Abstract
Chemoautotrophic canonical ammonia oxidizers (ammonia-oxidizing archaea (AOA) and ammonia-oxidizing bacteria (AOB)) and complete ammonia oxidizers (comammox Nitrospira) are accountable for ammonia oxidation, which is a fundamental process of nitrification in terrestrial ecosystems. However, the relationship between autotrophic nitrification and the active nitrifying populations during 15N-urea incubation has not been totally clarified. The 15N-labeled DNA stable isotope probing (DNA-SIP) technique was utilized in order to study the response from the soil nitrification process and the active nitrifying populations, in both acidic and neutral paddy soils, to the application of urea. The presence of C2H2 almost completely inhibited NO3−-N production, indicating that autotrophic ammonia oxidation was dominant in both paddy soils. 15N-DNA-SIP technology could effectively distinguish active nitrifying populations in both soils. The active ammonia oxidation groups in both soils were significantly different, AOA (NS (Nitrososphaerales)-Alpha, NS-Gamma, NS-Beta, NS-Delta, NS-Zeta and NT (Ca. Nitrosotaleales)-Alpha), and AOB (Nitrosospira) were functionally active in the acidic paddy soil, whereas comammox Nitrospira clade A and Nitrosospira AOB were functionally active in the neutral paddy soil. This study highlights the effective discriminative effect of 15N-DNA-SIP and niche differentiation of nitrifying populations in these paddy soils.
Key points
• 15N-DNA-SIP technology could effectively distinguish active ammonia oxidizers.
• Comammox Nitrospira clade A plays a lesser role than canonical ammonia oxidizers.
• The active groups in the acidic and neutral paddy soils were significantly different.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chemoautotrophic nitrifying microorganisms are key players in the global nitrogen cycle (Wang et al. 2020; Sun et al. 2022), and they were traditionally thought to involve ammonia-oxidizing bacteria (AOB) and ammonia-oxidizing archaea (AOA), together with nitrite-oxidizing bacteria (NOB) (Li et al. 2019; Lin et al. 2022; He et al. 2022). With the development of research, the finding of complete ammonia oxidizers (comammox Nitrospira), which can catalyze ammonia to nitrate in an independent microorganism, has demonstrated how they can be participants with autotrophic ammonia-oxidizers (Wang et al. 2019a; Li et al. 2019). Up to now, all given comammox Nitrospira belong to the sub-lineage II of the Nitrospira (Daims et al. 2015; He et al. 2022). Comammox Nitrospira can in turn be categorized into two divergent clades, namely clades A and B (Daims et al. 2015; van Kessel et al. 2015), while clade A is further classified into clades A.1, A.2.1, A.2.2, and A.3 (Li et al. 2021a; Sun et al. 2021b; He et al. 2022).
AOA and AOB are widely distributed in all kinds of habitats, which include soils (Ali et al. 2013; Tao et al. 2017), oceans (Wang et al. 2017; Qin et al. 2020), and freshwater (Park et al. 2022; Ren and Wang 2022) and play important roles in global nitrogen and carbon cycles. AOA contain four basal lineages, Ca. Nitrosocaldales (NC), Nitrososphaerales (NS), Ca. Nitrosotaleales (NT), and Nitrosopumilales (NP) (Alves et al. 2018). In soil, the activity of AOA and AOB is often linked to pH (Hu et al. 2014; Tago et al. 2015; Ying et al. 2017), perhaps because AOA are oligotrophic microorganisms that prefer acidic conditions (Wu et al. 2017; He et al. 2021), whereas AOB preferred soils with pH higher than 7 (Shen et al. 2012). The 13C-DNA-SIP technique is often adopted to study the activity of autotrophic ammonia oxidation (Pratscher et al. 2011; Xia et al. 2011; Pan et al. 2018). AOA, AOB, and comammox Nitrospira (nitrifying populations) exhibit differentiated activities (Wang et al. 2019a; Liu et al. 2021). Nevertheless, the technical limitation of 13C-DNA-SIP is to determine the autotrophic carbon-fixing growth of nitrifying populations by assimilation of 13C (Wu et al. 2013; Pornkulwat et al. 2018; Zhao et al. 2020). Hence, the 13C-DNA-SIP technique cannot directly determine the incorporation of N into the genome, although the changes in ammonia and nitrate concentrations are measured in order to characterize ammonia oxidation activity in such experiments (Zhang et al. 2010, 2018; Dong et al. 2019). An alternative technique is DNA-SIP based on 15N-substrates, which has employed isotopically labeled N incorporation into the genome to more directly determine active autotrophic microorganisms (Angel et al. 2018; Zhang et al. 2021).
The versatility of metabolism allows microorganisms to adapt to changes in substrate content or pH (Daims and Wagner 2018). Moreover, recent studies have shown that comammox Nitrospira responds positively to an increase in nitrogen substrates (Li et al. 2019, 2020; Osburn and Barrett 2020; Takahashi et al. 2020). Likewise, not all AOA were limited by high ammonia concentrations, and some AOA like Nitrosocosmicus AOA are similar to AOB and have been demonstrated to tolerate very high ammonium concentrations (Jung et al. 2021). In addition, kinetic analysis has shown that terrestrial AOA and AOB have similar ammonia affinities (Bello et al. 2021), which may be the cause for the vitality of AOB in acidic soils. More and more evidence has shown that nitrification in acidic soils is chiefly carried out by AOA (Gubry-Rangin et al. 2010; Zhang et al. 2012; Lu et al. 2012; Lu and Jia 2013). However, there are also AOB in acidic soils and the mechanism of their growth has been illustrated in low pH soils (Zhang et al. 2017; Huang et al. 2018; Séneca et al. 2020; Picone et al. 2021). In the presence of nitrification inhibitors (NIs) like C2H2, reducing competition for ammonia may also enhance any selective advantage provided by high ammonium (Ye et al. 2018).
The purpose of this experimentation was to investigate the relative contribution of nitrifying populations to nitrification, as well as their ecological niches in fertilized paddy soils. We set up a laboratory microcosm experiment to (1) measure the responses of nitrifying populations to the addition of nitrogen fertilizer and the nitrification inhibitor (C2H2) in both acidic and neutral paddy soils, (2) estimate the incorporation of 15N into the genome DNA of nitrifying population, and (3) analyze the active communities of 15N-incorporating nitrifiers.
Materials and methods
Soil sites
The acidic paddy soil (pHH2O 5.19) was collected in Yingtan (116° 55′ E, 28° 12′ N), Jiangxi Province, China. The mean annual precipitation (MAP) of the acidic paddy field is 1785 mm, and the mean average temperature (MAT) is 17.8 °C. The neutral paddy soil (pHH2O 7.09) was sampled in Baoshan (99° 15′ E, 25° 09′ N), Yunnan Province, China. The MAP of the neutral paddy field is 1000 mm, and the MAT is 16.0 °C. The soil samples were sampled from 0 to 20 cm using eight random soil cores (diam. 10 cm) and mixed into composite samples. Then, they were transported to the laboratory on ice for subsequent research and analysis. The determination methods for basic soil physicochemical properties were as described in previous studies (Li et al. 2021a, b; Yu et al. 2022; Zhang et al. 2022) and given in Table 1.
Soil incubation
Four treatments were set up: (1) control without adding any substrate, called CK; (2) 14N-urea treatment to provide key confirmation of active organisms in the 15N-treated samples; (3) 15N-urea treatment with 15N-labeled urea (99 atom%; Cambridge Isotope Laboratories, Inc., Andover, MA, USA); and (4) 15N-labeled urea and C2H2 treatment with 15N-labeled urea and 0.1% (v/v) C2H2. In each microcosm incubation system, 20 g of dry-weight soil (DWS) was placed in brown serum bottles (120 mL), equipped with rubber stoppers and aluminum caps for easy sealing. To avoid ammonia loss brought by the rapid hydrolysis of urea, urea was added four times, with each addition being 100 mg kg−1 N. Each treatment was sampled at 0 and 28 days, with three replicates per treatment. Aerobic conditions were maintained by opening the rubber plugs and replacing fresh air every 3 days. Incubation was continued for 28 days in a dark incubator at 25 °C. On the day of sampling, the soil was taken to detect the ammonium (NH4+-N) and nitrate (NO3−-N) contents. The soil NH4+-N and NO3−-N contents were determined by a continuous flow injection analyzer (FLA star 5000 Analyzer, Foss, Denmark) extracted by 1 mol L−1 KCl (soil/KCl, 1:10).
Nucleic acid extraction and SIP fractionation
After freeze-drying, about 0.5 g of soil was taken to extract genomic DNA using the Fast DNA SPIN Kit for Soil (MP Biomedicals, Santa Ana, CA, USA), then immediately aliquoted and placed at − 20 °C for subsequent experimentations.
In order to follow the smooth progress of the 15N-DNA-SIP experiment, the initial values of soil 15N (acidic paddy soil: 18.92, neutral paddy soil 9.08) were measured by isotope ratio mass spectrometer (Thermo Fisher Scientific, USA). The steps of DNA-SIP were implemented using a previously described method with improvements (Luo et al. 2021). Simply put, 3 µg DNA was added to the liquid mixture which was mixed with CsCl solution and gradient buffer. The liquid mixture was regulated to a refractive index of 1.4015 and centrifuged at 182,742 × g (45,000 rpm) for 48 h at 20 °C (Luo et al. 2021). After centrifugation, 16 × 330 µL was collected into 1.5 mL centrifuge tubes. Next, 100 µL liquor in each centrifuge tube was measured, in order to determine the buoyant densities.
The purification steps of each fraction of DNA were as follows: first, 550 µL PEG6000 (300 g polyethylene glycol 6000, 93.6 g NaCl) was added, heated at 37 °C for 1 h or placed at 25 °C for 2 h, then centrifuged at 13,000 × g for 0.5 h, removed the supernatant, and cleaned with 0.5 mL 70% alcohol twice; in the last procedure of fractionation, the purified DNA was dissolved in 30 µL of nuclease-free water.
Quantitative PCR (qPCR) analysis
QPCR was established on a Light Cycler 480 real-time PCR detection system (Roche480, USA) (Meng et al. 2020). The 515F/907R primer (Li et al. 2021a, b) was adopted for the bacterial 16S ribosomal(r) RNA gene, the CamoA-23F/616R (Meng et al. 2020) was adopted for the AOA amoA gene, and the amoA1F/2R (Meng et al. 2020) was adopted for AOB amoA gene. The six mixed primers comaA-244f_a-f/659r_a-f and comaB-244f_a-f/659r_a-f (Pjevac et al. 2017) were adopted for the comammox Nitrospira clade A (ComA) and comammox Nitrospira clade B (ComB) amoA genes, respectively (Table S1). The qPCR mixtures were 20 µL, containing 10 µL 2 × GoTaq® qPCR Master Mix (Promega), 0.5 M of each primer, and 2 µL of tenfold-diluted DNA template, and the part less than 20 µL was filled with nuclease-free water. The absolute quantitative PCR amplification conditions were as follows: The universal 16S rRNA gene was 95 °C for 3 min, followed by 35 cycles of 95 °C for 45 s, 56 °C for 45 s, and then 72 °C elongation for 5 min. The AOA were 95 °C for 2 min, followed by 40 cycles of 95 °C for 20 s, 55 °C for 20 s, and then 80 °C elongation for 30 s. The AOB were 95 °C for 2 min, followed by 40 cycles of 95 °C for 20 s, 57 °C for 30 s, and then 72 °C elongation for 30 s. ComA and ComB were 95 °C for 10 min, followed by 40 cycles of 94 °C for 30 s, 52 °C for 45 s, and then 72 °C elongation for 1 min. For qPCR, nuclease-free water was used as a negative control, the standard curve was established using a tenfold serial dilution of standard plasmid DNA, the amplification efficiencies were 93% to 99%, and the R2 was higher than 0.99.
MiSeq sequencing and bioinformation
The microbial communities were paired-end amplicon sequenced, except for the amoA gene of AOA which used the forward-sequence because of it being lower than 635 bp; others used the data of paired-end amplicon sequence. The Illumina MiSeq (PE300) platform (Illumina, San Diego, USA) was run by Guangdong Magigene Biotechnology Co., Ltd. (Guangdong, China). In order to denoise (error-correcting) Illumina amplicon reads into unique sequences (zero-radius operational taxonomic units; Zotus), the UNOISE3 algorithm in USEARCH, which was updated from the UNOISE2 algorithm (Edgar 2016), was used, and the sequences are taxonomically classified. Sequence reads are archived in the NCBI Sequence Read Archive, with the Bioproject number PRJNA880760.
Data analysis
The data of the nitrifying populations’ abundance was transformed to Log10 (base 10). In order to assess the differences, a one-way ANOVA (analysis of variance) was implemented. Graphs and histograms were drawn using the Origin 2017 software (Origin Lab Inc., USA). Phylogenetic analysis used the MEGA X version (Kumar et al. 2018). Bootstraps were based on 1000 replicated trees.
Results
Nitrification activity
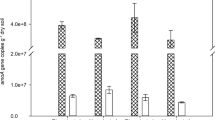
Soil nitrification was evaluated by the alterations in NH4+-N and NO3−-N contents (Fig. 1). The net nitrification rates for both soils were 2.10 and 5.24 mg kg−1 DWS day−1, respectively (Table 2). Urea fertilization caused a huge accumulation of NH4+-N in both soils (Fig. 1a). C2H2 additions significantly increased NH4+-N concentrations (P < 0.001) compared with the CK (Fig. 1a), Additionally, treated with urea alone was higher than the treated with urea and C2H2 at the significance level of 5% (Fig. 1a). Urea fertilization resulted in significant accumulation of NO3−-N in both soils (P < 0.001) with urea treatment (Fig. 1b). In both soils with added C2H2 inhibitor, NO3−-N concentrations were lower than those under urea alone at the significance level of 5% (Fig. 1b). The NH4+-N accumulation and NO3−-N production under C2H2 treatment indicated that autotrophic ammonia oxidation dominated in the both soils.
NH4+-N (a) and soil NO3−-N (b) concentrations following the microcosm incubation of the acidic and neutral paddy soils. Error bars indicate standard errors of triplicate samples. The urea treatment represents both 14N-urea and 15N-urea treatments as there was no difference between the two treatments
Changes in the abundance
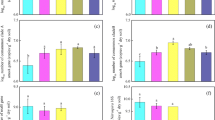
qPCR of the amoA genes was performed to count the population abundance of AOA, AOB, ComA, and ComB (Fig. 2). The percentage of comammox Nitrospira in the nitrifying population (mean value: 44.3%) was lower than that of the canonical ammonia oxidizers, except for the treatment with C2H2 inhibitor in both soils (Fig. 2). On the contrary, the percentage of ComA and ComB in the nitrifying population in the C2H2 treatment (the highest value: 83.5%) was higher than that of AOA and AOB, due to the activities of the latter being significantly inhibited (Fig. 2).
Compared with the CK, the utilization of urea increased the AOA abundance in the acidic soil (P < 0.05). However, the AOA abundance in the neutral soil was significantly decreased (P < 0.05) (Fig. 2a). The existence of C2H2 inhibited the abundance in both soils, compared with the CK (Fig. 2a). Compared with the CK, the AOB abundance was enhanced by adding urea in both soils (Fig. 2b). Furthermore, the existence of C2H2 inhibited the abundance of AOB (Fig. 2b). Comammox Nitrospira abundances in the neutral soil were not significantly impacted by urea addition (Fig. 2c and d). However, C2H2 caused a significant increase in the ComA abundance (P < 0.001), possibly due to the activities of canonical ammonia oxidizers being significantly inhibited, while the ComB was unaffected (Fig. 2c and d).
15N-labeling ofactive ammonia oxidizers
In order to distinguish the 15N-labeled DNA from the 14N-DNA, the activity of nitrifying populations was assessed by the quantification of amoA genes, which was isolated by density gradient centrifugation (Fig. 3). For the acidic soil, the peaks of AOA and AOB appeared in the 15N-labeled DNA ‘heavy fractions (HF)’ (fractions 7–9 and 6–7, respectively) from the 15N-urea microcosms when compared to those from the 14N-urea microcosms (Fig. 3a and c). For the neutral soil, the peaks of AOB and ComA also occurred in the 15N-labeled DNA “HF” (fractions 8–10 and 9–11, respectively) from the 15N-urea microcosms when compared to those from 14N-urea microcosms (Fig. 3b and. d). Although the active 15N-AOA appeared in DNA fractions 7–9, whereas the active 15N-AOB appeared in DNA fractions 6–7, the peak in HF for both occurred at a buoyant density (BD) of 1.7243 g mL−1 (Fig. 3a and c). Similarly, the active 15N-AOB appeared in DNA fractions 8–10, whereas active 15N-ComA appeared in DNA fractions 9–11; not only AOB but also ComA peaked at a BD of 1.7162 g mL−1 (Fig. 3b and d).
Quantitative PCR of gene copy numbers in the acidic soil of AOB (a) and AOA (c) and in the neutral soil of AOB (b) and comammox Nitrospira clade A (d) across the entire buoyant density gradient of the fractions from soil incubated with 14/15N-labeled urea at 28 days of incubation. The normalized data are the ratios of the gene copy number in each DNA gradient fraction to the maximum quantities from each treatment. Error bars show the standard errors of the triplicate microcosms
In the acidic soil, the percentage of gene copy number in the AOB and AOA HF to the sum in all 16 DNA gradient fractions was 90.6% and 87.1%, respectively (Table 2). In the neutral soil, the percentage of gene copy number in the AOB and AOA “heavy” DNA fractions to the sum in all sixteen DNA gradient fractions was 93.0% and 92.7%, respectively (Table 2). In the acidic soil, 15N-labeled AOB and AOA abundances were 3.15 × 107 and 2.13 × 108 copies g−1 DWS, respectively (Table 2). In the neutral soil, 15N-labeled AOB and ComA abundances were 1.33 × 108 and 3.26 × 107 copies g−1 DWS, respectively (Table 2).
Population community of active ammonia oxidizers
All AOB fell within the Nitrosospira genus of the class β-Proteobacteria (Fig. 4a and. b). All AOA fell within the order of Nitrososphaerales and Ca. Nitrosotaleales (Fig. 4c). The AOA consisted of clades NS-Alpha, NS-Beta, NS-Delta, NS-Zeta, NS-Gamma, and NT-Alpha in the acidic paddy soil; the clades NS-Alpha (38.4%), NS-Gamma (57.8%), NS-Zeta (3.6%), NT-Alpha (0.2%), NS-Beta (0.04%), and NS-Delta (0.01%) exhibited autotrophic growth during urea microcosm incubation (Fig. 4c). Furthermore, NS-Alpha and NS-Gamma were the main active AOA (Fig. 4c), as suggested by the 15N-DNA-SIP. All comammox Nitrospira belonged to the Nitrospira genus (Fig. 4d).
Compared with the CK, the dominant Zotus in the AOB-active community demonstrated high consistency and significant differences in both soils (P < 0.05) (Fig. 5a and. b). The dominant Zotus in the active community of AOA was basically consistent with that in the control, except that Zotu2 showed dominance in the acidic soil (Fig. 5c). The dominant Zotus in the active community of ComA in the neutral soil was extremely different from that in the control (Fig. 5d). Meanwhile, we compared the difference of active AOB in both soils, that is, the nitrification activities in both soils were dominated by different Zotus (Table S2). This means that the AOB in both soils were different, even though they belonged to the same genus level.
The AOA amoA phylogeny includes four basic lineages, which are consistent with the taxonomic orders of class Nitrososphaeria, including Nitrososphaerales (NS) and Ca. Nitrosotaleales (NT) (Fig. 4). The active AOA mostly belonged to the NS (Fig. 6a). The Nitrosocosmicus franklandus and Nitrososphaera viennensis clusters (NS-Alpha) and some uncultivated archaea (NS-Gamma) dominated the active AOA in the acidic paddy soils, accounting for up to 38.10% and 36.22%, respectively (Fig. 6a). Ca. Nitrosocosmicus arcticus, Ca. Nitrosocosmicus franklandus cluster (NS-Zeta), and Ca. Nitrosotalea devanaterra (NT-Alpha) had some activity, though they only account for a small proportion. Most ComA amoA gene sequences affiliated with Nitrospira II were observed in the neutral paddy soil microcosms. Regarding the active comammox Nitrospira communities, most of the comammox Nitrospira labeled with 15N were classified into comammox Nitrospira clades A2.1 and A2.2 (Fig. 6b). The active AOB communities were controlled by the Nitrosospira genus of β-proteobacteria and members within the Nitrosospira genus lineage accounted for 40.8% and 7.4% of the 15N-labeled AOB amoA genes in the both soils, respectively (Fig. 6c). Moreover, the Nitrosospira multiformis and Nitrosospira sp. Nsp17 appeared in the neutral soil, accounting for 3.3% and 4.1% of the total AOB, respectively (Fig. 6c). Numerous 15N-labeled AOB amoA gene sequences were joined to uncultured Nitrosospira in the acidic soil (Fig. 6c).
Discussion
Our results showed that active ammonia-oxidizing microorganisms changed significantly in the acidic and neutral soils under an intensive fertilization regime. Furthermore, since comammox Nitrospira were found, the growth and activity of each guild in ammonia oxidation under intensive fertilization regimes need to be re-evaluated. In this work, we utilized the 15N-DNA-SIP technique to uncover the activity of nitrifying populations.
An unsurprising finding was that AOA were higher than AOB in the acidic soil (Fig. 2a and b), especially after intensive fertilization. AOA were increased after urea application in the acidic soil at 5% significance level, whereas AOA were decreased after urea application in the neutral soil (Fig. 2a). Unsurprisingly, the AOA genome incorporated 15N during the incubation period in the acidic soil (Fig. 3a). Furthermore, the degree of AOA and AOB labeling suggested that AOA contributed significantly to the acidic soil (Fig. 3c), just as it was widely proposed that AOA play a significant role in autotrophic nitrification in acidic soils (Zhang et al. 2010; Yao et al. 2011; Li et al. 2018; Liu et al. 2021). A study based on soil pH gradient demonstrated AOA abundance and transcriptional activity, but the abundance and activity did not enhance with the reduction of pH (Nicol et al. 2008), and other results have elucidated that the AOA abundance was richer than that of AOB in acidic soils (Huang et al. 2012; Wu et al. 2017; Wang et al. 2019b). Nevertheless, the activity and growth of AOB in acid environments have been confirmed by the enrichment of AOB and the assimilation of 13CO2 (Huang et al. 2018; Picone et al. 2021; Bai et al. 2022). Indeed, under specific acidic conditions, AOB showed higher amoA gene abundance and transcriptional activity than AOA (Hayatsu et al. 2017), suggesting that pH is not a determining index in soil AOA and AOB niche under all conditions. A possible explanation for the continued activity of AOB in our acidic soil is that after urea application, urea hydrolysis can raise soil pH (Mehmood et al. 2018; Zhang et al. 2019), thereby creating an environment conducive to AOB growth, although subsequent nitrification can cause the pH to decrease again (Mehmood et al. 2018).
Another possible explanation for the activity of AOB is that the different ammonia oxidation pathways of AOB and AOA can affect the utilization of ammonium (Könneke et al. 2014; Kozlowski et al. 2016), accordingly influencing their abundance. The maximum activity (VMAX) and half-saturation constant (Km) of AOB were higher than that of AOA (Chen et al. 2018; Ouyang et al. 2018), which may lead to the rapid growth of AOB at high ammonium concentrations. In addition, the properties of the Rh-ammonia transporter may also be responsible for AOB abundance (Palomo et al. 2018), since ammonia levels in this study are millimolar at which AOB have a higher affinity and absorption capacity than AOA (Weidinger et al. 2007). The activity of AOB increases with the availability of ammonium (Ouyang et al. 2017), leading to a result in greater nitrification of AOB under conditions of high or saturated ammonia (Lin et al. 2021). For instance, AOA are more plentiful than AOB with lower ammonia contents in the open ocean (Wuchter et al. 2006; Agogué et al. 2008; Berg et al. 2015), whereas AOB response to ammonium was extremely significant in soil with higher ammonia concentrations (Verhamme et al. 2011; Palomo et al. 2018; French et al. 2021). This suggests that AOA were provided with much higher substrate affinity than AOB or comammox Nitrospira (Jung et al. 2021), that AOA are more plentiful in environments where ammonium is less available, and that AOB are more plentiful as the availability of ammonium increases (Herrmann et al. 2009; Auguet and Casamayor 2013; Jung et al. 2014). Hence, this research develops the understanding of the pH adaptability of AOB in paddy soil, particularly when ammonium concentrations are high.
ComA was not affected by fertilization, and apparently gave no labeling of 15N, while C2H2 had no effect (Fig. 2c), suggesting that comammox Nitrospira may be dormant. The same results also appeared in a 13CO2-labeled acidic soil (Liu et al. 2021), while the ComA abundance was not unaffected by urea (Liu et al. 2021; Feng et al. 2022). However, in another experiment with acidic forest soil, ComA incorporated 13CO2 (Li et al. 2019, 2020). Recent studies have revealed that the ComA is highly similar to the Rh-type transporters which were found in almost all β-AOB (Palomo et al. 2018; Gao et al. 2022). The Rh-type ammonia transporter can exhibit affinity for ammonium with high absorption capacity within the range of millimole (Palomo et al. 2018), which may favor the autotrophic growth of ComA at high ammonium concentrations. Therefore, ComA can undertake NH4+ to NO3− in acidic soil under specific circumstances. The activity of the neutral soil was verified by 15N. 15N was labeled, indicating that ComA participated in ammonia oxidation in the neutral soil (Fig. 3d). However, the low proportion of it labeled with 15N suggested that they played a lesser influence over autotrophic nitrification than canonical ammonia oxidizers (Table 2). Under high ammonium availability, the ability of ComA to absorb urea may lead to a change in competitiveness, which may be weaker compared to AOA and/or AOB (Liu et al. 2021), also having a Km (NH3) in comammox Nitrospira of only 63 nM (Kits et al. 2017). In the neutral soil, the ComA was observed with 15N-urea labeling (Gao et al. 2022), and this study supports evidence from previous observations that it grew autotrophically (Li et al. 2019; Zhao et al. 2020; Liu et al. 2021). All current comammox Nitrospira strains were isolated from aquatic systems, and comammox Nitrospira grows preferentially in neutral to slightly alkaline conditions (Daims et al. 2015; van Kessel et al. 2015; Li et al. 2019). A recent study provides direct and reliable evidence for nitrification that is partially due to neutral or basophilic ComA in soil. Since the average C/N ratio in the genome is about 2.1:1 (Cadisch et al. 2005) and the BD of the 15N-labeled fraction is only increased by about 0.02 g mL−1 (Cupples et al. 2007), we excluded the growth of ComB using 15N-urea in both soils. After nitrogen application, the autotrophic growth of ComB could not be detected (Fig. 2d), which might be the result that ComB only grows in paddy soils without nitrogen input (Wang et al. 2019a). Additionally, as weekly urea addition was unlikely to be the limiting factor for ComB growth, the absence of ComB growth suggests a potential inhibition, perhaps due to higher nitrogen input (Wang et al. 2019a; Jiang et al. 2020).
The AOB genome was more labeled with 15N than ComA in the neutral soil (Fig. 3b and. d). Additionally, AOB was significantly increased by applying urea, while urea application had no effect on ComA (Fig. 2b and. c). This indicated that AOB had a stronger contribution than ComA in the neutral soil. Although this method is a preliminary assessment of the relative contribution of ComA (Li et al. 2019), this finding still contributes to our understanding of the relative contribution of the nitrifying population.
AOA contain diverse organisms, including the NC, NS, NT, and NP. However, these organisms are extremely underrepresented in currently cultivated strains and genomes, compared to the total AOA yield (Alves et al. 2018). All AOA belonged to the class Nitrososphaeria (Fig. 4c); this also accords with earlier observations, which showed that Thaumarchaeota are the predominating lineage in acid soils (Subrahmanyam et al. 2014; Wu and Conrad 2014; Dai et al. 2018). NS-Alpha, NS-Beta, NS-Gamma, NS-Delta, NS-Zeta, and NT-Alpha exhibited autotrophic growth with different yields during urea miniature culture in the acidic soil (Fig. 4c). Only a few lineages of AOA occurred in acidic soils with pH below 7.0, including lineage NT and a few NS clades, particularly NS-Alpha and NS-Gamma (Alves et al. 2018), which was in agreement with our result that the AOA activity was dominated by NS-Alpha and NS-Gamma, as showed by the 15N-DNA-SIP (Fig. 4c). These results suggested that most AOA in the paddy soil showing autotrophic growth, which was identified using the amoA gene in the DNA-SIP, always originated from the AOA closely similar to the strains cultivated (Zhao et al. 2015; Alves et al. 2018; Pan et al. 2019). Moreover, Zotu2 dominated in the control treatment (Fig. 5c), Zotu1 and Zotu2 performed a dominant role in the 15N-urea treatment (Figs. 5c and. 6a), which came from NS-Gamma and NS-Alpha (Fig. 6a), respectively, indicating their dominant contribution to the growth of the AOA community. One interesting finding was that Zotu2 was consistently present and dominant in both control and 15N-urea treatments, suggesting that the clade of NS-Alpha could adapt to environmental changes and may maintain AOA diversity and activity in soils that are disturbed, for example, during intensive fertilization.
Phylogenetic analysis of AOA (a), comammox Nitrospira clade A (b), and AOB (c) amoA genes in 15N-labeled DNA from the 15N-urea-treated microcosms after an incubation period of 28 days. AS and NS represent acidic and neutral paddy soils, respectively. The designation “AS-AOA-HF–Zotu1-22,921–29.78%” indicates that Zotu1 contains 22,921 reads with 99% sequence similarity in the acidic paddy soil with the 15N-urea treatment. AS and NS mean acidic paddy soil and neutral paddy soil, respectively. Numbers with red, orange, yellow, green, blue, and purple colors represent sequences from different clades. NS and NT in notes mean Nitrososphaerales and Ca. Nitrosotaleales, respectively. The phylogeny of AOA, comammox Nitrospira, and AOB was generated using MEGA X with NJ-tree. Bootstraps are based on 1000 replicated trees. Bootstrap values are indicated at the branch nodes
In either acidic or neutral soils, the AOB was from the genus Nitrosospira, and no traces of Nitrosomonas and Nitrosococcus were found (Fig. 4a and. b). Interestingly, the dominant Zotus differed in both soils (Fig. 5a and. b, Table S2). The phylogenetic analysis further reflected that the dominant Zotus in the acidic and neutral soils belonged to different AOB clades, respectively (Fig. 6c). These findings suggested that AOB-active taxa in the acidic soils differed from those in the neutral soils at the OTU level. It can help us to understand the survival patterns and population dynamics of AOB in acidic and neutral soils and provide strong support for explaining the dominant role of AOB under high nitrogen input.
ComA dominated ammonia oxidation in the neutral paddy soil but may play a lesser role in nitrification than AOB. Additionally, the dominant Zotus in the 15N-urea treatment was different from that in the control, and the proportions of each Zotus were also similar (Fig. 5d), indicating the option of these Zotus for high nitrogen input. Based on phylogenetic analysis, ComA was previously divided into four subclades, clades A.1, A.2.1, A.2.2, and A.3 (Li et al. 2021a, b). Two comammox Nitrospira clusters were found in this study, ComA.2.1 and ComA.2.2 (Fig. 6b), commonly obtained from terrestrial ecosystems (Sun et al. 2021a, b; Yuan et al. 2021). Comammox Nitrospira community was predominated by ComA.2.2, which was different from other studies where ComA.2.1 dominated (Lin et al. 2022; He et al. 2022), probably because ComA.2.1 was biased towards relatively high pH (Lin et al. 2022; He et al. 2022). However, due to the lack of cultures, the survival mechanisms of ComA.2.1 and A.2.2 in agricultural ecosystems are still unclear and should be considered in future studies.
In this study, 15N-urea was used as a tracer to reveal the active taxa within both paddy soils. Despite the current high level of fertilization in the field, with the high demand for agricultural production, continued intensive fertilization may be expected to be applied to the soil in pursuit of high yields. Therefore, this experiment provides a prediction for future changes in the microbial populations following fertilizer application. It may be due to the continuous nitrification during the extraction process, the measured ammonium concentration may be lower than the original level of the soil, or the strong adsorption of the soil caused the ammonium concentration to be too low (Wang and Alva 2000; He et al. 2021; He et al. 2021). In addition, the combined effect of organic nitrogen mineralization in the soil and the oxidation of ammonium can also cause the measured ammonium concentration to be lower than the actual level (Wang et al. 2018). Furthermore, we elucidate differences in active taxa in both paddy soils at the OTU level. However, the sequencing primers and the specificity of the comammox Nitrospira gene under aerobic conditions need to be further developed and improved to make the characterization of the ammonia-oxidizing microbial community more direct. Meanwhile, the specific nitrogen cycle processes for comammox Nitrospira, AOA, and AOB coupling are still unclear, and more experiments are needed to further focus on functional genes and microbial metabolic processes related to the nitrogen cycle.
The study showed that urea significantly enhanced the abundance of canonical ammonia oxidizers in both soils, and the existence of C2H2 almost completely inhibited nitrification and the activities of these groups in both soils. 15N-DNA-SIP plays a useful role in distinguishing nitrifying populations and revealing their niche differentiation. ComA incorporated 15N into the genome, showing that it was active in the neutral soil. Furthermore, the low proportion of it labeled with 15N suggests that it plays a lesser role in autotrophic nitrification than canonical ammonia oxidizers. The growth of canonical ammonia oxidizers and ComA was detected, and it illuminated that canonical ammonia oxidizers dominated ammonia oxidation in the acidic soil, while AOB and ComA dominated in the neutral soil. In particular, although AOB grew autotrophically in both soils, phylogenetic analysis showed that AOB-active taxa differed from those at the OTU level. These findings develop our understanding of the nitrogen cycle and indicate that the ecological importance of nitrifying populations needs to be reassessed in environments impacted by nitrogen input.
Data availability
All datasets generated for this study are included in the article/supplementary material.
References
Agogué H, Brink M, Dinasquet J, Herndl GJ (2008) Major gradients in putatively nitrifying and non-nitrifying Archaea in the deep North Atlantic. Nature 456:788–791. https://doi.org/10.1038/nature07535
Ali TU, Kim M, Kim D-J (2013) Selective inhibition of ammonia oxidation and nitrite oxidation linked to N2O emission with activated sludge and enriched nitrifiers. J Microbiol Biotechnol 23:719–723. https://doi.org/10.4014/jmb.1302.02017
Alves RJE, Minh BQ, Urich T, von Haeseler A, Schleper C (2018) Unifying the global phylogeny and environmental distribution of ammonia-oxidising archaea based on amoA genes. Nat Commun 9:1517. https://doi.org/10.1038/s41467-018-03861-1
Angel R, Panhölzl C, Gabriel R, Herbold C, Wanek W, Richter A, Eichorst SA, Woebken D (2018) Application of stable-isotope labelling techniques for the detection of active diazotrophs: detecting diazotrophs with stable-isotope techniques. Environ Microbiol 20:44–61. https://doi.org/10.1111/1462-2920.13954
Auguet JC, Casamayor EO (2013) Partitioning of Thaumarchaeota populations along environmental gradients in high mountain lakes. FEMS Microbiol Ecol 84:154–164. https://doi.org/10.1111/1574-6941.12047
Bai X, Hu X, Liu J, Wei D, Zhu P, Cui X, Zhou B, Chen X, Liu J, Jin J, Liu X, Wang G (2022) Ammonia oxidizing bacteria dominate soil nitrification under different fertilization regimes in black soils of northeast China. Eur J Soil Biol 111:103410. https://doi.org/10.1016/j.ejsobi.2022.103410
Bello MO, Aigle A, Meng Y, Prosser JI, Gubry-Rangin C (2021) Preferential temperature and ammonia concentration for in-situ growth of Candidatus Nitrosocosmicus ammonia oxidising archaea. Soil Biol Biochem 162:108405. https://doi.org/10.1016/j.soilbio.2021.108405
Berg C, Vandieken V, Thamdrup B, Jürgens K (2015) Significance of archaeal nitrification in hypoxic waters of the Baltic Sea. ISME J 9:1319–1332. https://doi.org/10.1038/ismej.2014.218
Cadisch G, Espana M, Causey R, Richter M, Shaw E, Morgan JAW, Rahn C, Bending GD (2005) Technical considerations for the use of 15N-DNA stable-isotope probing for functional microbial activity in soils. Rapid Commun Mass Spectrom 19:1424–1428. https://doi.org/10.1002/rcm.1908
Chen X, Xia Y, Hu Y, Gunina A, Ge T, Zhang Z, Wu J, Su Y (2018) Effect of nitrogen fertilization on the fate of rice residue-C in paddy soil depending on depth: 13C amino sugar analysis. Biol Fertil Soils 54:523–531. https://doi.org/10.1007/s00374-018-1278-5
Cupples AM, Shaffer EA, Chee-Sanford JC, Sims GK (2007) DNA buoyant density shifts during 15N-DNA stable isotope probing. Microbiol Res 162:328–334. https://doi.org/10.1016/j.micres.2006.01.016
Dai Y, Wu Y, Ding Q, Zeng J, Li X, Zheng J, Lin X (2018) Oxygenated derivative is more influential than unsubstituted polycyclic aromatic hydrocarbon on ammonia-oxidizing archaea in an acidic soil. J Soils Sediments 18:2573–2580. https://doi.org/10.1007/s11368-018-1925-3
Daims H, Wagner M (2018) Nitrospira. Trends Microbiol 26:462–463. https://doi.org/10.1016/j.tim.2018.02.001
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M, Jehmlich N, Palatinszky M, Vierheilig J, Bulaev A, Kirkegaard RH, von Bergen M, Rattei T, Bendinger B, Nielsen PH, Wagner M (2015) Complete nitrification by Nitrospira bacteria. Nature 528:504–509. https://doi.org/10.1038/nature16461
Dong X, Zhang J, Qiu H, Zhang H, Luo C, Deng D, Shen Q, Jia Z (2019) Chronic nitrogen fertilization modulates competitive interactions among microbial ammonia oxidizers in a loess soil. Pedosphere 29:24–33. https://doi.org/10.1016/S1002-0160(18)60055-4
Edgar RC (2016) UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 081257. https://doi.org/10.1101/081257
Feng M, He Z, Fan J, Ge A, Jin S, Lin Y, He J (2022) Temperature has a strong impact on the abundance and community structure of comammox Nitrospira in an Ultisol. J Soils Sediments. https://doi.org/10.1007/s11368-022-03261-5
French E, Kozlowski JA, Bollmann A (2021) Competition between ammonia-oxidizing archaea and bacteria from freshwater environments. Appl Environ Microbiol 87:e01038-e1121. https://doi.org/10.1128/AEM.01038-21
Gao F, Li Y, Fan H, Xue J, Yao H (2022) Main environmental drivers of abundance, diversity, and community structure of comammox Nitrospira in paddy soils. Pedosphere 33:808–818. https://doi.org/10.1016/j.pedsph.2022.06.061
Gubry-Rangin C, Nicol GW, Prosser JI (2010) Archaea rather than bacteria control nitrification in two agricultural acidic soils. FEMS Microbiol Ecol 74:566–574. https://doi.org/10.1111/j.1574-6941.2010.00971.x
Hayatsu M, Tago K, Uchiyama I, Toyoda A, Wang Y, Shimomura Y, Okubo T, Kurisu F, Hirono Y, Nonaka K, Akiyama H, Itoh T, Takami H (2017) An acid-tolerant ammonia-oxidizing γ-proteobacterium from soil. ISME J 11:1130–1141. https://doi.org/10.1038/ismej.2016.191
He S, Li Y, Mu H, Zhao Z, Wang J, Liu S, Sun Z, Zheng M (2021) Ammonium concentration determines differential growth of comammox and canonical ammonia-oxidizing prokaryotes in soil microcosms. Appl Soil Ecol 157:103776. https://doi.org/10.1016/j.apsoil.2020.103776
He Z, Sun A, Jiao X, Ge A, Hu H, Jin S, Liu X, Lin Y, He J (2022) Fertilization has a greater effect than rhizosphere on community structures of comammox Nitrospira in an alkaline agricultural soil. Appl Soil Ecol 175:104456. https://doi.org/10.1016/j.apsoil.2022.104456
Herrmann M, Saunders AM, Schramm A (2009) Effect of lake trophic status and rooted macrophytes on community composition and abundance of ammonia-oxidizing prokaryotes in freshwater sediments. Appl Environ Microbiol 75:3127–3136. https://doi.org/10.1128/AEM.02806-08
Hu B, Liu S, Wang W, Li L, Lou L, Liu W, Tian G, Xu X, Zheng P (2014) pH-dominated niche segregation of ammonia-oxidising microorganisms in Chinese agricultural soils. FEMS Microbiol Ecol 90:290–299. https://doi.org/10.1111/1574-6941.12391
Huang R, Wu Y, Zhang J, Zhong W, Jia Z, Cai Z (2012) Nitrification activity and putative ammonia-oxidizing archaea in acidic red soils. J Soils Sediments 12:420–428. https://doi.org/10.1007/s11368-011-0450-4
Huang X, Zhao J, Su J, Jia Z, Shi X, Wright AL, Zhu-Barker X, Jiang X (2018) Neutrophilic bacteria are responsible for autotrophic ammonia oxidation in an acidic forest soil. Soil Biol Biochem 119:83–89. https://doi.org/10.1016/j.soilbio.2018.01.016
Jiang R, Wang J, Zhu T, Zou B, Wang D, Rhee S-K, An D, Ji Z, Quan Z (2020) Use of newly designed primers for quantification of complete ammonia-oxidizing (comammox) bacterial clades and strict nitrite oxidizers in the genus Nitrospira. Appl Environ Microbiol 86:e01775-e1820. https://doi.org/10.1128/AEM.01775-20
Jung M-Y, Park S-J, Kim S-J, Kim J-G, Sinninghe Damsté JS, Jeon CO, Rhee S-K (2014) A mesophilic, autotrophic, ammonia-oxidizing archaeon of thaumarchaeal group I.1a cultivated from a deep oligotrophic soil horizon. Appl Environ Microbiol 80:3645–3655. https://doi.org/10.1128/AEM.03730-13
Jung M-Y, Sedlacek CJ, Kits KD, Mueller AJ, Rhee S-K, Hink L, Nicol GW, Bayer B, Lehtovirta-Morley L, Wright C, de la Torre JR, Herbold CW, Pjevac P, Daims H, Wagner M (2021) Ammonia-oxidizing archaea possess a wide range of cellular ammonia affinities. ISME J. https://doi.org/10.1038/s41396-021-01064-z
Kits KD, Sedlacek CJ, Lebedeva EV, Han P, Bulaev A, Pjevac P, Daebeler A, Romano S, Albertsen M, Stein LY, Daims H, Wagner M (2017) Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle. Nature 549:269–272. https://doi.org/10.1038/nature23679
Könneke M, Schubert DM, Brown PC, Hügler M, Standfest S, Schwander T, Schada von Borzyskowski L, Erb TJ, Stahl DA, Berg IA (2014) Ammonia-oxidizing archaea use the most energy-efficient aerobic pathway for CO2 fixation. Proc Natl Acad Sci USA 111:8239–8244. https://doi.org/10.1073/pnas.1402028111
Kozlowski JA, Stieglmeier M, Schleper C, Klotz MG, Stein LY (2016) Pathways and key intermediates required for obligate aerobic ammonia-dependent chemolithotrophy in bacteria and Thaumarchaeota. ISME J 10:1836–1845. https://doi.org/10.1038/ismej.2016.2
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Li Y, Chapman SJ, Nicol GW, Yao H (2018) Nitrification and nitrifiers in acidic soils. Soil Biol Biochem 116:290–301. https://doi.org/10.1016/j.soilbio.2017.10.023
Li C, Hu H, Chen Q, Chen D, He J (2019) Comammox Nitrospira play an active role in nitrification of agricultural soils amended with nitrogen fertilizers. Soil Biol Biochem 138:107609. https://doi.org/10.1016/j.soilbio.2019.107609
Li C, Hu H, Chen Q, Chen D, He J (2020) Niche differentiation of clade A comammox Nitrospira and canonical ammonia oxidizers in selected forest soils. Soil Biol Biochem 149:107925. https://doi.org/10.1016/j.soilbio.2020.107925
Li C, Hu H, Chen Q, Yan Z, Thi Nguyen B-A, Chen D, He J (2021a) Niche specialization of comammox Nitrospira clade A in terrestrial ecosystems. Soil Biol Biochem 156:108231. https://doi.org/10.1016/j.soilbio.2021.108231
Li Y, Zhang Y, Chapman SJ, Yao H (2021b) Biological nitrification inhibition by sorghum root exudates impacts ammonia-oxidizing bacteria but not ammonia-oxidizing archaea. Biol Fertil Soils 57:399–407. https://doi.org/10.1007/s00374-020-01538-w
Lin Y, Hu H, Ye G, Fan J, Ding W, He Z, Zheng Y, He J (2021) Ammonia-oxidizing bacteria play an important role in nitrification of acidic soils: a meta-analysis. Geoderma 404:115395. https://doi.org/10.1016/j.geoderma.2021.115395
Lin Y, Fan J, Hu H, Duan C, Ye G, Wan S, He Z, Zheng Y, He J (2022) Differentiation of individual clusters of comammox Nitrospira in an acidic Ultisol following long-term fertilization. Appl Soil Ecol 170:104267. https://doi.org/10.1016/j.apsoil.2021.104267
Liu H, Hu H, Huang X, Ge T, Li Y, Zhu Z, Liu X, Tan W, Jia Z, Di H, Xu J, Li Y (2021) Canonical ammonia oxidizers, rather than comammox Nitrospira, dominated autotrophic nitrification during the mineralization of organic substances in two paddy soils. Soil Biol Biochem 156:108192. https://doi.org/10.1016/j.soilbio.2021.108192
Lu L, Jia Z (2013) Urease gene-containing Archaea dominate autotrophic ammonia oxidation in two acid soils: urea-linked archaeal ammonia oxidation in acid soil. Environ Microbiol 15:1795–1809. https://doi.org/10.1111/1462-2920.12071
Lu L, Han W, Zhang J, Wu Y, Wang B, Lin X, Zhu J, Cai Z, Jia Z (2012) Nitrification of archaeal ammonia oxidizers in acid soils is supported by hydrolysis of urea. ISME J 6:1978–1984. https://doi.org/10.1038/ismej.2012.45
Luo D, Meng X, Zheng N, Li Y, Yao H, Chapman SJ (2021) The anaerobic oxidation of methane in paddy soil by ferric iron and nitrate, and the microbial communities involved. Sci Total Environ 788:147773. https://doi.org/10.1016/j.scitotenv.2021.147773
Mehmood K, Baquy MA-A, Xu R (2018) Influence of nitrogen fertilizer forms and crop straw biochars on soil exchange properties and maize growth on an acidic Ultisol. Arch Agron Soil Sci 64:834–849. https://doi.org/10.1080/03650340.2017.1385062
Meng X, Li Y, Yao H, Wang J, Dai F, Wu Y, Chapman SJ (2020) Nitrification and urease inhibitors improve rice nitrogen uptake and prevent denitrification in alkaline paddy soil. Appl Soil Ecol 154:103665. https://doi.org/10.1016/j.apsoil.2020.103665
Nicol GW, Leininger S, Schleper C, Prosser JI (2008) The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environ Microbiol 10:2966–2978. https://doi.org/10.1111/j.1462-2920.2008.01701.x
Osburn ED, Barrett JE (2020) Abundance and functional importance of complete ammonia-oxidizing bacteria (comammox) versus canonical nitrifiers in temperate forest soils. Soil Biol Biochem 145:107801. https://doi.org/10.1016/j.soilbio.2020.107801
Ouyang Y, Norton JM, Stark JM (2017) Ammonium availability and temperature control contributions of ammonia oxidizing bacteria and archaea to nitrification in an agricultural soil. Soil Biol Biochem 113:161–172. https://doi.org/10.1016/j.soilbio.2017.06.010
Ouyang Y, Evans SE, Friesen ML, Tiemann LK (2018) Effect of nitrogen fertilization on the abundance of nitrogen cycling genes in agricultural soils: a meta-analysis of field studies. Soil Biol Biochem 127:71–78. https://doi.org/10.1016/j.soilbio.2018.08.024
Palomo A, Pedersen AG, Fowler SJ, Dechesne A, Sicheritz-Pontén T, Smets BF (2018) Comparative genomics sheds light on niche differentiation and the evolutionary history of comammox Nitrospira. ISME J 12:1779–1793. https://doi.org/10.1038/s41396-018-0083-3
Pan K, Gao J, Fan X, Li D, Dai H (2018) The more important role of archaea than bacteria in nitrification of wastewater treatment plants in cold season despite their numerical relationships. Water Res 145:552–561. https://doi.org/10.1016/j.watres.2018.08.066
Pan K, Gao J, Li D, Fan X (2019) The dominance of non-halophilic archaea in autotrophic ammonia oxidation of activated sludge under salt stress: a DNA-based stable isotope probing study. Bioresour Technol 291:121914. https://doi.org/10.1016/j.biortech.2019.121914
Park S, Cho K, Lee T, Lee E, Bae H (2022) Improved insights into the adaptation and selection of Nitrosomonas spp. for partial nitritation under saline conditions based on specific oxygen uptake rates and next generation sequencing. Sci Total Environ 822:153644. https://doi.org/10.1016/j.scitotenv.2022.153644
Picone N, Pol A, Mesman R, van Kessel MAHJ, Cremers G, van Gelder AH, van Alen TA, Jetten MSM, Lucker S, Op den Camp HJM (2021) Ammonia oxidation at pH 2.5 by a new gammaproteobacterial ammonia-oxidizing bacterium. ISME J 15:1150–1164. https://doi.org/10.1038/s41396-020-00840-7
Pjevac P, Schauberger C, Poghosyan L, Herbold CW, van Kessel MAHJ, Daebeler A, Steinberger M, Jetten MSM, Lücker S, Wagner M, Daims H (2017) AmoA-targeted polymerase chain reaction primers for the specific detection and quantification of comammox Nitrospira in the environment. Front Microbiol 8:1508. https://doi.org/10.3389/fmicb.2017.01508
Pornkulwat P, Kurisu F, Soonglerdsongpha S, Banjongproo P, Srithep P, Limpiyakorn T (2018) Incorporation of 13C-HCO3− by ammonia-oxidizing archaea and bacteria during ammonia oxidation of sludge from a municipal wastewater treatment plant. Appl Microbiol Biotechnol 102:10767–10777. https://doi.org/10.1007/s00253-018-9436-0
Pratscher J, Dumont MG, Conrad R (2011) Ammonia oxidation coupled to CO2 fixation by Archaea and bacteria in an agricultural soil. Proc Natl Acad Sci USA 108:4170–4175. https://doi.org/10.1073/pnas.1010981108
Qin W, Zheng Y, Zhao F, Wang Y, Urakawa H, Martens-Habbena W, Liu H, Huang X, Zhang X, Nakagawa T, Mende DR, Bollmann A, Wang B, Zhang Y, Amin SA, Nielsen JL, Mori K, Takahashi R, Virginia Armbrust E, Winkler M-KH, DeLong EF, Li M, Lee P-H, Zhou J, Zhang C, Zhang T, Stahl DA, Ingalls AE (2020) Alternative strategies of nutrient acquisition and energy conservation map to the biogeography of marine ammonia-oxidizing archaea. ISME J 14:2595–2609. https://doi.org/10.1038/s41396-020-0710-7
Ren M, Wang J (2022) Phylogenetic divergence and adaptation of Nitrososphaeria across lake depths and freshwater ecosystems. ISME J 16:1491–1501. https://doi.org/10.1038/s41396-022-01199-7
Séneca J, Pjevac P, Canarini A, Herbold CW, Zioutis C, Dietrich M, Simon E, Prommer J, Bahn M, Pötsch EM, Wagner M, Wanek W, Richter A (2020) Composition and activity of nitrifier communities in soil are unresponsive to elevated temperature and CO2, but strongly affected by drought. ISME J 14:3038–3053. https://doi.org/10.1038/s41396-020-00735-7
Shen J, Zhang L, Di H, He J (2012) A review of ammonia-oxidizing bacteria and archaea in Chinese soils. Front Microbiol 3:296. https://doi.org/10.3389/fmicb.2012.00296
Subrahmanyam G, Hu H, Zheng Y, Gattupalli A, He J, Liu Y (2014) Response of ammonia oxidizing microbes to the stresses of arsenic and copper in two acidic alfisols. Appl Soil Ecol 77:59–67. https://doi.org/10.1016/j.apsoil.2014.01.011
Sun D, Liu M, Hou L, Zhao M, Tang X, Zhao Q, Li J, Han P (2021a) Community structure and abundance of comammox Nitrospira in Chongming eastern intertidal sediments. J Soils Sediments 21:3213–3224. https://doi.org/10.1007/s11368-021-02940-z
Sun D, Zhao M, Tang X, Liu M, Hou L, Zhao Q, Li J, Gu J, Han P (2021b) Niche adaptation strategies of different clades of comammox Nitrospira in the Yangtze estuary. Int Biodeterior Biodegradation 164:105286. https://doi.org/10.1016/j.ibiod.2021.105286
Sun X, Zhao J, Zhou X, Bei Q, Xia W, Zhao B, Zhang J, Jia Z (2022) Salt tolerance-based niche differentiation of soil ammonia oxidizers. ISME J 16:412–422. https://doi.org/10.1038/s41396-021-01079-6
Tago K, Okubo T, Shimomura Y, Kikuchi Y, Hori T, Nagayama A, Hayatsu M (2015) Environmental factors shaping the community structure of ammonia-oxidizing bacteria and archaea in sugarcane field soil. Microbes Environ 30:21–28. https://doi.org/10.1264/jsme2.ME14137
Takahashi Y, Fujitani H, Hirono Y, Tago K, Wang Y, Hayatsu M, Tsuneda S (2020) Enrichment of comammox and nitrite-oxidizing Nitrospira from acidic soils. Front Microbiol 11:1737. https://doi.org/10.3389/fmicb.2020.01737
Tao R, Wakelin SA, Liang Y, Chu G (2017) Response of ammonia-oxidizing archaea and bacteria in calcareous soil to mineral and organic fertilizer application and their relative contribution to nitrification. Soil Biol Biochem 114:20–30. https://doi.org/10.1016/j.soilbio.2017.06.027
van Kessel MAHJ, Speth DR, Albertsen M, Nielsen PH, Op den Camp HJM, Kartal B, Jetten MSM, Lücker S (2015) Complete nitrification by a single microorganism. Nature 528:555–559. https://doi.org/10.1038/nature16459
Verhamme DT, Prosser JI, Nicol GW (2011) Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J 5:1067–1071. https://doi.org/10.1038/ismej.2010.191
Wang FL, Alva AK (2000) Ammonium adsorption and desorption in sandy soils. Soil Sci Soc Am J 64:1669–1674. https://doi.org/10.2136/sssaj2000.6451669x
Wang W, Jiang X, Zheng B, Chen J, Zhao L, Zhang B, Wang S (2018) Composition, mineralization potential and release risk of nitrogen in the sediments of Keluke Lake, a Tibetan Plateau freshwater lake in China. R Soc Open Sci 5:180612. https://doi.org/10.1098/rsos.180612
Wang Z, Cao Y, Zhu-Barker X, Nicol GW, Wright AL, Jia Z, Jiang X (2019a) Comammox Nitrospira clade B contributes to nitrification in soil. Soil Biol Biochem 135:392–395. https://doi.org/10.1016/j.soilbio.2019.06.004
Wang Z, Meng Y, Zhu-Barker X, He X, Horwath WR, Luo H, Zhao Y, Jiang X (2019b) Responses of nitrification and ammonia oxidizers to a range of background and adjusted pH in purple soils. Geoderma 334:9–14. https://doi.org/10.1016/j.geoderma.2018.07.038
Wang X, Wang S, Jiang Y, Zhou J, Han C, Zhu G (2020) Comammox bacterial abundance, activity, and contribution in agricultural rhizosphere soils. Sci Total Environ 727:138563. https://doi.org/10.1016/j.scitotenv.2020.138563
Wang J, Kan J, Zheng X, Xia Z, Zhang XC, Qian G, Miao Y, Leng X, Sun J (2017) Archaea dominate the ammonia-oxidizing community in deep-sea sediments of the Eastern Indian Ocean—from the equator to the Bay of Bengal. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.00415
Weidinger K, Neuhäuser B, Gilch S, Ludewig U, Meyer O, Schmidt I (2007) Functional and physiological evidence for a rhesus-type ammonia transporter in Nitrosomonas europaea. FEMS Microbiol Lett 273:260–267. https://doi.org/10.1111/j.1574-6968.2007.00805.x
Wu Y, Conrad R (2014) Ammonia oxidation-dependent growth of group I.1b Thaumarchaeota in acidic red soil microcosms. FEMS Microbiol Ecol 89:127–134. https://doi.org/10.1111/1574-6941.12340
Wu Y, Ke X, Hernández M, Wang B, Dumont MG, Jia Z, Conrad R (2013) Autotrophic growth of bacterial and archaeal smmonia oxidizers in freshwater sediment microcosms incubated at different temperatures. Appl Environ Microbiol 79:3076–3084. https://doi.org/10.1128/AEM.00061-13
Wu R, Meng H, Wang Y, Lan W, Gu J (2017) A more comprehensive community of ammonia-oxidizing archaea (AOA) revealed by genomic DNA and RNA analyses of amoA gene in subtropical acidic forest soils. Microb Ecol 74:910–922. https://doi.org/10.1007/s00248-017-1045-4
Wuchter C, Abbas B, Coolen MJL, Herfort L, van Bleijswijk J, Timmers P, Strous M, Teira E, Herndl GJ, Middelburg JJ, Schouten S, Sinninghe Damsté JS (2006) Archaeal nitrification in the ocean. Proc Natl Acad Sci USA 103:12317–12322. https://doi.org/10.1073/pnas.0600756103
Xia W, Zhang C, Zeng X, Feng Y, Weng J, Lin X, Zhu J, Xiong Z, Xu J, Cai Z, Jia Z (2011) Autotrophic growth of nitrifying community in an agricultural soil. ISME J 5:1226–1236. https://doi.org/10.1038/ismej.2011.5
Yao H, Gao Y, Nicol GW, Campbell CD, Prosser JI, Zhang L, Han W, Singh BK (2011) Links between ammonia oxidizer community structure, abundance, and nitrification potential in acidic soils. Appl Environ Microbiol 77:4618–4625. https://doi.org/10.1128/AEM.00136-11
Ye X, Han B, Li W, Zhang X, Zhang Y, Lin X, Zou H (2018) Effects of different irrigation methods on nitrous oxide emissions and ammonia oxidizers microorganisms in greenhouse tomato fields. Agric Water Manag 203:115–123. https://doi.org/10.1016/j.agwat.2018.03.012
Ying J, Li X, Wang N, Lan Z, He J, Bai Y (2017) Contrasting effects of nitrogen forms and soil pH on ammonia oxidizing microorganisms and their responses to long-term nitrogen fertilization in a typical steppe ecosystem. Soil Biol Biochem 107:10–18. https://doi.org/10.1016/j.soilbio.2016.12.023
Yu Y, Li X, Feng Z, Xiao M, Ge T, Li Y, Yao H (2022) Polyethylene microplastics alter the microbial functional gene abundances and increase nitrous oxide emissions from paddy soils. J Hazard Mater 432:128721. https://doi.org/10.1016/j.jhazmat.2022.128721
Yuan D, Zheng L, Tan Q, Wang X, Xing Y, Wang H, Wang S, Zhu G (2021) Comammox activity dominates nitrification process in the sediments of plateau wetland. Water Res 206:117774. https://doi.org/10.1016/j.watres.2021.117774
Zhang L, Offre PR, He J, Verhamme DT, Nicol GW, Prosser JI (2010) Autotrophic ammonia oxidation by soil thaumarchaea. Proc Natl Acad Sci USA 107:17240–17245. https://doi.org/10.1073/pnas.1004947107
Zhang L, Hu H, Shen J, He J (2012) Ammonia-oxidizing archaea have more important role than ammonia-oxidizing bacteria in ammonia oxidation of strongly acidic soils. ISME J 6:1032–1045. https://doi.org/10.1038/ismej.2011.168
Zhang M, Alves RJE, Zhang D, Han L, He J, Zhang L (2017) Time-dependent shifts in populations and activity of bacterial and archaeal ammonia oxidizers in response to liming in acidic soils. Soil Biol Biochem 112:77–89. https://doi.org/10.1016/j.soilbio.2017.05.001
Zhang L, Duff AM, Smith CJ (2018) Community and functional shifts in ammonia oxidizers across terrestrial and marine (soil/sediment) boundaries in two coastal bay ecosystems. Environ Microbiol 20:2834–2853. https://doi.org/10.1111/1462-2920.14238
Zhang Q, Li Y, He Y, Liu H, Dumont MG, Brookes PC, Xu J (2019) Nitrosospira cluster 3-like bacterial ammonia oxidizers and Nitrospira-like nitrite oxidizers dominate nitrification activity in acidic terrace paddy soils. Soil Biol Biochem 131:229–237. https://doi.org/10.1016/j.soilbio.2019.01.006
Zhang W, Wang X, Miao Y, Li Y, Zhang H, Niu L, Wang L (2021) Determining the effect of sertraline on nitrogen transformation through the microbial food web in sediments based on 15N-DNA-stable isotope probing. Environ Res 199:111347. https://doi.org/10.1016/j.envres.2021.111347
Zhang Y, Li X, Xiao M, Feng Z, Yu Y, Yao H (2022) Effects of microplastics on soil carbon dioxide emissions and the microbial functional genes involved in organic carbon decomposition in agricultural soil. Sci Total Environ 806:150714. https://doi.org/10.1016/j.scitotenv.2021.150714
Zhao J, Wang B, Jia Z (2015) Phylogenetically distinct phylotypes modulate nitrification in a paddy soil. Appl Environ Microbiol 81:3218–3227. https://doi.org/10.1128/AEM.00426-15
Zhao J, Meng Y, Drewer J, Skiba UM, Prosser JI, Gubry-Rangin C (2020) Differential ecosystem function stability of ammonia-oxidizing archaea and bacteria following short-term environmental perturbation. mSystems 5:e00309–20. https://doi.org/10.1128/mSystems.00309-20
Funding
This work was supported by the National Natural Science Foundation of China [grant numbers 42077036, 42277105, 42021005] and the Ningbo Municipal Bureau of Science and Technology [grant number 2021Z047, 2022Z168].
Author information
Authors and Affiliations
Contributions
FG, YL, and HY conceived and designed the research. FG and YL conducted the experiments. HF and DL contributed new reagents or analytical tools. FG, YL, SJC, and HY wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, F., Li, Y., Fan, H. et al. 15N-DNA stable isotope probing reveals niche differentiation of ammonia oxidizers in paddy soils. Appl Microbiol Biotechnol 108, 342 (2024). https://doi.org/10.1007/s00253-024-13170-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13170-x