Abstract
Prenyltransferases (PTs) from the dimethylallyl tryptophan synthase (DMATS) superfamily are known as efficient biocatalysts and mainly catalyze regioselective Friedel-Crafts alkylation of tryptophan and tryptophan-containing cyclodipeptides (CDPs). They can also use other unnatural aromatic compounds as substrates and play therefore a pivotal role in increasing structural diversity and biological activities of a broad range of natural and unnatural products. In recent years, several prenylated dimeric CDPs have been identified with wide range of bioactivities. In this study, we demonstrate the production of prenylated dimeric CDPs by chemoenzymatic synthesis with a known promiscuous enzyme EchPT1, which uses cyclo-l-Trp-l-Ala as natural substrate for reverse C2-prenylation. High product yields were achieved with EchPT1 for C3-N1′ and C3-C3′ linked dimers of cyclo-l-Trp-l-Trp. Isolation and structural elucidation confirmed the product structures to be reversely C19/C19′-mono- and diprenylated cyclo-l-Trp-l-Trp dimers. Our study provides an additional example for increasing structural diversity by prenylation of complex substrates with known biosynthetic enzymes.
Key points
• Chemoenzymatic synthesis of prenylated cyclo-l-Trp-l-Trp dimers
• Same prenylation pattern and position for cyclodipeptides and their dimers.
• Indole prenyltransferases such as EchPT1 can be widely used as biocatalysts.
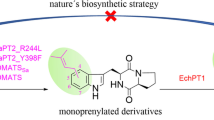
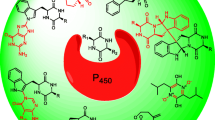
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Indole alkaloids derived from tryptophan-containing cyclodipeptides (CDPs) with a 2,5-diketopiperazine skeleton were isolated from various microorganisms and plants and are well-known for their structural diversity and pharmaceutical utility (Goetz et al. 2011; Li 2010; Lindel et al. 2012; Vinokurova et al. 2003; Xu et al. 2014). Among them, tryptophan-based dimeric diketopiperazine alkaloids have been identified in recent years and their biosynthetic pathways have been elucidated (Gerken and Walsh 2013; Harken and Li 2021; Kim and Movassaghi 2011; Kim and Movassaghi 2015). Typically, these natural products are biosynthesized from tryptophan and another amino acid such as tryptophan, proline, alanine, or valine. The two amino acids are mainly condensed by either a nonribosomal peptide synthetase in fungi or a cyclodipeptide synthase in bacteria, resulting in the formation of the two peptide bonds. The CDP core is then modified by cytochrome P450 to generate the homo- or heterodimer with unique bond connection (Gomes et al. 2019; Harken and Li 2021; Malit et al. 2021). Examples are the C6-N1′ linked aspergilazine A (1), the C3-C6′ linked naseseazine A (2), and tetratryptomycins A–C (3–5) with the symmetrical C3-C3′ and unsymmetrical C3-N1′ linkage (Fig. 1a) (Liu et al. 2020; Yu and Li 2019). The C3-C3′ linked (+)-WIN 64821 was isolated from Aspergillus versicolor and exhibits potential analgesic and anti-inflammatory activities by inhibition of substance P receptor (Fig. 1a) (Movassaghi et al. 2008; Tadano et al. 2013).
Recently, several tryptophan-containing dimeric CDPs carrying prenyl moieties were also isolated from different sources (Cai et al. 2019; Geng et al. 2017; Song et al. 2012). Interestingly, all these complex structures are C2-prenylated cyclo-l-Trp-l-Pro (cWP) or cyclo-l-Trp-l-Ala (cWA) dimers with different connections and symmetries. The cWP dimer brevianamide S exhibits selective antibacterial activity against Bacille Calmette-Guérin (BCG), which serves as a valuable lead for next-generation antitubercular drugs (Song et al. 2012). Another cWP dimer asperginulin A with an unprecedented 6/5/4/5/6 pentacyclic skeleton showed obvious toxicity in inhibiting settlement of the larvae of Balanus reticulatus (Cai et al. 2019). (+)/(−)-Uncarilin A, a pair of dimeric isoechinulin-type enantiomers with a symmetric four-membered core, was isolated from Uncaria rhynchophyl as a new type of melatonin receptors (Fig. 1b) (Geng et al. 2017). Attachment of prenyl moieties onto the indole nucleus usually increases the spectrum of the biological activities. However, in sharp contrast to the structural diversity of dimeric CDPs and CDP prenyltransferases (PTs), there are rare examples of prenylating enzymes toward CDP dimers.
The members of the dimethylallyl tryptophan synthase (DMATS) superfamily as important biocatalysts usually catalyze metal ion-independent Friedel-Crafts prenylations. They use predominantly tryptophan and other indole derivatives as prenyl acceptors but can also accept a broad spectrum of aromatic compounds for prenylation. They were therefore already used for structural modification of diverse small molecules (Fan et al. 2015). By using the fungal PT CdpC3PT and its mutants, Xu successfully obtained prenylated biflavonoids (Xu et al. 2021). In this study, we intended to get prenylated dimeric CDPs by these soluble PTs. Our previous studies demonstrated that the cyclo-l-Trp-l-Ala (cWA) C2-prenyltransferase EchPT1 is involved in the biosynthesis of echinulin (Fig. 2) (Wohlgemuth et al. 2017) and shows a high flexibility toward different substrates (Li et al. 2023; Wohlgemuth et al. 2018). Accordingly, we selected this enzyme and four additional DMATS PTs for prenylation of dimeric CDPs. One mono-and three diprenylated cyclo-l-Trp-l-Trp (cWW) dimers were obtained in high conversion yields.
Materials and methods
Chemicals
Dimethylallyl diphosphate (DMAPP) was chemically synthesized according to the method published previously (Woodside et al. 1988). Aspergilazine A (1), naseseazine A (2), and tetratryptomycins A–C (3–5) were isolated as described before (Liu et al. 2020; Yu and Li 2019).
Strains, plasmids, and culture conditions
Escherichia coli strain BL21 (DE3) pLysS (Invitrogen, Karlsruhe, Germany) and M15 (pREP4) (Qiagen, Hilden, Germany) were used for gene expression and cultivated at 37 °C in Terrific broth (TB) medium. The plasmids pVW90, pALF49, pPM37, pJW12, and pLW40 were used for overproduction of the proteins EchPT1, FgaPT2_R244L, FgaPT2_Y398F, 6-DMATSSa, and 7-DMATS, respectively (Fan and Li 2016; Kremer et al. 2007; Mai et al. 2016; Winkelblech and Li 2014; Wohlgemuth et al. 2017). To select the recombinant strains, ampicillin (50 μg/mL) and kanamycin (25 μg/mL) were added to the medium.
Escherichia coli ATCC 35218, Enterococcus faecalis DSM2570, Klebsiella pneumoniae DSM26371, Bacillus subtilis NCIB 3610, Bacillus circulans NRRL B-380, Staphylococcus aureus ATCC 29213, Staphylococcus delphini DSM20771, and Pseudomonas aeruginosa ATCC 27853 were used to evaluate the antibacterial activity.
Protein purification and enzyme assays
Recombinant EchPT1, FgaPT2_R244L, FgaPT2_Y398F, 6-DMATSSa, and 7-DMATS were purified by Ni-NTA affinity chromatography (Qiagen, Hilden) as described previously (Fan and Li 2016; Kremer et al. 2007; Mai et al. 2016; Winkelblech and Li 2014; Wohlgemuth et al. 2017). The purity of the five recombinant proteins was proven on 12% (w/v) SDS-PAGE (Li et al. 2023).
For enzyme reaction, standard assays (50 μL) contained Tris-HCl (50 mM, pH 7.5), CaCl2 (5 mM), dimeric CDP substrate (1 mM), DMAPP (1 mM), glycerol (0.5–5%, v/v), DMSO (2.5%, v/v), and the purified protein (7 μg). The reaction mixtures were incubated at 37 °C for 1 or 16 h and subsequently terminated with 50 μL MeOH. After centrifugation at 17,000 × g for 20 min, the enzyme reaction mixtures were analyzed on LCMS as described below.
Enzyme assays for product isolation were scaled up to a volume of 25 mL, containing Tris-HCl (50 mM, pH 7.5), CaCl2 (5 mM), DMAPP (1.5 mM), the respective dimeric CDP substrate (1 mM), and 5 mg purified EchPT1. The reaction mixtures were incubated at 37 °C for 16 h and extracted three times with two volumes of ethyl acetate each. The resulting organic phases were evaporated and dissolved in 1 ml MeOH for isolation.
The linearity of the EchPT1 reactions toward 3–5 was determined up to 360 min with 7 μg protein. To determine the kinetic parameters of EchPT1 toward the three cWW dimers, the enzyme assays (50 μL) contained Tris-HCl (50 mM, pH 7.5), CaCl2 (5 mM), DMAPP (1 mM), 7 μg EchPT1, and the cWW dimers at final concentrations of 0.01, 0.02, 0.05, 0.1, 0.2, 0.5, 1.0, and 2.0 mM. The reaction mixtures containing 3, 4, and 5 were incubated at 37 °C for 30, 30, and 20 min, respectively. For determination of the kinetic parameters of EchPT1 toward DMAPP in the presence of 3, 4, and 5, the reaction mixtures contained Tris-HCl (50 mM, pH 7.5), CaCl2 (5 mM), the respective dimeric CDP substrate (1 mM), 7 μg EchPT1, and DMAPP at final concentrations from 0.01 to 2.0 mM, which were incubated at 37 °C for 30, 30, and 20 min, respectively. The reactions were terminated with 50 μL MeOH and analyzed on HPLC as described below. All the assays were performed as duplicates. The conversion yields were calculated by comparing with the isolated products as standard or by the ratio of the peak areas in HPLC chromatograms. The data were analyzed by using Prism 8.01 (GraphPad Software).
LCMS and HPLC analysis of the enzyme products
LCMS analysis was performed as described previously (Zhou and Li 2021). The enzyme products were eluted at a flow rate of 0.25 mL/min with a linear gradient from 5 to 100% CH3CN in H2O in 10 min, followed by washing for 5 min and equilibration for 5 min. LCMS data were evaluated with DataAnalysis 4.2 software (Bruker Daltonik, Bremen, Germany).
For isolation of the target substances, semi-preparative HPLC was performed with an Agilent Eclipse XDB-C18 (250 × 9.4 mm, 5 μm) column. H2O (A) and CH3CN (B) were used as solvents at a flow rate of 2 mL/min. Compounds 3a2 and 5a2 were isolated with 80% B, 4a1 and 4a2 with 75% B. To determine the enzyme activities, an Agilent HPLC 1260 series equipped with a Multospher 120 RP-C18 (250 × 2 mm, 5 μm) column was used. H2O (A) and CH3CN (B) were used as mobile phase at a flow rate of 0.5 mL/min. The substances were eluted using a linear gradient from 5 to 100% B in A within 20 min.
Structural elucidation of the enzyme products by NMR analysis
NMR spectra were recorded on a 500 MHz Bruker AVIII spectrometer and processed with MestReNova 6.1.0 (Metrelab). All the samples were dissolved in DMSO-d6 for measurement. Chemical shifts were referred to those of the solvent signals at δH 2.50 ppm and δC 39.5 ppm. The NMR data are provided in Tables S1–S4 and spectra in Figs. S7–S26.
Antibacterial assays of the prenylated CDP dimers
The antibacterial activities of compounds 3a2, 4a1, 4a2, and 5a2 were evaluated by using agar disk-diffusion method as reported previously (Balouiri et al. 2016). The eight bacteria strains were spread onto LB agar medium. Filter paper disks of about 5 mm in diameter were placed on the agar surface and 5 μL of 2 mM DMSO solution of the test compounds were dropped onto the paper disks. The inhibition growth zones around the disks were observed after incubation at 37 °C for 16 h. Kanamycin was used as positive control and DMSO (5 μL) was used as a negative control. All assays were performed in duplicates.
Results
Acceptance of five dimeric CDPs by DMATS PTs with different activities
The five PTs EchPT1, FgaPT2_R244L, FgaPT2_Y398F, 6-DMATSSa, and 7-DMATS are responsible for the prenylation at C2, C4, C5, C6, and C7 of the indole ring of tryptophan and/or tryptophan-containing CDPs, respectively. To prove the acceptance of the tryptophan-containing dimeric CDPs by these enzymes, the recombinant proteins were incubated with cWP dimer apergilazine A (1) and cWA-cWP dimer naseseazine A (2) in the presence of DMAPP. LCMS analysis showed that formation of monoprenylated 1 with [M + H]+ ions at m/z 635.789 ± 0.005 and 2 at 607.735 ± 0.005 was only observed in the extracted ion chromatograms (EICs) of the reaction mixtures with EchPT1, FgaPT2_Y398F, and 7-DMATS (Figs. S1 and S2). Obviously, cWP-containing dimers are poor substrates of the tested PTs.
In comparison, cWW dimers like tetratryptomycins A–C (3–5) with the symmetrical C3-C3′ and the unsymmetrical C3-N1′ linkage were much better accepted, at least by two of the tested PTs. Incubation of 3–5 with the aforementioned five enzymes at 37 °C for 16 h and LCMS analysis showed that the C3-C3′ linked 3 and 5 were well consumed by EchPT1 with conversion yields for the sole products 3a2 and 5a2 at 33.6 ± 0.3 and 14.2 ± 0.2%, respectively. In the reaction mixture of the C3-N1′ linked 4 with EchPT1, the main product 4a1 with a conversion yield of 12.0 ± 0.2% was accompanied by the second product 4a2 with a conversion yield of 3.7 ± 0.3% (Fig. 3). Other four enzymes showed clearly lower catalytic activities toward 3–5 than EchPT1. FgaPT2_R244L showed a conversion yield of 8.8 ± 0.1% toward 4 and 6.5 ± 2.0% toward 5 for monoprenylated products. Almost no conversion of 3 and 4 was observed with FgaPT2_Y398F and 6-DMATSSa under the same conditions (Figs. S3–S5).
LCMS analysis of the acceptance of tetratryptomycins A–C (3–5) by EchPT1 for 1 and 16 h. UV absorptions at 280 nm are illustrated in black. The chromatograms depicted in blue and red refer to EICs of [M + H]+ of the monoprenylated at m/z 811.372 and those of the diprenylated products at m/z 879.434, respectively. A tolerance range of ± 0.005 was used for ion detection. All the assays were performed in duplicates. The conversion yields in percent are given in parenthesis after the product number as mean values. For better comparison, only the chromatograms between 8 and 16 min are illustrated
To detect monoprenylated products, we carried out incubations of 3–5 with EchPT1 in the presence of DMAPP for 1 h. As shown in Fig. 3, both monoprenylated (3a1, 4a1, and 5a1) with [M + H]+ ions at m/z 811.372 ± 0.005 and diprenylated products (3a2, 4a1, and 5a2) with [M + H]+ ions at m/z 879.434 ± 0.005 were clearly detected. Comparable product yields were calculated for the products of 3 and 5 in the range of 2.0–3.8%. The monoprenylated product 4a1 with a yield of 6.2 ± 0.2% was more accumulated in the reaction mixture of 4 than the diprenylated product 4a2 at 0.6 ± 0.1%. These results suggested that 3a1 and 5a1 were better accepted by EchPT1 than 4a1 for further prenylation.
To determine the relationship of mono- and diprenylated products in the reaction mixtures of the three cWW dimers with EchPT1, time dependence of their formation was determined. As shown in Fig. 4a, formation of 3a1 reached its maximum already in 5 min with a lower conversion yield than 3a2 and decreased slightly after that. In comparison, the formation of 3a2 increased continuously during the whole incubation process. Similar results were obtained for the formation of 5a1 and 5a2 with lower conversion yields than those for 3a1 and 3a2. However, the maximum conversion yield of 5a1 is higher than that of 5a2 in 5 min (Fig. 4c). The product yield of 4a1 reached its maximum after 30 min, approximately ten-folds of that of 4a2 and then decreased slightly, while the formation of 4a2 started at a lower level and kept steady increasing. The amount of 4a1 is about two-folds of that of 4a2 after incubation for 6 h (Fig. 4b). These results correspond well to the results after incubation for 16 h (Fig. 3) and proved again 3a1 and 5a1 are much better substrates for EchPT1 than 4a1.
Prenylation of cWW dimers by EchPT1 and structural elucidation of the prenylated derivatives
To verify the structures of the four prenylated cWW dimers 3a2, 4a1, 4a2, and 5a2, the assays of 3–5 and DMAPP were scaled up to 25 ml and incubated for 16 h. After extraction with ethyl acetate and isolation on HPLC, these products with UV absorption maxima at approximately 224 and 284 nm were obtained in high purity (Fig. S9). High-resolution mass spectrometric data proved again the monoprenylation in 4a1 by detection of the [M + H]+ ion at m/z 811.3713, which is 68 dalton larger than that of the substrate 4. In comparison, 3a2, 4a2, and 5a2 with [M + H]+ ions at m/z 879.434 ± 0.005 are diprenylated cWW dimers (Table S5).
The presence of the attached prenyl moieties in 3a2, 4a1, 4a2, and 5a2 was also confirmed by comparing their NMR data (Table S1–S4) with those of 3–5 (Liu et al. 2020). In the 1H NMR spectra, the signals for indole H-19 (3a2, 4a1, 4a2, and 5a2) and H-19′ (3a2, 4a2, and 5a2) disappeared. Instead, signals for one (4a1) or two (3a2, 4a2, and 5a2) reverse prenyl moieties can be observed by the characteristic chemical shifts and coupling patterns. A doublet of doublets with a chemical shift between 6.23 and 6.15 ppm was observed for H-28/H-28′. The coupling constants with the two protons at H-29/H-29′ also found as a doublet of doublets with a chemical shift around 5.11 to 5.00 ppm. The signals of the two methyl groups were detected at approximately 1.50 ppm (Figs. S7, S12, S17, and S22). These data indicate the attachment of the reverse prenyl moiety at C-19 (and C-19′) of the indole ring, which is also consistent with the EchPT1-catalyzed C2-prenylation at the indole ring for its natural substrate cWA.
In the 13C NMR spectra, the signals of C-19/C-19′ at the indole ring were observed at δc 141.4–141.5ppm (Figs. S8, S13, S18, and S23), which were comparable with the NMR data of the C2-prenylated CDPs in the literature (Li et al. 2023). Clear long-range correlations between H-28 (H-30/H-31) and C-19 (3a2, 4a1, 4a2, and 5a2) as well as H-28′ (H-30′/H-31′) and C-19′ (3a2, 4a2, and 5a2) were observed in the HMBC spectra, confirming the attachment of one or two reverse prenyl moieties at C-19 and C-19′ (Figs. S11, S16, S21, and S26). Taken together, their NMR data including 1H, 13C, 1H-1H COSY, HSQC, and HMBC proved unequivocally 4a1 as C19-prenylated tetratryptomycin B and 3a2, 4a2, and 5a2 as C19,C19′-diprenylated tetratryptomycins A, B, and C, respectively (Fig. 5). Prenylations at C-19/C-19′ do not change the stereochemistry of 3–5 and the absolute configurations of 3a2, 4a2, and 5a2 remain as those of their substrates (Liu et al. 2020).
Antibacterial activities of the obtained prenylated cWW dimers
After structural elucidation, the obtained prenylated products were screened for their inhibitory activities against eight bacterial strains. No inhibition was detected for the isolated compounds 3a2, 4a1, 4a2, and 5a2.
Determination of the kinetic parameters of EchPT1 toward tetratryptomycins and DMAPP
To further compare the catalytic efficiency of EchPT1 toward the dimeric derivatives 3–5 and DMAPP, kinetic parameters including Michaelis-Menten constants (KM) and turnover numbers (kcat) were determined for EchPT1. The most reactions followed the Michaelis-Menten kinetics, with the exception for 4a1 formation toward DMAPP and 4a2 formation toward 4 (Figs. S27–S30). Both of them fit well to a typical velocity equation with substrate inhibition (Figs. S28–S29). For the reactions of EchPT1 toward the prenyl acceptors 3–5, the highest KM at 0.25 mM was calculated for the formation of 4a1, significantly higher than that of the natural substrate cWA at 0.09 mM. Interestingly, the Michaelis-Menten constants for the formation of 3a2, 4a2, and 5a2 at 0.06, 0.01, and 0.05 mM, respectively, are somewhat lower than that of cWA (Table 1). The KM values of EchPT1 reaction toward DMAPP for the formation of the four products between 0.05 and 0.08 mM are also slightly lower than 0.18 mM in the presence of cWA. The turnover numbers from 0.002 to 0.14 s−1 and the kcat/KM values from 286 to 1286 s−1 M−1 were determined in the range of EchPT1 reactions toward most CDPs (Wohlgemuth et al. 2018). The lowest kcat/KM value of 4a2 at 25 s−1 M−1 is also in good consistence with the observed conversion yield depicted in Fig. 3.
Discussion
Prenyltransferases of the DMATS superfamily are soluble proteins and can be easily overproduced in Escherichia coli (Fan et al. 2015; Winkelblech et al. 2015). They show high flexibility toward aromatic prenyl acceptors and therefore contribute significantly to structural diversity of small molecules. Numerous studies in the last years have demonstrated that such PTs can be utilized as biocatalysts for the target structures (Chen et al. 2017; Mori et al. 2016; Ostertag et al. 2020), since they can efficiently prenylate various natural and unnatural substrates including indole, naphthalene, xanthone, flavonoid, and cyclodipeptide derivatives (Fan et al. 2015; Winkelblech et al. 2015). Previous investigations revealed that prenylated products often exhibit improved interactions with proteins and biological membranes compared with the original precursors (Botta et al. 2009; Wollinsky et al. 2012).
As mentioned in the introduction, EchPT1 as a member of the echinulin biosynthetic pathway was first identified in Aspergillus ruber and catalyzes the reverse C2-prenylation of cWA at the indole ring, followed by additional prenylations with EchPT2 as the second PT of the pathway (Wohlgemuth et al. 2017). As most members of the DMATS superfamily, this enzyme also shows high flexibility toward aromatic substrates and accepted other cyclodipeptides for C2-prenylation (Wohlgemuth et al. 2018). Our recent study revealed that this enzyme is much more promiscuous than reported before. It can even accept already prenylated CDPs as substrates and catalyze reverse C2-prenylation at the indole nucleus (Li et al. 2023), which differs clearly from the nature’s strategy with the first prenylation at C-2 of unprenylated CDPs. These results encouraged us to expand the substrate spectrum of EchPT1.
As proof of concept, our main objective in this study was chemoenzymatic synthesis of prenylated dimeric CDPs in vitro. To the best of our knowledge, there is no report on dimeric CDP prenylating enzymes in the literature prior to this study. Four new cWW dimers with C19/C19′-prenylations on the indole ring were successfully obtained by prenylation of tetratryptomycins A–C with the promiscuous cWA prenyltransferase EchPT1 in the presence of DMAPP. Our results demonstrated that even complex molecules can be modified by known enzymes. Therefore, for designed small molecule modification, it is worth to test other available biocatalysts. As presented in this study, cWP-containing dimers were poor substrates of the tested PTs. In this case, it would be interesting to test other available PTs or to get mutants of the known enzymes as demonstrated by Xu for prenylation of biflavonoids (Xu et al. 2021).
Data availability
All data generated during this study are included in this published article and its supplementary information file.
References
Balouiri M, Sadiki M, Ibnsouda SK (2016) Methods for in vitro evaluating antimicrobial activity: a review. J Pharm Anal 6:71–79
Botta B, Menendez P, Zappia G, de Lima RA, Torge R, Monachea GD (2009) Prenylated isoflavonoids: botanical distribution, structures, biological activities and biotechnological studies. An update (1995-2006). Curr Med Chem 16:3414–3468
Cai R, Jiang H, Xiao Z, Cao W, Yan T, Liu Z, Lin S, Long Y, She Z (2019) (-)- and (+)-Asperginulin A, a pair of indole diketopiperazine alkaloid dimers with a 6/5/4/5/6 pentacyclic skeleton from the mangrove endophytic fungus Aspergillus sp. SK-28. Org Lett 21:9633–9636
Chen R, Gao B, Liu X, Ruan F, Zhang Y, Lou J, Feng K, Wunsch C, Li S-M, Dai J, Sun F (2017) Molecular insights into the enzyme promiscuity of an aromatic prenyltransferase. Nat Chem Biol 13:226–234
Fan A, Li S-M (2016) Saturation mutagenesis on Arg244 of the tryptophan C4-prenyltransferase FgaPT2 leads to enhanced catalytic ability and different preferences for tryptophan-containing cyclic dipeptides. Appl Microbiol Biotechnol 100:5389–5399
Fan A, Winkelblech J, Li S-M (2015) Impacts and perspectives of prenyltransferases of the DMATS superfamily for use in biotechnology. Appl Microbiol Biotechnol 99:7399–7415
Geng CA, Huang XY, Ma YB, Hou B, Li TZ, Zhang XM, Chen JJ (2017) (+/-)-Uncarilins A and B, dimeric isoechinulin-type alkaloids from Uncaria rhynchophylla. J Nat Prod 80:959–964
Gerken T, Walsh CT (2013) Cloning and sequencing of the chaetocin biosynthetic gene cluster. Chembiochem 14:2256–2258
Goetz KE, Coyle CM, Cheng JZ, O'Connor SE, Panaccione DG (2011) Ergot cluster-encoded catalase is required for synthesis of chanoclavine-I in Aspergillus fumigatus. Curr Genet 57:201–211
Gomes NGM, Pereira RB, Andrade PB, Valentão P (2019) Double the chemistry, double the fun: Structural diversity and biological activity of marine-derived diketopiperazine dimers. Mar Drugs 17:551
Harken L, Li S-M (2021) Modifications of diketopiperazines assembled by cyclodipeptide synthases with cytochrome P450 enzymes. Appl Microbiol Biotechnol 105:2277–2285
Kim J, Movassaghi M (2011) Concise total synthesis and stereochemical revision of (+)-naseseazines A and B: regioselective arylative dimerization of diketopiperazine alkaloids. J Am Chem Soc 133:14940–14943
Kim J, Movassaghi M (2015) Biogenetically-inspired total synthesis of epidithiodiketopiperazines and related alkaloids. Acc Chem Res 48:1159–1171
Kremer A, Westrich L, Li S-M (2007) A 7-dimethylallyltryptophan synthase from Aspergillus fumigatus: overproduction, purification and biochemical characterization. Microbiology 153:3409–3416
Li S-M (2010) Prenylated indole derivatives from fungi: structure diversity, biological activities, biosynthesis and chemoenzymatic synthesis. Nat Prod Rep 27:57–78
Li W, Coby L, Zhou J, Li SM (2023) Diprenylated cyclodipeptide production by changing the prenylation sequence of the nature's synthetic machinery. Appl Microbiol Biotechnol 107:261–271
Lindel T, Marsch N, Adla SK (2012) Indole prenylation in alkaloid synthesis. Top Curr Chem 309:67–129
Liu J, Xie X, Li S-M (2020) Increasing cytochrome P450 enzyme diversity by identification of two distinct cyclodipeptide dimerases. Chem Commun 56:11042–11045
Mai P, Zocher G, Ludwig L, Stehle T, Li S-M (2016) Actions of tryptophan prenyltransferases toward fumiquinazolines and their potential application for the generation of prenylated derivatives by combining chemical and chemoenzymatic syntheses. Adv Synth Catal 358:1639–1653
Malit JJL, Liu W, Cheng A, Saha S, Liu LL, Qian PY (2021) Global genome mining reveals a cytochrome P450-catalyzed cyclization of crownlike cyclodipeptides with neuroprotective activity. Org Lett 23:6601–6605
Mori T, Zhang L, Awakawa T, Hoshino S, Okada M, Morita H, Abe I (2016) Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases. Nat Commun 7:10849
Movassaghi M, Schmidt MA, Ashenhurst JA (2008) Concise total synthesis of (+)-WIN 64821 and (-)-ditryptophenaline. Angew Chem Int Ed Engl 47:1485–1487
Ostertag E, Zheng L, Broger K, Stehle T, Li S-M, Zocher G (2020) Reprogramming substrate and catalytic promiscuity of tryptophan prenyltransferases. J Mol Biol 433:166726
Song F, Liu X, Guo H, Ren B, Chen C, Piggott AM, Yu K, Gao H, Wang Q, Liu M, Liu X, Dai H, Zhang L, Capon RJ (2012) Brevianamides with antitubercular potential from a marine-derived isolate of Aspergillus versicolor. Org Lett 14:4770–4773
Tadano S, Mukaeda Y, Ishikawa H (2013) Bio-inspired dimerization reaction of tryptophan derivatives in aqueous acidic media: three-step syntheses of (+)-WIN 64821, (-)-ditryptophenaline, and (+)-naseseazine B. Angew Chem Int Ed Engl 52:7990–7994
Vinokurova NG, Khmel'nitskaia II, Baskunov BP, Arinbasarov MU (2003) Occurrence of indole alkaloids among secondary metabolites of soil Aspergillus. Appl Biochem Microbiol 39:192–196
Winkelblech J, Fan A, Li S-M (2015) Prenyltransferases as key enzymes in primary and secondary metabolism. Appl Microbiol Biotechnol 99:7379–7397
Winkelblech J, Li S-M (2014) Biochemical investigations of two 6-DMATS enzymes from Streptomyces revealing novel features of L-tryptophan prenyltransferases. Chembiochem 15:1030–1039
Wohlgemuth V, Kindinger F, Li S-M (2018) Convenient synthetic approach for tri- and tetraprenylated cyclodipeptides by consecutive enzymatic prenylations. Appl Microbiol Biotechnol 102:2671–2681
Wohlgemuth V, Kindinger F, Xie X, Wang BG, Li S-M (2017) Two prenyltransferases govern a consecutive prenylation cascade in the biosynthesis of echinulin and neoechinulin. Org Lett 19:5928–5931
Wollinsky B, Ludwig L, Hamacher A, Yu X, Kassack MU, Li SM (2012) Prenylation at the indole ring leads to a significant increase of cytotoxicity of tryptophan-containing cyclic dipeptides. Bioorg Med Chem Lett 22:3866–3869
Woodside AB, Huang Z, Poulter CD (1988) Trisammonium geranyl diphosphate. Org Synth 66:211–215
Xu W, Gavia DJ, Tang Y (2014) Biosynthesis of fungal indole alkaloids. Nat Prod Rep 31:1474–1487
Xu Y, Li D, Tan G, Zhang Y, Li Z, Xu K, Li S-M, Yu X (2021) A single amino acid switch alters the prenyl donor specificity of a fungal aromatic prenyltransferase toward biflavonoids. Org Lett 23:497–502
Yu H, Li S-M (2019) Two cytochrome P450 enzymes from Streptomyces sp. NRRL S-1868 catalyze distinct dimerization of tryptophan-containing cyclodipeptides. Org Lett 21:7094–7098
Zhou J, Li S-M (2021) Conversion of viridicatic acid to crustosic acid by cytochrome P450 enzyme-catalysed hydroxylation and spontaneous cyclisation. Appl Microbiol Biotechnol 105:9181–9189
Funding
Open Access funding enabled and organized by Projekt DEAL. This project was financially funded by the DFG (INST 160/620-1 to S.-M. L.). Wen Li (201806220101), Jing Liu (201608310118), and Huili Yu (201306220024) are scholarship recipients from the China Scholarship Council.
Author information
Authors and Affiliations
Contributions
SL planned this project and WL carried out the experiments. XX did NMR analysis. JL and HY contributed to the substrate isolation. The manuscript was written through contributions of all authors. The authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, W., Xie, X., Liu, J. et al. Prenylation of dimeric cyclo-l-Trp-l-Trp by the promiscuous cyclo-l-Trp-l-Ala prenyltransferase EchPT1. Appl Microbiol Biotechnol 107, 6887–6895 (2023). https://doi.org/10.1007/s00253-023-12773-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12773-0