purification methods, efforts to obtain a crystalline product were either unsuccessful or inefficient. A similar reported process was based on the phosphorolysis of maltose with the MP of Bacillus sp. AHU2001 (Taguchi et al. 2017; Gao et al. 2019). The conversion rate of maltose into βGlc1P (30%) was comparable to the TP-based method. After purification, 40–57% of the product could be recovered with 96% purity. Due to the low thermostability of the MP, the reaction was carried out at 37 °C, which is much lower than the process temperature that was used in the reaction with TbTP (60 °C).
Interestingly, TP can also be used for the production of another sugar 1-phosphate, namely β-galactose 1-phosphate (βGal1P). Due to their relatively broad acceptor specificity, TPs are able to synthesize trehalose analogues such as lactotrehalose (α-glucopyranosyl-(1,1)-α-galactopyranoside) from βGlc1P and an alternative acceptor like galactose (see below). Due to the non-reducing, symmetric bond in lactotrehalose, the phosphorolysis of this compound can yield both βGlc1P and βGal1P, depending on whether the glucose or the galactose moiety was bound in the − 1 subsite of the enzyme. However, the wild-type TP from Caldanaerobacter subterraneus (CsTP) produces 23 times more βGlc1P than βGal1P. Via iterative saturation mutagenesis of three hotspots in the donor subsite, Chen et al. (2014) identified a triple mutant of CsTP (W371Y/L649G/A693Q) with a completely switched donor preference. This mutant produced 94 times more βGal1P than βGlc1P, with a specific activity that was 1.5 times higher than that of the wild-type enzyme. After further optimizing the reaction conditions, 50% of the valuable compound lactotrehalose could be converted into βGal1P. The product was obtained in high purity (> 99%) via chemical phosphate precipitation and anion exchange chromatography. However, the purification process was less efficient than for βGlc1P, with just 29% of the produced βGal1P being recovered. This strategy should be generally applicable to the synthesis of other β-glycosyl 1-phosphates, as long as the corresponding trehalose analogues can be produced as intermediate. This taps into the concept of glycodiversification, where sugar entities in natural small molecules are changed to alter their biological or pharmacokinetic properties (Yang et al. 2004; Thibodeaux et al. 2008). Access to more diverse sugar phosphates would make GPs interesting biocatalytic tools for this concept.
Coupled reactions for carbohydrate synthesis
The functional diversity of GH65 phosphorylases holds great potential for the synthesis of various glycosides when operating them in reverse phosphorolysis mode. However, they require βGlc1P as sugar donor. Even with the attempts to produce βGlc1P in a pure manner on lab-scale, it remains a very costly compound (~€30/mg) and is hardly commercially available. This issue can be circumvented using one-pot coupled reactions where the glycosyl donor, i.e. βGlc1P, is generated and consumed continuously (Table 4). This in-situ production of βGlc1P is much more practical, as there is no longer a need for isolation of reaction intermediates. Phosphate is recycled in the reaction, so only catalytic amounts must be added, which also facilitates downstream processing. When smartly selecting enzymes so that one can start from cheap bulk sugars to produce more valuable glycosides, this process is economically far more attractive. However, multi-step processes can quickly become more complex, especially in terms of selecting optimal process parameters (Sigg et al. 2024).
In the most obvious set-up, two GH65 GPs are combined. One of them operates in the phosphorolysis direction to provide βGlc1P and glucose, which are subsequently used as substrates for the other GP that works in synthesis direction. Trehalose, nigerose and kojioligosaccharides have been produced from maltose in that manner by combining an MP with a TP, NP and KP, respectively (Fig. 8a) (Murao et al. 1985; Yoshida et al. 1998; Chaen et al. 2001b; Nihira et al. 2014a; Bi et al. 2022). For the synthesis of trehalose 6-phosphate or 2-O-α-glucosylglycerol from maltose by the coupled reaction of an MP and a T6PP or GGP, it was necessary to add glucose 6-phosphate (Glc6P) or glycerol as external acceptors (Fig. 8b) (Taguchi et al. 2020; Zhang et al. 2020). In the case of trehalose 6-phosphate, it was possible to avoid the addition of Glc6P and instead produce it in situ from βGlc1P by including a β-phosphoglucomutase as a third enzyme in the reaction mix. Adding baker’s yeast to remove the glucose that otherwise accumulates as by-product also sped up the reaction (Fig. 8c).
Examples of coupled reactions of GH65 phosphorylases for the production of carbohydrates. Synthesis of a, e-g nigerose, b-c trehalose 6-phosphate, and d lactotrehalose, the galactose-containing trehalose analogue (Glcα1⟶1αGal). CBP cellobiose phosphorylase, Glc glucose, Glc6P glucose 6-phosphate, GP glycogen phosphorylase, IAm isoamylase, MP maltose phosphorylase, NP nigerose phosphorylase, Pi inorganic phosphate, SP sucrose phosphorylase, T6PP trehalose-6-phosphate phosphorylase, TP trehalose phosphorylase, XI xylose isomerase, α-PGM α-phosphoglucomutase, αGlc1P α-glucose 1-phosphate, β-PGM β-phosphoglucomutase, βGlc1P β-glucose 1-phosphate
Since many GH65 GPs have a rather relaxed acceptor scope, coupled reactions can also be employed for the production of various analogues of α-glucobioses, in which one of the glucose moieties is replaced by another monosaccharide. Nakai et al. (2010a) demonstrated the synthesis of various maltose analogues using N-acetyl glucosamine, glucosamine, mannose, l-fucose or xylose as acceptors, while Chaen et al. (2001a) produced a galactose-containing trehalose analogue, also referred to as lactotrehalose (Fig. 8d). Unfortunately, the activity of GH65 phosphorylases on alternative acceptors is typically lower than their activity on their preferred acceptor. However, this problem can be tackled via enzyme engineering, as demonstrated in a study of TbTP for the synthesis of lactotrehalose (Van der Borght et al. 2012). Semi-rational engineering of three hotspots in the acceptor subsite did not result in any improved variants, but via random mutagenesis using error-prone PCR three positions (A440, R448, N657) were identified that influenced the affinity for galactose. Mutating those positions decreased the Km value of TbTP for galactose two- to threefold, but the effects were non-additive. Combining the beneficial mutations even decreased the catalytic activity and thermostability. Mutant R448S was therefore considered the best option for the synthesis of lactotrehalose.
Although maltose and trehalose are relatively affordable and easily available sugars, it is also interesting to set up coupled reactions starting from other cheap sugar resources like cellobiose, sucrose or starch (Fig. 8e-g). Unfortunately, the GPs active on those substrates (cellobiose phosphorylase, sucrose phosphorylase and glycogen phosphorylase, resp.) produce αGlc1P and cannot be employed directly in a one-pot two-enzyme system with a βGlc1P-active GH65 phosphorylase. This hurdle can be overcome by including an α-phosphoglucomutase and a β-phosphoglucomutase, which can work together to convert αGlc1P and βGlc1P with Glc6P as intermediate. This strategy has been successfully employed for the synthesis of nigerose and its galactose-containing analogue (Nihira et al. 2014a). In the case of the sucrose-based system, a xylose isomerase was added to convert the released fructose into glucose, increasing the atom efficiency of the coupled reaction.
Concluding remarks
The extensive diversity of function among carbohydrate-active enzymes in the GH65 family presents great opportunities for further research. Some members of family GH65 are particularly appealing targets for evolutionary studies. A thorough investigation into the structure-function relationships in kojibiose phosphorylases, trehalose phosphorylases, and the corresponding hydrolases would shed light on how enzymes can switch from performing hydrolysis to catalyzing phosphorolysis, and vice versa. Similarly, mutational studies in the peculiar apramycin-5-phosphate synthase and its most closely related GP homologues could enhance our understanding of how the glycosyl donor preferred by GTs can be changed from expensive nucleotide-activated sugars to the corresponding glycosyl phosphates, which tend to be more easily available. Such studies could benefit from the use of ancestral protein reconstruction and resurrection, a technique that has proven useful for tracing the historical evolution of function in protein families.
Despite considerable efforts to identify the residues and structural elements responsible for the functional adaptations in this family’s evolution, success in leveraging this knowledge to exchange or engineer function between enzymes has been limited. Clearly, there is a lot left to learn. For instance, little is known about the dynamics of the four loops in and around the active site that appear to be indicative of substrate specificity, i.e. loops 3, 7, 11 and the N-loop. Similarly, the residues present in the network of evolutionarily correlated positions have been shown to be predictive of specificity, but it is currently unclear how those positions are involved in substrate binding or catalysis. A better understanding of those putative specificity determinants would enable protein engineers to design smarter libraries with a higher hit rate, focusing on the most appropriate mutational hotspots.
Finally, it is unlikely that we have uncovered the full functional diversity present in natural enzymes from the GH65 family. Novel substrate specificities continue to be discovered regularly, and the vast majority of family GH65 members are yet to be characterized experimentally. In particular, the specificity subgroups defined through analysis of the family’s correlation network should be a promising source of novel enzyme candidates. Indeed, the function of enzymes from ten of these subgroups is still unknown today.
The many phosphorolytic activities in family GH65 present substantial opportunities for their use in carbohydrate synthesis. However, the limited availability of thermostable GPs and the high cost of the glycosyl donor βGlc1P are critical bottlenecks that currently hamper their application on an industrial scale. Further expansion of the toolbox of useful GH65 enzymes through enzyme discovery and engineering would broaden the range of valuable carbohydrates and glycosides that can be obtained efficiently from affordable substrates.
References
Alizadeh P, Klionsky DJ (1996) Purification and biochemical characterization of the ATH1 gene product, vacuolar acid trehalase, from Saccharomyces cerevisiae FEBS Lett 391:273–278. https://doi.org/10.1016/0014-5793(96)00751-X
Andersen S, Møller MS, Poulsen J-CN, Pichler MJ, Svensson B, Lo Leggio L, Goh YJ, Abou Hachem M (2020) An 1,4-α-Glucosyltransferase defines a new maltodextrin catabolism scheme in Lactobacillus acidophilus. Appl Environ Microbiol 86:e00661–e00620. https://doi.org/10.1128/AEM.00661-20
Andersson U, Levander F, Rådström P (2001) Trehalose-6-phosphate phosphorylase is part of a novel metabolic pathway for trehalose utilization in Lactococcus lactis. J Biol Chem 276:42707–42713. https://doi.org/10.1074/jbc.M108279200
Awad FN (2021) Glycoside phosphorylases for carbohydrate synthesis: an insight into the diversity and potentiality. Biocatal Agric Biotechnol 31:101886. https://doi.org/10.1016/j.bcab.2020.101886
Bi R, Wu J, Su L, Xia W (2022) Efficient synthesis of nigerose by a novel nigerose phosphorylase from Anaerosporobacter mobilis Syst Microbiol Biomanuf 1:1–11. https://doi.org/10.1007/S43393-022-00122-7
Chaen H, Nakada T, Nishimoto T, Kuroda N, Fukuda S, Sugimoto T, Kurimoto M, Yoshio T (1999a) Purification and characterization of thermostable trehalose phosphorylase from Thermoanaerobium brockii J Appl Glycosci 46:399–405. https://doi.org/10.5458/jag.46.399
Chaen H, Yamamoto T, Nishimoto T, Nakada T, Fukuda S, Sugimoto T, Kurimoto M, Tsujisaka Y (1999b) Purification and characterization of a novel phosphorylase, kojibiose phosphorylase, from Thermoanaerobium brockii. J Appl Glycosci 46:423–429. https://doi.org/10.5458/jag.46.423
Chaen H, Nakada T, Mukai N, Nishimoto T, Fukuda S, Sugimoto T, Kurimoto M, Tsujisaka Y (2001a) Efficient enzymatic synthesis of disaccharide, α-d-galactosyl α-d-glucoside, by trehalose phosphorylase from Thermoanaerobacter brockii J Appl Glycosci 48:135–137. https://doi.org/10.5458/jag.48.135
Chaen H, Nishimoto T, Nakada T, Fukuda S, Kurimoto M, Tsujisaka Y (2001b) Enzymatic synthesis of kojioligosaccharides using kojibiose phosphorylase. J Biosci Bioeng 92:177–182. https://doi.org/10.1016/S1389-1723(01)80221-8
Chen C, Van Der Borght J, De Vreese R, D’Hooghe M, Soetaert W, Desmet T (2014) Engineering the specificity of trehalose phosphorylase as a general strategy for the production of glycosyl phosphates. Chem Commun 50:7834–7836. https://doi.org/10.1039/c4cc02202e
Curiel JA, Peirotén Á, Landete JM, Ruiz de la Bastida A, Langa S, Arqués JL (2021) Architecture insight of bifidobacterial α-l-fucosidases. Int J Mol Sci 22:8462. https://doi.org/10.3390/ijms22168462
D’Enfert C, Fontaine T (1997) Molecular characterization of the Aspergillus nidulans treA gene encoding an acid trehalase required for growth on trehalose. Mol Microbiol 24:203–216. https://doi.org/10.1046/j.1365-2958.1997.3131693.x
Davies GJ, Wilson KS, Henrissat B (1997) Nomenclature for sugar-binding subsites in glycosyl hydrolases. Biochem J 321:557–559. https://doi.org/10.1042/bj3210557
De Beul E, Jongbloet A, Franceus J, Desmet T (2021) Discovery of a kojibiose hydrolase by analysis of specificity-determining correlated positions in Glycoside Hydrolase family 65. Molecules 26:6321. https://doi.org/10.3390/MOLECULES26206321
Desmet T, Soetaert W (2011) Enzymatic glycosyl transfer: mechanisms and applications. Biocatal Biotransform 29:1–18. https://doi.org/10.3109/10242422.2010.548557
Drula E, Garron M-L, Dogan S, Lombard V, Henrissat B, Terrapon N (2022) The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577. https://doi.org/10.1093/nar/gkab1045
Egloff MP, Uppenberg J, Haalck L, van Tilbeurgh H (2001) Crystal structure of maltose phosphorylase from Lactobacillus brevis: unexpected evolutionary relationship with glucoamylases. Structure 9:689–697. https://doi.org/10.1016/S0969-2126(01)00626-8
Ehrmann MA, Vogel RF (1998) Maltose metabolism of Lactobacillus sanfranciscensis: cloning and heterologous expression of the key enzymes, maltose phosphorylase and phosphoglucomutase. FEMS Microbiol Lett 169:81–86. https://doi.org/10.1111/j.1574-6968.1998.tb13302.x
Franceus J, Desmet T (2020) Sucrose phosphorylase and related enzymes in Glycoside Hydrolase family 13: discovery, application and engineering. Int J Mol Sci 21:2526. https://doi.org/10.3390/ijms21072526
Franceus J, Verhaeghe T, Desmet T (2017) Correlated positions in protein evolution and engineering. J Ind Microbiol Biotechnol 44:687–695. https://doi.org/10.1007/s10295-016-1811-1
Franceus J, Decuyper L, D’hooghe M, Desmet T (2018) Exploring the sequence diversity in glycoside hydrolase family 13_18 reveals a novel glucosylglycerol phosphorylase. Appl Microbiol Biotechnol 102:3183–3191. https://doi.org/10.1007/s00253-018-8856-1
Franceus J, Lormans J, Cools L, D’hooghe M, Desmet T (2021) Evolution of phosphorylases from N-acetylglucosaminide hydrolases in family GH3. ACS Catal 11:6225–6233. https://doi.org/10.1021/ACSCATAL.1C00761
Franceus J, Lormans J, Desmet T (2022) Building mutational bridges between carbohydrate-active enzymes. Curr Opin Biotechnol 78:102804. https://doi.org/10.1016/j.copbio.2022.102804
Franceus J, Rivas-Fernández JP, Lormans J, Rovira C, Desmet T (2024) Evolution of phosphorylase activity in an ancestral glycosyltransferase. ACS Catal 14:3103–3114. https://doi.org/10.1021/acscatal.3c05819
Fujinaka H, Nakamura J, Murase D, Souno H, Kobayashi H (2004) Sugar phosphates and their use for improving the intestinal function. European Patent 1477173A1
Fujinaka H, Nakamura J, Kobayashi H, Takizawa M, Murase D, Tokimitsu I, Suda T (2007) Glucose 1-phosphate increases active transport of calcium in intestine. Arch Biochem Biophys 460:152–160. https://doi.org/10.1016/j.abb.2006.09.006
Gao Y, Saburi W, Taguchi Y, Mori H (2019) Biochemical characteristics of maltose phosphorylase MalE from Bacillus sp. AHU2001 and chemoenzymatic synthesis of oligosaccharides by the enzyme. Biosci Biotechnol Biochem 83:2097–2109. https://doi.org/10.1080/09168451.2019.1634516
Gu J, Xu Y, Nie Y (2023) Role of distal sites in enzyme engineering. Biotechnol Adv 63:108094. https://doi.org/10.1016/j.biotechadv.2023.108094
Hamazaki H, Hamazaki MH (2016) Catalytic site of human protein-glucosylgalactosylhydroxylysine glucosidase: three crucial carboxyl residues were determined by cloning and site-directed mutagenesis. Biochem Biophys Res Commun 469:357–362. https://doi.org/10.1016/j.bbrc.2015.12.005
Hamazaki H, Hotta K (1979) Purification and characterization of an α-glucosidase specific for hydroxylysine-linked disaccharide of collagen. J Biol Chem 254:9682–9687. https://doi.org/10.1016/s0021-9258(19)83570-6
Hidaka M, Honda Y, Kitaoka M, Nirasawa S, Hayashi K, Wakagi T, Shoun H, Fushinobu S (2004) Chitobiose phosphorylase from Vibrio proteolyticus, a member of Glycosyl Transferase family 36, has a clan GH-L-like (α/α)6 barrel fold. Structure 12:937–947. https://doi.org/10.1016/j.str.2004.03.027
Hidaka Y, Hatada Y, Akita M, Yoshida M, Nakamura N, Takada M, Nakakuki T, Ito S, Horikoshi K (2005) Maltose phosphorylase from a deep-sea Paenibacillus sp.: enzymatic properties and nucleotide and amino-acid sequences. Enzyme Microb Technol 37:185–194. https://doi.org/10.1016/j.enzmictec.2005.02.010
Huang Y, Niu B, Gao Y, Fu L, Li W (2010) CD-HIT suite: a web server for clustering and comparing biological sequences. Bioinformatics 26:680–682. https://doi.org/10.1093/BIOINFORMATICS/BTQ003
Hüwel S, Haalck L, Conrath N, Spener F (1997) Maltose phosphorylase from Lactobacillus brevis: purification, characterization, and application in a biosensor for ortho-phosphate. Enzyme Microb Technol 21:413–420. https://doi.org/10.1016/S0141-0229(97)00014-8
Inoue Y, Ishii K, Tomita T, Yatake T, Fukui F (2002a) Characterization of trehalose phosphorylase from Bacillus stearothermophilus SK-1 and nucleotide sequence of the corresponding gene. Biosci Biotechnol Biochem 66:1835–1843. https://doi.org/10.1271/bbb.66.1835
Inoue Y, Yasutake N, Oshima Y, Yamamoto Y, Tomita T, Miyoshi S, Yatake T (2002b) Cloning of the maltose phosphorylase gene from Bacillus sp. strain RK-1 and efficient production of the cloned gene and the trehalose phosphorylase gene from Bacillus stearothermophilus SK-1 in Bacillus subtilis. Biosci Biotechnol Biochem 66:2594–2599. https://doi.org/10.1271/bbb.66.2594
Isono N, Yagura S, Yamanaka K, Masuda Y, Mukai K, Katsuzaki H (2023) Enzymatic synthesis of β-d-fructofuranosyl α-d-glucopyranosyl-(1→2)-α-d-glucopyranoside using Escherichia coli glycoside phosphorylase YcjT. Biosci Biotechnol Biochem 87:1249–1253. https://doi.org/10.1093/bbb/zbad099
Jung JH, Seo DH, Holden JF, Park CS (2014) Identification and characterization of an archaeal kojibiose catabolic pathway in the hyperthermophilic Pyrococcus sp. strain ST04. J Bacteriol 196:1122–1131. https://doi.org/10.1128/JB.01222-13
Kabisch A, Otto A, König S, Becher D, Albrecht D, Schüler M, Teeling H, Amann RI, Schweder T (2014) Functional characterization of polysaccharide utilization loci in the marine Bacteroidetes ‘Gramella Forsetii’ KT0803. ISME J 8:1492–1502. https://doi.org/10.1038/ismej.2014.4
Kamogawa A, Yokobayashi K, Fukui T (1973) Properties of maltose phosphorylase from Lactobacillus brevis Agric Biol Chem 37:2813–2819. https://doi.org/10.1271/bbb1961.37.2813
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166. https://doi.org/10.1093/bib/bbx108
Kido Y, Maeno S, Tanno H, Kichise Y, Shiwa Y, Endo A (2021) Niche-specific adaptation of Lactobacillus helveticus strains isolated from malt whisky and dairy fermentations. Microb Genomics 7:000560. https://doi.org/10.1099/mgen.0.000560
Kuhaudomlarp S, Pergolizzi G, Patron NJ, Henrissat B, Field RA (2019) Unraveling the subtleties of β-(1→3)-glucan phosphorylase specificity in the GH94, GH149, and GH161 glycoside hydrolase families. J Biol Chem 294:6483–6493. https://doi.org/10.1074/jbc.RA119.007712
La Rosa SL, Leth ML, Michalak L, Hansen ME, Pudlo NA, Glowacki R, Pereira G, Workman CT, Arntzen MØ, Pope PB, Martens EC, Hachem MA, Westereng B (2019) The human gut Firmicute Roseburia intestinalis is a primary degrader of dietary β-mannans. Nat Commun 10:905. https://doi.org/10.1038/s41467-019-08812-y
Letunic I, Bork P (2021) Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296. https://doi.org/10.1093/NAR/GKAB301
Li A, Laville E, Tarquis L, Lombard V, Ropartz D, Terrapon N, Henrissat B, Guieysse D, Esque J, Durand J, Morgavi DP, Potocki-Veronese G (2020) Analysis of the diversity of the glycoside hydrolase family 130 in mammal gut microbiomes reveals a novel mannoside-phosphorylase function. Microb Genomics 6:1–14. https://doi.org/10.1099/mgen.0.000404
Li A, Benkoulouche M, Ladeveze S, Durand J, Cioci G, Laville E, Potocki-Veronese G (2022) Discovery and biotechnological exploitation of glycoside-phosphorylases. Int J Mol Sci 23:3043. https://doi.org/10.3390/ijms23063043
Liu Y, Wang Z, Yin Y, Cao Y, Zhao H, Xia Y (2007) Expression, purification, and characterization of recombinant Metarhizium anisopliae acid trehalase in Pichia pastoris. Protein Expr Purif 54:66–72. https://doi.org/10.1016/j.pep.2007.02.016
Liu N, Fosses A, Kampik C, Parsiegla G, Denis Y, Vita N, Fierobe H-P, Perret S (2019) In vitro and in vivo exploration of the cellobiose and cellodextrin phosphorylases panel in Ruminiclostridium cellulolyticum: implication for cellulose catabolism. Biotechnol Biofuels 12:208. https://doi.org/10.1186/s13068-019-1549-x
Luley-Goedl C, Nidetzky B (2010) Carbohydrate synthesis by disaccharide phosphorylases: reactions, catalytic mechanisms and application in the glycosciences. Biotechnol J 5:1324–1338. https://doi.org/10.1002/biot.201000217
Maruta K, Mukai K, Yamashita H, Kubota M, Chaen H, Fukuda S, Kurimoto M (2002) Gene encoding a trehalose phosphorylase from Thermoanaerobacter brockii ATCC 35047. Biosci Biotechnol Biochem 66:1976–1980. https://doi.org/10.1271/bbb.66.1976
Maruta K, Watanabe H, Nishimoto T, Kubota M, Chaen H, Fukuda S, Kurimoto M, Tsujisaka Y (2006) Acceptor specificity of trehalose phosphorylase from Thermoanaerobacter brockii: production of novel nonreducing trisaccharide, 6-O-α-d-galactopyranosyl trehalose. J Biosci Bioeng 101:385–390. https://doi.org/10.1263/JBB.101.385
Mittenbuhler K, Holzer H (1988) Purification and characterization of acid trehalase from the yeast suc2 mutant. J Biol Chem 263:8537–8543. https://doi.org/10.1016/s0021-9258(18)68511-4
Miyazaki T, Park EY (2020) Structure–function analysis of silkworm sucrose hydrolase uncovers the mechanism of substrate specificity in GH13 subfamily 17 exo-α-glucosidases. J Biol Chem 295:8784–8797. https://doi.org/10.1074/jbc.RA120.013595
Miyazaki T, Tanaka H, Nakamura S, Dohra H, Funane K (2022) Identification and characterization of dextran α-1,2-debranching enzyme from Microbacterium dextranolyticum. J Appl Glycosci 70:15–24. https://doi.org/10.5458/jag.jag.JAG-2022_0013
Mokhtari A, Blancato VS, Repizo GD, Henry C, Pikis A, Bourand A, de Fátima Álvarez M, Immel S, Mechakra-Maza A, Hartke A, Thompson J, Magni C, Deutscher J (2013) Enterococcus faecalis utilizes maltose by connecting two incompatible metabolic routes via a novel maltose 6′-phosphate phosphatase (MapP). Mol Microbiol 88:234–253. https://doi.org/10.1111/mmi.12183
Mukherjee K, Narindoshvili T, Raushel FM (2018) Discovery of a kojibiose phosphorylase in Escherichia coli K-12. Biochemistry 57:2857–2867. https://doi.org/10.1021/acs.biochem.8b00392
Murao S, Nagano H, Ogura S, Nishino T (1985) Enzymatic synthesis of trehalose from maltose. Agric Biol Chem 49:2113–2118. https://doi.org/10.1271/bbb1961.49.2113
Nakai H, Baumann MJ, Petersen BO, Westphal Y, Schols H, Dilokpimol A, Hachem MA, Lahtinen SJ, Duus JØ, Svensson B (2009) The maltodextrin transport system and metabolism in Lactobacillus acidophilus NCFM and production of novel α-glucosides through reverse phosphorolysis by maltose phosphorylase. FEBS J 276:7353–7365. https://doi.org/10.1111/j.1742-4658.2009.07445.x
Nakai H, Dilokpimol A, Hachem MA, Svensson B (2010a) Efficient one-pot enzymatic synthesis of α-(1→4)-glucosidic disaccharides through a coupled reaction catalysed by Lactobacillus acidophilus NCFM maltose phosphorylase. Carbohydr Res 345:1061–1064. https://doi.org/10.1016/j.carres.2010.03.021
Nakai H, Petersen BO, Westphal Y, Dilokpimol A, Abou Hachem M, Duus JØ, Schols HA, Svensson B (2010b) Rational engineering of Lactobacillus acidophilus NCFM maltose phosphorylase into either trehalose or kojibiose dual specificity phosphorylase. Protein Eng Des Sel 23:781–787. https://doi.org/10.1093/protein/gzq055
Nakamura S, Nihira T, Kurata R, Nakai H, Funane K, Park EY, Miyazaki T (2021) Structure of a bacterial α-1,2-glucosidase defines mechanisms of hydrolysis and substrate specificity in GH65 family hydrolases. J Biol Chem 297:101366. https://doi.org/10.1016/J.JBC.2021.101366
Nakamura S, Kurata R, Miyazaki T (2024) Structural insights into α-(1→6)-linkage preference of GH97 glucodextranase from Flavobacterium johnsoniae FEBS J. https://doi.org/10.1111/febs.17139
Nihira T, Nakai H, Chiku K, Kitaoka M (2012a) Discovery of nigerose phosphorylase from Clostridium phytofermentans Appl Microbiol Biotechnol 93:1513–1522. https://doi.org/10.1007/s00253-011-3515-9
Nihira T, Nakai H, Kitaoka M (2012b) 3-O-α-d-glucopyranosyl-l-rhamnose phosphorylase from Clostridium phytofermentans Carbohydr Res 350:94–97. https://doi.org/10.1016/j.carres.2011.12.019
Nihira T, Saito Y, Kitaoka M, Otsubo K, Nakai H (2012c) Identification of Bacillus selenitireducens MLS10 maltose phosphorylase possessing synthetic ability for branched α-d-glucosyl trisaccharides. Carbohydr Res 360:25–30. https://doi.org/10.1016/j.carres.2012.07.014
Nihira T, Saito Y, Chiku K, Kitaoka M, Ohtsubo K, Nakai H (2013) Potassium ion-dependent trehalose phosphorylase from halophilic Bacillus selenitireducens MLS10. FEBS Lett 587:3382–3386. https://doi.org/10.1016/j.febslet.2013.08.038
Nihira T, Miyajima F, Chiku K, Nishimoto M, Kitaoka M, Ohtsubo K, Nakai H (2014a) One pot enzymatic production of nigerose from common sugar resources employing nigerose phosphorylase. J Appl Glycosci 61:75–80. https://doi.org/10.5458/jag.jag.JAG-2013_012
Nihira T, Nishimoto M, Nakai H, Ohtsubo K, Kitaoka M (2014b) Characterization of two α-1,3-glucoside phosphorylases from Clostridium phytofermentans. J Appl Glycosci 61:59–66. https://doi.org/10.5458/jag.jag.JAG-2013_013
Nihira T, Saito Y, Ohtsubo K, Nakai H, Kitaoka M (2014c) 2-O-α-d-Glucosylglycerol phosphorylase from Bacillus selenitireducens MLS10 possessing hydrolytic activity on β-d-glucose 1-phosphate. PLoS ONE 9:e86548. https://doi.org/10.1371/journal.pone.0086548
Okada S, Yamamoto T, Watanabe H, Nishimoto T, Chaen H, Fukuda S, Wakagi T, Fushinobu S (2014) Structural and mutational analysis of substrate recognition in kojibiose phosphorylase. FEBS J 281:778–786. https://doi.org/10.1111/febs.12622
Osuna S (2021) The challenge of predicting distal active site mutations in computational enzyme design. WIREs Comput Mol Sci 11:e1502. https://doi.org/10.1002/wcms.1502
Parish CR, Cowden WB, Willenborg DO (1992) Phosphosugar-based anti-inflammatory and/or immunosuppressive drugs. US005506210A
Pedreño Y, Maicas S, Argüelles JC, Sentandreu R, Valentin E (2004) The ATC1 gene encodes a cell wall-linked acid trehalase required for growth on trehalose in Candida albicans J Biol Chem 279:40852–40860. https://doi.org/10.1074/jbc.M400216200
Plante OJ, Andrade RB, Seeberger PH (1999) Synthesis and use of glycosyl phosphates as glycosyl donors. Org Lett 1:211–214. https://doi.org/10.1021/ol9905452
Price MN, Dehal PS, Arkin AP (2010) FastTree 2 - approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:W320–W324. https://doi.org/10.1093/nar/gku316
Ronchera-Oms CL, Jiménez NV, Peidro J (1995) Stability of parenteral nutrition admixtures containing organic phosphates. Clin Nutr 14:373–380. https://doi.org/10.1016/S0261-5614(95)80055-7
Sakaguchi M (2020) Diverse and common features of trehalases and their contributions to microbial trehalose metabolism. Appl Microbiol Biotechnol 104:1837–1847. https://doi.org/10.1007/s00253-019-10339-7
Sánchez-Fresneda R, Martínez-Esparza M, Maicas S, Argüelles J-C, Valentín E (2014) In Candida parapsilosis the ATC1 gene encodes for an acid trehalase involved in trehalose hydrolysis, stress resistance and virulence. PLoS One 9:e99113. https://doi.org/10.1371/journal.pone.0099113
Sánchez-Fresneda R, Guirao-Abad JP, Martinez-Esparza M, Maicas S, Valentín E, Argüelles J-C (2015) Homozygous deletion of ATC1 and NTC1 genes in Candida parapsilosis abolishes trehalase activity and affects cell growth, sugar metabolism, stress resistance, infectivity and biofilm formation. Fungal Genet Biol 85:45–57. https://doi.org/10.1016/j.fgb.2015.10.007
Sato S, Fan P-H, Yeh Y-C, Liu H (2024) Complete in vitro reconstitution of the apramycin biosynthetic pathway demonstrates the unusual incorporation of a β-d-sugar nucleotide in the final glycosylation step. J Am Chem Soc 146:10103–10114. https://doi.org/10.1021/jacs.4c01233
Sernee MF, Ralton JE, Nero TL, Sobala LF, Kloehn J, Vieira-Lara MA, Cobbold SA, Stanton L, Pires DEV, Hanssen E, Males A, Ward T, Bastidas LM, Van Der Peet PL, Parker MW, Ascher DB, Williams SJ, Davies GJ, McConville MJ (2019) A family of dual-activity glycosyltransferase-phosphorylases mediates mannogen turnover and virulence in Leishmania Parasites. Cell Host Microbe 26:385-399e9. https://doi.org/10.1016/j.chom.2019.08.009
Shrestha P, Karmacharya J, Han S-R, Lee JH, Oh T-J (2023) Elucidation of bacterial trehalose-degrading trehalase and trehalose phosphorylase: physiological significance and its potential applications. Glycobiology 00:1–13. https://doi.org/10.1093/glycob/cwad084
Sigg A, Klimacek M, Nidetzky B (2024) Pushing the boundaries of phosphorylase cascade reaction for cellobiose production I: kinetic model development. Biotechnol Bioeng 121:580–592. https://doi.org/10.1002/bit.28602
Sun S, You C (2021) Disaccharide phosphorylases: structure, catalytic mechanisms and directed evolution. Synth Syst Biotechnol 6:23–31. https://doi.org/10.1016/j.synbio.2021.01.004
Taguchi Y, Saburi W, Imai R, Mori H (2017) Evaluation of acceptor selectivity of Lactococcus lactis ssp. lactis trehalose 6-phosphate phosphorylase in the reverse phosphorolysis and synthesis of a new sugar phosphate. Biosci Biotechnol Biochem 81:1512–1519. https://doi.org/10.1080/09168451.2017.1329620
Taguchi Y, Saburi W, Imai R, Mori H (2020) Efficient one-pot enzymatic synthesis of trehalose 6-phosphate using GH65 α-glucoside phosphorylases. Carbohydr Res 488:107902. https://doi.org/10.1016/j.carres.2019.107902
Thibodeaux CJ, Melançon CE III, Liu H (2008) Natural-product sugar biosynthesis and enzymatic glycodiversification. Angew Chem Int Ed 47:9814–9859. https://doi.org/10.1002/anie.200801204
Touhara KK, Nihira T, Kitaoka M, Nakai H, Fushinobu S (2014) Structural basis for reversible phosphorolysis and hydrolysis reactions of 2-O-α-glucosylglycerol phosphorylase. J Biol Chem 289:18067–18075. https://doi.org/10.1074/jbc.M114.573212
Van der Borght J, Desmet T, Soetaert W (2010) Enzymatic production of β-d-glucose-1-phosphate from trehalose. Biotechnol J 5:986–993. https://doi.org/10.1002/biot.201000203
Van der Borght J, Chen C, Hoflack L, Van Renterghem L, Desmet T, Soetaert W (2011) Enzymatic properties and substrate specificity of the trehalose phosphorylase from Caldanaerobacter subterraneus Appl Environ Microbiol 77:6939–6944. https://doi.org/10.1128/AEM.05190-11
Van der Borght J, Soetaert W, Desmet T (2012) Engineering the acceptor specificity of trehalose phosphorylase for the production of trehalose analogs. Biotechnol Prog 28:1257–1262. https://doi.org/10.1002/btpr.1609
Van Hoorebeke A, Stout J, Van der Meeren R, Kyndt J, Van Beeumen J, Savvides SN (2010) Crystallization and X-ray diffraction studies of inverting trehalose phosphorylase from Thermoanaerobacter Sp. Acta Crystallograph Sect F Struct Biol Cryst Commun 66:442–447. https://doi.org/10.1107/S1744309110005749
Weinstock KG, Bush D (1999) Nucleic acid sequences relating to Candida albicans for diagnostics and therapeutics. US Patent 6747137B1
Wilding M, Hong N, Spence M, Buckle AM, Jackson CJ (2019) Protein engineering: the potential of remote mutations. Biochem Soc Trans 47:701–711. https://doi.org/10.1042/BST20180614
Yamamoto T, Maruta K, Mukai K, Yamashita H, Nishimoto T, Kubota M, Fukuda S, Kurimoto M, Tsujisaka Y (2004) Cloning and sequencing of kojibiose phosphorylase gene from Thermoanaerobacter brockii ATCC35047. J Biosci Bioeng 98:99–106. https://doi.org/10.1016/s1389-1723(04)70249-2
Yamamoto T, Watanabe H, Nishimoto T, Aga H, Kubota M, Chaen H, Fukuda S (2006a) Acceptor recognition of kojibiose phosphorylase from Thermoanaerobacter brockii: syntheses of glycosyl glycerol and myo-inositol. J Biosci Bioeng 101:427–433. https://doi.org/10.1263/jbb.101.427
Yamamoto T, Yamashita H, Mukai K, Watanabe H, Kubota M, Chaen H, Fukuda S (2006b) Construction and characterization of chimeric enzymes of kojibiose phosphorylase and trehalose phosphorylase from Thermoanaerobacter brockii Carbohydr Res 341:2350–2359. https://doi.org/10.1016/j.carres.2006.06.024
Yamamoto T, Nishio-Kosaka M, Izawa S, Aga H, Nishimoto T, Chaen H, Fukuda S (2011) Enzymatic properties of recombinant kojibiose phosphorylase from Caldicellulosiruptor saccharolyticus ATCC43494. Biosci Biotechnol Biochem 75:1208–1210. https://doi.org/10.1271/bbb.110116
Yang J, Hoffmeister D, Liu L, Fu X, Thorson JS (2004) Natural product glycorandomization. Bioorg Med Chem 12:1577–1584. https://doi.org/10.1016/j.bmc.2003.12.046
Yoshida M, Nakamura N, Horikoshi K (1998) Production of trehalose by a dual enzyme system of immobilized maltose phosphorylase and trehalose phosphorylase. Enzyme Microb Technol 22:71–75. https://doi.org/10.1016/S0141-0229(97)00132-4
Zhang T, Yang J, Tian C, Ren C, Chen P, Men Y, Sun Y (2020) High-yield biosynthesis of glucosylglycerol through coupling phosphorolysis and transglycosylation reactions. J Agric Food Chem 68:15249–15256. https://doi.org/10.1021/acs.jafc.0c04851
Zilli DMW, Lopes RG, Alves SL, Barros LM, Miletti LC, Stambuk BU (2015) Secretion of the acid trehalase encoded by the CgATH1 gene allows trehalose fermentation by Candida Glabrata Microbiol Res 179:12–19. https://doi.org/10.1016/j.micres.2015.06.008
Funding
This work was supported by Research Foundation Flanders (FWO), grant number 1S17721N (doctoral scholarship for EDB).
Author information
Authors and Affiliations
Contributions
Conceptualization: EDB and JF; writing – original draft preparation: EDB; writing – review and editing: JF and TD; visualization: EDB and JF; supervision: TD. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(PDF 4.26 MB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
De Beul, E., Franceus, J. & Desmet, T. The many functions of carbohydrate-active enzymes in family GH65: diversity and application. Appl Microbiol Biotechnol 108, 476 (2024). https://doi.org/10.1007/s00253-024-13301-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13301-4