Abstract
Pseudogenes are defined as “non-functional” copies of corresponding parent genes. The cognition of pseudogenes continues to be refreshed through accumulating and updating research findings. Previous studies have predominantly focused on mammals, but pseudogenes have received relatively less attention in the field of microbiology. Given the increasing recognition on the importance of pseudogenes, in this review, we focus on several aspects of microorganism pseudogenes, including their classification and characteristics, their generation and fate, their identification, their abundance and distribution, their impact on virulence, their ability to recombine with functional genes, the extent to which some pseudogenes are transcribed and translated, and the relationship between pseudogenes and viruses. By summarizing and organizing the latest research progress, this review will provide a comprehensive perspective and improved understanding on pseudogenes in microorganisms.
Key points
• Concept, classification and characteristics, identification and databases, content, and distribution of microbial pseudogenes are presented.
• How pseudogenization contribute to pathogen virulence is highlighted.
• Pseudogenes with potential functions in microorganisms are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudogenes were first discovered as truncated and non-functional copies of the gene encoding 5S RNA in the genome of Xenopus laevis (Jacq et al. 1977). Many of pseudogenes have now been identified in all kingdoms of life, ranging from inferior organism to human and plants (Harrison et al. 2003; Torrents et al. 2003; Xie et al. 2019; Zhang et al. 2004). Pseudogenes are defined as defective nucleic acid sequences related to functional genes (or parent genes). In bacteria, pseudogenes, are defined here as “genes that have been silenced by one or more deleterious mutations” (Goodhead and Darby 2015). Relative to their functional homologs, pseudogenes are characterized by a variety of obvious defects in sequence, including insertion or deletion of nucleotides, frameshift, premature stop codons, and truncation. There are two ways to designate pseudogenes: One way is to mark the symbol “ψ” prior to the gene name, such as “ψfepE” (Hiyoshi et al. 2018), and the other way is to add the letter “P” after the gene name, such as “CYP4Z2P” (Zheng et al. 2016).
Initially, pseudogenes were regarded as “gene fossils” or “junk DNA,” but continued research has demonstrated biological function for many pseudogenes. Some pseudogenes produce a variety of functional RNA that play an important role in gene regulation and other physiological or pathological processes (Chen et al. 2020; Lai et al. 2019; Lou et al. 2020; Tam et al. 2008). These genes currently considered as pseudogenes, but these genes have potential functions. Some pseudogenes are also related to the occurrence, development, prognosis, and therapeutic targets of cancers (Chen et al. 2018; Kalyana-Sundaram et al. 2012; Kwon et al. 2021; Sisu 2021b; Sun et al. 2021).
Previous research on pseudogenes have mostly focused on mammals or plants, while studies or reviews on microbial pseudogenes are relatively rare. Initially, it was thought that microorganisms lacked or had very few pseudogenes. However, in recent years, the prevalence and importance of pseudogenes in microorganisms have gradually been recognized. In this review, we comprehensively summarize the history and recent progress in the field of microbial pseudogenes, including their concept, classification, characteristics, generation and fate, content, and distribution, as well as their impact on virulence. Additionally, consideration is also given to pseudogenes that may have potential functions. Further study of microbial pseudogenes will influence our understanding of pathogen virulence and molecular genetics.
Classification and characteristics of pseudogenes
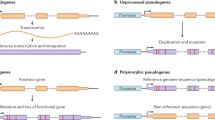
According to the sequence characteristics and mechanism of generation, pseudogenes can be divided into three major types; this includes duplicated pseudogenes, retro-pseudogenes, and circular RNA-derived pseudogenes.
Duplicated (or unprocessed) pseudogene
Pseudogene which arise from a gene duplication event and subsequent disabling mutation which is neither transcribed nor translated are called duplicated or unprocessed pseudogenes (Frankish and Harrow 2014). The deleterious mutations in nucleotide sequence include base deletion or insertion, premature stop codons and frameshift mutations, etc., which prevent the gene successful expression. Duplicated pseudogenes are very similar to parental genes in gene structure and retain much of the original sequence and structure, such as the promoter, intron, and exon (Fig. 1A) (Pei et al. 2012; Torrents et al. 2003).
Retropseudogene (or processed) pseudogene
Pseudogene created by a retro-transposition event and subsequent disabling mutation which is neither transcribed nor translated are called retropseudogene or processed pseudogenes (Frankish and Harrow 2014). DNA is transcribed into mRNA and then reverse-transcribed into cDNA and re-integrated into the genome; during this process, the gene function is lost due to the inappropriate insertion site or the mutation of the sequence, resulting in the formation of pseudogenes. The features of retropseudogenes are as follows: (1) lack the region of introns; (2) lack the region of promoter (the randomness of the position of the retroposition inserted into the genome); (3) the flank contains untranslated region (UTR) structure, and the 3′ end contains a polyA tail (derived from mRNA, so it has characteristic structure of mRNA) (Fig. 1B) (Pei et al. 2012; Torrents et al. 2003).
Circular RNA (CircRNA)–derived pseudogene
In addition to the above two pseudogenization mechanisms, researchers have recently discovered a new way of producing pseudogenes in mammals.
The pseudogene created by reverse transcription of circRNAs is called a circRNA-derived pseudogene (Dong et al. 2016). CircRNAs are a class of endogenous RNAs characterized by a covalently closed loop structure and without a 5′ cap and a 3′poly A tail (Verduci et al. 2021). Unlike linear RNAs, circRNAs are usually formed by reverse splicing events of exons or introns, that is, the downstream splicing site is connected to the upstream splicing site in reverse. Theoretically, a linear mRNA-derived pseudogene keeps the same sequential order (colinear) of exons as in its parent linear mRNA. In contrast, a circRNA-derived pseudogene can have an exon-exon junction in a reversed order (non-colinear). In addition, there is no polyA tail at the end of the sequence (Fig. 1C) (Dong et al. 2016; Li et al. 2018).
Generation and fate of pseudogenes in microorganisms
Generation of pseudogenes in microorganisms
DNA duplication and retro-transposition are the two main ways of generating pseudogenes in eukaryotes (Torrents et al. 2003). The formation of pseudogenes may differ between eukaryotes and prokaryotes. Phylogenomic analysis in prokaryotes indicated two mechanisms for pseudogene formation: (1) disruption of native genes and (2) failed horizontal gene transfer (HGT) events (Avni et al. 2018). The form of native gene disruption is diverse, including the point mutations, the presence of premature stop-codons, frame shift, etc. (Lerat and Ochman 2004). HGT is the process of an organism acquiring non-parental genetic information (Darmon and Leach 2014). HGT event between bacteria are common. HGT is one of the important mechanisms contributing to genetic diversity, microbial antibiotic resistance, survival, pathogenicity, and adaptation of bacteria (Darmon and Leach 2014; Soucy et al. 2015). It also plays an essential role in the evolution and speciation of bacteria (Darmon and Leach 2014). Liu et al. used a GC-content method to estimate horizontal transferred genes and reported that a large part of bacterial pseudogenes originated from failed HGT events (Liu et al. 2004).
Fate of pseudogenes in microorganisms
Once established in a genome, pseudogenes evolve over time. In eukaryotes, pseudogenes often persist for long time during evolution. Moreover, some pseudogenes may be shared by distant relative lineages, such as rodents and primates (Zhang et al. 2004). However, reports reveal that, unlike eukaryotes, in bacteria, pseudogenes are usually deleted relatively rapidly from genomes, suggesting that their presence is deleterious to some extent (Kuo and Ochman 2010). During evolution, the continuous accumulation of mutations is one of the reasons for the exclusion of pseudogenes. Energy-consuming transcription or the detrimental product is also one of the reasons for the rapidly elimination of pseudogenes. Many other pseudogenes persist and evolve over time. Overall, the different evolutionary processes and selective pressures lead to retention or clearing of pseudogenes from the genome of microorganism over time.
Identification of pseudogenes
Accurate identification and annotation of pseudogenes are the foundations and prerequisites of the research on pseudogene. Currently, the general process of pseudogene identification within the scope of genome is mainly as follows: (i) Firstly, a set of genome data is selected as the reference sequence; (ii) then, a group pseudogene is screened as candidate library based on their sequence homology with the parental gene, sequence defects, and the unique structural characteristics of pseudogenes, such as no exon structure and polyA tail structure; (iii) afterwards, incorrectly identified pseudogenes will be eliminated through different methods, such as manual inspection (Pei et al. 2012).
In fact, correctly distinguishing pseudogenes from parental genes is an extremely difficult process. It is worth noting that the misannotation of functional genes as pseudogenes can occur due to sequencing errors, which vary a lot depending on the sequencing method employed. By analyzing the mutation characteristics of known pseudogenes, a threshold can be deduced and utilized to distinguish between pseudogenes and functional genes.
Pseudogene identification tools
Several bioinformatics tools are now available to predict pseudogenes. At present, commonly pseudogene identification methods include the following: PseudoPipe (Zhang et al. 2006), which was developed by the Gerstein team at Yale University, is suitable for identification of pseudogenes in mammals; PseudoFinder (Zheng et al. 2007) (annotation pseudogene based on the gene homology) and Retrofinder (Zheng et al. 2007) (focus on annotation of processed pseudogenes), which were developed by the University of California; and HAVANA (Human and Vertebrate Analysis and Annotation), which focus on manual annotation.
Moreover, sideRETRO, a pipeline dedicated to identifying processed pseudogenes, was developed in recent years (Miller et al. 2021). Abrahamsson et al. reported the PΨFinder (P-psy-finder), a tool that can identify processed pseudogenes from DNA sequencing data and predicts their location in the genome (Abrahamsson et al. 2022).
Besides, CIRCpseudo (Dong et al. 2016), which was conceived by Yang team from Shanghai Institute of Biological Sciences, Chinese Academy of Sciences, may be used to discover and identify potential pseudogene sequences derived from circRNAs in the genome.
Lerat et al. designed a suite of programs named “Ψ-Φ” (short for Ψ-gene Finder) to identify pseudogenes in bacteria genomes (Lerat and Ochman 2004). Recently, researchers published a new open-source software Pseudofinder that can be used for identification and analysis pseudogenes in in bacterial and archaeal genomes (Syberg-Olsen et al. 2022). Furthermore, new pan-genomic analysis tools such as PEPPAN can help to analyze and identify pseudogenes in the bacterial genome (Zhou et al. 2020b).
At present, obtaining the whole genome sequence of bacteria is convenient, but the annotation of pseudogene is delayed (Goodhead and Darby 2015). The challenges of many annotation tools of genome lead to pseudogene misannotation or nonrecognition (Lerat and Ochman 2004; Zhou et al. 2020a). Although some genes were identified and annotated as “pseudogenes” due to their pseudogene-like features, they may produce viable transcripts and proteins. In addition, new evidence is emerging regarding the potential expression and implication of pseudogenes in gene regulation. In the strict sense, such “pseudogenes” cannot be called pseudogenes by the definition. The determination of their functionality relies on the research conducted by the corresponding laboratory. In recent years, the development of novel biological information resources, such as Prokaryotic Genome Annotation Pipeline (PGAP) and Reference Sequence (RefSeq) project, has greatly improved the quality of prokaryotic genome annotation and the ability to distinguish pseudogenes (Li et al. 2021; Sayers et al. 2020; Tatusova et al. 2016).
Pseudogene database
Some pseudogene databases have been built to collect, update, and publish pseudogene information, for example, psiDR (Pei et al. 2012), which was generated by GENCODE pseudogene resource project, is a consensus platform that integrates multiple pipelines. There are also other online available pseudogene databases, such as pseudoMap (Chan et al. 2013), PseudoFam (Lam et al. 2009), PseudoGene (pseudogene.org/), PseudoFuN (Johnson et al. 2019), and Dreambase (Zheng et al. 2018). Similar to eukaryotic genomes, the detection of pseudogenes within prokaryotic genomes relies on aligning them with parent genes and subsequently identifying sequence defects. Available databases for prokaryotic pseudogenes are mainly found in the “Prokaryote Pseudogene Information Site” (Liu et al. 2004).
In general, the factors affecting the quality of pseudogene annotations mainly include the following: (1) the quality of the reference genome. This is because pseudogenes may be incorrectly annotated if the reference genome is incorrectly annotated or contaminated; (2) quality of pseudogene identification pipeline and processes (Frankish and Harrow 2014). Different design principles and processes may lead to differences in the number of pseudogenes eventually identified and the accuracy of pseudogenes between different algorithms. Due to the different principles and processes of each pipeline, the results obtained by each pipeline may be various. For example, the number of pseudogenes eventually identified and the accuracy of annotation may be different. Furthermore, genome annotation may be challenging in some cases and might lead to confusing situations where real genes will be identified as pseudogenes and vice versa. However, there is so far no pipeline to detect pseudogenes that can take into account the requirement of both high-throughput and accuracy. The annotation accuracy can be improved by the following ways: selecting high-quality parent genome data as reference database and selecting multiple strategies combined with multiple algorithms for prediction. By summarizing and analyzing the limitations of existing algorithms, new prediction algorithms and processes can be improved and developed to help annotate, identify, and study more pseudogenes.
Pseudogenes in microorganisms
Pseudogenes have been reported and well-studied mostly in mammalian genomes, such as human and mouse (Torrents et al. 2003; Zhang et al. 2004). Limited concern has been devoted into pseudogene studies in microorganisms. Previously, pseudogenes are generally believed to be rare in microbial genomes. In fact, pseudogenes have been found in a variety of microorganisms. For instance, in Salmonella enterica subsp. enterica serovar Typhi (S. Typhi), Parkhill et al. identified 204 pseudogenes from a gene complement of 4599 genes (Parkhill et al. 2001). In Salmonella enterica subsp. enterica serovar Typhimurium (S. Typhimurium), several pseudogenes corresponding to genes are known to contribute to virulence (Parkhill et al. 2001). A total of 95 and 101 pseudogene candidates in whole-genome of Escherichia coli (E. coli) strains K-12 and O157, respectively, were reported (Homma et al. 2002). Besides, pseudogenes sequences have been observed within the genome of other bacteria species, such as Brucella (Bialer et al. 2021), Shigella (Cervantes-Rivera et al. 2020), Spirochete (Liu et al. 2022a), Staphylococcus (Chieffi et al. 2020), and Fungi (Oh et al. 2021; Shimizu et al. 2021). In addition, abundant pseudogenes have also been found in the obligate parasite Mycobacterium leprae (M. leprae) (Cole et al. 2001; Silva et al. 2022), Rickettsia (Andersson et al. 1998; Liu et al. 2004), and Anaplasma (Lin et al. 2023). Meanwhile, pseudogenes have also been reported in Archaea (Badel et al. 2020).
The content of pseudogenes
A comprehensive comparison and analysis data from 64 prokaryote genome sequences find a total of around 7000 pseudogene candidates, meanwhile indicating that pseudogenes are pervasive, ranging from 1 to 5% of the genome in most prokaryotes (Liu et al. 2004). These pseudogenes are associated with diverse protein families, such as ABC transporters, cytochrome P450 and PPE (proline-proline-glutamic acid) families (PF00067 and PF00823), and others that have a direct role in DNA transposition (Liu et al. 2004). The 64 genomes are divided into three categories by researchers: Archaea, pathogenic bacteria, and non-pathogenic bacteria, which include 3.6%, 3.9%, and 3.3% pseudogene in their genomes, respectively (Liu et al. 2004). However, another study indicated that pseudogenes appear to be more common in the genomes of pathogenic bacteria than in other non-pathogenic organisms (Lerat and Ochman 2005).
Intracellular lifestyle
Some intracellular parasites contain more pseudogenes. In the genome of M. leprae, an obligate intracellular organism causing human leprosy, has a high proportion of pseudogenes (36.5%) (Liu et al. 2004). M. leprae has a relatively small genome (3.3 Mbp) compared to other mycobacterial species. The formation of pseudogenes eliminated many important metabolic pathways involved in siderophore production, oxidative and respiratory chains, lipid biosynthesis and metabolism, catabolic systems, and their regulatory circuits (Cole et al. 2001; Gomez-Valero et al. 2007; Liu et al. 2004; Sugawara-Mikami et al. 2022). Neisseria meningitidis also harbors a large number of pseudogenes (12.4%) (Liu et al. 2004; Schoen et al. 2009). In Rickettsia, pseudogenes account for a large fraction of the genome (9.7%) (Andersson et al. 1998; Liu et al. 2004).
Host range
There are differences in the number and proportion of pseudogenes carried by pathogens with different host ranges. Salmonella is an important zoonotic pathogen with more than 2600 serotypes of different host ranges and clinical characteristics (Tanner and Kingsley 2018). Comparative genomic study of multiple Salmonella serotypes revealed that the number of pseudogenes is much higher in Salmonella with a narrow host range than that of Salmonella with a broad host range (Holt et al. 2009; McClelland et al. 2001). Johnson et al. reported that S. Typhi, a species of Salmonella restricted to humans, has a high proportion of pseudogenes (Johnson et al. 2018). One study reported that in Salmonella enterica subsp. enterica serovar Paratyphi A (S. Paratyphi A), a human-restricted pathogen that exists an average of 173 pseudogenes, while in S. Typhimurium, a general-host pathogen, approximately 30 pseudogenes were found (McClelland et al. 2004). Another study also showed a large number of genome degradation events through pseudogene formation in S. Paratyphi A (Jacob et al. 2023). A relatively high proportion of pseudogenes were also observed in the genome of Salmonella enterica subsp. houtenae (S. houtenae), an organism with host adaptation to reptiles (Hyeon et al. 2021). The observations obtained by comparing two Salmonella serotypes (S. Typhimurium and S. Enteritidis) with broad host ranges to two serotypes (S. Typhi and S. Pullorum) with limited host ranges are presented in Table 1. The analysis of the complete genome of Staphylococcus aureus (S. aureus) revealed the presence of 14 pseudogenes (range 2–30). In comparison, S. aureus subsp. anaerobius, an anaerobic subspecies restricted to sheep and goats, exhibited an average of 205 pseudogenes per genome (range 201–210) (Yebra et al. 2021).
Geographical environmental niche
In Yersinia pestis (Y. pestis), pseudogene accumulation may be a result of adaptive microevolution of the bacterium to different epidemic foci. For instance, pseudogene spectrum and genetic characteristics are different in Y. pestis strains from different epidemic foci (Tong et al. 2005). Based on this pseudogene distribution criteria, Tong et al. divided 260 Y. pestis isolates from different natural plague foci in China into eight genotypes (Tong et al. 2005). Therefore, pseudogene spectrum analysis can also be used as a typing technique for Y. pestis. Claesson et al. analyzed the genetic characterization of Aggregatibacter actinomycetemcomitans from periodontitis patients living in various geographical locations of Sweden (Claesson et al. 2022). They found that the genetic characteristics of isolates from different geographic areas were distinct. Moreover, based on the presence or absence of a point mutation in the pseudogene hbpA2 of the bacterium, the collected isolates can be divided into two types: North African or West African. Current research evidence suggests that pseudogenes have potential applications in bacterial classification.
The accumulation of bacterial pseudogenes has previously been suggested to be associated with the adaptation of pathogens to specific host or ecological niches (Goodhead et al. 2020; Key et al. 2020; Liu et al. 2004). This partly explains why some genomes have a higher number of pseudogenes compared to others. When bacteria adapt from free living life style to an intracellular lifestyle, some genes, such as genes involved with metabolism or defense, become inactivated because their functions are no longer required in highly specialized niches (Cole et al. 2001). Some obligate intracellular bacteria such as M. leprae and Rickettsia have much smaller genomes than their free-living counterparts; some pathways tend to have been deleted (Dagan et al. 2006; Goodhead and Darby 2015). In addition, bacteria accumulate mutations and generate numbers of pseudogenes during the long-term process of adapting to different hosts or habitats (Dagan et al. 2006; Goodhead and Darby 2015; Wu et al. 2022). For example, Liao et al. found that the genomic characteristics vary among Salmonella populations from different hosts and geographic origins (Liao et al. 2020). Wang et al. identified that a unique glutathione (GSH) utilization pathway is pseudogenized in host-adapted Francisella (pathogenic Francisella species), whereas this pathway remains functional in non-pathogenic Francisella species, which are known to inhabit the environment (Wang et al. 2023). This possibility suggests that bacteria tend to eliminate and discard “unnecessary” genes, by forming pseudogenes, as they adapt to novel environmental niches, for example, when transitioning from a free-living lifestyle to an intracellular lifestyle, shifting from multiple hosts to unique host, or adapting to different epidemic foci (Bawn et al. 2020; Kuo and Ochman 2010). In general, pseudogenes are pervasively distributed in various microorganisms with different proportions. The formation and accumulation of pseudogenes in microorganisms are likely to be associated with the adaptive evolution of pathogens to specific niches, while other parameters may also play a role.
Contribution of pseudogenization to pathogen virulence
Bacteria promote their virulence and environmental adaptation through gene acquisition and gene loss (Lawrence et al. 2001; Ochman and Moran 2001; Zhou et al. 2022). Acquisition of pathogenicity islands or virulence plasmids has been well studied. Pseudogenization is one of the most important mechanisms of gene loss, which is caused by the accumulation of nonsense mutations in protein-coding gene sequences (McCutcheon and Moran 2011). In the following section, we will discuss some mechanisms by which pseudogenization affects virulence in Salmonella (Fig. 2).
Evasion of the phagocyte respiratory burst
Polymorphonuclear neutrophil leukocytes (PMNs) play an important role in phagocytosis and the killing of microbes. PMN respiratory burst, also known as oxygen burst, is an oxygen-dependent killing mechanism used to eliminate phagocytized pathogens. The activation of respiratory burst activates the NADPH oxidase, and a large number of reactive oxygen species (ROS) are produced (Luo et al. 2016). The capsule structure (Vi antigen) of S. Typhi prevents IgM-dependent complement activation and C5a-mediated neutrophil chemotaxis (Hiyoshi et al. 2018). Salmonella lipopolysaccharide (LPS) is composed of lipid A, the core polysaccharide, and an O-specific polysaccharide chain (O-antigen) (Fig. 3A) (Seif et al. 2019). The side chain of the O-antigen consists of several repetitive oligosaccharide units and can be divided into three types: (1) short (S) O-antigen chain, containing 1–15 repeating units; (2) long (L) O-antigen chain, containing 16–35 repeat units; (3) very-long (VL) O-antigen chain, containing more than 100 repeat units (Fig. 3B) (Murray et al. 2003). The length regulator FepE controls the length of the O-antigen chain. If the fepE gene is inactivated, the strain can only synthesize S or L O-antigen chains, but not the VL O-antigen chain (Fig. 3C) (Hiyoshi et al. 2018).
How pseudogenization of fepE optimizes capsule-mediated respiratory burst evasion by S. Typhi. A Structural schematic of LPS. B Three types of LPS according to the number of O-antigen repeat units. C Pseudogenization of fepE gene is responsible for the lack of LPS that with VL O-antigen chain in S. Typhi. D A diagram showing the brief composition and structure of S. Typhi. E A local magnification of the circular area in Fig. 3D. The capsule shields underlying surface structures from IgM recognition. Repair of ψfepE results in the production of LPS with VL O-antigen chain, which is exposed outside of the capsule, thereby resulting in IgM binding
In S. Typhi, fepE gene is a pseudogene with an early termination codon in the coding gene sequence (Fig. 3C) (Crawford et al. 2013). After the pseudogene ψfepE from S. Typhi was repaired (the fepE gene from S. Typhimurium was transferred into S. Typhi), the mutant recovered the ability to synthesize LPS containing VL O-antigen chain. The presence of VL O-antigen chains stimulates a respiratory burst because parts of the O-antigen chain are exposed outside of the capsule (Fig. 3D) (Crawford et al. 2013).
S. Paratyphi A can also escape from PMN respiratory burst, despite the lack of a capsule (McClelland et al. 2004). However, a rough mutant of S. Paratyphi A (var. Durazzo strain ATCC11511) lacks the VL O-antigen chain that triggers a strong respiratory burst (Hiyoshi et al. 2018). S. Paratyphi A contains the O2 and O12 antigens, with trisaccharide (mannose-rhamnose-galactose, O12) as the backbone and paratose (O2) as the branching sugar. S. Typhi contains the O9 and O12 antigens, with trisaccharide (O12) as the backbone and tyvelose (O9) as the branching sugar (Hiyoshi et al. 2018). S. Typhimurium contains the O4 and O12 antigens, with trisaccharide (O12) as the backbone and abequose (O4) as the branching sugar. The rfbS gene encodes a CDP-paratose synthase to catalyze the synthesis of paratose (O2). The rfbE gene encodes a CDP-tyvelose-epimerase to catalyze the conversion of paratose (O2) to tyvelose (O9). The rfbJ gene encodes a CDP-abequose synthase to catalyze the synthesis of abequose (O4) (Fig. 4A) (Verma and Reeves 1989; Woo et al. 2001). In S. Paratyphi A, the rfbE gene is annotated as a pseudogene because of mutations in the coding sequence. As a result, the conversion of paratose (O2) to tyvelose (O9) is not catalyzed, resulting in the production of O-antigen chain containing O2 rather than O9 (Fig. 4 B and C). The O2 antigen chain is the key S. Paratyphi A factor for inhibiting the host respiratory burst, as IgM binds to the O4 or O9 antigen but not to the O2 antigen. Introducing the S. Typhi rfbE gene into S. Paratyphi A results in O9 expression and triggers a respiratory burst. Replacing the S. Paratyphi A rfbS and rfbE genes with the S. Typhimurium rfbJ gene results in O4 expression and also triggers a respiratory burst (Fig. 4D) (Hiyoshi et al. 2018).
Pseudogenization of rfbE gene mediated respiratory burst evasion by S. Paratyphi A. A Gene clusters responsible for O-antigen biosynthesis in the indicated Salmonella serovars. B Pseudogenization of rfbE gene results in the production of O2 antigen in S. Paratyphi A. C A diagram showing the brief composition and structure of S. Paratyphi A. D A local magnification of the circular area in C. The VL LPS of S. Paratyphi A contains the O2 antigen which is not recognized by IgM. Mutants containing the O9 antigen (ψrfbE repaired) or the O4 antigen can be recognized by IgM
Resistance to H2O2
The marT and misL genes are located in the Salmonella pathogenicity island (SPI-3). MarT is a transcriptional regulatory factor. MisL is a self-transporting adhesin that promotes the colonization of S. Typhimurium in the intestinal tract of mice. In S. Typhimurium, MarT acts as a transcriptional activator of MisL and directly promotes the transcription of misL (Dorsey et al. 2005). In S. Typhi, marT was annotated as pseudogene. The transfer of marT from S. Typhimurium into S. Typhi resulted in reduced tolerance to ROS (such as H2O2) (Ortega et al. 2016). SurV also promotes the tolerance of S. Typhimurium to H2O2 and is inhibited by MarT (Ortega et al. 2016; Retamal et al. 2010). However, the mechanism of how surV reduces ROS levels has not been fully clarified.
Cytotoxicity toward epithelial cells
Some Salmonella type III secretion system (T3SS) effectors affect host cell cytotoxicity by altering Salmonella-containing vacuole (SCV) biosynthesis. SseJ is a T3SS-2 effector that functions as a cholesterol acyltransferase/lipase that alters SCV lipid content (Kolodziejek et al. 2019; Kolodziejek and Miller 2015; Walch et al. 2021). In S. Typhi, sseJ is annotated as a pseudogene ψsseJ (Trombert et al. 2010). The S. Typhimurium sseJ gene was introduced into S. Typhi, which were referred to as S. Typhi-sseJS. Typhimurium. Polarized HT-29 epithelial cells monolayers were infected, and, compared with S. Typhi-WT, the ability of S. Typhi-sseJS. Typhimurium to destroy the HT-29 cell layer was reduced (Trombert et al. 2010). The destruction of the epithelial cell barrier function facilitates the spread of infections (Oscarsson et al. 2002). The functional sseJ appears to have a detrimental impact on the systemic infection of S. Typhi. Thus, the pseudogenization of sseJ enhances the cytotoxicity of S. Typhi towards epithelial cells and increases its virulence.
In a similar example, pseudogenization of sopD2 enhances the pathogenicity of S. Typhi by increasing its ability to damage epithelial cells. SopD2 is also a T3SS-2 effector involved in SCV biosynthesis (Schroeder et al. 2010). In S. Typhi, sopD2 is a pseudogene. When S. Typhimurium sopD2 was expressed in S. Typhi, the toxicity of S. Typhi to epithelial cells was reduced and the ability of S. Typhi to penetrate epithelial cell also declined (Trombert et al. 2011).
Systemic infection
S. Typhimurium has a wide range of hosts and generally causes limited gastroenteritis in humans and animals and scarcely causes systemic infections. Recently, typhoid S. Typhimurium strains causing systemic infections were isolated in sub-Saharan Africa (Feasey et al. 2012). The analysis of the isolates using multi-locus sequence typing (MLST) showed that their primary sequence type was sequence type 13 (ST313), whereas the typical S. Typhimurium strains associated with gastroenteritis mainly belonged to ST19 (Feasey et al. 2012; Kingsley et al. 2009). Compared with ST19, ST313 shows a certain degree of genomic degradation, including the formation of pseudogenes and gene deletions (Feasey et al. 2012; Kingsley et al. 2009). In a mouse infection model, ST313 S. Typhimurium has higher loads in tissues than that of ST19 type, and the same trend can also be observed in chickens (Parsons et al. 2013; Yang et al. 2015). S. Typhimurium D23580, a typical strain of ST313, can spread systemically, because of its ability to survive within the CD11b+ dendritic cells (DCs), and migrate with DCs from the intestinal tract through lymphatic system to lymphatic organs (Carden et al. 2017).
Some S. Typhimurium T3SS effector factors inhibit the migration of DCs. The sseI gene encodes a T3SS effector that inhibits the migration of DCs in isolates associated with gastroenteritis. The sseI gene is a pseudogene in the genome of D23580 (Carden et al. 2017). The expression of functional sseI in S. Typhimurium D23580 decreased the migration ability of DCs, reduced Salmonella-carrying DC cells in mesenteric lymph nodes and decreased the bacterial loads in mesenteric lymph nodes. When the sseI gene was deleted in S. Typhimurium ST19, the number of bacteria that migrated to mesenteric lymph nodes increased (Carden et al. 2017). Therefore, the pseudogenization of the sseI gene enhances the ability of ST313 S. Typhimurium to cause systemic infection.
Additionally, the sopA gene encodes SopA, a T3SS-1 effector that functions as a HECT-like E3 ubiquitin ligase and is associated with Salmonella-induced enterocolitis. The recent study demonstrated that the pseudogenization of the sopA gene favored the survival of human macrophages and facilitated systemic infection of S. Typhi (Ma et al. 2021).
Flagella formation
Flagella are important virulence factors of bacteria, which not only provide bacteria with mobility but also play crucial roles in adhesion, invasion, and biofilm formation during the process of bacterial pathogenicity (Duan et al. 2013, 2012; Zhou et al. 2013, 2015). It is thought that all Salmonella have flagella, except for Salmonella enterica subsp. enterica serovar Gallinarum (S. Gallinarum) and Salmonella enterica subsp. enterica serovar Pullorum (S. Pullorum). Although S. Gallinarum and S. Pullorum do not have flagellar structures on the bacterial surface, they possess a full set of flagellar-related coding genes. Some of those flagella-related coding genes are defined as pseudogenes in S. Gallinarum and S. Pullorum, while in other flagellated Salmonella genomes, the corresponding genes are functional and do not correspond to pseudogenes (Yang et al. 2020).
Among the many serotypes of Salmonella with flagella, we selected S. Enteritidis strain P125109 (GenBank Accession Number: AM933172.1), which has the nearest genetic relationships with S. Pullorum and S. Gallinarum, as the reference strain. Eight genomic sequences of S. Gallinarum and S. Pullorum, including S. Gallinarum str. 287/91 (AM933173.1), S. Gallinarum str. 9184 (CP019035.1), S. Gallinarum str. 9 (CM001153.1), S. Pullorum str. S06004 (CP006575.1), S. Pullorum str. ATCC9120 (CP012347.1), S. Pullorum S44987_1 (LK931482.1), S. Pullorum str. CDC1983-67 (CP003786.1), and S. Pullorum str. RKS5078 (CP003047.1), were obtained from the GenBank database. We compared the flagella-related gene sequences of S. Pullorum and S. Gallinarum with reference strain and then summarize nine pseudogenes associated with flagella synthesis in the genome of S. Pullorum and S. Gallinarum: ψflgK, ψflhB, ψflhA, ψflgI, ψcheM, ψfliN, ψfliL, ψmotB, and ψycgR (Table 2). If the gene is annotated as a pseudogene in the GenBank database, it is marked with “□”; and if it is a functional gene, it is marked with a “■.”The functions of each gene are listed under the gene name in Table 2, according to their annotation in the genomic database. Each S. Pullorum or S. Gallinarum strain contains at least two flagellar-related pseudogenes and up to six flagellar-related pseudogenes. Among them, ψflgK and ψflhB were pseudogenes in all S. Pullorum and S. Gallinarum strains.
FlgK, also known as HAP1 (hook-associated protein 1), is responsible for the effective connection between the flagellum hook and the flagellum in the process of flagella assembly (Makishima et al. 2001; Minamino et al. 2021). FlhB is located at the bottom of flagellar base and is responsible for controlling substrate export (Ferris et al. 2005; Halte and Erhardt 2021; Kuhlen et al. 2020). It remains unclear how the pseudogenization of the flagella genes benefits S. Pullorum and S. Gallinarum. It is known that the major structural protein of flagella, flagellin (FliC), is an agonist of Toll-like receptor 5 (TLR-5) (Duan et al. 2013). Given this, the non-flagella phenotype of S. Pullorum and S. Gallinarum probably confers an advantage in evading recognition by TLR-5 and attenuating the host immune response directed towards them.
In addition to Salmonella, there are also instances in which the pseudogenization of certain genes benefits the pathogenicity in other pathogens. The yadE gene is a pseudogene in Y. pestis and is functional in Yersinia pseudotuberculosis (Y. pseudotuberculosis). The yadE gene encodes YadE, a trimeric autotransporter that contributes to the stability of biofilm in Y. pseudotuberculosis. The expression of functional yadE in Y. pestis results in a reduced biofilm stability and altered production of Hms-ECM, which is important extracellular matrix for Y. pestis biofilm formation. The formation of biofilm is contributing to the survival, host interaction, and transmission of Yersinia. Therefore, the pseudogenization of yadE gene likely plays a role in the virulence of Y. pestis (Calder et al. 2020).
Functional pseudogenes
For ages, pseudogenes have been considered as dysfunctional copies of functional genes; however, emerging evidence suggests that many of them may be biologically active (Cheetham et al. 2020). Pseudogenes may play physiological roles, such as gene expression, gene regulation, generation of genetic (antibody, antigenic, and other) diversity, as proposed and reported in mammals, such as humans and mice (Balakirev and Ayala 2003; Sisu 2021a; Xu and Zhang 2016). In the field of microbiology, the evidence that pseudogenes are not “completely non-functional” is mainly related to the fact that pseudogenes are involved in recombination and improve genetic diversity. Additionally, in some cases, they can be transcribed and translated.
Contribution to gene genetic diversity
An important function of pseudogenes is to serve as a repository, providing material for improving gene sequence diversity. Chicken antibody (immunoglobulins, Igs) diversity is a classic example. Antibodies consist of two light chains (L) and two heavy chains (H), both of which contain a constant region (C) and a variable region (V), which is used to recognize foreign molecules (Ratcliffe 2006). The pseudogenes ψVL and ψVH are present upstream of the site of the coding nucleic acid sequence encoding the heavy and light chains of the antibody. Although the pseudogene itself cannot be expressed, it can be recombined with the variable region within the functional gene through insertion or replacement and expressed together with the functional gene, thereby increasing the diversity of the variable region coding gene (Vihinen 2014).
Antigenic variation is a strategy for the persistence of multiple microbial pathogens within hosts. Some pathogens use pseudogenes to produce antigenic variation in surface molecules, which is another example of using pseudogenes to significantly increase genetic diversity. For instance, pseudogene-mediated antigen variation is an important strategy for pathogenic microorganisms to evade host immune responses. Trypanosoma brucei (T. brucei) is a type of zoonotic parasite that causes sleeping sickness in human and Nagana disease in animals. T. brucei escapes the host’s immune system by periodically altering the variant surface glycoprotein (VSG) which covers the surface of the worm body. This variation relies on the large number (~ 2000) copies of vsg gene and the pseudogene form of vsg gene transcript (ψvsg) in the genome (Dakovic et al. 2023; Davies et al. 2021). As a result, the excessive level of recombination events between “vsg–vsg” and “vsg–ψvsg” continuously generates additional chimeric vsg genes and further enhances the diversity and richness of the vsg gene (Chandra et al. 2023; Faria et al. 2022).
Similar mechanisms have also been found in other pathogens. For example, both the major surface protein 2 (Msp2) of Anaplasma (Graca et al. 2019; Liu et al. 2019; Rejmanek et al. 2012) and the variable major protein (VMP) of Borrelia spp. (Gilmore et al. 2023; Restrepo et al. 1994; Schwartz et al. 2021) use the “functional gene-pseudogene” recombination mechanism to generate new variant antigens, thereby evading the immune recognition of hosts. In Pneumocystis jirovecii, which is an obligate pulmonary pathogen in human, pseudogenes are suggested to contribute to the generation of various mosaic msg genes, encoding the major surface glycoprotein, via integration into functional msg genes (Schmid-Siegert et al. 2017). Thus, it is obvious that “gene-pseudogene rearrangement” is an original genetic evolutionary mechanism, which contributes to the generation of genetic variability and diversity. Simultaneously, the presence of pseudogenes could serve as a reservoir of sequences for antigenic variation. Furthermore, in this case, the pseudogenes would not be eliminated or lost from the genome.
Some pseudogenes are transcribed or translated
Due to the development of next-generation sequencing (NGS) technologies, the cost of large-scale sequencing has largely reduced. It is easier to obtain high-throughput genome-wide sequence data (Ejigu and Jung 2020; Rehder et al. 2021). Pseudogenes have been discovered in large numbers from the genomics data of various organisms, with the aid of NGS technologies. Concurrently, data from transcriptomics and proteomics techniques are improving our understanding of whether pseudogenes are expressed at the RNA/protein level or not. Transcription and translation of pseudogenes have been widely reported and confirmed in mammals (Giuliani et al. 2022; Qian et al. 2022; Sisu et al. 2020), while the transcription or expression of pseudogenes has also been gradually observed and reported in microorganisms. RNA-seq technique was used to analyze pseudogenes in S. Typhi; results reveal that many pseudogenes are transcribed, despite at greatly reduced levels (Perkins et al. 2009). Feng et al. utilized RNA-seq and mass spectrometry technologies to describe the transcriptional and translational landscape in Salmonella species. They revealed that 101 out of 161 pseudogenes could be successfully translated in S. Paratyphi A and S. Typhi (Feng et al. 2022). Transcribed pseudogenes have also been observed in Shigella flexneri (Cervantes-Rivera et al. 2020; Chanin et al. 2019). Whole-genome analysis of M. leprae RNA expression demonstrated that pseudogenes and non-coding regions are not silent but strongly expressed (Akama et al. 2009). Gao et al. analyzed the proteome of Saccharomyces cerevisiae using mass spectrometry (MS) and provided evidence that pseudogenes can be translated (Gao et al. 2021). Goodhead et al. applied multiple “omic” strategies, combining genomics, transcriptomics, and proteomics techniques to analyze the transcriptional and translational landscape of pseudogenes in Sodalis, a Gram-negative, facultative endosymbiont bacterium. They have revealed that Sodalis pseudogenes are often transcribed, but at a significantly lower level than intact CDSs (Goodhead et al. 2020). Wen et al. reported the observation of a set of small interference RNAs (siRNAs) derived from pseudogenes of African T. brucei using high-throughput analysis. Authors then confirmed that the siRNA derived from pseudogenes in the T. brucei can regulate protein-coding gene expression by means of the RNA interference (Wen et al. 2011).
Some pseudogenes have a role in aspects of bacterial diseases pathogenicity. Adhesins are located on the bacterial surface and help bacteria to adhere to the surface of host cells (Duan et al. 2022; Patel et al. 2017). In Salmonella, the shdA gene encodes a novel non-pilus autotransporter adhesin in which the passenger domain binds to one or more extracellular matrix proteins (e.g., fibronectin and collagen) (Kingsley et al. 2004, 2002; Paxman et al. 2019). In S. Typhi, the deletion of a shdA gene fragment causes a termination codon to be prematurely generated, therefore, it was annotated as a pseudogene (ψshdA) (Urrutia et al. 2014). However, subsequent experiments have shown that the translation product of ψshdA can function as an active adhesin, based on the analysis of a ψshdA deletion as well as the detection of a translated protein (Urrutia et al. 2014). Furthermore, gene yqiG is annotated as a pseudogene (ψyqiG) in E. coli BW25113 genome due to the insertion of an insertion sequence (IS) element inside the gene sequence. However, subsequent experiments have shown that the product of pseudogene ψyqiG (YqiG protein) is functional and was identified as an essential protein in glycolysis pathway and hydrogen metabolism in E. coli BW25113 (Zakaria et al. 2018). Zhang et al. reveal that the pseudogene BMEA_B0173 plays a role in the virulence of the Brucella melitensis (Zhang et al. 2022). In a mouse infection model, the BMEA_B0173 deletion mutant exhibited increased colonization in the spleen compared to the wild-type strain.
Pseudogenes and viruses
Are there pseudogenes in the viruses?
Do the viruses contain pseudogenes or not? Some studies have reported the existence of defective mutated genes in a variety of viral genomes (Ceccaldi et al. 1998; Zhang et al. 1992); hence, these mutated genes, which, by definition, should be called pseudogenes. Aaskov et al. reported that there is a stop-codon mutation in the surface envelope (E) protein gene of dengue virus type 1 (DENV-1) in humans and mosquitoes from Myanmar (Aaskov et al. 2006). Hughes et al. found that mutations were present in the genome of the equine influenza virus (EIV) and cause stop codons (Hughes et al. 2012). Some studies have shown that there are large numbers of nonfunctional genes (pseudogenes) in human cytomegalovirus (HCMV) strains (Sijmons et al. 2014, 2015; Suarez et al. 2019). One study shows that approximately 75% of HCMV virus strains carry pseudogenes (Sijmons et al. 2015; Suarez et al. 2019). The researchers also point out that the evidence for pseudogenes was largely derived from strains isolated in cell culture. These mutations are substitutions that introduce premature stop codons or deletions or insertions that cause frame shifting (Suarez et al. 2019).
Pseudogenes and virus characterization
Do the pseudogenes contribute to alterations in virus characteristics? Up to now, the research about this aspect previously is not much. It has been reported that the mutations in the DENV-1 virus causing stop codons were likely to be long-term transmitted, and these mutations could provide the shift of viral fitness and then influence transmission dynamics (Aaskov et al. 2006). Here we attempt to explore the relationship between the gene defect mutations (pseudogenes) and the virulence of the virus, taking the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) as an example.
The outbreak of COVID-19, which was caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has a global impact. SARS-CoV-2 is an enveloped, positive-sense, single-stranded RNA virus and belongs to lineage B of the β-coronavirus genus. The virus genome encodes four structural proteins including spike (S), envelope (E), membrane (M), and nucleocapsid (N) proteins, as well as other non-structural or accessory proteins (Gordon et al. 2020). The genetic changes of the virus have attracted widespread and persistent attention around the world (Mannar et al. 2022). During the genomic surveillance of SARS-CoV-2 in the first year of the COVID-19 pandemic, a mutation that aspartic acid (D) was substituted by glycine (G) at position 614 (D614G) in the S protein was reported (Korber et al. 2020; Yurkovetskiy et al. 2020). The SARS-CoV-2 variant carrying the D614G mutation has then become the most prevalent form in the global pandemic. D614G shifts the conformation of the S protein and alters the affinity toward to the angiotensin-converting enzyme 2 (ACE2), which is the receptor of SARS-CoV-2 (Yurkovetskiy et al. 2020; Zhang et al. 2021a). D614G increased the infectivity of human lung cells or bat or pangolin ACE2 cells (Yurkovetskiy et al. 2020). The D614G variant was proved to have the ability of efficient replication in vitro and transmission in vivo (Hou et al. 2020; Shi and Xie 2021).
In addition, several defective mutations in the SARS-CoV-2 genome have been reported to date. Su et al. reported that there is a 382-nucleotide (nt) deletion (Δ382) in open reading frame ORF7b and ORF8 of SARS-CoV-2 (Su et al. 2020). This mutation resulted in truncation of ORF7 and non-transcription of ORF8 (Su et al. 2020). The Orf8 encodes an accessory protein and is one of the proteins that with least sequence similarity between SARS-CoV-2 and SARS-CoV (Gordon et al. 2020; Zhang et al. 2021b). ORF8 protein was be proved to mediate immune evasion through down-regulating major histocompability complex class Ι (MHC-Ι) (Zhang et al. 2021b). Recent studies revealed that ORF8 could mediate endoplasmic reticulum (ER) reshaping of host cell through forming mixed disulfide complexes with ER proteins (Liu et al. 2022b). Clinical symptom of patients infected with ∆382 variant viruses was milder, compared to patients infected with the wild viruses (Young et al. 2020). In addition, lower concentrations of proinflammatory cytokines were detected in plasma of patients infected with ∆382 variant viruses, compared to that of patients infected with the wild viruses (Young et al. 2020). In summary, it seems that ORF8 could be a candidate target for the development of treatments and vaccines for SARS-CoV-2.
Researchers reported that there is a premature stop codon in ORF3b gene at position 14 (E14*) of SARS-CoV-2 (Gordon et al. 2020). This mutation resulted in truncated product of ORF3b (Konno et al. 2020). This mutation resulted in truncated product of ORF3b (Konno et al. 2020). The Orf3 encodes one of the accessory proteins of SARS-CoV-2. Qu et al. reported that ORF3a mediated incomplete autophagy, which facilitates SARS-CoV-2 replication (Qu et al. 2021). The antibodies of protein ORF8 and ORF3b are accurate serological markers of early and late SARS-CoV-2 infections (Hachim et al. 2020). ORF3b of SARS-CoV-2 is a potent IFN-I antagonist, which inhibited the activation of human IFN-I (Konno et al. 2020). Besides, researchers reported that there are two premature stop codons at positions 41 (Q41*) and 44 (Q44*) in ORF9c of SARS-CoV-2 (Gordon et al. 2020). The biological significance of mutant in SARSCoV-2 ORF9c remains to be elucidated. Further research focused on defective gene variants (pseudogenes) of the virus may improve the understanding of the mechanisms of virus infection and could provide new insights in the development of targets for treatments and vaccines.
Virus-derived pseudogenes in mammals
The investigation and analysis of mammalian genomes suggest that some pseudogenes in mammals may be derived from viruses. For instance, a study analyzed the Mops condylurus genomic DNA samples and revealed that there is an Ebola virus nucleoprotein (NP)-derived pseudogene inserted in its genome (Hermida Lorenzo et al. 2021). Furthermore, filovirus-derived pseudogenes have also been reported in the genome of a wide variety of organisms including bats, marsupials, and rodents (Taylor et al. 2011). This is thought to originate from recombination events between viral genes and host genomic transposons during viral infection (Naville et al. 2016), while the significance and importance of the virus-derived pseudogenes remains to be determined.
Future perspectives
For a long time, pseudogenes were defined as “non-functional copies” sequences of functional gene in the genome. The prevalence and significance of pseudogenes in microorganisms have been increasingly acknowledged and have emerged as a focal point of research in recent years. In this review, we summarize the latest research progress on various aspects of microorganism pseudogenes (Table 3).
The formation of pseudogenes may be a result of the accumulation of bacterial adaptive evolution (Lawrence et al. 2001; Ochman and Davalos 2006). When the living environment of bacteria changes, such no longer required genes are prone to be lost from the genome by forming pseudogenes. This implies that the type and quantity of pseudogenes can potentially serve as indicators for inferring the evolutionary process of bacteria. Therefore, pseudogenes can be used as evolutionary records, providing very valuable materials for the study of bacterial evolution.
The abundance and distribution of pseudogenes could provide useful information to improve our understanding towards pathogens. For example, the abundance of pseudogenes may be associated with intracellular/free-living lifestyle or host range of the pathogens. And the distribution of pseudogenes (pseudogene profile) can be used for pathogen classification, for example, the classification of Y. pestis.
Furthermore, the pseudogenization of some genes contributes to the virulence of certain pathogenic microorganisms, since the complementation of functional parental genes reduces their virulence. It is like a story of “loss is gain,” where loss means the loss of function of the gene through the formation of a pseudogene, and gain means the increase in virulence of the pathogen. The further investigation focusing on pseudogenes holds the potential to enhance our comprehension of bacterial evolution and elucidate the underlying pathogenic mechanisms.
The potential functions of pseudogenes, particularly their contribution to sequence polymorphism, render them a reservoir of sequences for antigenic variation. In addition, greater emphasis should be placed on pseudogenes with potential for transcription or translation, and a more comprehensive exploration of functional pseudogenes is warranted.
Defective genetic mutations (pseudogenes) are also widespread viral genomes, some of which affect the pathogenic characteristic of the virus. More research that focuses on pseudogenes in virus may help to improve our understanding of the evolution and virulence of virus.
It is our hope that more attention and interest can be devoted to the research of pseudogenes in microorganisms. Many efforts and attempts are still required to acquire a comprehensive understanding on pseudogenes and their significance in microorganism, including but not limited to the following: (a) Define the updated and generally accepted concept of pseudogene. (b) Establish novel more accurate pipelines and standards for pseudogene identification. (c) Research on the generation and evolution mechanisms of pseudogenes. (d) Further study the significance of pseudogenization on pathogenicity of various pathogens. (e) Detection of transcribed and translated pseudogenes. (f) Investigation of potential regulatory functions of microbial pseudogenes. (g) Establish database of virus pseudogene and further research. We believe that advanced sequencing technique and bioinformatics tools, in conjunction with genomics, transcriptomics, and proteomics techniques, will improve the accuracy of pseudogene prediction and annotation to fully reveal the secret of pseudogenization in microorganisms.
Data Availability
Data will be made available on reasonable request.
References
Aaskov J, Buzacott K, Thu HM, Lowry K, Holmes EC (2006) Long-term transmission of defective RNA viruses in humans and Aedes mosquitoes. Science 311(5758):236–238. https://doi.org/10.1126/science.1115030
Abrahamsson S, Eiengard F, Rohlin A, Davila Lopez M (2022) PPsiFinder: a practical tool for the identification and visualization of novel pseudogenes in DNA sequencing data. BMC Bioinformatics 23(1):59. https://doi.org/10.1186/s12859-022-04583-4
Akama T, Suzuki K, Tanigawa K, Kawashima A, Wu H, Nakata N, Osana Y, Sakakibara Y, Ishii N (2009) Whole-genome tiling array analysis of Mycobacterium leprae RNA reveals high expression of pseudogenes and noncoding regions. J Bacteriol 191(10):3321–3327. https://doi.org/10.1128/JB.00120-09
Andersson SG, Zomorodipour A, Andersson JO, Sicheritz-Ponten T, Alsmark UC, Podowski RM, Naslund AK, Eriksson AS, Winkler HH, Kurland CG (1998) The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396(6707):133–140. https://doi.org/10.1038/24094
Avni E, Montoya D, Lopez D, Modlin R, Pellegrini M, Snir S (2018) A phylogenomic study quantifies competing mechanisms for pseudogenization in prokaryotes-The Mycobacterium leprae case. PLoS One 13(11):e0204322. https://doi.org/10.1371/journal.pone.0204322
Badel C, Da Cunha V, Forterre P, Oberto J (2020) Pervasive suicidal integrases in deep-sea Archaea. Mol Biol Evol 37(6):1727–1743. https://doi.org/10.1093/molbev/msaa041
Balakirev ES, Ayala FJ (2003) Pseudogenes: are they “junk” or functional DNA? Annu Rev Genet 37:123–151. https://doi.org/10.1146/annurev.genet.37.040103.103949
Bawn M, Alikhan NF, Thilliez G, Kirkwood M, Wheeler NE, Petrovska L, Dallman TJ, Adriaenssens EM, Hall N, Kingsley RA (2020) Evolution of Salmonella enterica serotype Typhimurium driven by anthropogenic selection and niche adaptation. PLoS Genet 16(6):e1008850. https://doi.org/10.1371/journal.pgen.1008850
Bialer MG, Ferrero MC, Delpino MV, Ruiz-Ranwez V, Posadas DM, Baldi PC, Zorreguieta A (2021) Adhesive functions or pseudogenization of type Va autotransporters in Brucella species. Front Cell Infect Microbiol 11:607610. https://doi.org/10.3389/fcimb.2021.607610
Calder JT, Christman ND, Hawkins JM, Erickson DL (2020) A trimeric autotransporter enhances biofilm cohesiveness in Yersinia pseudotuberculosis but not in Yersinia pestis. J Bacteriol 202(20). https://doi.org/10.1128/JB.00176-20
Carden SE, Walker GT, Honeycutt J, Lugo K, Pham T, Jacobson A, Bouley D, Idoyaga J, Tsolis RM, Monack D (2017) Pseudogenization of the secreted effector gene sseI confers rapid systemic dissemination of S. Typhimurium ST313 within migratory dendritic cells. Cell Host Microbe 21(2):182–194. https://doi.org/10.1016/j.chom.2017.01.009
Ceccaldi PE, Fayet J, Conzelmann KK, Tsiang H (1998) Infection characteristics of rabies virus variants with deletion or insertion in the pseudogene sequence. J Neurovirol 4(1):115–119. https://doi.org/10.3109/13550289809113489
Cervantes-Rivera R, Tronnet S, Puhar A (2020) Complete genome sequence and annotation of the laboratory reference strain Shigella flexneri serotype 5a M90T and genome-wide transcriptional start site determination. BMC Genomics 21(1):285. https://doi.org/10.1186/s12864-020-6565-5
Chan WL, Yang WK, Huang HD, Chang JG (2013) pseudoMap: an innovative and comprehensive resource for identification of siRNA-mediated mechanisms in human transcribed pseudogenes. Database (Oxford) 2013:bat001. https://doi.org/10.1093/database/bat001
Chandra M, Dakovic S, Foti K, Zeelen JP, van Straaten M, Aresta-Branco F, Tihon E, Lubbehusen N, Ruppert T, Glover L, Papavasiliou FN, Stebbins CE (2023) Structural similarities between the metacyclic and bloodstream form variant surface glycoproteins of the African trypanosome. Plos Neglect Trop D 17(2):e0011093. https://doi.org/10.1371/journal.pntd.0011093
Chanin RB, Nickerson KP, Llanos-Chea A, Sistrunk JR, Rasko DA, Kumar DKV, de la Parra J, Auclair JR, Ding J, Li K, Dogiparthi SK, Kusber BJD, Faherty CS (2019) Shigella flexneri adherence factor expression in in vivo-like conditions. mSphere 4(6). https://doi.org/10.1128/mSphere.00751-19
Cheetham SW, Faulkner GJ, Dinger ME (2020) Overcoming challenges and dogmas to understand the functions of pseudogenes. Nat Rev Genet 21(3):191–201. https://doi.org/10.1038/s41576-019-0196-1
Chen B, Wang C, Zhang J, Zhou Y, Hu W, Guo T (2018) New insights into long noncoding RNAs and pseudogenes in prognosis of renal cell carcinoma. Cancer Cell Int 18:157. https://doi.org/10.1186/s12935-018-0652-6
Chen X, Wan L, Wang W, Xi WJ, Yang AG, Wang T (2020) Re-recognition of pseudogenes: from molecular to clinical applications. Theranostics 10(4):1479–1499. https://doi.org/10.7150/thno.40659
Chieffi D, Fanelli F, Cho GS, Schubert J, Blaiotta G, Franz CMAP, Bania J, Fusco V (2020) Novel insights into the enterotoxigenic potential and genomic background of Staphylococcus aureus isolated from raw milk. Food Microbiol 90:103482. https://doi.org/10.1016/j.fm.2020.103482
Claesson R, Oscarsson J, Johansson A (2022) Carriage of the JP2 genotype of Aggregatibacter actinomycetemcomitans by periodontitis patients of various geographic origin, living in Sweden. Pathogens 11(11). https://doi.org/10.3390/pathogens11111233
Cole ST, Eiglmeier K, Parkhill J, James KD, Thomson NR, Wheeler PR, Honore N, Garnier T, Churcher C, Harris D, Mungall K, Basham D, Brown D, Chillingworth T, Connor R, Davies RM, Devlin K, Duthoy S, Feltwell T, Fraser A, Hamlin N, Holroyd S, Hornsby T, Jagels K, Lacroix C, Maclean J, Moule S, Murphy L, Oliver K, Quail MA, Rajandream MA, Rutherford KM, Rutter S, Seeger K, Simon S, Simmonds M, Skelton J, Squares R, Squares S, Stevens K, Taylor K, Whitehead S, Woodward JR, Barrell BG (2001) Massive gene decay in the leprosy bacillus. Nature 409(6823):1007–1011. https://doi.org/10.1038/35059006
Crawford RW, Wangdi T, Spees AM, Xavier MN, Tsolis RM, Baumler AJ (2013) Loss of very-long O-antigen chains optimizes capsule-mediated immune evasion by Salmonella enterica serovar Typhi. mBio 4(4). https://doi.org/10.1128/mBio.00232-13
Dagan T, Blekhman R, Graur D (2006) The “domino theory” of gene death: gradual and mass gene extinction events in three lineages of obligate symbiotic bacterial pathogens. Mol Biol Evol 23(2):310–316. https://doi.org/10.1093/molbev/msj036
Dakovic S, Zeelen JP, Gkeka A, Chandra M, van Straaten M, Foti K, Zhong J, Vlachou EP, Aresta-Branco F, Verdi JP, Papavasiliou FN, Stebbins CE (2023) A structural classification of the variant surface glycoproteins of the African trypanosome. PLoS Negl Trop Dis 17(9):e0011621. https://doi.org/10.1371/journal.pntd.0011621
Darmon E, Leach DR (2014) Bacterial genome instability. Microbiol Mol Biol Rev 78(1):1–39. https://doi.org/10.1128/MMBR.00035-13
Davies C, Ooi CP, Sioutas G, Hall BS, Sidhu H, Butter F, Alsford S, Wickstead B, Rudenko G (2021) TbSAP is a novel chromatin protein repressing metacyclic variant surface glycoprotein expression sites in bloodstream form Trypanosoma brucei. Nucleic Acids Res 49(6):3242–3262. https://doi.org/10.1093/nar/gkab109
Dong R, Zhang XO, Zhang Y, Ma XK, Chen LL, Yang L (2016) CircRNA-derived pseudogenes. Cell Res 26(6):747–750. https://doi.org/10.1038/cr.2016.42
Dorsey CW, Laarakker MC, Humphries AD, Weening EH, Baumler AJ (2005) Salmonella enterica serotype Typhimurium MisL is an intestinal colonization factor that binds fibronectin. Mol Microbiol 57(1):196–211. https://doi.org/10.1111/j.1365-2958.2005.04666.x
Duan Q, Zhou M, Zhu X, Bao W, Wu S, Ruan X, Zhang W, Yang Y, Zhu J, Zhu G (2012) The flagella of F18ab Escherichia coli is a virulence factor that contributes to infection in a IPEC-J2 cell model in vitro. Vet Microbiol 160(1–2):132–140. https://doi.org/10.1016/j.vetmic.2012.05.015
Duan Q, Zhou M, Zhu L, Zhu G (2013) Flagella and bacterial pathogenicity. J Basic Microbiol 53(1):1–8. https://doi.org/10.1002/jobm.201100335
Duan Q, Pang S, Feng L, Liu J, Lv L, Li B, Liang Y, Zhu G (2022) Heat-labile enterotoxin enhances F4-producing enterotoxigenic E. coli adhesion to porcine intestinal epithelial cells by upregulating bacterial adhesins and STb enterotoxin. Vet Res 53(1):88. https://doi.org/10.1186/s13567-022-01110-4
Ejigu GF, Jung J (2020) Review on the computational genome annotation of sequences obtained by next-generation sequencing. Biology (Basel) 9(9). https://doi.org/10.3390/biology9090295
Faria J, Briggs EM, Black JA, McCulloch R (2022) Emergence and adaptation of the cellular machinery directing antigenic variation in the African trypanosome. Curr Opin Microbiol 70:102209. https://doi.org/10.1016/j.mib.2022.102209
Feasey NA, Dougan G, Kingsley RA, Heyderman RS, Gordon MA (2012) Invasive non-typhoidal Salmonella disease: an emerging and neglected tropical disease in Africa. Lancet 379(9835):2489–2499. https://doi.org/10.1016/S0140-6736(11)61752-2
Feng Y, Wang Z, Chien KY, Chen HL, Liang YH, Hua X, Chiu CH (2022) “Pseudo-pseudogenes” in bacterial genomes: proteogenomics reveals a wide but low protein expression of pseudogenes in Salmonella enterica. Nucleic Acids Res 50(9):5158–5170. https://doi.org/10.1093/nar/gkac302
Ferris HU, Furukawa Y, Minamino T, Kroetz MB, Kihara M, Namba K, Macnab RM (2005) FlhB regulates ordered export of flagellar components via autocleavage mechanism. J Biol Chem 280(50):41236–41242. https://doi.org/10.1074/jbc.M509438200
Frankish A, Harrow J (2014) GENCODE pseudogenes. Methods Mol Biol 1167:129–155. https://doi.org/10.1007/978-1-4939-0835-6_10
Gao Y, Ping L, Duong D, Zhang C, Dammer EB, Li Y, Chen P, Chang L, Gao H, Wu J, Xu P (2021) Mass-spectrometry-based near-complete draft of the Saccharomyces cerevisiae proteome. J Proteome Res 20(2):1328–1340. https://doi.org/10.1021/acs.jproteome.0c00721
Gilmore RD, Armstrong BA, Brandt KS, Van Gundy TJ, Hojgaard A, Lopez JE, Kneubehl AR (2023) Analysis of variable major protein antigenic variation in the relapsing fever spirochete, Borrelia miyamotoi, in response to polyclonal antibody selection pressure. PLoS ONE 18(2):e0281942. https://doi.org/10.1371/journal.pone.0281942
Giuliani A, Bui TT, Helmy M, Selvarajoo K (2022) Identifying toggle genes from transcriptome-wide scatter: A new perspective for biological regulation. Genomics 114(1):215–228. https://doi.org/10.1016/j.ygeno.2021.11.027
Gomez-Valero L, Rocha EP, Latorre A, Silva FJ (2007) Reconstructing the ancestor of Mycobacterium leprae: the dynamics of gene loss and genome reduction. Genome Res 17(8):1178–1185. https://doi.org/10.1101/gr.6360207
Goodhead I, Darby AC (2015) Taking the pseudo out of pseudogenes. Curr Opin Microbiol 23:102–109. https://doi.org/10.1016/j.mib.2014.11.012
Goodhead I, Blow F, Brownridge P, Hughes M, Kenny J, Krishna R, McLean L, Pongchaikul P, Beynon R, Darby AC (2020) Large-scale and significant expression from pseudogenes in Sodalis glossinidius - a facultative bacterial endosymbiont. Microb Genom 6(1). https://doi.org/10.1099/mgen.0.000285
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Huttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, Lyu J, Mathy CJP, Perica T, Pilla KB, Ganesan SJ, Saltzberg DJ, Rakesh R, Liu X, Rosenthal SB, Calviello L, Venkataramanan S, Liboy-Lugo J, Lin Y, Huang XP, Liu Y, Wankowicz SA, Bohn M, Safari M, Ugur FS, Koh C, Savar NS, Tran QD, Shengjuler D, Fletcher SJ, O’Neal MC, Cai Y, Chang JCJ, Broadhurst DJ, Klippsten S, Sharp PP, Wenzell NA, Kuzuoglu-Ozturk D, Wang HY, Trenker R, Young JM, Cavero DA, Hiatt J, Roth TL, Rathore U, Subramanian A, Noack J, Hubert M, Stroud RM, Frankel AD, Rosenberg OS, Verba KA, Agard DA, Ott M, Emerman M, Jura N, von Zastrow M, Verdin E, Ashworth A, Schwartz O, d’Enfert C, Mukherjee S, Jacobson M, Malik HS, Fujimori DG, Ideker T, Craik CS, Floor SN, Fraser JS, Gross JD, Sali A, Roth BL, Ruggero D, Taunton J, Kortemme T, Beltrao P, Vignuzzi M, Garcia-Sastre A, Shokat KM, Shoichet BK, Krogan NJ (2020) A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583(7816):459–468. https://doi.org/10.1038/s41586-020-2286-9
Graca T, Ku PS, Silva MG, Turse JE, Hammac GK, Brown WC, Palmer GH, Brayton KA (2019) Segmental variation in a duplicated msp2 pseudogene generates Anaplasma marginale antigenic variants. Infect Immun 87(2). https://doi.org/10.1128/IAI.00727-18
Hachim A, Kavian N, Cohen CA, Chin AWH, Chu DKW, Mok CKP, Tsang OTY, Yeung YC, Perera R, Poon LLM, Peiris JSM, Valkenburg SA (2020) ORF8 and ORF3b antibodies are accurate serological markers of early and late SARS-CoV-2 infection. Nat Immunol 21(10):1293–1301. https://doi.org/10.1038/s41590-020-0773-7
Halte M, Erhardt M (2021) Protein export via the type III secretion system of the bacterial flagellum. Biomolecules 11(2). https://doi.org/10.3390/biom11020186
Harrison PM, Milburn D, Zhang Z, Bertone P, Gerstein M (2003) Identification of pseudogenes in the Drosophila melanogaster genome. Nucleic Acids Res 31(3):1033–1037. https://doi.org/10.1093/nar/gkg169
Hermida Lorenzo RJ, Cadar D, Koundouno FR, Juste J, Bialonski A, Baum H, Garcia-Mudarra JL, Hakamaki H, Bencsik A, Nelson EV, Carroll MW, Magassouba N, Gunther S, Schmidt-Chanasit J, Munoz Fontela C, Escudero-Perez B (2021) Metagenomic snapshots of viral components in guinean bats. Microorganisms 9(3). https://doi.org/10.3390/microorganisms9030599
Hiyoshi H, Wangdi T, Lock G, Saechao C, Raffatellu M, Cobb BA, Baumler AJ (2018) Mechanisms to evade the phagocyte respiratory burst arose by convergent evolution in typhoidal Salmonella serovars. Cell Rep 22(7):1787–1797. https://doi.org/10.1016/j.celrep.2018.01.016
Holt KE, Thomson NR, Wain J, Langridge GC, Hasan R, Bhutta ZA, Quail MA, Norbertczak H, Walker D, Simmonds M, White B, Bason N, Mungall K, Dougan G, Parkhill J (2009) Pseudogene accumulation in the evolutionary histories of Salmonella enterica serovars Paratyphi A and Typhi. BMC Genomics 10:36. https://doi.org/10.1186/1471-2164-10-36
Homma K, Fukuchi S, Kawabata T, Ota M, Nishikawa K (2002) A systematic investigation identifies a significant number of probable pseudogenes in the Escherichia coli genome. Gene 294(1–2):25–33. https://doi.org/10.1016/s0378-1119(02)00794-1
Hou YJ, Chiba S, Halfmann P, Ehre C, Kuroda M, Dinnon KH 3rd, Leist SR, Schafer A, Nakajima N, Takahashi K, Lee RE, Mascenik TM, Graham R, Edwards CE, Tse LV, Okuda K, Markmann AJ, Bartelt L, de Silva A, Margolis DM, Boucher RC, Randell SH, Suzuki T, Gralinski LE, Kawaoka Y, Baric RS (2020) SARS-CoV-2 D614G variant exhibits efficient replication ex vivo and transmission in vivo. Science 370(6523):1464–1468. https://doi.org/10.1126/science.abe8499
Hughes J, Allen RC, Baguelin M, Hampson K, Baillie GJ, Elton D, Newton JR, Kellam P, Wood JL, Holmes EC, Murcia PR (2012) Transmission of equine influenza virus during an outbreak is characterized by frequent mixed infections and loose transmission bottlenecks. PLoS Pathog 8(12):e1003081. https://doi.org/10.1371/journal.ppat.1003081
Hyeon JY, Helal ZH, Polkowski R, Vyhnal K, Mishra N, Kim J, Risatti GR, Lee DH (2021) Genomic features of Salmonella enterica subspecies houtenae serotype 45:g,z51:- isolated from multiple abdominal abscesses of an african fat-tailed gecko, United States, 2020. Antibiotics (Basel) 10(11). https://doi.org/10.3390/antibiotics10111322
Jacob JJ, Pragasam AK, Vasudevan K, Velmurugan A, Priya Teekaraman M, Priya Thirumoorthy T, Ray P, Gupta M, Kapil A, Bai SP, Nagaraj S, Saigal K, Chandola TR, Thomas M, Bavdekar A, Ebenezer SE, Shastri J, De A, Dutta S, Alexander AP, Koshy RM, Jinka DR, Singh A, Srivastava SK, Anandan S, Dougan G, John J, Kang G, Veeraraghavan B, Mutreja A (2023) Genomic analysis unveils genome degradation events and gene flux in the emergence and persistence of S. Paratyphi A lineages. Plos Pathog 19(4):e1010650. https://doi.org/10.1371/journal.ppat.1010650
Jacq C, Miller JR, Brownlee GG (1977) A pseudogene structure in 5S DNA of Xenopus laevis. Cell 12(1):109–120. https://doi.org/10.1016/0092-8674(77)90189-1
Johnson R, Mylona E, Frankel G (2018) Typhoidal Salmonella: distinctive virulence factors and pathogenesis. Cell Microbiol 20(9):e12939. https://doi.org/10.1111/cmi.12939
Johnson TS, Li S, Franz E, Huang Z, Dan Li S, Campbell MJ, Huang K, Zhang Y (2019) PseudoFuN: deriving functional potentials of pseudogenes from integrative relationships with genes and microRNAs across 32 cancers. Gigascience 8(5). https://doi.org/10.1093/gigascience/giz046
Kalyana-Sundaram S, Kumar-Sinha C, Shankar S, Robinson DR, Wu YM, Cao X, Asangani IA, Kothari V, Prensner JR, Lonigro RJ, Iyer MK, Barrette T, Shanmugam A, Dhanasekaran SM, Palanisamy N, Chinnaiyan AM (2012) Expressed pseudogenes in the transcriptional landscape of human cancers. Cell 149(7):1622–1634. https://doi.org/10.1016/j.cell.2012.04.041
Key FM, Posth C, Esquivel-Gomez LR, Hubler R, Spyrou MA, Neumann GU, Furtwangler A, Sabin S, Burri M, Wissgott A, Lankapalli AK, Vagene AJ, Meyer M, Nagel S, Tukhbatova R, Khokhlov A, Chizhevsky A, Hansen S, Belinsky AB, Kalmykov A, Kantorovich AR, Maslov VE, Stockhammer PW, Vai S, Zavattaro M, Riga A, Caramelli D, Skeates R, Beckett J, Gradoli MG, Steuri N, Hafner A, Ramstein M, Siebke I, Losch S, Erdal YS, Alikhan NF, Zhou ZM, Achtman M, Bos K, Reinhold S, Haak W, Kuhnert D, Herbig A, Krause J (2020) Emergence of human-adapted Salmonella enterica is linked to the Neolithization process. Nat Ecol Evol 4(3):324-+. https://doi.org/10.1038/s41559-020-1106-9
Kingsley RA, Santos RL, Keestra AM, Adams LG, Baumler AJ (2002) Salmonella enterica serotype Typhimurium ShdA is an outer membrane fibronectin-binding protein that is expressed in the intestine. Mol Microbiol 43(4):895–905. https://doi.org/10.1046/j.1365-2958.2002.02805.x
Kingsley RA, Abi Ghanem D, Puebla-Osorio N, Keestra AM, Berghman L, Baumler AJ (2004) Fibronectin binding to the Salmonella enterica serotype Typhimurium ShdA autotransporter protein is inhibited by a monoclonal antibody recognizing the A3 repeat. J Bacteriol 186(15):4931–4939. https://doi.org/10.1128/JB.186.15.4931-4939.2004
Kingsley RA, Msefula CL, Thomson NR, Kariuki S, Holt KE, Gordon MA, Harris D, Clarke L, Whitehead S, Sangal V, Marsh K, Achtman M, Molyneux ME, Cormican M, Parkhill J, MacLennan CA, Heyderman RS, Dougan G (2009) Epidemic multiple drug resistant Salmonella Typhimurium causing invasive disease in sub-Saharan Africa have a distinct genotype. Genome Res 19(12):2279–2287. https://doi.org/10.1101/gr.091017.109
Kolodziejek AM, Miller SI (2015) Salmonella modulation of the phagosome membrane, role of SseJ. Cell Microbiol 17(3):333–341. https://doi.org/10.1111/cmi.12420
Kolodziejek AM, Altura MA, Fan J, Petersen EM, Cook M, Brzovic PS, Miller SI (2019) Salmonella translocated effectors recruit OSBP1 to the phagosome to promote vacuolar membrane integrity. Cell Rep 27(7):2147-2156 e5. https://doi.org/10.1016/j.celrep.2019.04.021
Konno Y, Kimura I, Uriu K, Fukushi M, Irie T, Koyanagi Y, Sauter D, Gifford RJ, Consortium U-C, Nakagawa S, Sato K (2020) SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is increased by a naturally occurring elongation variant. Cell Rep 32(12):108185. https://doi.org/10.1016/j.celrep.2020.108185
Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, de Silva TI, Sheffield C-GG, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, Montefiori DC (2020) Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182(4):812–827 e19. https://doi.org/10.1016/j.cell.2020.06.043
Kuhlen L, Johnson S, Zeitler A, Baurle S, Deme JC, Caesar JJE, Debo R, Fisher J, Wagner S, Lea SM (2020) The substrate specificity switch FlhB assembles onto the export gate to regulate type three secretion. Nat Commun 11(1):1296. https://doi.org/10.1038/s41467-020-15071-9
Kuo CH, Ochman H (2010) The extinction dynamics of bacterial pseudogenes. PLoS Genet 6(8). https://doi.org/10.1371/journal.pgen.1001050
Kwon J, Liu YV, Gao C, Bassal MA, Jones AI, Yang J, Chen Z, Li Y, Yang H, Chen L, Di Ruscio A, Tay Y, Chai L, Tenen DG (2021) Pseudogene-mediated DNA demethylation leads to oncogene activation. Sci Adv 7(40):eabg1695. https://doi.org/10.1126/sciadv.abg1695
Lai YX, Li JY, Zhong LT, He X, Si XY, Sun YL, Chen YM, Zhong JY, Hu YL, Li B, Liao WJ, Liu C, Liao YL, Xiu JC, Bin JP (2019) The pseudogene PTENP1 regulates smooth muscle cells as a competing endogenous RNA. Clin Sci 133(13):1439–1455. https://doi.org/10.1042/Cs20190156
Lam HY, Khurana E, Fang G, Cayting P, Carriero N, Cheung KH, Gerstein MB (2009) Pseudofam: the pseudogene families database. Nucleic Acids Res 37(Database issue):D738–43. https://doi.org/10.1093/nar/gkn758
Lawrence JG, Hendrix RW, Casjens S (2001) Where are the pseudogenes in bacterial genomes? Trends Microbiol 9(11):535–540. https://doi.org/10.1016/s0966-842x(01)02198-9
Lerat E, Ochman H (2004) Psi-Phi: exploring the outer limits of bacterial pseudogenes. Genome Res 14(11):2273–2278. https://doi.org/10.1101/gr.2925604
Lerat E, Ochman H (2005) Recognizing the pseudogenes in bacterial genomes. Nucleic Acids Res 33(10):3125–3132. https://doi.org/10.1093/nar/gki631
Li X, Yang L, Chen LL (2018) The biogenesis, functions, and challenges of circular RNAs. Mol Cell 71(3):428–442. https://doi.org/10.1016/j.molcel.2018.06.034
Li W, O’Neill KR, Haft DH, DiCuccio M, Chetvernin V, Badretdin A, Coulouris G, Chitsaz F, Derbyshire MK, Durkin AS, Gonzales NR, Gwadz M, Lanczycki CJ, Song JS, Thanki N, Wang J, Yamashita RA, Yang M, Zheng C, Marchler-Bauer A, Thibaud-Nissen F (2021) RefSeq: expanding the Prokaryotic Genome Annotation Pipeline reach with protein family model curation. Nucleic Acids Res 49(D1):D1020–D1028. https://doi.org/10.1093/nar/gkaa1105
Liao J, Orsi RH, Carroll LM, Wiedmann M (2020) Comparative genomics reveals different population structures associated with host and geographic origin in antimicrobial-resistant Salmonella enterica. Environ Microbiol 22(7):2811–2828. https://doi.org/10.1111/1462-2920.15014
Lin ZT, Du LF, Zhang MZ, Han XY, Wang BH, Meng J, Yu FX, Zhou XQ, Wang N, Li C, Wang XY, Liu J, Gao WY, Ye RZ, Xia LY, Sun Y, Jia N, Jiang JF, Zhao L, Cui XM, Zhan L, Cao WC (2023) Genomic characteristics of emerging intraerythrocytic Anaplasma capra and high prevalence in goats, China. Emerg Infect Dis 29(9):1780–1788. https://doi.org/10.3201/eid2909.230131
Liu Y, Harrison PM, Kunin V, Gerstein M (2004) Comprehensive analysis of pseudogenes in prokaryotes: widespread gene decay and failure of putative horizontally transferred genes. Genome Biol 5(9):R64. https://doi.org/10.1186/gb-2004-5-9-r64
Liu ZJ, Peasley AM, Yang JF, Li YQ, Guan GQ, Luo JX, Yin H, Brayton KA (2019) The Anaplasma ovis genome reveals a high proportion of pseudogenes. Bmc Genom 20:69. https://doi.org/10.1186/s12864-018-5374-6
Liu P, Li Y, Ye Y, Chen J, Li R, Zhang Q, Li Y, Wang W, Meng Q, Ou J, Yang Z, Sun W, Gu W (2022a) The genome and antigen proteome analysis of Spiroplasma mirum. Front Microbiol 13:996938. https://doi.org/10.3389/fmicb.2022.996938
Liu P, Wang X, Sun Y, Zhao H, Cheng F, Wang J, Yang F, Hu J, Zhang H, Wang CC, Wang L (2022b) SARS-CoV-2 ORF8 reshapes the ER through forming mixed disulfides with ER oxidoreductases. Redox Biol 54:102388. https://doi.org/10.1016/j.redox.2022.102388
Lou WY, Ding BS, Fu PF (2020) Pseudogene-derived lncRNAs and their miRNA sponging mechanism in human cancer. Front Cell Dev Biol 8:85. https://doi.org/10.3389/fcell.2020.00085
Luo B, Wang J, Liu Z, Shen Z, Shi R, Liu YQ, Liu Y, Jiang M, Wu Y, Zhang Z (2016) Phagocyte respiratory burst activates macrophage erythropoietin signalling to promote acute inflammation resolution. Nat Commun 7:12177. https://doi.org/10.1038/ncomms12177
Ma SS, Liu XQ, Ma S, Jiang LY (2021) SopA inactivation or reduced expression is selected in intracellular Salmonella and contributes to systemic Salmonella infection. Res Microbiol 172(2):103795. https://doi.org/10.1016/j.resmic.2020.103795
Makishima S, Komoriya K, Yamaguchi S, Aizawa SI (2001) Length of the flagellar hook and the capacity of the type III export apparatus. Science 291(5512):2411–2413. https://doi.org/10.1126/science.1058366
Mannar D, Saville JW, Sun Z, Zhu X, Marti MM, Srivastava SS, Berezuk AM, Zhou S, Tuttle KS, Sobolewski MD, Kim A, Treat BR, Da Silva Castanha PM, Jacobs JL, Barratt-Boyes SM, Mellors JW, Dimitrov DS, Li W, Subramaniam S (2022) SARS-CoV-2 variants of concern: spike protein mutational analysis and epitope for broad neutralization. Nat Commun 13(1):4696. https://doi.org/10.1038/s41467-022-32262-8
McClelland M, Sanderson KE, Spieth J, Clifton SW, Latreille P, Courtney L, Porwollik S, Ali J, Dante M, Du F, Hou S, Layman D, Leonard S, Nguyen C, Scott K, Holmes A, Grewal N, Mulvaney E, Ryan E, Sun H, Florea L, Miller W, Stoneking T, Nhan M, Waterston R, Wilson RK (2001) Complete genome sequence of Salmonella enterica serovar Typhimurium LT2. Nature 413(6858):852–856. https://doi.org/10.1038/35101614
McClelland M, Sanderson KE, Clifton SW, Latreille P, Porwollik S, Sabo A, Meyer R, Bieri T, Ozersky P, McLellan M, Harkins CR, Wang C, Nguyen C, Berghoff A, Elliott G, Kohlberg S, Strong C, Du F, Carter J, Kremizki C, Layman D, Leonard S, Sun H, Fulton L, Nash W, Miner T, Minx P, Delehaunty K, Fronick C, Magrini V, Nhan M, Warren W, Florea L, Spieth J, Wilson RK (2004) Comparison of genome degradation in Paratyphi A and Typhi, human-restricted serovars of Salmonella enterica that cause typhoid. Nat Genet 36(12):1268–1274. https://doi.org/10.1038/ng1470
McCutcheon JP, Moran NA (2011) Extreme genome reduction in symbiotic bacteria. Nat Rev Microbiol 10(1):13–26. https://doi.org/10.1038/nrmicro2670
Miller TLA, Orpinelli Rego F, Buzzo JLL, Galante PAF (2021) sideRETRO: a pipeline for identifying somatic and polymorphic insertions of processed pseudogenes or retrocopies. Bioinformatics 37(3):419–421. https://doi.org/10.1093/bioinformatics/btaa689
Minamino T, Morimoto YV, Kinoshita M, Namba K (2021) Multiple roles of flagellar export chaperones for efficient and robust flagellar filament formation in Salmonella. Front Microbiol 12:756044. https://doi.org/10.3389/fmicb.2021.756044
Murray GL, Attridge SR, Morona R (2003) Regulation of Salmonella Typhimurium lipopolysaccharide O antigen chain length is required for virulence; identification of FepE as a second Wzz. Mol Microbiol 47(5):1395–1406. https://doi.org/10.1046/j.1365-2958.2003.03383.x
Naville M, Warren A, Haftek-Terreau Z, Chalopin D, Brunet F, Levin P, Galiana D, Volff JN (2016) Not so bad after all: retroviruses and long terminal repeat retrotransposons as a source of new genes in vertebrates. Clin Microbiol Infec 22(4):312–323. https://doi.org/10.1016/j.cmi.2016.02.001
Ochman H, Davalos LM (2006) The nature and dynamics of bacterial genomes. Science 311(5768):1730–1733. https://doi.org/10.1126/science.1119966
Ochman H, Moran NA (2001) Genes lost and genes found: evolution of bacterial pathogenesis and symbiosis. Science 292(5519):1096–1099. https://doi.org/10.1126/science.1058543
Oh SH, Schliep K, Isenhower A, Rodriguez-Bobadilla R, Vuong VM, Fields CJ, Hernandez AG, Hoyer LL (2021) Using genomics to shape the definition of the agglutinin-like sequence (ALS) family in the Saccharomycetales. Front Cell Infect Microbiol 11:794529. https://doi.org/10.3389/fcimb.2021.794529
Ortega AP, Villagra NA, Urrutia IM, Valenzuela LM, Talamilla-Espinoza A, Hidalgo AA, Rodas PI, Gil F, Calderon IL, Paredes-Sabja D, Mora GC, Fuentes JA (2016) Lose to win: marT pseudogenization in Salmonella enterica serovar Typhi contributed to the surV-dependent survival to H2O2, and inside human macrophage-like cells. Infect Genet Evol 45:111–121. https://doi.org/10.1016/j.meegid.2016.08.029
Oscarsson J, Westermark M, Lofdahl S, Olsen B, Palmgren H, Mizunoe Y, Wai SN, Uhlin BE (2002) Characterization of a pore-forming cytotoxin expressed by Salmonella enterica serovars Typhi and Paratyphi A. Infect Immun 70(10):5759–5769. https://doi.org/10.1128/IAI.70.10.5759-5769.2002
Parkhill J, Dougan G, James KD, Thomson NR, Pickard D, Wain J, Churcher C, Mungall KL, Bentley SD, Holden MTG, Sebaihia M, Baker S, Basham D, Brooks K, Chillingworth T, Connerton P, Cronin A, Davis P, Davies RM, Dowd L, White N, Farrar J, Feltwell T, Hamlin N, Haque A, Hien TT, Holroyd S, Jagels K, Krogh A, Larsen TS, Leather S, Moule S, O’Gaora P, Parry C, Quail M, Rutherford K, Simmonds M, Skelton J, Stevens K, Whitehead S, Barrell BG (2001) Complete genome sequence of a multiple drug resistant Salmonella enterica serovar Typhi CT18. Nature 413(6858):848–852. https://doi.org/10.1038/35101607
Parsons BN, Humphrey S, Salisbury AM, Mikoleit J, Hinton JC, Gordon MA, Wigley P (2013) Invasive non-typhoidal Salmonella Typhimurium ST313 are not host-restricted and have an invasive phenotype in experimentally infected chickens. PLoS Negl Trop Dis 7(10):e2487. https://doi.org/10.1371/journal.pntd.0002487
Patel S, Mathivanan N, Goyal A (2017) Bacterial adhesins, the pathogenic weapons to trick host defense arsenal. Biomed Pharmacother 93:763–771. https://doi.org/10.1016/j.biopha.2017.06.102
Paxman JJ, Lo AW, Sullivan MJ, Panjikar S, Kuiper M, Whitten AE, Wang G, Luan CH, Moriel DG, Tan L, Peters KM, Phan MD, Gee CL, Ulett GC, Schembri MA, Heras B (2019) Unique structural features of a bacterial autotransporter adhesin suggest mechanisms for interaction with host macromolecules. Nat Commun 10(1):1967. https://doi.org/10.1038/s41467-019-09814-6
Pei B, Sisu C, Frankish A, Howald C, Habegger L, Mu XJ, Harte R, Balasubramanian S, Tanzer A, Diekhans M, Reymond A, Hubbard TJ, Harrow J, Gerstein MB (2012) The GENCODE pseudogene resource. Genome Biol 13(9):R51. https://doi.org/10.1186/gb-2012-13-9-r51
Perkins TT, Kingsley RA, Fookes MC, Gardner PP, James KD, Yu L, Assefa SA, He M, Croucher NJ, Pickard DJ, Maskell DJ, Parkhill J, Choudhary J, Thomson NR, Dougan G (2009) A strand-specific RNA-Seq analysis of the transcriptome of the typhoid bacillus Salmonella Typhi. PLoS Genet 5(7):e1000569. https://doi.org/10.1371/journal.pgen.1000569
Qian SH, Chen L, Xiong YL, Chen ZX (2022) Evolution and function of developmentally dynamic pseudogenes in mammals. Genome Biol 23(1):235. https://doi.org/10.1186/s13059-022-02802-y
Qu Y, Wang X, Zhu Y, Wang W, Wang Y, Hu G, Liu C, Li J, Ren S, Xiao MZX, Liu Z, Wang C, Fu J, Zhang Y, Li P, Zhang R, Liang Q (2021) ORF3a-mediated incomplete autophagy facilitates severe acute respiratory syndrome coronavirus-2 replication. Front Cell Dev Biol 9:716208. https://doi.org/10.3389/fcell.2021.716208
Ratcliffe MJ (2006) Antibodies, immunoglobulin genes and the bursa of fabricius in chicken B cell development. Dev Comp Immunol 30(1–2):101–118. https://doi.org/10.1016/j.dci.2005.06.018
Rehder C, Bean LJH, Bick D, Chao E, Chung W, Das S, O’Daniel J, Rehm H, Shashi V, Vincent LM, Committee ALQA (2021) Next-generation sequencing for constitutional variants in the clinical laboratory, 2021 revision: a technical standard of the American College of Medical Genetics and Genomics (ACMG). Genet Med 23(8):1399–1415. https://doi.org/10.1038/s41436-021-01139-4
Rejmanek D, Foley P, Barbet A, Foley J (2012) Evolution of antigen variation in the tick-borne pathogen Anaplasma phagocytophilum. Mol Biol Evol 29(1):391–400. https://doi.org/10.1093/molbev/msr229
Restrepo BI, Carter CJ, Barbour AG (1994) Activation of a vmp pseudogene in Borrelia hermsii: an alternate mechanism of antigenic variation during relapsing fever. Mol Microbiol 13(2):287–299. https://doi.org/10.1111/j.1365-2958.1994.tb00423.x
Retamal P, Castillo-Ruiz M, Villagra NA, Morgado J, Mora GC (2010) Modified intracellular-associated phenotypes in a recombinant Salmonella Typhi expressing S. Typhimurium SPI-3 sequences. PLoS One 5(2):e9394. https://doi.org/10.1371/journal.pone.0009394
Sayers EW, Beck J, Brister JR, Bolton EE, Canese K, Comeau DC, Funk K, Ketter A, Kim S, Kimchi A, Kitts PA, Kuznetsov A, Lathrop S, Lu Z, McGarvey K, Madden TL, Murphy TD, O’Leary N, Phan L, Schneider VA, Thibaud-Nissen F, Trawick BW, Pruitt KD, Ostell J (2020) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 48(D1):D9–D16. https://doi.org/10.1093/nar/gkz899
Schmid-Siegert E, Richard S, Luraschi A, Muhlethaler K, Pagni M, Hauser PM (2017) Mechanisms of surface antigenic variation in the human pathogenic fungus Pneumocystis jirovecii. Mbio 8(6):e01470-17. https://doi.org/10.1128/mBio.01470-17
Schoen C, Tettelin H, Parkhill J, Frosch M (2009) Genome flexibility in Neisseria meningitidis. Vaccine 27(Suppl 2):B103–B111. https://doi.org/10.1016/j.vaccine.2009.04.064
Schroeder N, Henry T, de Chastellier C, Zhao W, Guilhon AA, Gorvel JP, Meresse S (2010) The virulence protein SopD2 regulates membrane dynamics of Salmonella-containing vacuoles. PLoS Pathog 6(7):e1001002. https://doi.org/10.1371/journal.ppat.1001002
Schwartz I, Margos G, Casjens SR, Qiu WG, Eggers CH (2021) Multipartite genome of lyme disease Borrelia: structure, variation and prophages. Curr Issues Mol Biol 42:409–454. https://doi.org/10.21775/cimb.042.409
Seif Y, Monk JM, Machado H, Kavvas E, Palsson BO (2019) Systems biology and pangenome of Salmonella O-antigens. mBio 10(4). https://doi.org/10.1128/mBio.01247-19
Shi AC, Xie X (2021) Making sense of spike D614G in SARS-CoV-2 transmission. Sci China Life Sci 64(7):1062–1067. https://doi.org/10.1007/s11427-020-1893-9
Shimizu N, Katagiri T, Matsumoto A, Matsuda Y, Arai H, Sasaki N, Abe K, Katase T, Ishida H, Kusumoto KI, Takeuchi M, Yamagata Y (2021) Oryzapsins, the orthologs of yeast yapsin in Aspergillus oryzae, affect ergosterol synthesis. Appl Microbiol Biotechnol 105(21–22):8481–8494. https://doi.org/10.1007/s00253-021-11639-7
Sijmons S, Thys K, Corthout M, Van Damme E, Van Loock M, Bollen S, Baguet S, Aerssens J, Van Ranst M, Maes P (2014) A method enabling high-throughput sequencing of human cytomegalovirus complete genomes from clinical isolates. PLoS ONE 9(4):e95501. https://doi.org/10.1371/journal.pone.0095501
Sijmons S, Thys K, Mbong Ngwese M, Van Damme E, Dvorak J, Van Loock M, Li G, Tachezy R, Busson L, Aerssens J, Van Ranst M, Maes P (2015) High-throughput analysis of human cytomegalovirus genome diversity highlights the widespread occurrence of gene-disrupting mutations and pervasive recombination. J Virol 89(15):7673–7695. https://doi.org/10.1128/JVI.00578-15
Silva FJ, Santos-Garcia D, Zheng X, Zhang L, Han XY (2022) Construction and analysis of the complete genome sequence of leprosy agent Mycobacterium lepromatosis. Microbiol Spectr 10(3):e0169221. https://doi.org/10.1128/spectrum.01692-21
Sisu C (2021a) GENCODE Pseudogenes. Methods Mol Biol 2324:67–82. https://doi.org/10.1007/978-1-0716-1503-4_5
Sisu C (2021b) Pseudogenes as biomarkers and therapeutic targets in human cancers. Methods Mol Biol 2324:319–337. https://doi.org/10.1007/978-1-0716-1503-4_20
Sisu C, Muir P, Frankish A, Fiddes I, Diekhans M, Thybert D, Odom DT, Flicek P, Keane TM, Hubbard T, Harrow J, Gerstein M (2020) Transcriptional activity and strain-specific history of mouse pseudogenes. Nat Commun 11(1):3695. https://doi.org/10.1038/s41467-020-17157-w
Soucy SM, Huang J, Gogarten JP (2015) Horizontal gene transfer: building the web of life. Nat Rev Genet 16(8):472–482. https://doi.org/10.1038/nrg3962
Su YCF, Anderson DE, Young BE, Linster M, Zhu F, Jayakumar J, Zhuang Y, Kalimuddin S, Low JGH, Tan CW, Chia WN, Mak TM, Octavia S, Chavatte JM, Lee RTC, Pada S, Tan SY, Sun L, Yan GZ, Maurer-Stroh S, Mendenhall IH, Leo YS, Lye DC, Wang LF, Smith GJD (2020) Discovery and genomic characterization of a 382-nucleotide deletion in ORF7b and ORF8 during the early evolution of SARS-CoV-2. mBio 11(4). https://doi.org/10.1128/mBio.01610-20
Suarez NM, Wilkie GS, Hage E, Camiolo S, Holton M, Hughes J, Maabar M, Vattipally SB, Dhingra A, Gompels UA, Wilkinson GWG, Baldanti F, Furione M, Lilleri D, Arossa A, Ganzenmueller T, Gerna G, Hubacek P, Schulz TF, Wolf D, Zavattoni M, Davison AJ (2019) Human cytomegalovirus genomes sequenced directly from clinical material: variation, multiple-strain infection, recombination, and gene loss. J Infect Dis 220(5):781–791. https://doi.org/10.1093/infdis/jiz208
Sugawara-Mikami M, Tanigawa K, Kawashima A, Kiriya M, Nakamura Y, Fujiwara Y, Suzuki K (2022) Pathogenicity and virulence of Mycobacterium leprae. Virulence 13(1):1985–2011. https://doi.org/10.1080/21505594.2022.2141987
Sun M, Wang Y, Zheng C, Wei Y, Hou J, Zhang P, He W, Lv X, Ding Y, Liang H, Hon CC, Chen X, Xu H, Chen Y (2021) Systematic functional interrogation of human pseudogenes using CRISPRi. Genome Biol 22(1):240. https://doi.org/10.1186/s13059-021-02464-2
Syberg-Olsen MJ, Garber AI, Keeling PJ, McCutcheon JP, Husnik F (2022) Pseudofinder: detection of pseudogenes in prokaryotic genomes. Mol Biol Evol 39(7). https://doi.org/10.1093/molbev/msac153
Tam OH, Aravin AA, Stein P, Girard A, Murchison EP, Cheloufi S, Hodges E, Anger M, Sachidanandam R, Schultz RM, Hannon GJ (2008) Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453(7194):534–538. https://doi.org/10.1038/nature06904
Tanner JR, Kingsley RA (2018) Evolution of Salmonella within hosts. Trends Microbiol 26(12):986–998. https://doi.org/10.1016/j.tim.2018.06.001
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44(14):6614–6624. https://doi.org/10.1093/nar/gkw569
Taylor DJ, Dittmar K, Ballinger MJ, Bruenn JA (2011) Evolutionary maintenance of filovirus-like genes in bat genomes. BMC Evol Biol 11:336. https://doi.org/10.1186/1471-2148-11-336
Tong Z, Zhou D, Song Y, Zhang L, Pei D, Han Y, Pang X, Li M, Cui B, Wang J, Guo Z, Qi Z, Jin L, Zhai J, Du Z, Wang J, Wang X, Yu J, Wang J, Huang P, Yang H, Yang R (2005) Pseudogene accumulation might promote the adaptive microevolution of Yersinia pestis. J Med Microbiol 54(Pt 3):259–268. https://doi.org/10.1099/jmm.0.45752-0
Torrents D, Suyama M, Zdobnov E, Bork P (2003) A genome-wide survey of human pseudogenes. Genome Res 13(12):2559–2567. https://doi.org/10.1101/gr.1455503
Trombert AN, Berrocal L, Fuentes JA, Mora GC (2010) S. Typhimurium sseJ gene decreases the S. Typhi cytotoxicity toward cultured epithelial cells. BMC Microbiol 10:312. https://doi.org/10.1186/1471-2180-10-312
Trombert AN, Rodas PI, Mora GC (2011) Reduced invasion to human epithelial cell lines of Salmonella enterica serovar Typhi carrying S. Typhimurium sopD2. FEMS Microbiol Lett 322(2):150–6. https://doi.org/10.1111/j.1574-6968.2011.02347.x
Urrutia IM, Fuentes JA, Valenzuela LM, Ortega AP, Hidalgo AA, Mora GC (2014) Salmonella Typhi shdA: pseudogene or allelic variant? Infect Genet Evol 26:146–152. https://doi.org/10.1016/j.meegid.2014.05.013
Verduci L, Tarcitano E, Strano S, Yarden Y, Blandino G (2021) CircRNAs: role in human diseases and potential use as biomarkers. Cell Death Dis 12(5):468. https://doi.org/10.1038/s41419-021-03743-3
Verma N, Reeves P (1989) Identification and sequence of rfbS and rfbE, which determine antigenic specificity of group A and group D Salmonellae. J Bacteriol 171(10):5694–5701. https://doi.org/10.1128/jb.171.10.5694-5701.1989
Vihinen M (2014) Contribution of pseudogenes to sequence diversity. Methods Mol Biol 1167:15–24. https://doi.org/10.1007/978-1-4939-0835-6_2
Walch P, Selkrig J, Knodler LA, Rettel M, Stein F, Fernandez K, Vieitez C, Potel CM, Scholzen K, Geyer M, Rottner K, Steele-Mortimer O, Savitski MM, Holden DW, Typas A (2021) Global mapping of Salmonella enterica-host protein-protein interactions during infection. Cell Host Microbe 29(8):1316–1332 e12. https://doi.org/10.1016/j.chom.2021.06.004
Wang Y, Ledvina HE, Tower CA, Kambarev S, Liu E, Charity JC, Kreuk LSM, Tang Q, Chen Q, Gallagher LA, Radey MC, Rerolle GF, Li Y, Penewit KM, Turkarslan S, Skerrett SJ, Salipante SJ, Baliga NS, Woodward JJ, Dove SL, Peterson SB, Celli J, Mougous JD (2023) Discovery of a glutathione utilization pathway in Francisella that shows functional divergence between environmental and pathogenic species. Cell Host Microbe 31(8):1359–1370 e7. https://doi.org/10.1016/j.chom.2023.06.010
Wen YZ, Zheng LL, Liao JY, Wang MH, Wei Y, Guo XM, Qu LH, Ayala FJ, Lun ZR (2011) Pseudogene-derived small interference RNAs regulate gene expression in African Trypanosoma brucei. Proc Natl Acad Sci U S A 108(20):8345–8350. https://doi.org/10.1073/pnas.1103894108
Woo PC, Fung AM, Wong SS, Tsoi HW, Yuen KY (2001) Isolation and characterization of a Salmonella enterica serotype Typhi variant and its clinical and public health implications. J Clin Microbiol 39(3):1190–1194. https://doi.org/10.1128/JCM.39.3.1190-1194.2001
Wu Y, Hao T, Qian X, Zhang X, Song Y, Yang R, Cui Y (2022) Small insertions and deletions drive genomic plasticity during adaptive evolution of Yersinia pestis. Microbiol Spectr 10(3):e0224221. https://doi.org/10.1128/spectrum.02242-21
Xie JB, Li Y, Liu XM, Zhao YY, Li BL, Ingvarsson PK, Zhang DQ (2019) Evolutionary origins of pseudogenes and their association with regulatory sequences in plants. Plant Cell 31(3):563–578. https://doi.org/10.1105/tpc.18.00601
Xu JR, Zhang JZ (2016) Are human translated pseudogenes functional? Mol Biol Evol 33(3):755–760. https://doi.org/10.1093/molbev/msv268
Yang J, Barrila J, Roland KL, Kilbourne J, Ott CM, Forsyth RJ, Nickerson CA (2015) Characterization of the invasive, multidrug resistant non-typhoidal Salmonella strain D23580 in a murine model of infection. PLoS Negl Trop Dis 9(6):e0003839. https://doi.org/10.1371/journal.pntd.0003839
Yang Y, Wang P, Xia P, Yang B, Dai P, Hong T, Li J, Meng X, El Qaidi S, Zhu G (2020) Rapid detection of flagellated and non-flagellated Salmonella by targeting the common flagellar hook gene flgE. Appl Microbiol Biotechnol 104(22):9719–9732. https://doi.org/10.1007/s00253-020-10925-0
Yebra G, Haag AF, Neamah MM, Wee BA, Richardson EJ, Horcajo P, Granneman S, Tormo-Mas MA, de la Fuente R, Fitzgerald JR, Penades JR (2021) Radical genome remodelling accompanied the emergence of a novel host-restricted bacterial pathogen. PLoS Pathog 17(5):e1009606. https://doi.org/10.1371/journal.ppat.1009606
Young BE, Fong SW, Chan YH, Mak TM, Ang LW, Anderson DE, Lee CY, Amrun SN, Lee B, Goh YS, Su YCF, Wei WE, Kalimuddin S, Chai LYA, Pada S, Tan SY, Sun L, Parthasarathy P, Chen YYC, Barkham T, Lin RTP, Maurer-Stroh S, Leo YS, Wang LF, Renia L, Lee VJ, Smith GJD, Lye DC, Ng LFP (2020) Effects of a major deletion in the SARS-CoV-2 genome on the severity of infection and the inflammatory response: an observational cohort study. Lancet 396(10251):603–611. https://doi.org/10.1016/S0140-6736(20)31757-8
Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y, Baum A, Diehl WE, Dauphin A, Carbone C, Veinotte K, Egri SB, Schaffner SF, Lemieux JE, Munro JB, Rafique A, Barve A, Sabeti PC, Kyratsous CA, Dudkina NV, Shen K, Luban J (2020) Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183(3):739–751 e8. https://doi.org/10.1016/j.cell.2020.09.032
Zakaria MA, Yusoff MZM, Zakaria MR, Hassan MA, Wood TK, Maeda T (2018) Pseudogene product YqiG is important for pflB expression and biohydrogen production in Escherichia coli BW25113. 3 Biotech 8(10):435. https://doi.org/10.1007/s13205-018-1461-2
Zhang Y, Nelson M, Van Etten JL (1992) A single amino acid change restores DNA cytosine methyltransferase activity in a cloned chlorella virus pseudogene. Nucleic Acids Res 20(7):1637–1642. https://doi.org/10.1093/nar/20.7.1637
Zhang Z, Carriero N, Gerstein M (2004) Comparative analysis of processed pseudogenes in the mouse and human genomes. Trends Genet 20(2):62–67. https://doi.org/10.1016/j.tig.2003.12.005
Zhang Z, Carriero N, Zheng D, Karro J, Harrison PM, Gerstein M (2006) PseudoPipe: an automated pseudogene identification pipeline. Bioinformatics 22(12):1437–1439. https://doi.org/10.1093/bioinformatics/btl116
Zhang J, Cai Y, Xiao T, Lu J, Peng H, Sterling SM, Walsh RM Jr, Rits-Volloch S, Zhu H, Woosley AN, Yang W, Sliz P, Chen B (2021a) Structural impact on SARS-CoV-2 spike protein by D614G substitution. Science 372(6541):525–530. https://doi.org/10.1126/science.abf2303
Zhang G, Dong H, Feng Y, Jiang H, Wu T, Sun J, Wang X, Liu M, Peng X, Zhang Y, Zhang X, Zhu L, Ding J, Shen X (2022) The pseudogene BMEA_B0173 deficiency in Brucella melitensis contributes to m-epitope formation and potentiates virulence in a mice infection model. Curr Microbiol 79(12):378. https://doi.org/10.1007/s00284-022-03078-y
Zhang Y, Chen Y, Li Y, Huang F, Luo B, Yuan Y, Xia B, Ma X, Yang T, Yu F, Liu J, Liu B, Song Z, Chen J, Yan S, Wu L, Pan T, Zhang X, Li R, Huang W, He X, Xiao F, Zhang J, Zhang H (2021b) The ORF8 protein of SARS-CoV-2 mediates immune evasion through down-regulating MHC-Iota. Proc Natl Acad Sci U S A 118(23). https://doi.org/10.1073/pnas.2024202118
Zheng D, Frankish A, Baertsch R, Kapranov P, Reymond A, Choo SW, Lu Y, Denoeud F, Antonarakis SE, Snyder M, Ruan Y, Wei CL, Gingeras TR, Guigo R, Harrow J, Gerstein MB (2007) Pseudogenes in the ENCODE regions: consensus annotation, analysis of transcription, and evolution. Genome Res 17(6):839–851. https://doi.org/10.1101/gr.5586307
Zheng L, Li X, Meng X, Chou J, Hu J, Zhang F, Zhang Z, Xing Y, Liu Y, Xi T (2016) Competing endogenous RNA networks of CYP4Z1 and pseudogene CYP4Z2P confer tamoxifen resistance in breast cancer. Mol Cell Endocrinol 427:133–142. https://doi.org/10.1016/j.mce.2016.03.012
Zheng LL, Zhou KR, Liu S, Zhang DY, Wang ZL, Chen ZR, Yang JH, Qu LH (2018) dreamBase: DNA modification, RNA regulation and protein binding of expressed pseudogenes in human health and disease. Nucleic Acids Res 46(D1):D85–D91. https://doi.org/10.1093/nar/gkx972
Zhou MX, Yang Y, Chen PL, Hu HJ, Hardwidge PR, Zhu GQ (2015) More than a locomotive organelle: flagella in Escherichia coli. Appl Microbiol Biot 99(21):8883–8890. https://doi.org/10.1007/s00253-015-6946-x
Zhou L, Yu H, Wang K, Chen T, Ma Y, Huang Y, Li J, Liu L, Li Y, Kong Z, Zheng Q, Wang Y, Gu Y, Xia N, Li S (2020a) Genome re-sequencing and reannotation of the Escherichia coli ER2566 strain and transcriptome sequencing under overexpression conditions. BMC Genomics 21(1):407. https://doi.org/10.1186/s12864-020-06818-1
Zhou Z, Charlesworth J, Achtman M (2020b) Accurate reconstruction of bacterial pan- and core genomes with PEPPAN. Genome Res 30(11):1667–1679. https://doi.org/10.1101/gr.260828.120
Zhou MX, Duan QD, Zhu XF, Guo ZY, Li YC, Hardwidge PR, Zhu GQ (2013) Both flagella and F4 fimbriae from F4ac(+) enterotoxigenic Escherichia coli contribute to attachment to IPEC-J2 cells in vitro. Vet Res 44:30. https://doi.org/10.1186/1297-9716-44-30
Zhou WY, Wen H, Li YJ, Gao YJ, Zheng XF, Yuan L, Zhu GQ, Yang ZQ (2022) Whole-genome analysis reveals that bacteriophages promote environmental adaptation of Staphylococcus aureus via gene exchange, acquisition, and loss. Viruses 14(6):1199. https://doi.org/10.3390/v14061199
Funding
This study was supported by grants from the Zhong Ze Zhen Xing Program of Jiangsu Province (grant number JBGS (2021)111), the High End Talent Program for International Collaboration (G2022014150L), and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
YY and PW conceived and drafted the manuscript of this review. QS and PH critically reviewed and edited the manuscript. JH and GZ supervised and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yi Yang and Pengzhi Wang contributed equally to this work and share first authorship.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Y., Wang, P., Qaidi, S.E. et al. Loss to gain: pseudogenes in microorganisms, focusing on eubacteria, and their biological significance. Appl Microbiol Biotechnol 108, 328 (2024). https://doi.org/10.1007/s00253-023-12971-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12971-w