Abstract
Marine bacteria living in association with marine sponges have proven to be a reliable source of biologically active secondary metabolites. However, no studies have yet reported natural products from Microbacterium testaceum spp. We herein report the isolation of a M. testaceum strain from the sponge Tedania brasiliensis. Molecular networking analysis of bioactive pre-fractionated extracts from culture media of M. testaceum enabled the discovery of testacosides A–D. Analysis of spectroscopic data and chemical derivatizations allowed the identification of testacosides A–D as glycoglycerolipids bearing a 1-[α-glucopyranosyl-(1 → 3)-(α-mannopyranosyl)]-glycerol moiety connected to 12-methyltetradecanoic acid for testacoside A (1), 14-methylpentadecanoic acid for testacoside B (2), and 14-methylhexadecanoic acid for testacosides C (3) and D (4). The absolute configuration of the monosaccharide residues was determined by 1H-NMR analysis of the respective diastereomeric thiazolidine derivatives. This is the first report of natural products isolated from cultures of M. testaceum.
Key points
• The first report of metabolites produced by Microbacterium testaceum.
• 1-[α-Glucopyranosyl-(1 → 3)-(α-mannopyranosyl)]-glycerol lipids isolated and identified.
• Microbacterium testaceum strain isolated from the sponge Tedania brasiliensis.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine sponges (Porifera) are the oldest animals on Earth. Widely distributed all over the world, marine sponges harbor highly diverse and dense microbial communities, which may represent up to 60% of sponge biomass (Webster and Thomas 2016; Hentschel et al. 2012; Van Soest et al. 2012). Although the ecological roles of such microbe-sponge associations are yet poorly known, associated microorganisms are fundamental to maintain sponge health and defense against predators, since it has been shown that many sponge-associated bacteria and cyanobacteria produce various bioactive secondary metabolites (Balskus 2014). In return, sponge-associated microorganisms grow in a nutrient-rich environment (Abdelmohsen et al. 2014; Flórez et al. 2015). Evidences demonstrating the microbial origin of secondary metabolites isolated from marine sponges (Nakashima et al. 2016; Storey et al. 2020; Tianero et al. 2019; Piel et al. 2004; Agarwal et al. 2017) confer to these microorganisms the potential for biotechnological applications towards a sustainable and reliable source of pharmacologically bioactive molecules (Radjasa et al. 2011; Blockley et al. 2017; Brinkmann et al. 2017).
For example, bacteria isolated from marine sponges have been reported to produce antimicrobial agents (Almeida et al. 2023), including in fish aquacultures (Duan et al 2020), as well as several bioactive agents of medical importance (Karthikeyan et al 2022). Also, many metabolites produced by sponge-associated bacteria are of biotechnological interest to the cosmetic industry, such as carotenoids, lipids, melanin, anti-oxidant secondary metabolites, and as biosurfactants (Esposito et al 2021). Therefore, marine sponge-associated bacteria are currently considered of significant biotechnological potential for the production and development of several biotechnological products using green processes, positively impacting bioeconomy (Laport 2017; Romano S, 2018; Brinkmann et al 2017).
Traditional methods for the investigation of bacterial secondary metabolites are labor-intensive and time-consuming, often leading to the re-isolation of known compounds (Berlinck et al. 2019, 2022). Consequentially, an array of new approaches and strategies that enables reliable dereplication of microbial metabolic output have been developed (Gaudêncio and Pereira 2015; Ito and Masubuchi 2014; Wolfender et al. 2019; Beniddir et al. 2021). LC–MS-based metabolomic tools have been applied as effective strategies for the early stage prioritization of microbial bioactive metabolites (Bauermeister et al. 2022; Wang et al. 2016; Quinn et al. 2017).
We have previously investigated the chemistry of the marine sponge Tedania brasiliensis and discovered a series of bromopyrrole derivatives (Parra et al. 2018). We then decided to perform an extensive isolation of T. brasiliensis-associated cultivable bacteria aiming to obtain bacterial strains to yield media extracts potentially presenting brominated compounds. While the details of this screening searching for brominated metabolites will be reported elsewhere, few bacterial strains were detected to produce bioactive metabolites, among which a strain of the bacterium Microbacterium testaceum.
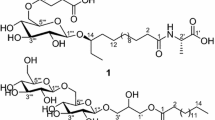
Thus, the aim of the present report was to investigate the bioactive compounds produced by M. testaceum isolated from the sponge T. brasiliensis. Analysis of M. testaceum media extracts by LC-HRMS/MS indicated a series of related but previously unreported compounds, present in a bioactive fraction. Further scaling up the bacterial culture enabled the MS-guided isolation of four new glycoglycerolipids, named testacosides A–D (1–4) (Fig. 1). Herein, we report the isolation, identification, and absolute configuration of testacosides A–D, the first metabolites isolated from cultures of a M. testaceum strain.
Materials and methods
General experimental procedures
Optical rotations were recorded on a Polartronic H Schmidt + Haensch polarimeter. Ultraviolet spectra were recorded on a Shimadzu UV-3600 spectrophotometer. Infrared spectra were recorded on a Shimadzu IRAffinnity-1 Fourier transform infrared spectrophotometer on a silicon plate. NMR spectra were obtained at 25 °C on a Bruker AvanceIII of 14.1 T equipment with a 5-mm cryoprobe, operating at 600.23 MHz for 1H and 150.94 MHz for 13C, using the residual signal solvent as internal standard. High resolution mass spectrometry (HRMS) data were acquired on a Waters Xevo G2-XS Q-TOF Mass Spectrometer, connected to a Waters Acquity UPLC-H Class system, equipped with a Waters Acquity UPLC BEH C18 column (dimensions 2.1 × 100 mm, 1.7 μm). The mobile phase consisted of a gradient of MeCN (Sigma) (+ 0.1% HCO2H) in H2O (Sigma) (+ 0.1% HCO2H), starting from 10% of MeCN to 100% MeCN in 7 min, maintained in 100% MeCN for 2 min, returning to 10% MeCN in 0.1 min, and reconditioning for 0.9 min, with a flow rate of 0.500 mL min−1. The column was maintained at 40 °C, and the samples were maintained at 15 °C. Mass spectrometry data acquisition was performed in centroid MS mode, positive resolution mode from 0 to 10 min, with an electrospray ionization (ESI) source and mass range from 100 to 2000 Da, with a scan time of 0.2 s−1. The ESI positive mode conditions were 1.2 kV capillary voltage, 30 V cone voltage, 100 °C source temperature, 450 °C desolvation temperature, 50 L h−1 cone gas flow, and 750 L h−1 desolvation gas flow. A 200 pg mL−1 solution of leucine enkephalin (Sigma) was used for internal calibration, infused by the lock-mass probe with a flow rate of 10 μL min−1. HPLC–PDA-ELSD-MS analyses were performed on a Waters Alliance 2695 control system coupled online with a Waters 2996 photodiode array detector (PDA), connected sequentially to a Waters 2424 evaporative light scattering detector (ELSD) and a Waters Micromass ZQ2000 low resolution mass spectrometry (MS) detector equipped with a ESI interface. PDA detector was set to detect in the wavelength range from 190 to 600 nm. The ELSD was operated with gain 100, nebulizer heating mode at 60%, drift tube temperature of 80 ± 5 °C, and N2 gas pressure of 50 psi. The MS detector was adjusted with the following parameters: capillary voltage 3.00 kV, source temperature 100 °C, desolvation temperature 350 °C, and simultaneous positive and negative ion detection from 100 to 1500 Da. Cone and desolvation gas flow rates are 50 and 350 L h−1, respectively. Chromatographic analyses were performed employing a Waters C18 X-Terra column (dimensions 250 × 4.6 mm; 5.0 μm), using a mobile phase with a flow rate of 1 mL min−1 consisting of (A) Milli-Q grade H2O (+ 0.1% HCO2H) and (B) 1:1 MeOH/MeCN (+ 0.1 HCO2H) as follows: 0–1.0 min hold at 10% B, then a linear gradient to 100% B from 1.0 to 18.0 min, hold at 100% B from 18 to 22 min, followed to 10% B at 22–23 min, and finally column equilibration from 23 to 30 min at 10% B. Volume injection of 20 μL of 2.0 mg mL−1 sample solutions was employed. Data acquisition and processing were performed employing the Empower 2.0 software.
Bacterial strain isolation and identification
The bacterial strain M. testaceum was isolated from specimens of the marine sponge T. brasiliensis collected between 9 and 38-m depth by SCUBA diving at Arraial do Cabo, state of Rio de Janeiro in January 2017. The strain M. testaceum was identified by 16S gene sequencing. The 16S rRNA gene sequencing analyses were performed by the University of Illinois at Chicago Research Resources Center, Sequencing Core, Chicago, USA. The partial 16S rRNA gene sequence (1063 nucleotides [nt]) was submitted to the Basic Local Alignment Search Tool (https://blast.ncbi.nlm.nih.gov) and EzTaxon server (http://eztaxon-e.ezbiocloud.net/; Kim et al. 2012), aligned with 16S rRNA gene sequences of the most closely related Microbacterium species using CLUSTAL W and phylogenetic trees were constructed by using MEGA 7.0 (Kumar et al. 2016). Phylogenetic trees were generated using the maximum-parsimony, minimum-evolution (Rzhetsky and Nei 1992), and neighbor-joining (Saitou and Nei 1987) algorithms drawn from the MEGA 7; an evolutionary distance matrix for the neighbor-joining analysis was prepared using the p-distance method (Nei and Kumar 2000) and is in the units of the number of base differences per site. The robustness of the inferred tree topologies was evaluated after 1000 bootstrap replicates (Felsenstein 1985). The analysis involved 12 nucleotide sequences. All positions with less than 95% site coverage were eliminated. That is, fewer than 5% alignment gaps, missing data, and ambiguous bases were allowed at any position. There were a total of 1029 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Gene sequences of the identified bacteria were obtained from BLASTN search in the GenBank database with the highest sequence similarities against type strains. GenBank accession number is OQ134920. A reference sample of the strain identified as M. testaceum has been deposited in the Brazilian Collection of Environmental and Industrial Microorganisms (CBMAI) under the accession number CBMAI 2824.
Extraction and isolation
M. testaceum strain was reactivated from frozen stocks in marine agar plates (Difco 2216) incubated at 28 °C for 2 days. A seed culture was inoculated by pouring individual colonies into 250 mL Schott flasks containing 100 mL of YEME medium (4 g yeast, 4 g glucose, and 10 g malt extract per liter of artificial sea water ASW) at 28 °C in a rotary shaker at 180 rpm for 3 days. Artificial sea water was prepared as follows: 30.0 g NaCl, 9.68 g MgCl2⋅6H2O, 3.47 g Na2SO4, 1.36 g CaCl2⋅2H2O, 0.610 KCl, 0.170 g NaHCO3, 0.1 g KBr, 0.030 g H3BO3, 0.040 g SrCl2⋅6H2O, and 0.140 g Na2HPO4 in 1 L of deionized water. Schott flasks (500 mL, 300 × 200 mL) containing YEME medium and 10 g of Diaion® HP-20 resin were inoculated with 2 mL aliquots of the seed culture and shaken at 28 °C and 180 rpm for 7 days. The resin was separated by vacuum filtration through cheesecloth, washed with deionized water, and extracted with MeOH (4 × 1 L) and acetone (4 × 1 L), then concentrated under reduced pressure to yield 14.3 g of a dry extract. Separately, the culture broth was extracted with an equal volume of EtOAc. Then, the EtOAc fraction was concentrated in vacuum to yield 4.6 g of an organic extract. Both HP-20 resin and EtOAc extracts showed similar chemical profiles by HPLC-MS and were pooled together. The pooled extract was fractionated by C18 reversed-phase column chromatography using a step gradient of H2O/MeOH from 20% MeOH to 100% MeOH to give five fractions. Fraction 5 (1.0 g) was further fractionated by chromatography on a Sephadex LH-20 column eluted with MeOH, yielding fractions 5A to 5H. Fraction 5C (185.1 mg) was purified by reversed-phase HPLC chromatography using a semipreparative C18 InertSustain column (250 × 10 mm, 5 μm) eluted in isocratic mode with H2O/MeCN 47:53 (0.1% formic acid) at flow rate of 4 mL min−1 and ELS detection, to give fraction 5C2 (testacoside A, 3.2 mg), fraction 5C3 (testacoside B, 2.5 mg), fraction 5C7 (testacoside C, 5.4 mg), and fraction 5C8 (testacoside D, 2.2 mg). Fraction 5D (668.9 mg) was purified by reversed-phase HPLC using a semipreparative C18 InertSustain column (250 × 10 mm, 5 μm) eluted in isocratic mode with H2O/MeCN 18:82 (0.1% formic acid) at flow rate of 4 mL min−1 and ELS detection, to give fraction 5D2 (testacoside A, 16.8 mg), fraction 5D3 (testacoside B, 0.8 mg), and 5D6 (testacoside C, 12.6 mg). NMR data of compounds 1-4 are provided in Tables 1 and 2 and NMR spectra in the supplementary information Figures S1-S30.
Testacoside A (1): clear colorless glass (0.0200 g); [α]25D 62.0 (c 0.002, CHCl3); IR (neat) vmax 3360, 1676 cm−1; 1H and 13C NMR data in Table 1; HRESIMS m/z 641.3733 [M + H]+ (calcd for C30H57O14, 641.3748).
Testacoside B (2): clear colorless glass (0.0033 g); [α]25D 72.9 (c 0.004, CHCl3); IR (neat) vmax 3344, 1678 cm−1; 1H and 13C NMR data in Table 1; HRESIMS m/z 655.3882 [M + H]+ (calcd for C31H59O14, 655.3905).
Testacoside C (3): clear colorless glass (0.0180 g); [α]25D + 72.4 (c 0.004, CHCl3); IR (neat) vmax 3369, 1678 cm−1; 1H and 13C NMR data in Table 1; HRESIMS m/z 669.4041 [M + H]+ (calcd for C32H61O14, 669.4061).
Testacoside D (4): clear colorless glass (0.0022 g); [α]25D 62.6 (c 0.002, CHCl3); IR (neat) vmax 3381, 1678 cm−1; 1H and 13C NMR data in Table 2; HRESIMS m/z 691.3864 [M + Na]+ (calcd for C32H60NaO14, 691.3881).
Acetylation of testacosides A–D (1–4)
Approximately 0.5 mg of each compound was dissolved in 1.5 mL of freshly distilled pyridine and 1.5 mL of freshly distilled acetic anhydride at room temperature for 18 h. The reaction mixture was evaporated under a N2 stream and the residue was partitioned between H2O and EtOAc. The organic phase was washed with H2O three times, dried under reduced pressure to give a yellow solid, for further analyses by NMR and HRMS. Data are provided in Tables S1-S2 and spectra in the supplementary information Figures S31-S56.
Testacoside A peracetate (5): clear yellow glass (0.0014 g); [α]25D 43.7 (c 0.001, CHCl3); IR (neat) vmax 1747 cm−1; 1H and 13C NMR data in Table S1; HRESIMS m/z 999.4453 [M + Na]+ (calcd for C46H72NaO22, 999.4407).
Testacoside B peracetate (6): clear yellow glass (0.0010 g); [α]25D 36.9 (c 0.001, CHCl3); IR (neat) vmax 1749 cm−1; 1H and 13C NMR data in Table S1; HRESIMS m/z 1013.4573 [M + Na]+ (calcd for C47H74NaO22, 1013.4564).
Testacoside C peracetate (7): clear yellow glass (0.0017 g); [α]25D 45.7 (c 0.002, CHCl3); IR (neat) vmax 1747 cm−1; 1H and 13C NMR data in Table S1; HRESIMS m/z 1027.4727 [M + Na]+ (calcd for C48H76NaO22, 1027.4720).
Testacoside D peracetate (8): clear yellow glass (0.0010 g); [α]25D + 38.0 (c 0.001, CHCl3); IR (neat) vmax 1749 cm−1; 1H and 13C NMR data in Table S2; HRESIMS m/z 1027.4731 [M + Na]+ (calcd for C48H76NaO22, 1027.4720).
Determination of the absolute configuration of sugar residues of 2 and 4
Separately, thiazolidine derivatives of D-glucose and D-mannose standards were prepared by weighting about 2.0 mg of each D-monosaccharide and 4.0 mg of L-cysteine methyl ester hydrochloride, dissolved in 120 μL of pyridine-d5 in NMR tubes. Reaction mixtures were heated at 60 °C for 1 h and then left at room temperature overnight before 1H NMR analysis. Additionally, a reaction with a 1:1 mixture of D-glucose and D-mannose was carried out under the abovementioned conditions. Testacosides B (2) and D (4) (1.0 mg each) were dissolved in 2 M trifluoroacetic acid (0.5 mL) and stirred at 100 °C for 8 h. After the completion of hydrolysis, the TFA was evaporated in vacuo to obtain the hydrolysate, redissolved in deionized H2O (3 mL), and partitioned with CHCl3 (3 mL, × 3). The aqueous phase containing the sugar residues was dissolved in 120 μL of pyridine-d5 and mixed with 1.0 mg of L-cysteine methyl ester hydrochloride to prepare the thiazolidine derivatives from the isolated compounds, as described above. Comparison of 1H NMR shifts and J coupling constants with standard derivatives was performed to establish the absolute configuration of glucose and mannose as the D-enantiomers.
GNPS molecular networking analysis
Tandem MS data were acquired on a data-dependent acquisition (DDA) mode with a ramp collision energy (CE) of low CE from 6 to 9 eV and high CE from 60 to 80 eV, for fragmentation of the three most abundant ions. The raw data were converted to mzXML data format using MSConvert (ProteoWizard) (Chambers et. al. 2012; Holman et al. 2014). A molecular network was created following the Classical Molecular Networking protocol on the GNPS platform (Aron et al. 2020). Consensus spectra were generated using the MS-Cluster algorithm with both precursor mass and fragment ion tolerance of 0.02 Da. The network was created with a minimum cosine score of 0.7, a Network TopK of 10, minimum 4 matched fragment ions, and minimum cluster size of 4. Library spectra search options for input data were applied with a score threshold of 0.7 and a minimum of 4 shared fragment ions to be considered as a match. Cytoscape 3.7.0 was used to visualize the network (Shannon et al. 2003). GNPS library spectra search was applied with the same parameters as the input data.
Antiplasmodial in vitro assay against Plasmodium facilparum parasites
Plasmodium falciparum 3D7 strain parasites (chloroquine sensitive) were cultured as previously described (Trager and Jensen 1976). Freshly sorbitol synchronized ring stages of the parasites (Lambros and Vanderberg 1979) were incubated with the samples at 50 μg mL−1 for non-purified fractions and a twofold serial dilution from 10 μM to 0.156 μM for pure compounds, previously solubilized in 0.05% DMSO. Sodium artesunate was used as antiplasmodial positive control. Each assay was performed in triplicate. The activity was measured using the SYBR green assay (Smilkstein et al. 2004). Briefly, the plates were centrifuged at 700 g for 5 min at room temperature to remove the culture medium, washed once with PBS, and incubated for 30 min with lysis buffer solution (20 mM Tris base, 5 mM EDTA, 0.0008% v/v Triton X-100, 0.008% w/v saponin, pH 7.5) and SYBR green DNA stain 0.002% v/v. Plates were incubated at room temperature for 30 min. The fluorescence of uninfected erythrocytes was considered as background. Fluorescence was measured on a SpectraMAX Gemini EM plate reader fluorimeter (485-nm excitation, 535-nm emission). Antiplasmodial activity is calculated relative to the parasite growth control (no compound added, 100% viability) and positive control (0% viability). Results are reported as percent inhibition for fractions and percent viability for compounds.
Cytotoxicity assay
Cancer cell line MCF-7 (breast adenocarcinoma) and normal breast epithelial cell line MCF-10A were seeded in complete medium containing DMEM/HAM’s F10 medium (1:1, v/v), supplemented with 10% fetal bovine serum and the antibiotic mixture penicillin/streptomycin (10 mL L−1) plus kanamycin sulfate (10 mg mL−1) in 96-well plates (5000 cells well−1) at 37 °C under 5% CO2 for 24 h to allow cell attachment. Fractions were added to the cell cultures at concentrations ranging from 0.25 to 250 μg mL−1. After 24 h of incubation, the cell plate was washed with PBS; then, a DMEM without phenol red plus XTT/electron solution was added to each well and incubated for 4 h, after which the absorbance was measured at 492 nm and 690 nm. Cell viability was directly proportional to absorbance based on the cleavage tetrazolium XTT by metabolically active cells forming an orange formazan dye, and was compared with the negative control. The inhibitory concentration (IC50) was calculated by a nonlinear regression analysis using GraphPad Prism (3.0).
Anti-leishmania in vitro assays
Anti-leishmania activity was evaluated in Leishmania (L.) infantum and Leishmania (V.) braziliensis promastigotes, as previously described (Parra et al. 2018). The anti parasitic activity was assessed using an MTT viability assay, after 24-h incubation with the fractions at concentrations of 50 μg mL−1 and 100 μg mL−1, and inhibition percentage values were calculated.
Antibacterial assay
The antibacterial activity was tested against Escherichia coli 25,922, Escherichia coli 35,218, Staphylococcus aureus 33,591, S. aureus 25,923, 700,603, Klebsiella pneumoniae id-146/19, Acinetobacter baumannii 19,606, Acinetobacter baumannii 261/16, Pseudomonas aeruginosa 27,853, Pseudomonas aeruginosa S.6065/06, Enterococcus faecalis 51,299, Enterococcus faecalis 29,212, and Enterobacter cloacae 003/21 (NDM +), and determined by the serial microdilution method in 96-well plates, accordingly with the Clinical and Laboratory Standards Institute protocols (CLSI 2018). Briefly, bacterial strains were initially cultivated in triptone soy agar (TSA) plates at 35 °C for 24 h. Subsequently, a standardized microbial suspension adjusted to 0.5 McFarland (1.5 × 108 CFU mL−1) scale was prepared in sterile saline solution and diluted 1:150 in cation-adjusted Mueller Hinton broth. Stock solutions of the compounds were prepared in DMSO at an initial concentration of 100 μM, from which twofold serial dilutions were prepared diluting with cation-adjusted Mueller Hinton broth in the range of 100 to 0.8 μM. Fifty microliters of the bacterial suspension was added to each well of the 96-well microtiter plate. The culture plates were incubated at 35 °C for 20 h. Optical density measurements were made at 630 nm with a microplate reader to obtain growth inhibition values. Vancomycin and polymyxin were used as positive controls. Cell viability was determined by MTT assay.
Results
Extraction and isolation of compounds 1–4
M. testaceum media were first adsorbed onto HP-20 and then extracted with EtOAc. Both extracts were analyzed by HPLC–MS and showed similar chemical profiles. The extracts were pooled and fractionated by C18 reversed-phase column chromatography to give five fractions. The fractions were screened for in vitro antiplasmodial activity against P. falciparum, anti-leishmanial activity against Leishmania infantum and L. braziliensis promastigotes, as well as cytotoxic activity against human cancer cell lines MCF-7 (breast cancer) and the non-cancerous breast cell line MCF-10A. The fraction eluted with 100% MeOH from the C18 chromatography column was active against P. falciparum with a 99% inhibition at 50 μg mL−1, moderately active at 100 μg mL−1 against L. infantum and L. braziliensis promastigotes with cell viability values of 44.6% and 29.6%, respectively, and displayed moderate cytotoxic activity against MCF-7 and MCF-10A cell lines, with IC50 values of 32.9 ± 2.2 μg mL−1 and 55.4 ± 2.8 μg mL−1, respectively.
The C18 column chromatography fractions were subjected to molecular networking (MN) analysis (Fig. 2a) (Aron et al. 2020). Our attention was directed to a cluster of ions with m/z 663.42, 677.44, and 691.46, present in the most active fraction (F5, Fig. 2b). Mass differences of 14 and 28 Da between the MN nodes indicated that these were homologous compounds. Automated comparison of ions m/z 663.42, 677.44, and 691.46 with spectral libraries at the GNPS platform did not annotate any known compound with our experimental data. Manual dereplication was then performed with the Dictionary of Natural Products database, resulting in no matches. These results directed the efforts towards the isolation of the compounds highlighted in the network, guided by MS analyses and leading to the isolation of testacosides A–D (1–4).
Molecular networking analysis of Microbacterium testaceum media extract and C18 column fractions. a Molecular network of all nodes present only in media extract and C18 fractions. Pie charts represent the distributions of the compounds in the fractions. b Cluster corresponding to the compounds present in the most active C18 fraction F5 (eluted with 100% MeOH)
Structural elucidation of testacosides A–D (1–4)
Testacoside A (1) was obtained as a clear colorless glass, with a molecular formula of C30H56O14 deduced by HRESIMS, which showed a [M + H]+ ion at m/z 641.3733 and a [M + Na]+ ion at m/z 663.3556 (Fig. S1), indicating 3 degrees of unsaturation. The 1H NMR spectrum of testacoside A (1) (Table 1, Fig. S2) revealed a group of resonances between δH 3.2 and 5.1, typical of oxymethine protons, and a series of aliphatic methylene and methyl proton resonances between δH 0.9 and 2.4, suggesting an aliphatic hydrocarbon fragment. Analysis of 13C NMR, DEPT-135 and HSQC spectra (Figs. S3-S5) indicated two sugar moieties based on typical anomeric signals at δC 102.4 (δH 4.74) and δC 102.2 (δH 5.13), in addition to thirteen oxygenated methine and methylene carbon resonances between δC 62.9 and 82.0. A signal at δC 175.7 was assigned to a carbonyl carbon, completing the unsaturation number calculated by the molecular formula. These NMR features indicated that testacoside A was a glycolipid.
Analysis of HSQC, COSY, and HMBC (Figs. S5-S7) spectra allowed the assignment of the planar structure of the sugar and aliphatic units (Fig. 3). Two spin systems were identified for the sugar residue 1, from H-1′ (δH 4.74) to H-3′ (δH 3.84), the second from H-5′ (δH 3.78) to H2-6′ (δH 4.40/4.23) and connected through C-4′ (δH 3.86, δC 67.7) by HMBC correlations 3JCH H-4′/C-6′ and H2-6′/C-4′, and 2JCH H-4′/C-5′ and H-5′/C-4′. In the case of sugar residue 2, spin systems from H-1″ (δH 5.13) to H-5″ (δH 3.87) and the other corresponding to methylene H2-6″ protons (δH 3.85/3.64) were connected by HMBC 2JCH H-5″/C-6″, and 3JCH H2-6″/C-4″ and H-4″/C-6″ correlations. The chemical shifts of protons H-5′ (δH 3.78) and H-5″ (δH 3.87), and long-range couplings 3JCH H-1′/C-5′, H-1″/C-5″ and H-5″/C-1″, confirmed both sugar residues as hexopyranoses. Correlations between H-3′ (δH 3.84)/C-1″ (δC 102.2) and H-1″ (δH 5.13)/C-3′ (δC 81.7) unambiguously confirmed the 1″ → 3′-glycosidic linkage between sugar residues.
Vicinal couplings between oxymethine H-2 (δH 3.80, δC 72.3) and diasterotopic oxymethylenes H2-1 (δH 3.77/3.42, δC 70.3) and H2-3 (δH 3.58/3.53, δC 64.4) indicated the presence of a glycerol unit, connected to the anomeric carbon C-1′ by observing 3JCH H-1′/C-1 and H2-1/C-1′ correlations. The structure of the acyl moiety of testacoside A (1) was established as follows. The carbonyl carbon C1‴ at δC 175.7 showed 2JCH and 3JCH correlations with methylenes at δH 2.36 and δH 1.63, respectively, confirming the presence of fatty acid chain which, in agreement with the un-assigned signals in the 13C NMR spectrum, consisted of 15 carbon atoms. The HSQC spectrum showed a triplet methyl group (δH 0.88, δC 11.9) and a doublet methyl group (δH 0.86, δC 19.8), typical of a branched anteiso chain, corresponding to the 12-methyltetradecanoid acid. Finally, correlation of diasterotopic protons’ H2-6′ with the ester carbonyl carbon C-1‴ confirmed the connection of the fatty acid moiety at C6′.
Analysis of vicinal 3JHH coupling constants of testacoside A (1) indicated large coupling constants of ca. 9.5 Hz between JH2″-H3″, JH3″-H4″, and JH4″-H5″, in addition to the JH1″-H2″ of 3.8 Hz, indicating that sugar 2 was an α-glucopyranose. The small coupling constant JH1′-H2′ of 1.6 Hz typical of an equatorial-equatorial arrangement established an α-anomeric configuration for the sugar moiety attached to the glycerol portion. However, the overlapping of 1H signals prevented the measurement of the other coupling constants, hampering to establish the relative configuration of this sugar residue. Then, testacoside A peracetate derivative 5 was prepared with the aim to resolve the 1H signals of the two sugar moieties (Table S1). The coupling constants JH2′-H3′ of 3.8 Hz, JH3′-H4′ of 9.7 Hz, and JH4′-H5′ of 10.0 Hz indicated an equatorial-axial, axial-axial, and axial-axial arrangement, respectively, which, in addition to the small coupling constant of the anomeric proton JH1′-H2′ of 1.5 Hz, typical of an equatorial-equatorial orientation, proved that the sugar residue connected to glycerol in 1 was α-mannopyranose. Analysis of NMR and HRESIMS data of testacoside A peracetate derivative 5 (Figs. S31-S37) confirmed the identification of 1 as 1-[α-glucopyranosyl-(1 → 3)-(6-O-acyl-α-mannopyranosyl)]-glycerol.
Analysis of the MS/MS data of 1 showed the fragment ions [M − glycerol]+ at m/z 549.3217, [M − glucose + H2O]+ at m/z 479.3220, [M − glucose]+ at m/z 461.3117, and [M − glucose − glycerol]+ at m/z 387.2766 (Fig. S8).
Analysis of NMR data of testacosides B (2) and C (3) (Table 1, Figs. S9-S23), as well as of their respective peracetylated derivatives 6 and 7 (Table S1, Figs. S38-S50), indicated almost the same structural features to those of testacoside A, with subtle differences in their acyl moieties. In the case of testacoside B (2), the HSQC spectrum (Fig. S12) disclosed a methyl doublet (δH 0.88, δC 23.8) characteristic of a branched iso fatty acid chain, indicating its structure as 14-methylpentadecanoic acid. As for testacoside C (3) (Fig. S20), a methyl triplet (δH 0.88, δC 11.9) and a methyl doublet (δH 0.86, δC 19.8) were observed, indicating an anteiso 14-mehtylhexadecanoic acid. Testacoside B (2) displayed a [M + H]+ ion at m/z 655.3882 and a [M + Na]+ ion at m/z 677.3707 (Fig. S14), consistent with the molecular formula C31H58O14 according to its HRMS analysis. Testacoside C (3) presented the molecular formula C32H60O14 based on its HRMS analysis, which showed a [M + H]+ ion at m/z 669.4041 and an [M + Na]+ ion at m/z 691.3864 (Fig. S22). These results confirmed testacosides B and C as analogs of testacoside A with one and two additional CH2 carbons, respectively, in their fatty acid chains. Such structures were validated by analysis of HRMS/MS data, which showed the same fragmentation pattern of testacoside A with + 14 Da for testacoside B (2) (Fig. S15) and + 28 Da for and testacoside C (3) (Fig. S23).
Testacoside D (4) presented the molecular formula C32H60O14, determined from its [M + Na]+ ion at m/z 691.3864, isomeric to testacoside C (3) (Fig. S24). Analysis of its NMR data (Table 2, Figs. S25-S29) and of its peracetylated derivative 8 (Table S2, Figs. S51-S56) indicated the same sugar, glycerol, and fatty acid units. The strong long-range correlation 3JCH H2-3/C-1‴ demonstrated that the fatty acid chain was connected to the C-3 position of the glycerol unit, instead of the C-6′ position of the mannose residue, establishing the structure of testacoside D (4) as 1-[α-glucopyranosyl-(1 → 3)-(α-mannopyranosyl)]-3-O-acylglycerol (Fig. 3). The chemical shift of the geminal protons H2-6′ (δH 3.84/3.73) indicated that C-6′ was not substituted. Analysis of the HRMS/MS spectrum confirmed the linkage of the fatty acid chain by means of the fragment ion [M − glucose − mannose]+ at m/z 345.3005 (Fig. S30).
Determination of the absolute configuration of sugar residues of 2 and 4
A recent method reported the determination of the absolute stereochemistry of sugar enantiomers by 1H NMR (Inagaki et al. 2021). In applying this procedure, acid hydrolysis of testacosides B (2) and D (4) yielded the free monosaccharides, which were subsequently reacted with L-cysteine methyl ester hydrochloride, directly in the NMR tube in pyridine-d5, to give their corresponding thiazolidine derivatives. The 1H-NMR measured chemical shifts and JHH coupling constants of the relevant enantio-dependent proton were consistent with those of the enantiomerically pure D-glucose and D-mannose thiazolidine derivatives (Fig. 4). The absolute configuration for testacosides A (1) and C (3) could not be established by this method because their corresponding thiazolidine derivatives’ sugar moieties degraded during the 1H NMR analysis, for no obvious reason. However, considering that NMR data of sugar moieties of 1 and 3 were practically identical to those of 2 and 4, as well as similar values of optical rotation were recorded, we propose the D-configuration for both sugar residues of 1 and 3.
Discussion
Testacosides A–C were inactive against P. falciparum at concentration of 10 μM, as well as when tested in antimicrobial activity against a panel of multi-resistant ESKAPE bacterial strains at concentration of 60 μM.
Testacosides A–D (1–4) are similar to glycoglycerolipids isolated from cultures of the gram-negative bacterium Flavobacterium marinotypicum (Yagi and Maruyama 1999) and from cultures of the sponge associated bacterium Microbacterium sp. (Wicke et al. 2000a, b). Reports of secondary metabolites produced by Microbacterium spp. are scarce and include the cytotoxic peptaibols microbacterins A and B produced in culture by the deep sea bacterium M. sediminis sp. nov. YLB-01(T) (Liu et al. 2015). The α- and γ-pyrone micropyrones A and B produced in culture by an endophytic Microbacterium sp. did not display antibacterial activity against S. aureus and methicillin-resistant S. aureus (Xu et al. 2021). Other studies reported the production of carotenoids by Microbacterium spp. (Reis-Mansur et al. 2019; Mandakovic et al. 2020).
Glycoglycerolipids related to testacosides have been isolated from cultures of a wide variety of bacteria obtained from different sources, such as the pathogen Rothia mucilaginosa (Pasciak et al. 2004), Bacillus pumilus (Ramm et al. 2004), Microccocus luteus (Pakkiri et al. 2004), and the marine-derived Bacillus licheniformis 09IDYM23 (Tareq et al. 2015). Unusual glycoglycerolipids diacylated at both 3 and 6′ positions have already been isolated from bacteria. Such are the cases of lutoside from the sponge-associated bacterium M. luteus (Bultel-Poncé, et al. 1995) and the polar glycolipid 1-[α-mannopyranosyl-(1 → 3)-(6-O-acyl-α-mannopyranosyl)]-3-O-acylglycerol from Arthrobacter atrocyaneus (Niepel, et al. 1997). Glycoglycerolipids were also reported from cyanobacteria (Shirahashi et al. 1993), algae (Morimoto, et al. 1995), lichens (Sassaki et al. 1999), and marine organisms (Cheng-Sánchez and Sarabia 2018).
The glycoglycerolipid di-O-12-methyl-tetradecanoyl-3- O-beta-d-galactopyranosyl-sn-glycerol has been previously reported from Microbacterium sp. M874 and has been related to avoid cell damage by radicalar oxygen and heat (Nakata 2000). Lipids comprising of 1-O-acyl-3-[α-glucopyranosyl-(1–3)-(6-O-acyl-α-mannopyranosyl)]glycerol connected to 14-methyl-hexadecanoic acid and 12-methyl-tetradecanoic acid moieties at C-6 of the mannose unit and glycerol have been isolated from cultures of Microbacterium sp. DSM 12583 obtained from the marine sponge Halichondria panicea (Wicke et al 2000a, b). Antiviral activity against herpes simplex virus types 1 and 2 have been reported for synthetic monoglycosyl diglycerides (Janwitayanuchit et al. 2003) as well as very mild antimicrobial activity (Cateni et al 2007, 2008). The chemical structure of glycoglycerolipids significantly impacts the biological activity and functions of these compounds, which are of biological and medical importance (Pagano et al. 2016). Glycoglycerolipids are some of the most abundant glycolipids in plants, animals, and bacteria, exerting essential roles in cell membranes and as chemical protecting agents against biological pathogens and physical damage (Jala et al. 2022). Thus, glycoglycerolipids, such as testacosides, are of considerable biological importance for maintenance of cell integrity under healthy and stressed conditions, being biotechnologically relevant agents such as biosurfactants, biomaterials, and biologically active compounds (Orive-Milla et al 2020).
In summary, a series of four new glycoglycerolipids were isolated from cultures produced by T. brasiliensis-associated bacterium M. testaceum (Figure S57). Dereplication with molecular networking along with biological activity results enabled us to direct the isolation of these compounds, demonstrating the feasibility of combining chemical and biological information at the early stages of the biodiscovery process. This is the first report of secondary metabolites produced in culture by a M. testaceum species.
Data availability
All data generated and analyzed are included in the supplementary information file.
References
Abdelmohsen UR, Bayer K, Hentschel U (2014) Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat Prod Rep 31:381–399. https://doi.org/10.1039/c3np70111e
Agarwal V, Blanton JM, Podell S, Taton A, Schorn MA, Busch J, Lin Z, Schmidt EW, Jensen PR, Paul VJ, Biggs JS, Golden JW, Allen EE, Moore BS (2017) Metagenomic discovery of polybrominated diphenyl ether biosynthesis by marine sponges. Nat Chem Biol 13:537–543. https://doi.org/10.1038/nchembio.2330
Almeida JF, Marques M, Oliveira V, Egas C, Mil-Homens D, Viana R, Cleary DFR, Huang YM, Fialho AM, Teixeira MC, Gomes NCM, Costa R, Keller-Costa T (2023) Marine sponge and octocoral-associated bacteria show versatile secondary metabolite biosynthesis potential and antimicrobial activities against human pathogens. Mar Drugs 21:34. https://doi.org/10.3390/md21010034
Aron AT, Gentry EC, McPhail KL, Nothias LF, Nothias-Esposito M, Bouslimani A, Petras D, Gauglitz JM, Sikora N, Vargas F, van der Hooft JJJ, Ernst M, Bin KK, Aceves CM, Caraballo-Rodríguez AM, Koester I, Weldon KC, Bertrand S, Roullier C, Sun K, Tehan RM, Boya PCA, Christian MH, Gutiérrez M, Ulloa AM, Tejeda-Mora JA, Mojica-Flores R, Lakey-Beitia J, Vásquez-Chaves V, Zhang Y, Calderón AI, Tayler N, Keyzers RA, Tugizimana F, Ndlovu N, Aksenov AA, Jarmusch AK, Schmid R, Truman AW, Bandeira N, Wang M, Dorrestein PC (2020) Reproducible molecular networking of untargeted mass spectrometry data using GNPS. Nat Protoc 15:1954–1991. https://doi.org/10.1038/s41596-020-0317-5
Balskus EP (2014) Sponge symbionts play defense. Nat Chem Biol 10:611–612. https://doi.org/10.1038/nchembio.1588
Bauermeister A, Mannochio-Russo H, Costa-Lotufo LV, Jarmusch AK, Dorrestein PC (2022) Mass spectrometry-based metabolomics in microbiome investigations. Nat Rev Microbiol 20:143–160. https://doi.org/10.1038/s41579-021-00621-9
Beniddir MA, Kang KB, Genta-Jouve G, Huber F, Rogers S, van der Hooft JJJ (2021) Advances in decomposing complex metabolite mixtures using substructure- and network-based computational metabolomics approaches. Nat Prod Rep 38:1967–1993. https://doi.org/10.1039/D1NP00023C
Berlinck RGS, Monteiro AF, Bertonha AF, Bernardi DI, Gubiani JR, Slivinski J, Michaliski LF, Tonon LAC, Venancio VA, Freire VF (2019) Approaches for the isolation and identification of hydrophilic, light-sensitive, volatile and minor natural products. Nat Prod Rep 36:981–1004. https://doi.org/10.1039/c9np00009g
Berlinck RGS, Crnkovic CM, Gubiani JR, Bernardi DI, Ióca LP, Quintana-Bulla JI (2022) The isolation of water-soluble natural products—challenges, strategies and perspectives. Nat Prod Rep 39:596–669. https://doi.org/10.1039/D1NP00037C
Blockley A, Elliott DR, Roberts AP, Sweet M (2017) Symbiotic microbes from marine invertebrates: driving a new era of natural product drug discovery. Diversity 9:49. https://doi.org/10.3390/d9040049
Brinkmann CM, Marker A, Kurtböke DI (2017) An overview on marine sponge-symbiotic bacteria as unexhausted sources for natural product discovery. Diversity 9:40. https://doi.org/10.3390/d9040040
Bultel-Poncé V, Debitus C, Blond A, Cerceau C, Guyot M (1997) Lutoside: an acyl-1-(acyl-6′-mannobiosyl)-3-glycerol isolated from the sponge-associated bacterium Micrococcus luteus. Tetrahedron Lett 38:5805–5808. https://doi.org/10.1016/S0040-4039(97)01283-5
Cateni F, Bonivento P, Procida G, Zacchigna M, Favretto LG, Scialino G, Banfi E (2007) Chemoenzymatic synthesis and antimicrobial activity evaluation of monoglucosyl diglycerides. Bioorg Med Chem 15:815–826. https://doi.org/10.1016/j.bmc.2006.10.045
Cateni F, Bonivento P, Procida G, Zacchigna M, Favretto LG, Scialino G, Banfi E (2008) Chemoenzymatic synthesis and antimicrobial activity evaluation of monogalactosyl diglycerides. Eur J Med Chem 43:210–221. https://doi.org/10.1016/j.ejmech.2007.03.012
Chambers MC, MacLean B, Burke R, Amode D, Ruderman DL, Neumann S, Gatto L, Fischer B, Pratt B, Egertson J, Hoff K, Kessner D, Tasman N, Shulman N, Frewen B, Baker TA, Brusniak M-Y, Paulse C, Creasy D, Flashner L, Kani K, Moulding C, Seymour SL, Nuwaysir LM, Lefebvre B, Kuhlmann F, Roark J, Rainer P, Detlev S, Hemenway T, Huhmer A, Langridge J, Connolly B, Chadick T, Holly K, Eckels J, Deutsch EW, Moritz RL, Katz JE, Agus DB, MacCoss M, Tabb DL, Mallick P (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30:918–920. https://doi.org/10.1038/nbt.2377
Cheng-Sánchez I, Sarabia F (2018) Chemistry and biology of bioactive glycolipids of marine origin. Mar Drugs 16:294. https://doi.org/10.3390/md16090294
CLSI (2018) Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. 11th ed. CLSI standard M07. Wayne, PA: Clinical and Laboratory Standards Institute.
Duan Y, Petzold M, Saleem-Batcha R, Teufel R (2020) Bacterial tropone natural products and derivatives: overview of their biosynthesis, bioactivities, ecological role and biotechnological potential. ChemBioChem 21:2384–2407. https://doi.org/10.1002/cbic.201900786
Esposito R, Ruocco N, Viel T, Federico S, Zupo V, Costantini M (2021) Sponges and their symbionts as a source of valuable compounds in cosmeceutical field. Mar Drugs 19:444. https://doi.org/10.3390/md19080444
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.2307/2408678
Flórez LV, Biedermann PHW, Engl T, Kaltenpoth M (2015) Defensive symbioses of animals with prokaryotic and eukaryotic microorganisms. Nat Prod Rep 32:904–936. https://doi.org/10.1039/C5NP00010F
Gaudêncio SP, Pereira F (2015) Dereplication: racing to speed up the natural products discovery process. Nat Prod Rep 32:779–810. https://doi.org/10.1039/C4NP00134F
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10:641–654. https://doi.org/10.1038/nrmicro2839
Holman JD, Tabb DL, Mallick P (2014) Employing ProteoWizard to convert raw mass spectrometry data. Curr Protoc Bioinformatics 46:13.24.1–13.24.9. https://doi.org/10.1002/0471250953.bi1324s46
Inagaki M, Iwakuma R, Kawakami S, Otsuka H, Rakotondraibe HL (2021) Detecting and differentiating monosaccharide enantiomers by 1H NMR spectroscopy. J Nat Prod 84:1863–1869. https://doi.org/10.1021/acs.jnatprod.0c01120
Ito T, Masubuchi M (2014) Dereplication of microbial extracts and related analytical technologies. J Antibiot (tokyo) 67:353–360. https://doi.org/10.1038/ja.2014.12
Jala RCR, Vudhgiri S, Kumar CG (2022) A comprehensive review on natural occurrence, synthesis and biological activities of glycolipids. Carbohyd Res 516:108556. https://doi.org/10.1016/j.carres.2022.108556
Janwitayanuchit W, Suwanborirux K, Patarapanich C, Pummangura S, Lipipun V, Vilaivan T (2003) Synthesis and anti-herpes simplex viral activity of monoglycosyl diglycerides. Phytochem 64:1253–1264. https://doi.org/10.1016/j.phytochem.2003.09.008
Karthikeyan A, Joseph A, Nair BG (2022) Promising bioactive compounds from the marine environment and their potential effects on various diseases. J Gen Engineer Biotechnol 20:14. https://doi.org/10.1186/s43141-021-00290-4
Kim O-S, Cho Y-J, Lee K, Yoon S-H, Kim M, Na H, Park S-C, Jeon YS, Lee J-H, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721. https://doi.org/10.1099/ijs.0.038075-0
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lambros C, Vanderberg JP (1979) Synchronization of Plasmodium falciparum erythrocytic stages in culture. J Parasitol 65:418–420
Laport MS (2017) Isolating bacteria from sponges: why and how? Curr Pharm Biotechnol 18:1224–1236. https://doi.org/10.2174/1389201019666180329111327
Liu D, Lin H, Proksch P, Tang X, Shao Z, Lin W (2015) Microbacterins A and B, new peptaibols from the deep sea actinomycete Microbacterium sediminis sp. nov. YLB-01(T). Org Lett 17:1220–1223. https://doi.org/10.1021/acs.orglett.5b00172
Mandakovic D, Cintolesi Á, Maldonado J, Mendoza SN, Aïte M, Gaete A, Saitua F, Allende M, Cambiazo V, Siegel A, Maass A, González M, Latorre M (2020) Genome-scale metabolic models of Microbacterium species isolated from a high altitude desert environment. Sci Rep 10:5560. https://doi.org/10.1038/s41598-020-62130-8
Morimoto T, Nagatsu A, Murakami N, Sakakibara J, Tokuda H, Nishino H, Iwashima A (1995) Anti-tumour-promoting glyceroglycolipids from the green alga, Chlorella vulgaris. Phytochemistry 40:1433–1437. https://doi.org/10.1016/0031-9422(95)00458-J
Nakashima Y, Egami Y, Kimura M, Wakimoto T, Abe I (2016) Metagenomic analysis of the sponge discodermia reveals the production of the cyanobacterial natural product kasumigamide by ‘Entotheonella.’ PLoS One 11
Nakata, K (2000) High resistance to oxygen radicals and heat is caused by a galactoglycerolipid in Microbacterium sp M874. J Biochem 127: 731–737. https://doi.org/10.1093/oxfordjournals.jbchem.a022664.
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Niepel T, Meyer H, Wray V, Abraham W (1997) A new type of glycolipid, 1-[α-mannopyranosyl-(1α-3)-(6-O-acyl-α-mannopyranosyl)]-3-O-acylglycerol, from Arthrobacter atrocyaneus. Tetrahedron 53:3593–3602. https://doi.org/10.1016/S0040-4020(97)00079-3
Orive-Milla N, Delmulle T, de Mey M, Faijes M, Planas A (2020) Metabolic engineering for glycoglycerolipids production in E. coli: tuning phosphatidic acid and UDP-glucose pathways. Metabol Engineer 61:106–119. https://doi.org/10.1016/j.ymben.2020.05.010
Pagano D, Cutignano A, Manzo E, Tinto F, Fontana A (2016) Glycolipids synthesis: improved hydrazinolysis conditions for preparation of 1,2-polyunsaturated fatty acyl-β-monogalactosyl-glycerols. Carbohyd Res 424:21–23. https://doi.org/10.1016/j.carres.2016.02.005
Pakkiri LS, Wolucka BA, Lubert EJ, Waechter CJ (2004) Structural and topological studies on the lipid-mediated assembly of a membrane-associated lipomannan in Micrococcus luteus. Glycobiology 14:73–81. https://doi.org/10.1093/glycob/cwh012
Parra LLL, Bertonha AF, Severo IRM, Aguiar ACC, De Souza GE, Oliva G, Guido RVC, Grazzia N, Costa TR, Miguel DC, Gadelha FR, Ferreira AG, Hajdu E, Romo D, Berlinck RGS (2018) Isolation, derivative synthesis, and structure-activity relationships of antiparasitic bromopyrrole alkaloids from the marine sponge Tedania brasiliensis. J Nat Prod 81:188–202. https://doi.org/10.1021/acs.jnatprod.7b00876
Paściak M, Holst O, Lindner B, Mierzchała M, Grzegorzewicz A, Mordarska H, Gamian A (2004) Structural and serological characterization of the major glycolipid from Rothia mucilaginosa. Biochim Biophys Acta - Gen Subj 1675:54–61. https://doi.org/10.1016/j.bbagen.2004.08.004
Piel J, Hui D, Wen G, Butzke D, Platzer M, Fusetani N, Matsunaga S (2004) Antitumor polyketide biosynthesis by an uncultivated bacterial symbiont of the marine sponge Theonella swinhoei. Proc Natl Acad Sci 101:16222–16227. https://doi.org/10.1073/pnas.0405976101
Quinn RA, Nothias LF, Vining O, Meehan M, Esquenazi E, Dorrestein PC (2017) Molecular networking as a drug discovery, drug metabolism, and precision medicine strategy. Trends Pharmacol Sci 38:143–154. https://doi.org/10.1016/j.tips.2016.10.011
Radjasa OK, Vaske YM, Navarro G, Vervoort HC, Tenney K, Linington RG, Crews P (2011) Highlights of marine invertebrate-derived biosynthetic products: their biomedical potential and possible production by microbial associants. Bioorganic Med Chem 19:6658–6674. https://doi.org/10.1016/j.bmc.2011.07.017
Ramm W, Schatton W, Wagner-Döbler I, Wray V, Nimtz M, Tokuda H, Enjyo F, Nishino H, Beil W, Heckmann R, Lurtz V, Lang S (2004) Diglucosyl-glycerolipids from the marine sponge-associated Bacillus pumilus strain AAS3: their production, enzymatic modification and properties. Appl Microbiol Biotechnol 64:497–504. https://doi.org/10.1007/s00253-003-1471-8
Reis-Mansur MCPP, Cardoso-Rurr JS, Silva JVMA, de Souza GR, Cardoso V da S, Mansoldo FRP, Pinheiro Y, Schultz J, Lopez Balottin LB, da Silva AJR, Lage C, dos Santos EP, Rosado AS, Vermelho AB (2019) Carotenoids from UV-resistant Antarctic Microbacterium sp. LEMMJ01. Sci Rep 9:9554. https://doi.org/10.1038/s41598-019-45840-6.
Romano S (2018) Ecology and biotechnological potential of bacteria belonging to the genus Pseudovibrio. Appl Environm Microbiol 84:e02516-e2517. https://doi.org/10.1128/AEM.02516-17
Rzhetsky A, Masatoshi N (1992) A simple method for estimating and testing minimum-evolution trees. Mol Biol Evol 9:945. https://doi.org/10.1093/oxfordjournals.molbev.a040771
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sassaki GL, Machado MJ, Tischer CA, Gorin PAJ, Iacomini M (1999) Glycosyldiacylglycerolipids from the lichen Dictyonema glabratum. J Nat Prod 62:844–847. https://doi.org/10.1021/np980547f
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Shirahashi H, Murakami N, Watanabe M, Nagatsu A, Sakakibara J, Tokuda H, Nishino H, Iwashima A (1993) Studies on glycolipids. Part VIII. Isolation and identification of anti-tumor-promoting principles from the fresh-water cyanobacterium Phormidium tenue. Chem Pharm Bull 41:1664–1666. https://doi.org/10.1248/cpb.41.1664
Smilkstein M, Sriwilaijaroen N, Kelly JX, Wilairat P, Riscoe M (2004) Simple and inexpensive fluorescence-based technique for high-throughput antimalarial drug screening. Antimicrob Agents Chemother 48:1803–1806. https://doi.org/10.1128/AAC.48.5.1803-1806.2004
Storey MA, Andreassend SK, Bracegirdle J, Brown A, Keyzers RA, Ackerley DF, Northcote PT, Owen JG (2020) Metagenomic exploration of the marine sponge Mycale hentscheli uncovers multiple polyketide-producing bacterial symbionts. mBio 11:e02997–19. https://doi.org/10.1128/mBio.02997-19
Tareq FS, Lee HS, Lee YJ, Lee JS, Shin HJ (2015) Ieodoglucomide C and ieodoglycolipid, new glycolipids from a marine-derived bacterium Bacillus licheniformis 09IDYM23. Lipids 50:513–519. https://doi.org/10.1007/s11745-015-4014-z
Tianero MD, Balaich JN, Donia MS (2019) Localized production of defence chemicals by intracellular symbionts of Haliclona sponges. Nat Microbiol 4:1149–1159. https://doi.org/10.1038/s41564-019-0415-8
Trager W, Jensen JB (1976) Human malaria parasites in continuous culture. Science (80-) 193:673–675. https://doi.org/10.1126/science.781840
van Soest RWM, Boury-Esnault N, Vacelet J, Dohrmann M, Erpenbeck D, de Voogd NJ, Santodomingo N, Vanhoorne B, Kelly M, Hooper JNA (2012) Global diversity of sponges (Porifera). PLoS One 7. https://doi.org/10.1371/journal.pone.0035105
Wang M, Carver JJ, Phelan VV, Sanchez LM, Garg N, Peng Y, Nguyen DD, Watrous J, Kapono CA, Luzzatto-Knaan T, Porto C, Bouslimani A, Melnik AV, Meehan MJ, Liu WT, Crüsemann M, Boudreau PD, Esquenazi E, Sandoval-Calderón M, Kersten RD, Pace LA, Quinn RA, Duncan KR, Hsu CC, Floros DJ, Gavilan RG, Kleigrewe K, Northen T, Dutton RJ, Parrot D, Carlson EE, Aigle B, Michelsen CF, Jelsbak L, Sohlenkamp C, Pevzner P, Edlund A, McLean J, Piel J, Murphy BT, Gerwick L, Liaw CC, Yang YL, Humpf HU, Maansson M, Keyzers RA, Sims AC, Johnson AR, Sidebottom AM, Sedio BE, Klitgaard A, Larson CB, Boya CAP, Torres-Mendoza D, Gonzalez DJ, Silva DB, Marques LM, Demarque DP, Pociute E, O’Neill EC, Briand E, Helfrich EJN, Granatosky EA, Glukhov E, Ryffel F, Houson H, Mohimani H, Kharbush JJ, Zeng Y, Vorholt JA, Kurita KL, Charusanti P, McPhail KL, Nielsen KF, Vuong L, Elfeki M, Traxler MF, Engene N, Koyama N, Vining OB, Baric R, Silva RR, Mascuch SJ, Tomasi S, Jenkins S, Macherla V, Hoffman T, Agarwal V, Williams PG, Dai J, Neupane R, Gurr J, Rodríguez AMC, Lamsa A, Zhang C, Dorrestein K, Duggan BM, Almaliti J, Allard PM, Phapale P, Nothias LF, Alexandrov T, Litaudon M, Wolfender JL, Kyle JE, Metz TO, Peryea T, Nguyen DT, VanLeer D, Shinn P, Jadhav A, Müller R, Waters KM, Shi W, Liu X, Zhang L, Knight R, Jensen PR, Palsson B, Pogliano K, Linington RG, Gutiérrez M, Lopes NP, Gerwick WH, Moore BS, Dorrestein PC, Bandeira N (2016) Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol 34:828–837. https://doi.org/10.1038/nbt.3597
Webster NS, Thomas T (2016) The Sponge Hologenome Mbio 7:e00135-e216. https://doi.org/10.1128/mBio.00135-16
Wicke C, Hüners M, Wray V, Nimtz M, Bilitewski U, Lang S (2000a) Production and structure elucidation of glycoglycerolipids from a marine sponge-associated Microbacterium species. J Nat Prod 63:621–626. https://doi.org/10.1021/np990313b
Wolfender J-L, Nuzillard J-M, van der Hooft JJJ, Renault J-H, Bertrand S (2019) Accelerating metabolite identification in natural product research: toward an ideal combination of liquid chromatography–high-resolution tandem mass spectrometry and NMR profiling, in silico databases, and chemometrics. Anal Chem 91:704–742. https://doi.org/10.1021/acs.analchem.8b05112
Xu Y-T, Luo Y-C, Xue J-H, Li Y-P, Dong L, Li W-J, Zhou Z-Y, Wei X-Y (2021) Micropyrones A and B, two new α -pyrones from the actinomycete Microbacterium sp. GJ312 isolated from Glycyrrhiza uralensis Fisch. Nat Prod Res 37:462–467. https://doi.org/10.1080/14786419.2021.1979546
Yagi H, Maruyama A (1999) A novel monoacyldiglycosyl−monoacylglycerol from Flavobacterium marinotypicum. J Nat Prod 62:631–632. https://doi.org/10.1021/np9804280
Wicke C, Huners H, Wray V, Nimtz M, Bilitewski U, Lang S (2000b) Production and structure elucidation of glycoglycerolipids from a marine sponge-associated Microbacterium species. J Nat Prod 63:621–626. https://doi.org/10.1021/np990313b
Funding
JIQB received a PhD Scholarship from FAPESP (Grant No. 2016/16033–3). LACT received a Postdoctoral Scholarship from FAPESP (Grant No. 2016/14375–4). LFM received a PhD Scholarship from FAPESP (Grant No. 2019/07894–3). RGSB received research funding grants from FAPESP (No. 2013/50228–8 and No. 2019/17721–9).
Fundação de Amparo à Pesquisa do Estado de São Paulo, 2016/16033-3, Jairo I. Quintana-Bulla, 2016/14375-4, Luciane A. C. Tonon, 2019/07894-3, Lamonielli F. Michaliski, 2013/50228-8, Roberto G. S. Berlinck, 2019/17721-9, Roberto G. S. Berlinck.
Author information
Authors and Affiliations
Contributions
JIQB and RGSB conceived and planned the experiments. JIQB conducted experiments. LACT and LFM contributed to taxonomical identification of bacterial strain. EH collected the sponge samples. AGF optimized and recorded NMR data. JIQB and RGSB wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Quintana-Bulla, J.I., Tonon, L.A.C., Michaliski, L.F. et al. Testacosides A–D, glycoglycerolipids produced by Microbacterium testaceum isolated from Tedania brasiliensis. Appl Microbiol Biotechnol 108, 112 (2024). https://doi.org/10.1007/s00253-023-12870-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12870-0