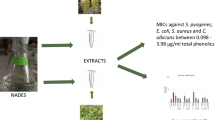
Abstract
The increasing antibiotic resistance towards a panel of microorganisms is one of the public health concerns. For this reason, the search for alternatives to the widely used antibiotic has been undertaken. In the era of sustainable chemistry, deep eutectic solvents (DESs) have emerged as promising antimicrobial agents. These solvents possess several advantages such as low volatility, low flammability, ease of preparation, and typically low cost of production. These properties make DES suitable for various applications, including extraction of biomolecules and preparation of cosmetics. Natural DESs (NADESs) are special category of DESs prepared from natural sources, which matched the recent trends of leaning back to nature, and decreasing dependence on synthetic precursors. NADES can be prepared by heating and stirring, freeze-drying, evaporation, grinding, and ultrasound-assisted and microwave-assisted synthesis. Utilizing NADESs as an alternative to traditional antibiotics, which become ineffective over time due to bacterial resistance, holds great promise for these reasons. This review aims to discuss the antimicrobial properties of multiple NADESs, including antibacterial and antifungal activities. To the best of our knowledge, this review is the first literature survey of the antimicrobial activities of NADESs.
Key points
• Natural deep eutectic solvents are promising antimicrobial alternative to antibiotics
• NADES holds high potential for their activity against bacterial resistance
• NADES have also substantial antifungal activities
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial infection is one of the greatest challenges facing the world, due to its serious clinical manifestations. Many researchers have been doing their best to overcome this problem in the last few decades (Silva et al. 2019). One of their interventions was the discovery of antibiotics, which was a turning point in medical therapies for combating microbial infections. However, each discovery was followed by the rise of antibiotic resistance, leading to the emergence of multidrug-resistant bacteria. The World Health Organization (WHO) has published a list of the most serious bacteria that are resistant to current treatments and require the development of new antibiotics to combat resistance.
Efforts have been made in research to meet the urgent demand for novel treatment approaches other than antibiotics. Biological alternatives to antibiotics against serious pathogens have been investigated. Antimicrobial proteins, bacteriophages, probiotics, and plant-based compounds are among the most promising alternatives (Łojewska and Sakowicz 2021). It has been known for centuries that plants possess natural antibacterial properties, in addition to their effectiveness against antibiotic-resistant microorganisms (Ng et al. 2021). They also contain volatile oil compounds (VOCs). These VOCs are molecules that are biosynthesized by primary and secondary metabolic pathways. VOCs include chemical classes such as alcohols, esters, aliphatic and aromatic hydrocarbons, terpenes, nitrogen, and sulfur compounds (Garrido et al. 2020). Additionally, the development of new green solvents that exhibit better properties than organic solvents is a key feature in green chemistry (Silva et al. 2019). Examples of these green solvents are room-temperature ionic liquids (RTILs) and deep eutectic solvents (DES). RTILs are very promising replacements for traditional volatile organic solvents as antibacterial treatments (Fernandes et al. 2022). They are composed of an organic cation and an inorganic or organic anion. They are attractive in research due to their versatility for an increasing number of technological applications. The main advantages of RTILs are high mobility, low melting points, negligible vapor pressure, thermal stability, low toxicity, large electrochemical window, and no flammability (Fernandes et al. 2022). However, experts frequently question their biodegradability and biological toxicity (Usmani et al. 2023).
On the other side, DESs have emerged as good alternatives to traditional organic solvents as well as their counterparts, ionic liquids (ILs) (Silva et al. 2019). DESs are mixtures of different ingredients that transition to liquid due to a massive depression in the melting point and charge delocalization generated by hydrogen bonding (Usmani et al. 2023). Moreover, another green solvent that has been developed is natural deep eutectic solvents (NADESs). NADESs are a new revolutionary class of green media that have now emerged as significant endeavor (Mouden et al. 2017), prepared from natural combinations of sugars, polyalcohols, sugar-based alcohols, amino acids, and organic acids. These liquid supermolecules are made up of economically viable and readily available constituents such as quaternary ammonium salts (e.g., choline chloride or betaine), hydrogen bond acceptors (HBA), and naturally derived uncharged hydrogen bond donors (HBD) (such as amines, sugars, acids, and alcohols). These solvents have numerous benefits, including easy synthesis, tunable physicochemical qualities, low toxicity, high biodegradability, solute sustainability and stabilization, and a low melting point (Usmani et al. 2023). NADESs have been widely used as media for chemical and enzymatic reactions, essential oil extraction, nanocarrier for drug delivery, and antioxidant and antimicrobial agents (Liu et al. 2018). In this mini-review, we will discuss the antibacterial, and the antifungal, activities of NADESs towards a panel of microorganisms. The novel and critical aspects of addressing the global challenge of microbial infections using these natural materials are elaborated. This review addresses a pressing need for innovative solutions in the fight against microbial infections. As far as we know, this review represents the first comprehensive survey of the antimicrobial potentials and activities of NADESs in the literature.
NADESs preparation
NADES are typically prepared using one of six physical methods: heating and stirring, freeze-drying, evaporation, grinding, and ultrasound-assisted and microwave-assisted synthesis, as show in Fig. 1. Heating and stirring is the most commonly used method in NADESs preparation, due to its simplicity and low cost (Florindo et al. 2014; Castro et al. 2018). Another technique is vacuum evaporation, which involves heating a mixture of NADES components under reduced pressure to remove excess water and obtain a homogeneous liquid (Dai et al. 2013). Ultrasound-assisted synthesis utilizes sound waves to create cavitation, promoting the development of NADES (Rutkowska et al. 2017). Microwave-based methods subject the precursors to microwave energy, leading to molecular agitation and faster NADES formation (Gomez et al. 2018). These methods have been employed by different researchers, each with specific variations in the procedures and conditions. Overall, these diverse synthesis techniques contribute to the preparation and development of NADES. More information about these preparation methods can be find in recent reviews (Mišan et al. 2020; El Achkar et al. 2021; Omar and Sadeghi 2022; Wu et al. 2022).
Different methods for the preparation of natural deep eutectic solvents from malic acid and choline chloride, with the permission of Wu et al. (2022)
Techniques used to evaluate antimicrobial activity of NADESs
Different methods were used to evaluate the antimicrobial activity of NADESs against a broad category of microorganisms, including disc diffusion, well diffusion, broth dilution, Microtox assay, drop plate, and Fourier-transform infrared spectroscopy (FTIR) bioassay (Marchel et al. 2022). Among these methods, disc diffusion method was the most commonly used due to its ease of use (Marchel et al. 2022). In the next sections, we will briefly discuss each method.
The disc diffusion method
The disc diffusion method, also known as the Kirby-Bauer test or the agar diffusion test, was first applied to determine the sensitivity of microorganisms to antibiotics (Jorgensen and Turnidge 2009). Recently, this method has been expanded to study the antimicrobial effect of some chemical compounds such as ILs (Ventura et al. 2012) and DES (Hayyan et al. 2013). Nowadays, it is being used to evaluate the antimicrobial toxicity of NADESs (Marchel et al. 2022).
In this test, a filter paper disc that has been impregnated with the testing substance is placed on the surface of an agar plate that has previously been uniformly swabbed with microorganisms. Then, the plate is put in an incubator with optimum growth conditions (temperature, time). During the incubation time, the microorganisms are allowed to grow, and the tested compound diffuses from the agar plate, inhibiting the growth of the microorganism. The result is obtained by measuring a clear zone of inhibition and comparing it with clinical laboratory standard institutes (Jorgensen and Turnidge 2009).
The main benefits of the disk diffusion method are that it is a cost-effective procedure, easy to perform, and easy to evaluate. On the other hand, the main disadvantage of this approach is that it only allows determining if the tested agent is toxic or non-toxic to the microorganism (Marchel et al. 2022). Therefore, it is recommended to determine the minimum inhibitory concentration (MIC) for the same pair of tested compound and microorganism in order to obtain accurate results.
The well diffusion method is another technique, similar to the disk diffusion method (Juneidi et al. 2015). In the well diffusion method, the liquid antimicrobial agents are added to wells in the agar plate, instead of using paper or plastic disks, as in the disks diffusion method. Accordingly, the concentration of the antimicrobial agent depends on the volume added to the wells, which affects the consistency of the results. This may explain why the disk diffusion method is much more commonly used than the well diffusion method. The results found in the literature for the antibacterial activity of NADESs against microorganisms using the disk diffusion method are summarized in Table 1.
Agar and broth dilution technique
It has been reported that agar or broth dilution methods are among the most commonly used techniques to evaluate the antimicrobial activity of NADESs (Marchel et al. 2022). These methods allow for the determination of MIC of the studied antimicrobial agent.
In the agar dilution technique, a predetermined inoculum of microorganisms is directly applied onto nutrient agar plates containing different concentrations of the tested antimicrobial agent (Wiegand et al. 2008). The plates are then incubated under optimum conditions (e.g., time, temperature) for the growth of the tested microorganism. After the incubation period, the results are observed. Colonies on the plates indicate the growth of the microorganism, while the plate with the lowest concentration of the studied molecule where the bacteria did not grow specifies its MIC value (Wiegand et al. 2008).
The main advantage of the agar dilution method is that it is suitable for evaluating a large number of bacterial isolates against a small number of antimicrobial agents with limited concentrations (Kadlec et al. 2015). However, the main disadvantages of this method are that it is time-consuming and requires a large number of plates with varying concentrations of the antimicrobial agent (Kadlec et al. 2015). This makes it cost-ineffective due to the requirement of a substantial amount of the tested antimicrobial agent with different concentrations (Kadlec et al. 2015).
In the broth dilution method, the bacteria are grown in liquid nutrient medium containing the antibiotic in increasing concentrations (usually in twofold dilution series). A certain number of microbial cells are then inoculated into the medium. The main advantages of the broth dilution method are its ease of application, the ability to test the sensitivity of microorganisms to several chemicals simultaneously (Wiegand et al. 2008), and obtaining quantitative results. However, the disadvantages of this method are similar to those of the agar dilution method (Kadlec et al. 2015).
Microtox assay
In this assay, the toxicity of different substances can be determined by using bioluminescent bacteria Aliivibrio fischeri, which are non-pathogenic marine microorganisms that naturally luminesce as part of their metabolism (Johnson 2005). The toxic agent disrupts the respiratory process of these bacteria, resulting in a decrease in light output (Johnson 2005). The main advantages of this assay are its rapidity, ease of use, and sensitivity. In addition, a strain of bioluminescent bacteria is available in a lyophilized vial format, which increases their shelf life and usability (Johnson 2005). The in vitro toxicity of the NADESs was evaluated using the Microtox assay, for the first time, in the work of De Morais et al. (2015) who used cholinium chloride as hydrogen bond acceptor and organic acids as hydrogen bond donor. The results indicated that the four carboxylic acids (acetic, citric, lactic, and glycolic acids) presented a moderate toxicity (De Morais et al. 2015), in the following ascending order: acetic acid < lactic acid < citric acid < glycolic acid. It was observed that the effective concentration (EC50), in mg/L, obtained after 5 min was 30.9 for choline chloride: glycolic acid at molar ratio of 1:2 (De Morais et al. 2015). However, the level of toxicity of these different acids towards Aliivibrio fischeri was similar, except for acetic acid, being 4 to 5 times less than the other acids. Figure 2 illustrates the values of EC50, after different durations of exposure of the marine bacteria Vibrio fischeri to different NADES. Although organic acids resulted in lower EC50, and hence more antimicrobial efficiency, NADES are usually more potent than its starting compounds. These results show how Microtox assay can be beneficial to compare between newly prepared NADES.
Values of median EC50, in mg L−1, obtained after 5, 15, and 30 min of exposure of the marine bacteria Vibrio fischeri (Microtox.® toxicity test) to different chemical compounds, with the permission from De Morais et al. (2015)
Drop plate method
In this method, a 24-well plates for serial dilutions are used, followed by drop plating on agar in a 4 × 4 format using an automatic spiral plater (Chen et al. 2003; Wikene et al. 2017). After that, the plates are left to dry for a few minutes and then placed into an incubator for 18–20 h (37 °C) (Chen et al. 2003; Wikene et al. 2017). After incubation, viable colony forming units (CFUs) are counted and compared to control samples. Figure 3 illustrates the schematic of the 6 × 6 drop plate method. The bottom photograph depicts 6 drops × six 1 to 10 serial dilutions of Campylobacter jejuni. In the study conducted by Wikene and his coworkers, the drop plate method was used to assess the toxicity of NADESs. The tested NADESs included citric acid/sucrose and glucose/malic acid, while the model microorganisms included Escherichia coli and Enterococcus faecalis. Notably, the results unveiled that the 100-fold dilutions of the aforementioned NADESs exhibited minimal toxicity towards the bacterial strains, as evidenced by the absence of a significant reduction in colony-forming units (CFUs) relative to the untreated control samples (Wikene et al. 2015a). Additionally, this method was employed to evaluate the toxic effect of glucose/sucrose and choline chloride/maleic acid NADESs on Escherichia coli (Wikene et al. 2015b). The results revealed that the carbohydrate-based NADESs were non-toxic to Escherichia coli, with no significant reduction in the viable bacterial count (Wikene et al. 2015b). In addition, the antimicrobial properties of choline chloride/xylitol, malic acid/fructose/glucose, and citric acid/sucrose NADESs against three strains of bacteria were explored using the drop plate method [35]. The results showed that citric acid/sucrose NADES was non-toxic to all three bacterial strains (Escherichia coli, Enterococcus faecalis, and Staphylococcus aureus) (Wikene et al. 2016), which supported the findings of Wikene et al. (2015a).
Results of the drop plate method for studying Campylobacter jejuni (1 to 10 serial dilutions). With the permission of Chen et al. (2003)
FTIR-based biological assay
Another method used to evaluate the antimicrobial activity of NADESs depends on using FTIR spectroscopy, which is an instrumental technique used to elucidate the chemical composition of a substance by measuring the absorption, emission, or reflection of IR radiation. This technique is based on the principle that different chemical bonds vibrate at specific frequencies when exposed to IR. Thus, FTIR spectroscopy can be utilized to assess any changes in the molecular structure of the microorganisms or the NADESs themselves after exposure. FTIR is also capable of detecting changes in cell metabolites (Corte et al. 2010). This allows for a metabolomics analysis to be obtained. Since cells under stress exhibit rapid changes in their metabolites, this bioassay can estimate the toxicity level by measuring the variation in FTIR spectra of the cells upon exposure to the chemicals. Additionally, it provides metabolic indexes that can be used to classify and quantify the toxicity of the tested agent. The yeast strain of Saccharomyces cerevisiae cells has been commonly used as biosensor cells in this method. The main advantages of this approach are its speed and reproducibility (Corte et al. 2010). FTIR spectra are usually acquired using the potassium bromide (KBr) disc method in transmission mode in the range of 4000–400 cm−1. Sample pellets containing KBr were prepared by grinding in a mortar and pestle (Biswas et al. 2019). Background spectra from the KBr disc were subtracted from the spectra of the KBr and sample discs. The FTIR spectra of the untreated strains exhibit characteristic bands corresponding to biomolecules: lipids (3000–2800 cm−1), proteins/amides I and II (1700–1500 cm−1), phospholipids/DNA/RNA (1500–1185 cm−1), polysaccharides (1185–900 cm−1), and the fingerprint region (900–600 cm−1). The spectral analysis should reveal significant differences between untreated and treated samples, particularly in the absorbance of macromolecules such as proteins, nucleic acids, carbohydrates, and phospholipids, as shown in Fig. 4. These changes help elucidate the mechanism of action of the studied antimicrobial agent (Biswas et al. 2019).
A FTIR spectra of A. baumannii (ATCC) before (blue) and after treatment with 32 µg/mL of chlorhexidine (CHX, red), B before treatment, and C after treatment with 32 µg/mL CHX. The experiment was performed in triplicates. With the permission of Biswas et al. (2019)
Electron microscopy techniques
Electron microscopy serves as a powerful investigative tool in the exploration of the underlying mechanism of action associated with NADES as antimicrobial agents. The high resolution of scanning electron microscopy (SEM) provides researchers with valuable insight into the nature of interactions between NADES and microbes. A prominent application of SEM in these studies involves the close examination of their impact on the ultrastructure of microorganisms. Through the testing of NADES on the microorganism, followed by subsequent analysis using electron microscopy, researchers are able to report any observable alterations in cellular morphology or organelle structure, such as disrupting the integrity of the cell membrane, changing membrane permeability, or inducing a leakage of cellular contents. In this context, SEM plays an important role in elucidating the intracellular targets of antimicrobials by visualizing the internal structure of microorganisms. The SEM images in these experiments help determining the specific sites of action of NADES as antimicrobials. In addition, electron microscopy offers invaluable insights into the localization and distribution of these agents within microorganisms. Researchers can effectively employ electron-dense markers to label NADES, thereby facilitating the tracking of their uptake and subcellular localization (Durand et al. 2021). This information significantly aids in determining whether antimicrobial agents accumulate in specific organelles or interact with particular cellular components, thereby contributing to an enhanced understanding of their mechanism of action. Figure 5 illustrates the TEM images of Escherichia coli cells before and after treatment with the tested DES.
TEM images of Escherichia coli cells before any treatment A and after treatment with a suspension of L-menthol/oleic acid, 1:1. With the permission of Cao et al. (2020)
Antibacterial activities of NADESs
Antibacterial activity of NADESs against gram positive bacteria
Several NADESs were tested on gram-positive cocci, including Staphylococcus aureus, which can lead to inflammatory diseases such as skin infections, pneumonia, endocarditis, septic arthritis, osteomyelitis, and abscesses. It can also cause toxic shock syndrome (TSST-1), scalded skin syndrome (exfoliative toxin), and food poisoning (enterotoxin). Methicillin-resistant Staphylococcus aureus (MRSA), Methicillin-resistant Staphylococcus epidermis (MRSE), and Enterococcus faecalis are among the strains that have received great attention from the WHO due to their serious complications (Jubeh et al. 2020). Streptococcus pneumoniae was also used to evaluate the antibacterial activity of NADESs.
Among the gram-positive rod-shaped bacteria, spore-forming Clostridium perfringens (Wojeicchowski et al. 2021) and Bacillus subtilis (Mouffok et al. 2023) were tested. Listeria monocytogenes, a non-spore-forming gram-positive rod, was also included in the evaluation. Listeria monocytogenes can cause neonatal meningitis, meningitis in immunocompromised patients, gastroenteritis, and septicemia.
The best NADES that demonstrated large inhibition zones for MRSA, MRSE, and Staphylococcus aureus was menthol/lauric acid at a molar ratio of 4:1 (Oliveira et al. 2023). The inhibition zone sizes were 16.70 ± 0.47mm, 17.16 ± 0.62mm, and 14.17 ± 0.62mm, respectively, as measured by the disc diffusion method. The MIC for these bacteria, determined by the broth micro dilution assay using the same NADES, was 1250 µg/mL for MRSA and MRSE, and 625 µg/mL for Staphylococcus aureus. The results of the disc diffusion assay were compared to the positive control, gentamicin antibiotic, which showed inhibition zones of 19 mm for MRSA, 27.83 ± 0.62 mm for MRSE, and 36.33 ± 0.58 mm for Staphylococcus aureus (Oliveira et al. 2023).
It was also observed that capric acid/lauric acid at a molar ratio of 2:1 exhibited strong antimicrobial activity against MRSA and MRSE, with inhibition zone sizes of 16.50 ± 0.41 mm and 20.00 ± 0.82 mm, respectively. The inhibition zone for Staphylococcus aureus with the same NADES was 15.67 ± 0.58 mm. MIC values for these three strains were all equal to 625 µg/mL. Similar results were obtained for gentamicin as the positive control (Silva et al. 2019). These findings show the high potential of fatty acid-based NADES as antibacterial agents.
The largest inhibition zone for Staphylococcus aureus was achieved using choline chloride/oxalic acid at a molar ratio of 1:1, with an inhibition zone of 73 ± 3 mm (Radošević et al. 2018). Choline chloride/acetic acid at a molar ratio of 1:2 produced a strong inhibition zone of 50 ± 5 mm for spore-forming gram-positive bacteria Bacillus subtilis used (Mouffok et al. 2023). The MIC for Bacillus subtilis using the same NADES was 0.012 ± 0.002 µL/mL, as determined by the broth micro dilution method. Clostridium perfringens (Wojeicchowski et al. 2021), another spore-forming gram-positive bacteria associated with gas gangrene and food poisoning, showed an inhibition zone of 13–17 mm when treated with choline chloride/1,2-propanediol at a molar ratio of 1:2. Choline chloride/acetic acid at a molar ratio of 1:2 was found to be the best NADES for Listeria monocytogenes and Enterococcus faecalis, with inhibition zones of 40 ± 2 mm and 50 ± 5 mm, respectively (Mouffok et al. 2023).
Previous studies have shown that NADESs with strong inhibition zones often contain choline chloride as the hydrogen bond acceptor (HBA). This can be attributed to the interaction between the cholinium cation in NADES and the polysaccharide chains of peptidoglycan through hydrogen bonding or electrostatic forces, resulting in cell wall damage (Wen et al. 2015). Organic acids are primarily used as the hydrogen bond donor (HBD) in NADESs, and their ability to lower the pH below optimal values (pH = 6.5–7.5) for bacterial growth contributes to the inhibition of bacterial growth by organic acid-based NADESs (Zhao et al. 2015). Table 1 compares the antibacterial effects of the reported NADES.
Antibacterial activity of NADESs against gram-negative bacteria
As mentioned earlier, the WHO prioritized gram-negative bacteria in their list due to the presence of lipopolysaccharides on their outer cell wall. Gram-negative bacteria are often more resistant to antibiotics compared to gram-positive bacteria. These bacteria are associated with serious and life-threatening infections such as bloodstream, surgical site, complicated urinary tract, and lung infections (Rangarajan and Venkataraman 2020). They also contribute to increased mortality and morbidity rates. Some of the most challenging bacteria to treat include Escherichia coli, Klebsiella, Acinetobacter, Pseudomonas species, Proteus mirabilis, Salmonella typhimurium, and Serratia marcescens (Rangarajan and Venkataraman 2020). These bacteria exhibit resistance to multiple drugs and show increasing resistance against commonly available antibiotics. Additionally, they have the ability to develop new resistance mechanisms and transfer genetic material, making other bacteria drug-resistant as well. Therefore, it is crucial to search for alternatives to antibiotics as their effectiveness diminishes over time.
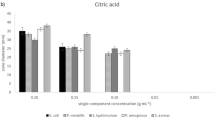
Different NADESs were tested on a panel of gram-negative bacteria, and the inhibition zone was determined. According to the literature, citric-based NADESs demonstrated the highest toxicity towards gram-negative bacteria. Specifically, citric acid/fructose/glycerol at a molar ratio of 1:1:1 was tested on Escherichia coli, Pseudomonas aeruginosa, Proteus mirabilis, and Salmonella typhimurium, resulting in inhibition zones of 50 ± 4 mm, 51 ± 4 mm, 81 ± 2 mm, and 55 ± 1 mm, respectively. This was reported as the largest inhibition zone achieved for these bacteria using NADESs (Radošević et al. 2018). In the same study, choline chloride/oxalic acid at a molar ratio of 1:1 was tested on various gram-positive and gram-negative bacteria (Radošević et al. 2018). It was observed that gram-positive bacteria exhibited larger inhibition zones compared to gram-negative bacteria. This can be attributed to the additional outer lipopolysaccharide membrane present in gram-negative bacteria, making them less permeable than gram-positive bacteria. Consequently, choline chloride/oxalic acid had a lesser inhibitory effect on Escherichia coli (47 ± 2 mm), Proteus mirabilis (49 ± 1 mm), Salmonella typhimurium (45 ± 4 mm), and Pseudomonas aeruginosa (50 ± 2 mm) compared to Staphylococcus aureus (73 ± 3 mm) (Radošević et al. 2018). For Klebsiella pneumoniae and Serratia marcescens, the inhibition zones were 50 ± 5 mm and 45 ± 5 mm, respectively. These results were obtained using choline chloride/acetic acid at a molar ratio of 1:2. The MIC values for Klebsiella pneumoniae and Serratia marcescens were 0.10 ± 0.05 µL/mL and 0.001 ± 0.005 µL/mL, respectively. Figure 6 illustrates the antibacterial potential of garlic extract and the NADESs used.
The antibacterial potential of garlic extract and the NADESs used with the permission of Mouffok et al. (2023)
It was also observed that these acid-based NADESs exhibited the strongest inhibition zones against bacteria compared to other NADESs (Mouffok et al. 2023). Therefore, it can be concluded that NADESs containing acids demonstrated higher antibacterial activity. Similar results and explanations apply to gram-positive bacteria as well (Zhao et al. 2015). Acids act as starting materials in NADESs and contribute to their antibacterial activities by creating an acidic environment, disrupting microbial membranes, and causing protein denaturation.
Mechanism of action of NADES against bacteria and fungi
There are several factors that explain how these NADES induce toxicity against different microorganisms. Factors that affect NADES toxicity could be divided according to the structure of the microorganisms and the chemical composition of the starting materials of certain NADES (Marchel et al. 2022). In addition, the method of preparation of NADES could also affect their toxicity.
According to the structure of microorganisms, gram-negative bacteria have extra lipopolysaccharides on their outer cell wall. It was observed that the toxicity of NADES decreases in gram-negative compared to gram-positive bacteria, which do not have lipopolysaccharides (Radošević et al. 2018). For example, when choline chloride/oxalic acid in a molar ratio of 1:1 was tested on both gram-positive and negative bacteria, the inhibition zone was larger in gram-positive bacteria, supporting this assumption (Radošević et al. 2018). It was also observed that the toxicity of the same NADES was lower in fungi compared to bacteria. Fungi have a two-layered cell wall mainly composed of chitin and glucans, which makes it difficult for the NADES to penetrate their cell wall. This difference in fungal structure could explain why choline chloride/oxalic acid/glycerol and choline chloride/citric acid/glycerol were found to be toxic to bacteria but not to yeast Candida albicans (Jangir et al. 2020).
In addition to factors related to the chemical structure of the starting material of the tested NADES, it was mentioned previously that NADES consist of a hydrogen bond donor and a hydrogen bond acceptor, each of which affects NADES toxicity. Regarding the hydrogen bond donor, it was found that organic acid-based NADES showed strong antimicrobial activity. This can be explained by the fact that bacteria and fungi have their optimum pH for growth, which is 6.5–7.5 for bacteria (Marchel et al. 2022) and 5.0–9.0 for fungi (Nevarez et al. 2009). Since organic acid-based NADES have pH values below 3, this high acidity increases their toxicity towards microbial cells. The acidic pH denatures proteins located on the microorganism’s cell wall, negatively affecting cell activity. This negative effect of pH theory is supported by the study of De Morais et al. (2015), who observed that the pH values of organic acid-based NADESs were lower than 3, resulting in protein denaturation and decreased Aliivibrio fischeri cell activity.
Continuing with the hydrogen bond donor, it was also reported that carbohydrate-based NADES have a negative effect on microbial cells. The highly viscous nature of carbohydrate-containing NADESs increases their toxic effect. Additionally, these NADESs have high osmotic pressure properties, which lead to negative dehydration effects that rapidly remove water from the microbial cell, causing cell dehydration and lysis. Moreover, high concentrations of the tested NADES generate high osmotic pressure, resulting in microbial cell dehydration. For example, in the work of Redovniković’s group, high concentrations of choline chloride/ethylene glycol and choline chloride/glucose caused high osmotic pressure and decreased viability of baker’s yeast cells (Cvjetko Bubalo et al. 2015). Therefore, high concentrations of carbohydrate-based NADES would increase their toxicity.
Regarding the hydrogen bond acceptor, it was observed that most of the HBA were choline chloride. This can be explained by the fact that choline chloride has a delocalized cation, and the higher toxicity of choline chloride-based NADESs is often attributed to the interaction of cholinium cation side chains and head groups with cellular membrane groups (Modica-Napolitano and Aprille 2001). It was also assumed that the accumulation of positively charged cations, such as cholinium, enhances electrostatic interactions with the negatively charged bilayer on the surface of cell membranes, leading to cell wall disruption (Wen et al. 2015). Finally according to method of preparation, it was found that heating and stirring method might result in impurities. The presence of impurities can change some of the mixture properties (e.g., by increasing their viscosities) and indirectly intensifying toxic effect of these NADESs (Florindo et al. 2014). Figure 7 summarizes different mechanisms of action of NADESs as antibacterial agents.
Antifungal activity of NADESs
Fungal infections are one of the most significant public health concerns (Reddy et al. 2022). They can have severe complications in individuals, especially those with various disorders, including Covid-19. Fungal infections can lead to life-threatening mycoses and death when they occur as coinfections with Covid-19. The severity of fungal infections varies depending on the fungal species and the type of infection (Reddy et al. 2022). These infections range from superficial, cutaneous, subcutaneous, mucosal, to systemic infections. However, systemic infections are often diagnosed late, leading to increased mortality rates. One of the most important pathogens associated with serious fungal infections is Candida spp., which are part of the human microbiota and can cause opportunistic infections in healthy individuals. They can also cause life-threatening infections such as invasive candidiasis in immunocompromised people, including those infected with human immunodeficiency virus (HIV) or receiving chemotherapy for cancer, as well as patients on immunosuppressive drugs (Reddy et al. 2022).
In addition to opportunistic and systemic infections, fungal pathogens such as Candida, Aspergillus, Fusarium, Mucorales, and molds can cause healthcare-associated infections (HAIs) in patients with underlying diseases (Perlroth et al. 2007). Due to increasing resistance to current antifungal medications, alternative strategies must be considered for effective antifungal therapy (Reddy et al. 2022). One of these alternative strategies is the use of natural products (Reddy et al. 2022). Several NADES were tested against different panels of fungi to evaluate their antifungal activity. Among the tested NADES, choline chloride/oxalic acid at a molar ratio of 1:1 exhibited a strong inhibition zone against Candida albicans (Radošević et al. 2018). The inhibition zone was measured as 48 ± 3 mm using the disc diffusion method (Radošević et al. 2018). This result supports the finding that organic acid-based NADES have stronger antimicrobial activity compared to alcohol-, amine-, and sugar-based NADES (Zhao et al. 2015). This can be explained by the introduction of an additional hydroxyl group to organic acids, which increases their antimicrobial activity as the hydrogen bond donor of the NADES (Zhao et al. 2015).
Other applications of NADES
In addition to the antimicrobial activity of NADES, they are also effective against biofilms, which represent a significant obstacle to the effective treatment of various infections in skin and soft tissues (Nystedt et al. 2023). NADES are also cytotoxic against tumor cells (Popović et al. 2023). Besides, it was reported that some NADES have a role in the extraction and purification of virus like particles (Marchel et al. 2020). Moreover they play an important role in antibacterial photodynamic therapy. These applications prove that NADES are promising alternatives to currently used materials not only as antimicrobials, but also in other fields of science.
Conclusion
Natural deep eutectic solvents (NADES) are considered one of the most promising classes of alternative organic solvents. Their properties, including low flammability, non-toxicity, low volatility, low cost, biodegradability, and ease of preparation, make them widely used in diverse fields, ranging from pharmaceuticals to energy. The fact that NADES exhibit antimicrobial properties against a panel of microorganisms opens up possibilities for their use as alternatives to classical antibiotics. Their antimicrobial activity has been observed to be strong and efficient when based on organic acids. However, it has been noted that the toxicity of NADES varies according to the type of bacteria and cell line, which can be explained by the interaction of these systems with bacterial cell membranes. Currently, it is necessary to evaluate their biological activities, such as antimicrobial effects, cytotoxicity, and antioxidant properties, in order to identify the best solvent for large-scale industrial use. Rapidly acquiring knowledge about their toxicity is crucial for exploring their potential in various applications, particularly in biomedicine and the pharmaceutical industry.
Data availability
This manuscript has no associated data.
Abbreviations
- ChCl :
-
Choline chloride
- CHX :
-
Chlorhexidine
- DES :
-
Deep eutectic solvents
- DNA :
-
Deoxyribonucleic acid
- EC50 :
-
Effective concentration
- FTIR :
-
Fourier-transform infrared spectroscopy
- GCE :
-
Garlic clove extract
- HBA :
-
Hydrogen bond acceptor
- HBD :
-
Hydrogen bond donor
- ILs :
-
Ionic liquids
- KBr :
-
Potassium bromide
- MAS :
-
Microwave-assisted synthesis
- MIC :
-
Minimum inhibitory concentration
- MRSA :
-
Methicillin-resistant Staphylococcus aureus
- MRSE :
-
Methicillin-resistant Staphylococcus epidermis
- NADES :
-
Natural DES
- NR :
-
Not reported
- PCA :
-
Principle component analysis
- RNA :
-
Ribonucleic acid
- RTILs :
-
Room-temperature ionic liquids
- SEM :
-
Scanning electron microscopy
- TSST-1 :
-
Toxic shock syndrome
- UAS :
-
Ultrasound-assisted synthesis
- VOCs :
-
Volatile oil compounds
- WHO :
-
World Health Organization
References
Al-Akayleh F, Khalid R, Hawash D, Al-Kaissi E, Al-Adham I, Al-Muhtaseb N, Jaber N, Al-Remawi M, Collier PJ (2022) Antimicrobial potential of natural deep eutectic solvents. Lett Appl Microbiol 75:607–615. https://doi.org/10.1111/lam.13699
Alsaud N, Shahbaz K, Farid M (2021) Antioxidant and antibacterial evaluation of Manuka leaves (Leptospermum scoparium) extracted by hydrophobic deep eutectic solvent. Chem Eng Res Des 174:96–106. https://doi.org/10.1016/j.cherd.2021.08.004
Biswas D, Tiwari M, Tiwari V (2019) Molecular mechanism of antimicrobial activity of chlorhexidine against carbapenem-resistant Acinetobacter baumannii. PLoS One 14:1–17. https://doi.org/10.1371/journal.pone.0224107
Cao J, Wu R, Dong Q, Zhao L, Cao F, Su E (2020) Effective release of intracellular enzymes by permeating the cell membrane with hydrophobic deep eutectic solvents. ChemBioChem 21:672–680. https://doi.org/10.1002/cbic.201900502
Castro VIB, Craveiro R, Silva JM, Reis RL, Paiva A, Ana AR (2018) Natural deep eutectic systems as alternative nontoxic cryoprotective agents. Cryobiology 83:15–26. https://doi.org/10.1016/j.cryobiol.2018.06.010
Chen CY, Nace GW, Irwin PL (2003) A 6x6 drop plate method for simultaneous colony counting and MPN enumeration of Campylobacter jejuni, Listeria monocytogenes, and Escherichia coli. J Microbiol Methods 55:475–479. https://doi.org/10.1016/S0167-7012(03)00194-5
Corte L, Rellini P, Roscini L, Fatichenti F, Cardinali G (2010) Development of a novel, FTIR (Fourier transform infrared spectroscopy) based, yeast bioassay for toxicity testing and stress response study. Anal Chim Acta 659:258–265. https://doi.org/10.1016/j.aca.2009.11.035
Cvjetko Bubalo M, Mazur M, Radošević K, Radojčić Redovniković I (2015) Baker’s yeast-mediated asymmetric reduction of ethyl 3-oxobutanoate in deep eutectic solvents. Process Biochem 50:1788–1792. https://doi.org/10.1016/j.procbio.2015.07.015
Dai Y, van Spronsen J, Witkamp G-J, Verpoorte R, Choi YH (2013) Natural deep eutectic solvents as new potential media for green technology. Anal Chim Acta 766:61–68. https://doi.org/10.1016/j.aca.2012.12.019
De Morais P, Gonçalves F, Coutinho JAP, Ventura SPM (2015) Ecotoxicity of cholinium-based deep eutectic solvents. ACS Sustain Chem Eng 3:3398–3404. https://doi.org/10.1021/acssuschemeng.5b01124
Durand E, Villeneuve P, Bourlieu-lacanal C, Carrière F (2021) Natural deep eutectic solvents: hypothesis for their possible roles in cellular functions and interaction with membranes and other organized biological systems. Advances in Botanical Research. Elsevier, Amsterdam, Netherlands, pp 133–158
El Achkar T, Greige-Gerges H, Fourmentin S (2021) Basics and properties of deep eutectic solvents: a review. Environ Chem Lett 19:3397–3408. https://doi.org/10.1007/s10311-021-01225-8
Fernandes MM, Carvalho EO, Correia DM, Esperança JMSS, Padrão J, Ivanova K, Hoyo J, Tzanov T, Lanceros-Mendez S (2022) Ionic liquids as biocompatible antibacterial agents: a case study on structure-related bioactivity on Escherichia coli. ACS Appl Bio Mater 5:5181–5189. https://doi.org/10.1021/acsabm.2c00615
Florindo C, Oliveira FS, Rebelo LPN, Fernandes AM, Marrucho IM (2014) Insights into the synthesis and properties of deep eutectic solvents based on cholinium chloride and carboxylic acids. ACS Sustain Chem Eng 2:2416–2425. https://doi.org/10.1021/sc500439w
Garrido A, Atencio LA, Bethancourt R, Bethancourt A, Guzmán H, Gutiérrez M, Durant-Archibold AA (2020) Antibacterial activity of volatile organic compounds produced by the octocoral-associated bacteria Bacillus sp. Bo53 and Pseudoalteromonas sp. ga327. Antibiotics 9:1–10. https://doi.org/10.3390/antibiotics9120923
Gomez FJV, Espino M, Fernández MA, Silva MF (2018) A greener approach to prepare natural deep eutectic solvents. ChemistrySelect 3:6122–6125. https://doi.org/10.1002/SLCT.201800713
Hayyan M, Hashim MA, Hayyan A, Al-Saadi MA, AlNashef IM, Mirghani MES, Saheed OK (2013) Are deep eutectic solvents benign or toxic? Chemosphere 90:2193–2195. https://doi.org/10.1016/j.chemosphere.2012.11.004
Jangir AK, Lad B, Dani U, Shah N, Kuperkar K (2020) In vitro toxicity assessment and enhanced drug solubility profile of green deep eutectic solvent derivatives (DESDs) combined with theoretical validation. RSC Adv 10:24063–24072. https://doi.org/10.1039/c9ra10320a
Johnson BT (2005) Microtox® acute toxicity test. Small-Scale Freshw Toxic Investig Vol 1. Toxic Test Methods 1:69–105. https://doi.org/10.1007/1-4020-3120-3_2
Jorgensen JH, Turnidge JD (2009) Kirby-Bauer disk diffusion susceptibility test protocol. Am Soc Microbiol 15:55–63
Jubeh B, Breijyeh Z, Karaman R (2020) Resistance of gram-positive bacteria to current antibacterial agents and overcoming approaches. Molecules 25:1–22. https://doi.org/10.3390/molecules25122888
Juneidi I, Hayyan M, Hashim MA (2015) Evaluation of toxicity and biodegradability for cholinium-based deep eutectic solvents. RSC Adv 5:83636–83647. https://doi.org/10.1039/c5ra12425e
Kadlec K, Wendlandt S, Feßler AT, Schwarz S (2015) Methods for the detection of antimicrobial resistance and the characterization of Staphylococcus aureus isolates from food-producing animals and food of animal origin. In: Antimicrobial Resistance and Food Safety. Elsevier, 207–232
Liang Y, Pan Z, Chen Z, Fei Y, Zhang J, Yuan J, Zhang L, Zhang J (2020) Ultrasound-assisted natural deep eutectic solvents as separation-free extraction media for hydroxytyrosol from olives. ChemistrySelect 5:10939–10944. https://doi.org/10.1002/slct.202002026
Liu Y, Friesen JB, McAlpine JB, Lankin DC, Chen S-N, Pauli GF (2018) Natural deep eutectic solvents: properties, applications, and perspectives. J Nat Prod 81:679–690. https://doi.org/10.1021/acs.jnatprod.7b00945
Łojewska E, Sakowicz T (2021) An alternative to antibiotics: selected methods to combat zoonotic foodborne bacterial infections. Curr Microbiol 78:4037–4049. https://doi.org/10.1007/s00284-021-02665-9
Mano F, Martins M, Sá-Nogueira I, Barreiros S, Borges JP, Reis RL, Duarte ARC, Paiva A (2017) Production of electrospun fast-dissolving drug delivery systems with therapeutic eutectic systems encapsulated in gelatin. AAPS PharmSciTech 18:2579–2585. https://doi.org/10.1208/s12249-016-0703-z
Mao S, Li K, Hou Y, Liu Y, Ji S, Qin H, Lu F (2018) Synergistic effects of components in deep eutectic solvents relieve toxicity and improve the performance of steroid biotransformation catalyzed by Arthrobacter simplex. J Chem Technol Biotechnol 93:2729–2736. https://doi.org/10.1002/jctb.5629
Marchel M, Niewisiewicz J, Coroadinha AS, Marrucho IM (2020) Purification of virus-like particles using aqueous biphasic systems composed of natural deep eutectic solvents. Sep Purif Technol 252:117480. https://doi.org/10.1016/j.seppur.2020.117480
Marchel M, Cieśliński H, Boczkaj G (2022) Deep eutectic solvents microbial toxicity: current state of art and critical evaluation of testing methods. J Hazard Mater 425:127963. https://doi.org/10.1016/j.jhazmat.2021.127963
Mišan A, Nađpal J, Stupar A, Pojić M, Mandić A, Verpoorte R, Choi YH (2020) The perspectives of natural deep eutectic solvents in agri-food sector. Crit Rev Food Sci Nutr 60:2564–2592. https://doi.org/10.1080/10408398.2019.1650717
Modica-Napolitano JS, Aprille JR (2001) Delocalized lipophilic cations selectively target the mitochondria of carcinoma cells. Adv Drug Deliv Rev 49:63–70. https://doi.org/10.1016/S0169-409X(01)00125-9
Mouden S, Klinkhamer PGL, Choi YH, Leiss KA (2017) Towards eco-friendly crop protection: natural deep eutectic solvents and defensive secondary metabolites. Phytochem Rev 16:935–951. https://doi.org/10.1007/s11101-017-9502-8
Mouffok A, Bellouche D, Debbous I, Anane A, Khoualdia Y, Boublia A, Darwish AS, Lemaoui T, Benguerba Y (2023) Synergy of garlic extract and deep eutectic solvents as promising natural antibiotics: experimental and COSMO-RS. J Mol Liq 375:121321. https://doi.org/10.1016/j.molliq.2023.121321
Nevarez L, Vasseur V, Le Madec A, Le Bras MA, Coroller L, Leguérinel I, Barbier G (2009) Physiological traits of Penicillium glabrum strain LCP 08.5568, a filamentous fungus isolated from bottled aromatised mineral water. Int J Food Microbiol 130:166–171. https://doi.org/10.1016/j.ijfoodmicro.2009.01.013
Ng WJ, Shit CS, Ee KY, Chai TT (2021) Plant natural products for mitigation of antibiotic resistance. In: Sustainable agriculture reviews, Springer Nature Cham, Switzerland, 57–91. https://doi.org/10.1007/978-3-030-58259-3_3
Nystedt HL, Grønlien KG, Rolfsnes RR, Winther-Larsen HC, Løchen Økstad OA, Tønnesen HH (2023) Neutral natural deep eutectic solvents as anti-biofilm agents. Biofilm 5. https://doi.org/10.1016/j.bioflm.2023.100114
Oliveira F, Silva E, Matias A, Silva JM, Reis RL, Duarte ARC (2023) Menthol-based deep eutectic systems as antimicrobial and anti-inflammatory agents for wound healing. Eur J Pharm Sci 182:106368. https://doi.org/10.1016/j.ejps.2022.106368
Omar KA, Sadeghi R (2022) Physicochemical properties of deep eutectic solvents: a review. J Mol Liq 360:119524. https://doi.org/10.1016/j.molliq.2022.119524
Perlroth J, Choi B, Spellberg B (2007) Nosocomial fungal infections: epidemiology, diagnosis, and treatment. Med Mycol 45:321–346. https://doi.org/10.1080/13693780701218689
Popović BM, Gligorijević N, Aranđelović S, Macedo AC, Jurić T, Uka D, Mocko-Blažek K, Serra AT (2023) Cytotoxicity profiling of choline chloride-based natural deep eutectic solvents. RSC Adv 13:3520–3527. https://doi.org/10.1039/d2ra07488e
Radošević K, Čanak I, Panić M, Markov K, Bubalo MC, Frece J, Srček VG, Redovniković IR (2018) Antimicrobial, cytotoxic and antioxidative evaluation of natural deep eutectic solvents. Environ Sci Pollut Res 25:14188–14196. https://doi.org/10.1007/s11356-018-1669-z
Rangarajan R, Venkataraman R (2020) Antibiotics targeting gram-negative bacteria. In: Drug discovery targeting drug-resistant bacteria. Academic Press, Cambridge, Massachusetts, USA, pp 39–70. https://doi.org/10.1016/B978-0-12-818480-6.00003-5
Reddy GKK, Padmavathi AR, Nancharaiah YV (2022) Fungal infections: pathogenesis, antifungals and alternate treatment approaches. Curr Res Microb Sci 3:100137. https://doi.org/10.1016/j.crmicr.2022.100137
Rodríguez-Juan E, López S, Abia R, JG Muriana F, Fernández-Bolaños J, García-Borrego A (2021) 150x10^3 mg/L. J Mol Liq 337. https://doi.org/10.1016/j.molliq.2021.116343
Rutkowska M, Namieśnik J, Konieczka P (2017) Ultrasound-assisted extraction. In: The application of green solvents in separation processes. Elsevier, Amsterdam, pp 301–324. https://doi.org/10.1016/B978-0-12-805297-6.00010-3
Silva JM, Silva E, Reis RL, Duarte ARC (2019) A closer look in the antimicrobial properties of deep eutectic solvents based on fatty acids. Sustain Chem Pharm 14:100192. https://doi.org/10.1016/j.scp.2019.100192
Syed UT, Leonardo I, Lahoz R, Gaspar FB, Huertas R, Crespo MTB, Arruebo M, Crespo JG, Sebastian V, Brazinha C (2020) Microengineered membranes for sustainable production of hydrophobic deep eutectic solvent-based nanoemulsions by membrane emulsification for enhanced antimicrobial activity. ACS Sustain Chem Eng 8:16526–16536. https://doi.org/10.1021/acssuschemeng.0c05612
Usmani Z, Sharma M, Tripathi M, Lukk T, Karpichev Y, Gathergood N, Singh BN, Thakur VK, Tabatabaei M, Gupta VK (2023) Biobased natural deep eutectic system as versatile solvents: structure, interaction and advanced applications. Sci Total Environ 881:163002. https://doi.org/10.1016/J.SCITOTENV.2023.163002
Ventura SPM, de Barros RLF, Sintra T, Soares CMF, Lima ÁS, Coutinho JAP (2012) Simple screening method to identify toxic/non-toxic ionic liquids: agar diffusion test adaptation. Ecotoxicol Environ Saf 83:55–62. https://doi.org/10.1016/j.ecoenv.2012.06.002
Wen Q, Chen JX, Tang YL, Wang J, Yang Z (2015) Assessing the toxicity and biodegradability of deep eutectic solvents. Chemosphere 132:63–69. https://doi.org/10.1016/j.chemosphere.2015.02.061
Wiegand I, Hilpert K, Hancock REW (2008) Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3:163–175. https://doi.org/10.1038/nprot.2007.521
Wikene KO, Bruzell E, Tønnesen HH (2015a) Improved antibacterial phototoxicity of a neutral porphyrin in natural deep eutectic solvents. J Photochem Photobiol B Biol 148:188–196. https://doi.org/10.1016/j.jphotobiol.2015.04.022
Wikene KO, Bruzell E, Tønnesen HH (2015b) Characterization and antimicrobial phototoxicity of curcumin dissolved in natural deep eutectic solvents. Eur J Pharm Sci 80:26–32. https://doi.org/10.1016/j.ejps.2015.09.013
Wikene KO, Rukke HV, Bruzell E, Tønnesen HH (2016) Physicochemical characterisation and antimicrobial phototoxicity of an anionic porphyrin in natural deep eutectic solvents. Eur J Pharm Biopharm 105:75–84. https://doi.org/10.1016/j.ejpb.2016.06.001
Wikene KO, Rukke HV, Bruzell E, Tønnesen HH (2017) Investigation of the antimicrobial effect of natural deep eutectic solvents (NADES) as solvents in antimicrobial photodynamic therapy. J Photochem Photobiol B Biol 171:27–33. https://doi.org/10.1016/j.jphotobiol.2017.04.030
Wojeicchowski JP, Marques C, Igarashi-Mafra L, Coutinho JAP, Mafra MR (2021) Extraction of phenolic compounds from rosemary using choline chloride – based deep eutectic solvents. Sep Purif Technol 258. https://doi.org/10.1016/j.seppur.2020.117975
Wu K, Ren J, Wang Q, Nuerjiang M, Xia X, Bian C (2022) Research progress on the preparation and action mechanism of natural deep eutectic solvents and their application in food. Foods 11:3528. https://doi.org/10.3390/foods11213528
Zhao BY, Xu P, Yang FX, Wu H, Zong MH, Lou WY (2015) Biocompatible deep eutectic solvents based on choline chloride: characterization and application to the extraction of rutin from Sophora japonica. ACS Sustain Chem Eng 3:2746–2755. https://doi.org/10.1021/acssuschemeng.5b00619
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
FRM proposed the mini-review idea and structure. HMB performed the literature review. HMB, TS and FRM drafted the manuscript. TS and FRM critically reviewed the draft. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bedair, H.M., Samir, T.M. & Mansour, F.R. Antibacterial and antifungal activities of natural deep eutectic solvents. Appl Microbiol Biotechnol 108, 198 (2024). https://doi.org/10.1007/s00253-024-13044-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13044-2