Abstract
Microorganisms produce extracellular polymeric substances (EPS, also known as exopolysaccharides) of diverse composition and structure. The biochemical and biophysical properties of these biopolymers enable a wide range of industrial applications. EPS from cyanobacteria are particularly versatile as they incorporate a larger number and variety of building blocks and adopt more complex structures than EPS from other organisms. However, the genetic makeup and regulation of EPS biosynthetic pathways in cyanobacteria are poorly understood. Here, we measured the effect of changing culture media on titre and composition of EPS released by Synechocystis sp. PCC 6803, and we integrated this information with transcriptomic data. Across all conditions, daily EPS productivity of individual cells was highest in the early growth phase, but the total amount of EPS obtained from the cultures was highest in the later growth phases due to accumulation. Lowering the magnesium concentration in the media enhanced per-cell productivity but the produced EPS had a lower total sugar content. Levels of individual monosaccharides correlated with specific culture media components, e.g. xylose with sulfur, glucose and N-acetyl-galactosamine with NaCl. Comparison with RNA sequencing data suggests a Wzy-dependent biosynthetic pathway and a protective role for xylose-rich EPS. This multi-level analysis offers a handle to link individual genes to the dynamic modulation of a complex biopolymer.
Key points
• Synechocystis exopolysaccharide amount and composition depends on culture condition
• Production rate and sugar content can be modulated by Mg and S respectively
• Wzy-dependent biosynthetic pathway and protective role proposed for xylose-rich EPS
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Extracellular polymeric substances (EPS, also referred to as exopolysaccharides) are complex glycans secreted by various microorganisms including bacteria, microalgae and fungi (Freitas et al. 2017). These structurally diverse, high molecular weight polymers serve many functions in nature including protection against environmental stresses (starvation, desiccation, radiation, predation), interactions (surface adhesion, colony/biofilm formation, symbiosis), motility and infectivity (Costa et al. 2018; Kehr and Dittmann 2015; Rossi and De Philippis 2015).
Cyanobacterial EPS are more complex than those produced by other organisms. Depending on species and condition, they contain up to 13 different sugars as well as different modifications (anionic, hydrophobic and peptidic moieties), branching and linkage types (De Philippis et al. 2001; Panoff et al. 1988; Pereira et al. 2009). This diversity in EPS building blocks translates into tremendous structural complexity and functional variation which has enabled cyanobacteria to colonise some of the most extreme habitats on the planet (Bhatnagar and Bhatnagar 2019; Rossi and De Philippis 2015). The anionic, sulphated, or amphipathic characteristics of cyanobacterial EPS also result in chelating, emulsifying, gelling and immuno-modulating properties thus enabling a wide range of applications in food and healthcare (pharmaceutical, nutraceutical, cosmetics) or in wastewater and construction industries (De Philippis and Vincenzini 1998; Pereira et al. 2019b).
Not only are cyanobacterial EPS promising products, but the photosynthetic cyanobacteria themselves are also promising production platforms as light-powered cell factories (Camsund and Lindblad 2014; Cassier-Chauvat et al. 2021; Santos-Merino et al. 2019). In terms of manufacturing EPS, the most cost effective and energy efficient EPS product to harvest are the released exopolymeric substances (RPS) secreted into the surrounding environment, which maintain no or only very loose linkage to the cell surface. RPS can either be readily harvested in bulk from batch cultures at the end of the growth period or regularly ‘milked’ from the culture media of continuous cultivation systems (Cruz et al. 2020; Zydney 2016). It is therefore convenient that the model cyanobacterium Synechocystis sp. PCC 6803, in which the greatest engineering efforts have been made so far, maintains high EPS production rates in the late growth phase and into stationary phase (Panoff et al. 1988). Furthermore, EPS influence cell flocculation properties and can be used to enhance cell/media separation with no effect on cell viability thus enabling retention of the total biomass for long-term EPS production (Allen et al. 2019; Sun et al. 2020).
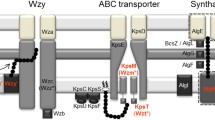
Three main pathways for EPS assembly and export have been found in bacteria and seem to be well conserved: Wzy-, ATP-binding cassette (ABC) transporter-, and synthase-dependent pathways (Pereira et al. 2009; Schmid 2018). EPS production starts with synthesis and activation of the sugar residues in the cytoplasm followed by assembly and polymerisation of sugar repeat units at the plasma membrane and final export across the outer membrane. Genes involved in EPS production can be categorised into three groups: 1) pathway genes for general sugar metabolism and not specific to EPS biosynthesis, 2) transferase genes for linkage of specific sugar residues, and 3) saccharide-processing genes for assembly, polymerisation, and export of EPS (Reeves et al. 1996). The modular nature of EPS biosynthesis is well suited to synthetic biology approaches: components of the pathways could be assembled and re-shuffled to adjust the types, order and frequency of sugars incorporated, as well as modifications, linkages, and branching, thus generating bespoke polymers with desirable properties (Pereira et al. 2019b). To develop this approach, it is paramount to understand the precise link between individual genes, enzymatic functions and secreted products.
Cyanobacterial genomes contain genes for all core functions of the EPS production machinery but, in contrast to the well-organised operons of other bacteria, EPS-related genes in cyanobacteria occur as multiple copies scattered throughout the genomes (Kehr and Dittmann 2015; Pereira et al. 2009; Pereira et al. 2015). Assignment of genes and assembly of complete pathways have therefore remained challenging. Initial investigations in the model cyanobacterium Synechocystis sp. PCC 6803 have confirmed the involvement of Wzy and ABC transporter pathways in EPS production (Fisher et al. 2013; Jittawuttipoka et al. 2013) and a biosynthetic pathway has been identified for the sulphated EPS synechan (Maeda et al. 2021). There is also good evidence that EPS production is regulated at the transcriptional level through the action of tyrosine and histidine kinase phosphorylation systems and alternative sigma factors such as SigF (Flores et al. 2019; Maeda et al. 2021; Pereira et al. 2019a).
EPS production often increases under stress. It is likely that, under conditions that limit biomass production, EPS synthesis provides a sink for excess electrons and photosynthetically assimilated carbon. For example, EPS production in several cyanobacterial strains was reported to increase under nutrient limitation (N, P, S, Mg, Ca) (De Philippis et al. 1991; Kharwar et al. 2021; Myklestad 1995) and oxidative stress (Hu et al. 2022), and in response to various abiotic factors including light quality and intensity, temperature, pH, salinity, and aeration (Bhatnagar and Bhatnagar 2019; Delattre et al. 2016). However, little is known whether environmental factors also alter the final composition of EPS. Monitoring the dynamic characteristics of the released polymer and integrating this information with changes in gene expression could potentially help to unravel the biosynthetic pathways. In this study, we therefore investigate how EPS change in response to different environmental conditions in the model cyanobacterium Synechocystis sp. PCC 6803. In the first part of the study, we evaluated factors that are of practical importance; we measured RPS production (titre and composition) in different growth phases of the cultures and we confirmed that they were not toxic for mammalian cells. In the second part of the study, we compared RPS production of cultures grown in different media (control, low S, low Mg, and 300 mM NaCl) and we integrated the data with previously obtained transcriptomics data for the same conditions. This multi-level analysis revealed novel correlations between environment, EPS sugar usage and genes, thus offering a basis for manipulating EPS composition in a model cyanobacterium.
Materials and methods
Culture conditions
Synechocystis sp. PCC 6803 was used in all experiments apart from the glucose experiments, which were carried out with the glucose-tolerant strain Synechocystis sp. PCC 6803-GT. Cultures were grown at 30°C with photoperiod 18-h/6-h light/dark, light intensity 120 ± 15 μmol photons/m2/s and sparged with humidified ambient air. 20 ml cultures were set up from glycerol stocks maintained at -80°C and gradually scaled up to 1.5 L cultures in Bijou bottles with a working volume of 60–75% of the bottle capacity. Cultures were grown in full BG11 medium (Stanier et al. 1971) for control conditions, in BG11 with 12.5% of the specified nutrients for low nutrient conditions (Madsen et al. 2021) or in BG11 with 300 mM NaCl or glucose added. For low light conditions, light intensity was reduced to 35 or 80 μmol photons/m2/s. Growth was monitored by measuring optical density at 730 nm (OD730) in a Lambda 45 UV/VIS Spectrophotometer (PerkinElmer, Waltham, MA, USA). To ensure measurements were performed in the linear range of the spectrophotometer, cultures were diluted in fresh media to OD <1, the sample OD was measured, and the culture OD was calculated by multiplying the sample OD with the respective dilution factor.
RPS harvest
EPS harvest was performed at various times during culture growth to reflect different growth phases: 1) “early” at the beginning of rapid growth phase after initial lag phase, 2) “late” at the end of the rapid growth phase when culture growth slows down due to emerging limitations, and 3) “stationary” when culture density no longer increased, or decreased. The harvest days to reflect these phases were decided for each culture based on preliminary growth curves and regular OD measurements during growth. ODs at harvest varied because growth differed between cultures and conditions. At the end of the experiment only those samples that could be assigned to one of the three growth phases were included in the analysis. Details on harvest days and ODs are provided in supplemental Table S1 and supplemental Figure S1.
For RPS harvest, supernatant of cultures was collected by centrifugation at 4000 g for 10 min at 4 °C. The supernatant was vacuum filtered through a 1.2 μm cellulose ester filter (Sigma-Aldrich, St. Louis, MO, USA) and subsequently dialysed in 8 kDa cellulose dialysis tubing (Thermo Fisher Scientific, Waltham, MA, USA) against ELGA water (1:10) for 48 h with five water changes and constant slow stirring. The dialysed product was lyophilised at -60 °C and below 20 mTor pressure using a VirTis sentry 2.0 freeze dryer (SP Industries, Warminster, PA, USA). Dry RPS samples were weighed, and this value was used to determine RPS production rates. Water-soluble RPS stocks were suspended at 10 mg/ml in ELGA water and insoluble precipitates were removed by centrifugation at 8000 g for 2 min at 21 °C.
RPS production analysis
The area under the growth curve (AUGC) was used to normalise RPS production titres to the total number of cells that were available for production up to the time point of harvest. It was calculated using Eq. 1:
where OD is the culture density as optical density at 730 nm, D is the culture age in days, and n represents day of OD measurement. This equation approximates the area under the growth curve with the sum of areas of rectangles where one side is time between OD measurements and the other side is the linear average of two consecutively measured ODs. In an alternative approach we tried to fit established growth models (e.g. Gompertz) to the measured ODs to calculate integrals. However, goodness of fits varied between cultures, and we therefore decided to use Equation 1 as the simplest approximation for all cultures based on the measured OD values.
RPS compositional analysis
Protein concentration was determined using a bicinchoninic acid (BCA) assay with bovine serum albumin (BSA) as reference (Thermo Fisher Scientific). Carbohydrate concentration was determined using the phenol-sulfuric acid method with glucose as reference (Dubois et al. 1956). Sulphate concentration was determined using the sodium rhodizonate assay with H2SO4 as reference (Terho and Hartiala 1971).
Monosaccharide composition was determined by methanolysis/tri-methylsilane derivatisation followed by quantitative analysis using a Shimadzu GC-2014 Gas Chromatograph equipped with a Flame-Ionisation Detector, Zebron ZB-5 MS column, Shimadzu FocusLiner (Shimadzu, Kyoto, Japan) and 300 °C splitless injection of 1 μl samples. References were included for each of the reported monosaccharides (Sigma-Aldrich).
High-performance liquid chromatography (HPLC) - size exclusion chromatography (SEC) was performed using an Alliance 2695 HPLC system equipped with a Shodex SB806M aqueous GFC column (Shodex) and refractive index detector (Waters, Milford, MA, USA) with dextran and heparin as references (Sigma-Aldrich). The mobile phase was 0.1 mM EDTA, 5 mM Tris pH 7, 0.09% NaCl at a flow rate of 0.5 ml/min.
RNA-sequencing
The transcriptomics experiment was previously published by our group (Madsen et al. 2021) and the sequencing dataset is available from the European Nucleotide Archive (PRJEB40560). In brief, mRNA was harvested from two growth phases (early and late) of Synechocystis spp. PCC 6803 cultures grown in control (BG11) and five nutrient limited conditions (12.5% N, P, K, Mg or S in BG11 background) in three biological replicates (independently grown cultures). Data are presented as fragments per kilobase of gene per million reads mapped (FPKM). Significant differences between conditions and time points were determined using Cuffdiff (Trapnell et al. 2012).
Statistical analyses
Statistical analysis of multiple comparisons was carried out in SigmaPlot software using one-way ANOVA with Tukey post hoc analysis, or Kruskal-Wallis One Way Analysis of Variance on Ranks with Dunn’s post-hoc analysis, a non-parametric test that does not require normal distribution. Pearson correlation analysis for multicomponent comparisons was performed in R software, using the cor.test function.
Results
Released extracellular polymeric substances (RPS) in different growth phases
RPS production rates in different growth phases
To compare RPS production at different stages of culture growth, RPS samples were harvested by dialysing and lyophilising the supernatant of batch cultures. A total of 54 samples were obtained during the “early” exponential, “late” transition to stationary, and “stationary” growth phases from cultures grown in a range of different media including BG11 (control), BG11 with low (12.5%) concentrations of N, K, P, Mg or S (see Madsen et al. 2021), and BG11 with NaCl or glucose added, as well as low light conditions. Information on all samples, ODs and harvest days is provided in Table S1. The variety of cultures used for this analysis was intentionally broad to extract consistent differences between growth phases. Figure 1 shows RPS production in the different growth phases. RPS titre (dry weight per culture volume; mg/L, Fig. 1a), daily productivity (titre normalised to culture age in days; mg/L/day, Fig. 1b) and RPS titre per cell (titre normalised to OD, mg/L/OD, Fig. 1c) were low in early growth, increased in late growth and remained high in the stationary phase. These measures indicate that synthesis of RPS is faster than degradation and therefore RPS accumulate in the culture media of batch cultures. However, the values obtained for daily RPS titre per cell (mg/L/OD/day, Fig. 1d) and for titre over area under the growth curve (mg/L/AUGC, Fig. 1e) show that the net daily RPS production rates per cell are highest in the early growth phase followed by significant decreases in later growth phase. These results indicate that Synechocystis cells are more productive during early growth compared to the later growth phases (for assignment into growth phases see Fig. 1f and supplemental Table S1). In other words, if the initial production rates were maintained over the entire growth period the final titre of RPS would be expected to be higher than the measured value. This has implications for larger scale production where continuous milking of a continuous culture in early exponential phase could potentially achieve higher cumulative yields than harvest from batch culture in stationary phase.
RPS production in different growth phases. RPS production is expressed as (A) dry weight per culture volume (titre), (B) titre per day, (C) titre per cell (using OD730 as a proxy for cell density), (D) titre per cell per day, and (E) titre per area under the growth curve (AUGC, see Eq. 1). RPS was harvested by dialysing and lyophilising the supernatant of Synechocystis sp. PCC 6803 cultures in three different growth phases: early growth (n=17), late growth (n=23) and stationary phase (n=14). Different letters indicate significant difference at adjusted p-values of p<0.005 (a,b) and p<0.05 (c) as determined by Kruskal-Wallis ANOVA with Dunn’s post hoc multiple comparison analysis. Harvest periods in a schematic growth curve are shown in (F). Cultures were grown in a range of conditions. Information on conditions, growth curves and harvest days for each culture are provided in Table S1
RPS composition in different growth phases
We next set out to compare the composition of water-soluble RPS between early and late Synechocystis sp. PCC 6803 cultures grown in control BG11 media. Water-soluble RPS were isolated from dry total RPS samples by resuspending in water (10 mg/ml) and removing insoluble particulate matter. EPS are complex polymers comprised of a sugar backbone decorated with various non-sugar components such as acidic and anionic side groups, proteins and DNA (De Philippis and Vincenzini 1998; Pereira et al. 2009). Of particular interest are sulphated EPS as they are associated with various bioactive and immunomodulatory properties (Raposo et al. 2013). Figure 2 shows that sulphate content (detected with a sodium rhodizonate assay) was higher in the early growth phase than the late growth phase with the latter being below the detection limit (<3.8% by weight). By contrast, protein content was low in the early growth phase and high in the late growth phase. EPS from early samples could not be detected by HPLC-SEC and generated poor GC-FID signals. Late samples generated complex HPLC-SEC profiles with EPS of varying sizes and good GC-FID signals as described in detail below. Combined, the ability to identify EPS in late but not early samples, despite starting with the same amount of dry total RPS material, suggests differences in solubility and/or amenity to acid hydrolysis and therefore differences in the content and/or structure of RPS produced during different stages of growth.
RPS composition in different growth phases. Sulphate and protein content of water-soluble RPS harvested during early and late growth of Synechocystis sp. PCC 6803 cultivated in control BG11 media. Sodium rhodizonate and BCA assays were used to determine sulphate and protein, respectively. Data are means ± S.E.M. of three independent cultures
Effect of cyanobacterial RPS on mammalian cells
For potential applications in health-related industries, it is important to assess whether cyanobacterial EPS samples are safe for use in mammalian cells. Figure 3 shows the metabolic activity (as ATP levels) of baby hamster kidney (BHK) cells treated with water-soluble RPS samples isolated from Synechocystis cultures grown in control conditions. RPS samples harvested during early or late growth phases had little impact on the metabolic activity of BHK cells. While relative ATP levels were slightly lower in cells treated with early RPS samples than in cells treated with late RPS samples, they were similar to ATP levels in cells treated with fucoidan, a sulphated polysaccharide with low cytotoxicity (Nagaoka et al. 2000), and significantly higher than in cells treated with doxorubicin hydrochloride (Dox), an anthracycline antibiotic with high cytotoxicity (Thorn et al. 2011). BHK cells treated with late RPS samples showed a slightly higher ATP level than those treated with fucoidan. In summary, the water-soluble RPS of Synechocystis sp. PCC 6803 is not toxic to mammalian cells independent of whether they are isolated from early or late-stage cultures.
Metabolic activity of mammalian cells treated with RPS. ATP levels (relative to the HBSS vector control) after 24 h treatment of mammalian baby hamster kidney (BHK) cells with water-soluble RPS harvested during early and late growth of Synechocystis sp. PCC 6803 cultivated in control BG11 media. Fucoidan serves as negative control showing low cytotoxicity. Doxorubicin hydrochloride (Dox) serves as positive control showing high cytotoxicity. Data are means ± S.E.M. of three technical replicates from two independent cyanobacterial cultures. Different letters indicate significant difference (p<0.05; one-way ANOVA with Tukey post hoc analysis)
RPS production in different environmental conditions
Cyanobacterial EPS production often increases under environmental stress (Delattre et al. 2016; Myklestad 1995). To identify nutritional conditions that may alter EPS production, we mined a previously obtained transcriptomics dataset of Synechocystis sp. PCC 6803 grown in different media (Madsen et al. 2021) for genes with published function in EPS biosynthesis. Three genes of a Wzy-dependent pathway (Jittawuttipoka et al. 2013) showed opposite responses during the transition to the late growth phase with up-regulation in low Mg (Fig. S1a) and down-regulation in low S (Fig. S1b). We therefore used cultures grown in BG11 with 12.5% of the original Mg or S concentration (low Mg, low S) for further EPS analyses. We also included samples grown with high salinity (BG11 with 300 mM NaCl added) as increased EPS production under salt stress and its requirement for salt tolerance had been reported by others (Ozturk and Aslim 2010) (Jittawuttipoka et al. 2013). Fig. S2 shows growth curves of the cultures and photographs of dry RPS samples from the different cultures. RPS samples were harvested in late exponential phase.
RPS production rates in different environmental conditions
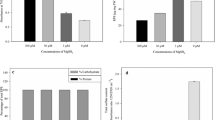
Figure 4 shows RPS production rates in the different culture media as titre normalised to time and/or culture density. Total culture productivity, measured as RPS titre (mg/L, Fig. 4a) and RPS titre normalised to culture age (mg/L/day, Fig. 4b), showed similar RPS production rates across the different conditions. “Per cell” productivity obtained from titres normalised to culture density either alone (mg/L/OD, Fig. 4c) or in combination with culture age (mg/L/OD/day, Fig. 4d) and from area under the growth curve (mg/L/AUGC, Fig. 4e), showed significant increases under nutrient limitation (low Mg and low S) compared to the control condition. Adding salt (300 mM NaCl) to the culture medium did not improve RPS production. In summary, nutrient limitation enhances the productivity of individual cells, but overall productivity of the culture remains unchanged due to lower culture density compared to control.
RPS production rates in different media. Total RPS of Synechocystis sp. PCC 6803 grown to the late growth phase in control (n=6), 12.5% Mg (n=4), 12.5% S (n=4) and 300 mM NaCl (n=4) in BG11 background. RPS production is expressed as (A) dry weight per culture volume (titre), (B) titre per day, (C) titre per cell (using OD730 as a proxy for cell density), (D) titre per cell per day, and (E) titre per area under the growth curve (AUGC, see Eq. 1). Different letters indicate significant difference (p<0.05; one-way ANOVA with Tukey post hoc analysis)
RPS composition in different environmental conditions
We next characterised the composition of the water-soluble fraction of late RPS samples harvested from the different conditions. Figure 5 shows that the RPS produced in control media were mostly comprised of sugar (55%) and protein (25%). Sulphur content was below the detection limit of the sodium rhodizonate assay (<3.8%). Sugar content decreased significantly when Mg levels in the culture media were low. Figure 6 shows that, of the monosaccharides detected by GC-FID, there were six “core” monosaccharides present in all conditions: glucose was the most common followed by mannose, fucose, xylose, rhamnose and N-acetyl-glucosamine. Also common were N-acetyl-galactosamine and galactose, detected in 93.8 and 87.5% of samples respectively. Glucuronic acid was detected in 25% of samples. Arabinose, iduronic acid and galacturonic acid were not detected in any of the samples tested. Correlation analysis of 19 variables including culture media components, RPS composition, and RPS production rates shows a very strong correlation between four of the core monosaccharides (mannose, fucose, xylose and rhamnose) and N-acetyl-galactosamine, which were consistently present at high levels when glucose was low (Table 1). Decreasing Mg levels in the culture media strongly enhances the productivity of individual cells (mg/L/OD, mg/L/OD/day, mg/L/AUGC), but the RPS had a lower sugar content, suggesting a tradeoff between RPS production rate and total sugar content. Low S correlated with low protein and low xylose contents. High NaCl in the media resulted in higher levels of glucose and lower levels of N-acetyl-galactosamine. In fact, N-acetyl-galactosamine was detected in all conditions except high NaCl. The HPLC-SEC refractive index chromatograms in Fig. 7 show that all conditions produced very high molecular weight RPS, evident as a peak at the resolution limit of the column (≥5 MDa, ~12.4 min) with a broad shoulder representing a mixture of molecules outside of the resolution of the dextran standard (>1.4 MDa, <16 min). All conditions also produced a mixture of smaller RPS molecules (<36 kDa, >18.5 min) evident as multiple peaks prior to the buffer peak (~22 min). In some samples, a distinct, symmetrical peak occurred at 15.7 min, suggesting an increase in either a single molecule or a uniform mixture of molecules. The HPLC profiles showed some consistent differences, e.g. presence of peaks in the 20-22 min range of low S or high NaCl samples compared to control, but these fractions as well as the larger RPS would require additional separation for deconvolution. Further analytical techniques will be needed to link culture conditions with specific polymer structures.
Protein and sugar content of RPS in different media. Protein (BCA assay, n=4) and sugar content (phenol-sulfuric assay, n=3) of water-soluble RPS from Synechocystis sp. PCC 6803 grown to the late growth phase in control, 12.5% Mg, 12.5% S and 300 mM NaCl in BG11 background. Data are means ± S.E.M. Different letters indicate significant difference (p<0.05; two-way ANOVA with Tukey post-hoc analysis)
Relative monosaccharide content of RPS in different media. Average monosaccharide levels (relative to total recovery, GC-FID) of water-soluble RPS from Synechocystis sp. PCC 6803 grown to the late growth phase in control (n=5), 12.5% Mg (n=4), 12.5% S (n=3), and 300 mM NaCl (n=4) in BG11 background. Monosaccharides are Glc glucose, Man mannose, Fuc fucose, Xyl xylose, Rha rhamnose, GlcNAc N-acetyl-glucosamine, Gal galactose, GalNAc N-acetyl-galactosamine, GlcA glucuronic acid
Molecular weight profiles of RPS in different media. HPLC-SEC refractive index chromatograms of water-soluble RPS from Synechocystis sp. PCC 6803 grown to the late growth phase in (A) control, (B) 12.5% Mg, (C) 12.5% S, and (D) 300 mM NaCl in BG11 background. Different colours represent independent cultures
Transcriptional analysis of RPS production
The modularity of EPS biosynthetic pathways offers opportunities for bioengineering bespoke polymers (Pereira et al. 2019b), however this requires detailed knowledge of the EPS production machinery and the genes that encode them. In Synechocystis sp. PCC 6803, a few genes have been identified which underpin the production of specific EPS products e.g. synechan (Maeda et al. 2021) or the incorporation of specific EPS components e.g. fucose (Mohamed et al. 2005). In an attempt to assign functions to EPS-related genes, we performed Pearson correlation analysis comparing different parameters of RPS production (environmental conditions, production rates, composition) to RNAseq data for EPS-related genes (average gene expression level) during the late growth phase (Madsen et al. 2021). A list of 436 genes with annotated functions in EPS biosynthesis or carbohydrate metabolism was compiled from previous publications (Fisher et al. 2013; Flores et al. 2019; Maeda et al. 2021; Pereira et al. 2015) (Lombard et al. 2014). Table S2 lists the genes with normalised transcript levels and p-values of all pairwise comparisons (Madsen et al. 2021). The correlation analysis identified 260 and 110 genes with strong correlations (p<0.05) to total sugar and xylose content, respectively. The gene lists (with associated Pearson correlation and transcriptomic data) can be found in Table S3 for total sugar and Table S4 for xylose content. Table 2 shows the number of genes of a particular EPS-related functional annotation category in each gene list. Interestingly, Pfam domains associated with Wzy-dependent pathways are generally negatively correlated with total sugar content (Wzb, Wzc, Wzx, Wzy) and positively correlated with xylose content (Wza, Wzc, Wzx, Wzy). This suggests that Wzy-dependent pathways may be important for the integration of specific monosaccharides into EPS. Furthermore, this analysis points towards a Wzy-dependent biosynthetic pathway for xylose-rich RPS, summarised in Fig. 8. The proposed pathway is comprised of 7 glycosyltransferases (sll1534, sll1566, sll5048, sll5050, slr1050, slr1166 and slr2120), one Wzx flippase (slr1543), 3 Wzy polymerases (sll0737, sll5047, slr0728), and one outer membrane complex comprised of Wzc (sll0923) and Wza (sll1581). Some of the potentially regulatory genes that correlated with xylose content have previously been reported to be involved in stress responses and included 6 genes encoding histidine kinase sensor and/or response regulators responding to acid (slr1759, slr1909) (Chen et al. 2016; Michel et al. 2009; Nodop et al. 2006), heavy metal (sll0649) (Chen et al. 2014b), butanol (slr1037) (Chen et al. 2014a) and Ci stress (slr0312) (Wang et al. 2004), as well as hybrid sensor and regulator sll5060 of unknown function (Xu and Wang 2019). In summary, the correlation analysis suggests 1) a Wzy-dependent biosynthesis and export pathway and 2) a protective role for xylose-rich RPS.
Proposed xylose rich RPS production pathway. Schematic diagram shows genes with strong correlation (p<0.05) to xylose content of water-soluble RPS from Synechocystis sp. PCC 6803. These genes encode EPS machinery for (1) assembly of repeat sugar units by glycosyltransferases, (2) transfer across the inner membrane by flippase Wzx, (3) polymerisation of repeat units by polymerase Wzy, and (4) transfer across the outer membrane by Wza/Wzc complex
Discussion
Extracellular polymeric substances are versatile bio-based polymers that can offer sustainable alternatives for many industries. The modular nature of EPS biosynthesis has great potential for the rational design of biomaterials with new and bespoke properties (Pereira et al. 2019b). However, plug-and-play approaches require a comprehensive understanding of the biosynthetic genes, machinery and pathways. While genes involved in cyanobacterial EPS production have been identified, functional understanding of the encoded proteins and organisation into biosynthetic pathways is still very limited (Fisher et al. 2013; Maeda et al. 2021; Pereira et al. 2015). Given that cyanobacteria adjust EPS amount and composition to protect from changing conditions (De Philippis and Vincenzini 1998), we characterised RPS produced by a single cyanobacterium in different growth phases and culture media and compared with transcriptomic data to begin unravelling the genes underpinning EPS production and its regulation. We identified culture media components that modulate both the production rate and composition of RPS as well as a potential biosynthetic pathway for xylose-rich RPS in Synechocystis sp. PCC 6803.
Responses to changes in growth phase
EPS production generally increases in the stationary phase (Delattre et al. 2016; Panoff et al. 1988). Indeed, we observed an increase in overall EPS production in the late growth and stationary phases of Synechocystis sp. PCC 6803 batch cultures (Fig. 1a). However, we found that the daily productivity of individual cells was in fact highest during the early growth phase (Fig. 1d, e). The decrease in cell productivity during the later growth phases was masked by the high cell density—a feature also observed in the efficient EPS producer Cyanothece sp. CCY 0110 where the total number of cells contributes more to overall productivity than the productivity of individual cells (Mota et al. 2013). Properties of RPS produced in the different growth phases also differed, evident in their amenability (or lack thereof) to different analytical techniques. Samples harvested during the early growth phase could not be detected by liquid or gas chromatography while late samples yielded strong signals, suggesting differences in RPS content, structure and/or physicochemical characteristics. Increasing culture volume, and therefore amount of dry RPS material harvested, could help to improve the resolution of early stage RPS products. While saccharide content could not be compared between growth phases in this study, analysis of sulphate content showed that sulphated RPS were only produced in the early growth phase (Fig. 2). Furthermore, we observed different levels of sulphation across one replicate of early RPS samples grown in different culture media (data not shown). In this study, however, we opted to focus on RPS harvested during the late growth phase given their amenability to more detailed compositional analysis and did not detect sulphate in any of the late samples examined. Sulphated RPS are associated with bioactive and immunomodulatory properties (Raposo et al. 2013), and we showed that the RPS of Synechocystis sp. PCC 6803 is not toxic to mammalian cells (Fig. 3). The RPS of Synechocystis sp. PCC 6803 is therefore a good candidate for the identification and production of new treatments for healthcare applications. A biosynthetic pathway for a sulphated RPS, synechan, was recently proposed for Synechocystis sp. PCC 6803 (Maeda et al. 2021). Comparison of sulphate content in early RPS samples with gene expression levels in different culture media would confirm the gene(s) underpinning sulphated RPS production and provide insights to the function and regulation of genes involved in the biosynthesis of synechan and other sulphated RPS. Protein content was found to be higher in the late growth phase than in the early growth phase (Fig. 2). This could be due to an increased N:C availability when N-supply in the media is still sufficient but C assimilation is limiting because of insufficient CO2 supply or shading (Kim et al. 2010).
Responses to changes in the environment
Culture media offer a cheap and simple solution to control product synthesis in microbial factories. EPS are particularly amenable to this approach given their role in forming a dynamic shield whose amount and composition adjusts to suit changing conditions (De Philippis and Vincenzini 1998). In this study, we investigated just three media components (Mg, S and NaCl) and showed that all three components could be used to modulate RPS production in Synechocystis sp. PCC 6803. Firstly, productivity of individual cells was significantly improved under nutrient limitation (Fig. 4cc, d, e), particularly low Mg, and is consistent with the general role of the anionic EPS to capture and store essential nutrients in nature (De Philippis et al. 1991; Delattre et al. 2016). While decreasing levels of individual nutrients boosted “per cell” productivity, the concomitant decrease in culture density negated any improvements in overall productivity, i.e. productivity of the total biomass (Fig. 4a, b). The limited nutrients were supplied at a very low level in this study (12.5% relative to the control medium) leading to a large decrease in culture density (e.g. 50% in low Mg; Fig. S2). Fine tuning nutrient levels may achieve high cell productivity at a relatively high culture density to improve overall productivity for industrial manufacturing processes. Several other nutrients are also reported to affect EPS production in cyanobacteria including P, Ca, and N and could be optimised for RPS production (Qian et al. 2022; Singh et al. 2016; Zevin et al. 2015). It is important to note, however, that altering nutrient levels can lead to significant changes in saccharidic composition, both in terms of total sugar content (e.g. by Mg; Fig. 5) and the amount of individual monosaccharides (e.g. by S and NaCl; Fig. 6) present in the EPS (Table 1). The next step will be to identify which polymers are the main contributors to the observed changes. Note that there is no evidence that Synechocystis produces cellulose (Nobles et al., 2001) and unlike some other cyanobacteria (Zhao et al. 2015) its genome does not contain the CesA gene for cellulose synthase. In addition to media composition, other conditions known to affect EPS production in cyanobacteria could offer additional options to increase production, including light (intensity, duration, wavelengths), temperature, aeration and/or agitation (Delattre et al. 2016; Mota et al. 2013; Soule et al. 2016). EPS composition is also influenced by abiotic factors, and it is therefore essential to ensure that the optimised cultivation conditions do not compromise product integrity for industrial manufacturing.
EPS production pathways
Biosynthetic pathways can be engineered to decouple product synthesis from other cellular processes, e.g. growth or stress response (Burg et al. 2016). Product integrity can thus be maintained irrespective of, e.g. culture media composition, by placing genes encoding EPS production machinery under the control of regulatory elements, i.e. promoters, that do not respond to changes in nutrient supply. Subsequent tweaking of expression levels of individual components can balance the flux within the EPS biosynthesis pathway, and wider metabolic engineering can channel resources/alleviate bottlenecks into this pathway to further improve productivity (Angermayr et al. 2015; Vijayakumar et al. 2020). Entirely new EPS biosynthetic pathways can be rationally designed by combining specific genes to determine EPS composition and structure thus creating bespoke polymers with selected properties (Pereira et al. 2019b). Alternatively, random combinations of sugars, modifications and linkages can be generated using gene shuffling approaches to generate new polymers and may in fact yield more genetically stable expression constructs for cyanobacteria (Taylor et al. 2021). However, pathway engineering requires detailed knowledge of EPS production machinery and its regulation. Cyanobacterial genes have been identified but functional annotation, biochemical characterisation, and assignment to biosynthetic pathways is limited (Fisher et al. 2013; Maeda et al. 2021; Pereira et al. 2015). In this study, we attempted to begin assigning specific functions to EPS-related genes by comparing RPS composition with transcriptomic data. While we did not identify genes underpinning specific EPS features, we did identify a candidate pathway for the production of RPS with high xylose content, summarised in Fig. 8. This may generate a product with repeating units of 7 monosaccharides assembled by 7 glycosyltransferases, one or more of which will presumably catalyse the transfer of “activated” xylose nucleotide monomers to the growing polysaccharide chain. This is similar to the 8 monosaccharide repeats assembled by 8 glycosyltransferases in synechan biosynthesis (Maeda et al. 2021). Repeat sugar units are then transferred across the inner membrane by a single flippase (Wzx), assembled into larger chains by up to three polymerases (Wzy) and exported by an outer membrane transport complex (Wza/Wzc). This pathway was originally characterized in E. coli (Whitfield and Paiment 2003) but has yet to be linked to specific polymeric products within cyanobacterial RPS. In Synechocystis 6803, this pathway includes 5 genes of unknown function (sll1534, slr0728, slr1050, slr1543, slr2120), including the Wzx flippase, which provide a good starting point for targeted knockout analyses.
To conclude; this study provides insights into how environmental conditions, notably culture media, can be used to control EPS production and composition in a cyanobacterium and further proposes a new EPS biosynthesis pathway. Characterisation of more EPS features in more environmental conditions, paired with “omics and knockout analyses,” will generate a more detailed picture of EPS biosynthesis, its regulation and the machinery involved. This knowledge base will improve industrial production and enable approaches using both random and carefully selected combinations of EPS production machinery to generate polymers with new and exciting properties for industry.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. The RNAseq data is deposited at ENA, https://www.ebi.ac.uk/ena under project accession number PRJEB40560.
References
Allen R, Rittmann BE, Curtiss R 3rd (2019) Axenic biofilm formation and aggregation by Synechocystis sp. strain PCC 6803 are induced by changes in nutrient concentration and require cell surface structures. Appl Environ Microbiol 85(7)
Angermayr SA, Gorchs Rovira A, Hellingwerf KJ (2015) Metabolic engineering of cyanobacteria for the synthesis of commodity products. Trends Biotechnol 33(6):352–361
Bhatnagar M, Bhatnagar A (2019) Diversity of polysaccharides in cyanobacteria. In: Satyanarayana T, Johri BN, Das SK (eds) Microbial Diversity in Ecosystem Sustainability and Biotechnological Applications: Volume 1. Microbial Diversity in Normal & Extreme Environments. Springer Singapore, Singapore, pp 447–496
Burg JM, Cooper CB, Ye Z, Reed BR, Moreb EA, Lynch MD (2016) Large-scale bioprocess competitiveness: the potential of dynamic metabolic control in two-stage fermentations. Curr Opin Chem Eng 14:121–136
Camsund D, Lindblad P (2014) Engineered transcriptional systems for cyanobacterial biotechnology. Front Bioeng Biotechnol 2:40
Cassier-Chauvat C, Blanc-Garin V, Chauvat F (2021) Genetic, genomics, and responses to stresses in cyanobacteria: biotechnological implications. Genes (Basel) 12(4)
Chen L, Ren Q, Wang J, Zhang W (2016) Slr1909, a Novel two-component response regulator involved in acid tolerance in Synechocystis sp. PCC 6803. Stress and Environmental Regulation of Gene Expression and Adaptation in Bacteria, pp 935–943
Chen L, Wu L, Wang J, Zhang W (2014a) Butanol tolerance regulated by a two-component response regulator Slr1037 in photosynthetic Synechocystis sp. PCC 6803. Biotechnol Biofuels 7(1):89
Chen L, Zhu Y, Song Z, Wang J, Zhang W (2014b) An orphan response regulator Sll0649 involved in cadmium tolerance and metal homeostasis in photosynthetic Synechocystis sp. PCC 6803. J Proteomics 103:87–102
Costa OYA, Raaijmakers JM, Kuramae EE (2018) Microbial extracellular polymeric substances: ecological function and impact on soil aggregation. Front Microbiol 9:1636
Cruz D, Vasconcelos V, Pierre G, Michaud P, Delattre C (2020) Exopolysaccharides from cyanobacteria: strategies for bioprocess development. Appl Sci 10(11):3763
De Philippis R, Sili C, Paperi R, Vincenzini M (2001) Exopolysaccharide-producing cyanobacteria and their possible exploitation: A review. J Appl Phycol 13:293–299
De Philippis R, Sili C, Tassinato G, Vincenzini M, Materassi R (1991) Effects of growth conditions on exopolysaccharide production by Cyanospira capsulata. Bioresour Technol 38:101–104
De Philippis R, Vincenzini M (1998) Exocellular polysaccharides from cyanobacteria and their possible applications. FEMS Microbiol Rev 22:151–175
Delattre C, Pierre G, Laroche C, Michaud P (2016) Production, extraction and characterization of microalgal and cyanobacterial exopolysaccharides. Biotechnol Adv 34(7):1159–1179
Dubois M, Guilles KA, Hamilton JK, Rebers PA, Smith F (1956) Colorimetric method for the determination of sugars and related substances. Anal Chem 28(3):350–356
Fisher ML, Allen R, Luo Y, Curtiss R 3rd. (2013) Export of extracellular polysaccharides modulates adherence of the cyanobacterium Synechocystis. PLoS One 8(9):e74514
Flores C, Santos M, Pereira SB, Mota R, Rossi F, De Philippis R, Couto N, Karunakaran E, Wright PC, Oliveira P, Tamagnini P (2019) The alternative sigma factor SigF is a key player in the control of secretion mechanisms in Synechocystis sp. PCC 6803. Environ Microbiol 21(1):343–359
Freitas F, Torres CAV, Reis MAM (2017) Engineering aspects of microbial exopolysaccharide production. Bioresour Technol 245(Pt B):1674–1683
Hu X, Luo K, Ji K, Wang L, Chen W (2022) ABC transporter slr0982 affects response of Synechocystis sp. PCC 6803 to oxidative stress caused by methyl viologen. Res Microbiol 173(1-2):103888
Jittawuttipoka T, Planchon M, Spalla O, Benzerara K, Guyot F, Cassier-Chauvat C, Chauvat F (2013) Multidisciplinary evidences that Synechocystis PCC6803 exopolysaccharides operate in cell sedimentation and protection against salt and metal stresses. PLoS One 8(2):e55564
Kehr JC, Dittmann E (2015) Biosynthesis and function of extracellular glycans in cyanobacteria. Life (Basel) 5(1):164–180
Kharwar S, Bhattacharjee S, Mishra AK (2021) Disentangling the impact of sulfur limitation on exopolysaccharide and functionality of Alr2882 by in silico approaches in Anabaena sp. PCC 7120. Appl Biochem Biotechnol 193(5):1447–1468
Kim HW, Vannela R, Zhou C, Harto C, Rittmann BE (2010) Photoautotrophic nutrient utilization and limitation during semi-continuous growth of Synechocystis sp. PCC6803. Biotechnol Bioeng 106:553–563
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42(Database issue):D490–D495
Madsen MA, Hamilton G, Herzyk P, Amtmann A (2021) Environmental regulation of PndbA600, an auto-inducible promoter for two-stage industrial biotechnology in cyanobacteria. Front Bioeng Biotechnol 8:619055
Maeda K, Okuda Y, Enomoto G, Watanabe S, Ikeuchi M (2021) Biosynthesis of a sulfated exopolysaccharide, synechan, and bloom formation in the model cyanobacterium Synechocystis sp. strain PCC 6803. Elife 10
Michel KP, Schroder AK, Zimmermann M, Brandt S, Pistorius EK, Frankenberg-Dinkel N, Staiger D (2009) The hybrid histidine kinase Slr1759 of the cyanobacterium Synechocystis sp. PCC 6803 contains FAD at its PAS domain. Arch Microbiol 191(6):553–559
Mohamed HE, van de Meene AM, Roberson RW, Vermaas WF (2005) Myxoxanthophyll is required for normal cell wall structure and thylakoid organization in the cyanobacterium Synechocystis sp. strain PCC 6803. J Bacteriol 187(20):6883–6892
Mota R, Guimarães R, Büttel Z, Rossi F, Colica G, Silva CJ, Santos C, Gales L, Zille A, De Philippis R, Pereira SB, Tamagnini P (2013) Production and characterization of extracellular carbohydrate polymer from Cyanothece sp. CCY 0110. Carbohydr Polym 92(2):1408–1415
Myklestad SM (1995) Release of extracellular products by phytoplankton with special emphasis on polysaccharides. Sci Total Environ 165(1):155–164
Nagaoka M, Shibata H, Kimura-Takagi I, Hashimoto S, Aiyama R, Ueyama S, Yokokura T (2000) Anti-ulcer effects and biological activities of polysaccharides from marine algae. Biofactors 12(1-4):267–274
Nobles DR, Romanovicz DK, Brown RM Jr (2001) Cellulose in cyanobacteria. Origin of vascular plant cellulose synthase? Plant Physiol 127:529–542
Nodop A, Suzuki I, Barsch A, Schroder AK, Niehaus K, Staiger D, Pistorius EK, Michel KP (2006) Physiological and molecular characterization of a Synechocystis sp. PCC 6803 mutant lacking histidine kinase Slr1759 and response regulator Slr1760. Z Naturforsch C. J Biosci 61(11-12):865–878
Ozturk S, Aslim B (2010) Modification of exopolysaccharide composition and production by three cyanobacterial isolates under salt stress. Environ Sci Pollut Res Int 17(3):595–602
Panoff J-M, Priem B, Morvan H, Joset F (1988) Sulphated exopolysaccharides produced by two unicellular strains of cyanobacteria, Synechocystis PCC 6803 and 6714. Arch Microbiol 150:558–563
Pereira S, Zille A, Micheletti E, Moradas-Ferreira P, De Philippis R, Tamagnini P (2009) Complexity of cyanobacterial exopolysaccharides: composition, structures, inducing factors and putative genes involved in their biosynthesis and assembly. FEMS Microbiol Rev 33(5):917–941
Pereira SB, Mota R, Vieira CP, Vieira J, Tamagnini P (2015) Phylum-wide analysis of genes/proteins related to the last steps of assembly and export of extracellular polymeric substances (EPS) in cyanobacteria. Sci Rep 5:14835
Pereira SB, Santos M, Leite JP, Flores C, Eisfeld C, Buttel Z, Mota R, Rossi F, De Philippis R, Gales L, Morais-Cabral JH, Tamagnini P (2019a) The role of the tyrosine kinase Wzc (Sll0923) and the phosphatase Wzb (Slr0328) in the production of extracellular polymeric substances (EPS) by Synechocystis PCC 6803. Microbiologyopen 8(6):e00753
Pereira SB, Sousa A, Santos M, Araujo M, Serodio F, Granja P, Tamagnini P (2019b) Strategies to obtain designer polymers based on cyanobacterial extracellular polymeric substances (EPS). Int J Mol Sci 20(22)
Qian L, Ye X, Xiao J, Lin S, Wang H, Liu Z, Ma Y, Yang L, Zhang Z, Wu L (2022) Nitrogen concentration acting as an environmental signal regulates cyanobacterial EPS excretion. Chemosphere 291(Pt 2):132878
Raposo MF, de Morais RM, Bernardo de Morais AM (2013) Bioactivity and applications of sulphated polysaccharides from marine microalgae. Mar Drugs 11(1):233–252
Reeves PR, Hobbs M, Valvano MA, Skurnik M, Whitfield C, Coplin D, Kido N, Klena J, Maskell D, Raetz CR, Rick PD (1996) Bacterial polysaccharide synthesis and gene nomenclature. Trends Microbiol 4(12):495–503
Rossi F, De Philippis R (2015) Role of cyanobacterial exopolysaccharides in phototrophic biofilms and in complex microbial mats. Life (Basel) 5(2):1218–1238
Santos-Merino M, Singh AK, Ducat DC (2019) New Applications of Synthetic Biology Tools for Cyanobacterial Metabolic Engineering. Front Bioeng Biotechnol 7:33
Schmid J (2018) Recent insights in microbial exopolysaccharide biosynthesis and engineering strategies. Curr Opin Biotechnol 53:130–136
Singh S, Verma E, Niveshika, Tiwari B, Mishra AK (2016) Exopolysaccharide production in Anabaena sp. PCC 7120 under different CaCl2 regimes. Physiol Mol Biol Plants 22(4):557–566
Soule T, Shipe D, Lothamer J (2016) Extracellular polysaccharide production in a scytonemin-deficient mutant of nostoc punctiforme under UVA and oxidative stress. Curr Microbiol 73(4):455–462
Stanier RY, Kunisawa R, Mandel M, Cohen-Bazire G (1971) Purification and properties of unicellular blue-green algae (order Chroococcales). Bacteriol Rev 35(2):171–205
Sun F, Zhang H, Qian A, Yu H, Xu C, Pan R, Shi Y (2020) The influence of extracellular polymeric substances on the coagulation process of cyanobacteria. Sci Total Environ 720:137573
Taylor GM, Hitchcock A, Heap JT (2021) Combinatorial assembly platform enabling engineering of genetically stable metabolic pathways in cyanobacteria. Nucleic Acids Res 49(21):e123–e123
Terho TT, Hartiala K (1971) Method for determination of the sulfate content of glycosaminoglycans. Anal Biochem 41(2):471–476
Thorn CF, Oshiro C, Marsh S, Hernandez-Boussard T, McLeod H, Klein TE, Altman RB (2011) Doxorubicin pathways: pharmacodynamics and adverse effects. Pharmacogenet Genomics 21(7):440–446
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578
Vijayakumar S, Rahman PKSM, Angione C (2020) A hybrid flux balance analysis and machine learning pipeline elucidates metabolic adaptation in cyanobacteria. iScience 23(12):101818
Wang H-L, Postier BL, Burnap RL (2004) Alterations in global patterns of gene expression in Synechocystis sp. PCC 6803 in response to inorganic carbon limitation and the inactivation of ndhR, a LysR Family Regulator*. J Biol Chem 279(7):5739–5751
Whitfield C, Paiment A (2003) Biosynthesis and assembly of Group 1 capsular polysaccharides in Escherichia coli and related extracellular polysaccharides in other bacteria. Carbohydr Res 338(23):2491–2502
Xu W, Wang Y (2019) Sequences, domain architectures, and biological functions of the serine/threonine and histidine kinases in Synechocystis sp. PCC 6803. Appl Biochem Biotechnol 188(4):1022–1065
Zevin AS, Nam T, Rittmann B, Krajmalnik-Brown R (2015) Effects of phosphate limitation on soluble microbial products and microbial community structure in semi-continuous Synechocystis-based photobioreactors. Biotechnol Bioeng 112(9):1761–1769
Zhao C, Li Z, Li T, Zhang Y, Bryant DA, Zhao J (2015) High-yield production of extracellular type-I cellulose by the cyanobacterium Synechococcus sp. PCC 7002. Cell Discov 1:15004
Zydney AL (2016) Continuous downstream processing for high value biological products: a review. Biotechnol Bioeng 113(3):465–475
Acknowledgements
We thank Craig Carr, University of Glasgow, for technical support and we are grateful to Dr. Pawel Herzyk, University of Glasgow, for helping with mathematics during revision.
Funding
This work was funded by the Biotechnology and Biological Sciences Research Council (BB/R505195/1, IBCarb-BIV-0316 and BB/R019894/1) and by Glycomar Ltd (in conjunction with BIV-0316).
Author information
Authors and Affiliations
Contributions
AA, CDB, CM, and MAM conceived and designed the research. MAM and SS conducted experiments. CDB and CM supervised and contributed to the EPS compositional experiments. Data analysis was carried out by MAM, AA, and JDT with contributions from SS and CM. MAM wrote the manuscript with support from AA. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplemental information
ESM 1
Fig. S1 RNA levels of selected genes in cultures grown in different growth media (PPTX 47 kb)
ESM 2
Fig. S2 Growth and RPS extracts of Synechocystis sp. PCC 6803 in different media (PPTX 212 kb)
ESM 3
Table S1 Growth curves and RPS harvest days of Synechocystis cultures included in Figure 1. (XLSX 171 kb)
ESM 4
Table S2 EPS-related genes of Synechocystis sp. PCC 6803 (XLSX 420 kb)
ESM 5
Table S3 Genes with strong correlation to total sugar content of water-soluble RPS produced during the late growth phase by Synechocystis sp. PCC 6803 (XLSX 279 kb)
ESM 6
Table S4 Genes with strong correlation to xylose content of water-soluble RPS produced during the late growth phase by Synechocystis sp. PCC 6803 (XLSX 132 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Madsen, M.A., Semerdzhiev, S., Twigg, J.D. et al. Environmental modulation of exopolysaccharide production in the cyanobacterium Synechocystis 6803. Appl Microbiol Biotechnol 107, 6121–6134 (2023). https://doi.org/10.1007/s00253-023-12697-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12697-9