Abstract
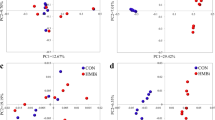
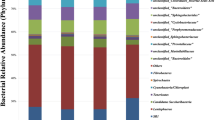
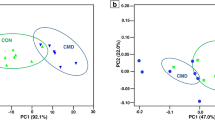
This study aimed to evaluate the effects of partial reducing rumen-protected Lys (RPLys) on rumen fermentation and microbial composition in heifers. Three ruminal fistulated Holstein Friesian bulls were used to determine the effective degradability of RPLys using an in situ method at incubation times of 0, 2, 6, 12, 16, 24, 36, and 48 h. Thereafter, 36 Holstein heifers at 90 days of age were assigned to one of two dietary treatments: a theoretically balanced amino acid diet (PC group; 1.21% Lys, 0.4% Met) or a 30% Lys-reduced diet (PCLys group, 0.85% Lys, 0.4% Met). Rumen fluid samples from five heifers in each group were extracted using esophageal tubing on day 90 to determine pH, microprotein, ammonia, volatile fatty acids, and microbial communities. Results showed that the effective ruminal degradability was 25.76%. Furthermore, differences in rumen fermentation parameters and alpha diversity of the microbiota between the two groups were not significant, but beta diversity was significant. Based upon relative abundance analysis, short-chain fatty acid–producing bacteria, including Sharpea, Syntrophococcus, [Ruminococcus]_gauvreauii_group, Acetitomaculum, and [Eubacterium]_nadotum_group belonging to Firmicutes, were significantly decreased in the PCLys group. Spearman’s analysis revealed a positive correlation between the butyrate molar proportion and the relative abundance of butyrate-producing bacteria such as [Eubacterium]_nadotum_group, Coprococcus_1, Ruminococcaceae_UCG_013, Pseudoramibacter, and Lachnospiraceae_UCG_010. Phylogenetic Investigation of Communities by Reconstruction of Unobserved States analysis further validated that RPLys deduction influenced energy metabolism. Together, our findings highlight the role of RPLys or Lys in butyrate-producing bacteria. However, the number of bacteria affected by Lys was very limited and insufficient to alter rumen fermentation.
Key Points • Reducing 30% Lys via rumen-protected Lys did not affect rumen fermentation parameters and alpha diversity of microbiota of Holstein heifers. It meant that the ruminal fermentation pattern was not changed. • Reducing 30% Lys via rumen-protected lysine significantly decreased relative abundance of short-chain fatty acid–producing bacteria belonging to Firmicutes. • Functions of microorganisms were changed by reducing 30% Lys via rumen-protected Lys, especially amino acid metabolism. It may affect the amino acid composition of microprotein. |






Similar content being viewed by others
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Abe M, Iriki T, Funaba M (1997) Lysine deficiency in postweaned calves fed corn and corn gluten meal diets. J Anim Sci 75:1974–1984
Abe M, Iriki T, Funaba M, Onda S (1998) Limiting amino acids for a corn and soybean meal diet in weaned calves less than three months of age. J Anim Sci 76:628–636
AOAC (2012) Official methods of analysis of the association of offical analytical chemists, 19th edn. AOAC International, Gaithersburg
Ayudthaya SPN, van de Weijer AHP, van Gelder AH, Stams AJM, de Vos WM, Plugge CM (2018) Organic acid production from potato starch waste fermentation by rumen microbial communities from Dutch and Thai dairy cows. Biotechnol Biofuels 11:13
Barros T, Quaassdorff MA, Aguerre MA, Colmenero JJO, Wattiaux MA (2017) Effects of dietary crude protein concentration on late-lactation dairy cow performance and indicators of nitrogen utilization. J Dairy Sci 100:5434–5448
Bi Y, Zeng S, Zhang R, Diao Q, Tu Y (2018) Effects of dietary energy levels on rumen bacterial community composition in Holstein heifers under the same forage to concentrate ratio condition. BMC Microbiol 18:69
Bin P, Azad MAK, Liu G, Zhu D, Kim SW, Yin Y (2018) Effects of different levels of methionine on sow health and plasma metabolomics during late gestation. Food Funct 9:4979–4988
Bladen HA, Bryant MP, Doetsch RN (1961) A study of bacterial species from the rumen which produce ammonia from protein hydrolyzate. Appl Microbiol 9:175–180
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Charles GS, Glen BA (2017) A 100-year review: protein and amino acid nutrition in dairy cows. J Dairy Sci 100:10094–10112
Chen J, Bittinger K, Charlson ES, Hoffmann C, Lewis J, Wu GD, Collman GR, Bushman DF, Li H (2012) Associating microbiome composition with environmental ovariates using generalized UniFrac distances. Bioinformatics 28:2106–2113
Ciarlo E, Heinonen T, Herderschee J, Fenwick C, Mombelli M, Le Roy D, Roger T (2016) Impact of the microbial derived short chain fatty acid propionate on host susceptibility to bacterial and fungal infections in vivo. Sci Rep 6:37944
Clark JH, Klusmeyer TH, Cameron MR (1992) Microbial protein synthesis and flows of nitrogen fractions to the duodenum of dairy cows. J Dairy Sci 75:2304–2323
Domingo MC, Huletsky A, Boissinot M, Bernard KA, Picard FJ, Bergeron MG (2008) Ruminococcus gauvreauii sp. nov., a glycopeptide-resistant species isolated from a human faecal specimen. Int J Syst Evol Microbiol 58:1393–1397
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2000
Elwakeel EA, Amachawadi RG, Nour AM, Nasser MEA, Nagaraja TG, Titgemeyer EC (2013) In vitro degradation of lysine by ruminal fluid-based fermentations and by Fusobacterium necrophorum. J Dairy Sci 96:495–505
Evans E, Lamont J, Leclerc H (2015) Methods to evaluate rumen protected lysine for dairy cows. Open J Anim Sci 5:495–499
Garner MR, Flint JF, Russell JB (2002) Allisonella histaminiformans gen. nov., sp. nov. A novel bacterium that produces histamine, utilizes histidine as its sole energy source, and could play a role in bovine and equine laminitis. Syst Appl Microbiol 25:498–506
Gorka P, Kowalski ZM, Pietrzak P, Kotunia A, Jagusiak W, Holst JJ, Guilloteau P, Zabielski R (2011) Effect of method of delivery of sodium butyrate on rumen development in newborn calves. J Dairy Sci 94:5578–5588
Guilloteau P, Zabielski R, David JC, Blum JW, Morisset JA, Biernat M, Wolinski J, Laubitz D, Hamon Y (2009) Sodium-butyrate as a growth promoter in milk replacer formula for young calves. J Dairy Sci 92:1038–1049
Han IK, Ha JK, Lee SS, Ko YG, Lee HS (1996) Effect of supplementing rumen-protected lysine and methionine on ruminal characteristics and nutrient digestibility in sheep. Asian Australas J Anim Sci 9:223–230
Hill GB, Ayers OM, Kohan AP (1987) Characteristics and sites of infection of Eubacterium nodatum, Eubacterium timidum, Eubacterium brachy, and other asaccharolytic eubacteria. J Clin Microbiol 25:1540–1545
Hristov AN, Bannink A, Crompton LA, Huhtanen P, Kreuzer M, McGee M, Noziere P, Reynolds CK, Bayat AR, Yanez-Ruiz DR, Dijkstra J, Kebreab E, Schwarm A, Shingfield KJ, Yu Z (2019) Invited review: nitrogen in ruminant nutrition: a review of measurement techniques. J Dairy Sci 102:5811–5852
Hullar MA, Fu BC (2014) Diet, the gut microbiome, and epigenetics. Cancer J 20:170–175
Ishimaru S, Elsabagh M, Saiki S, Haruno A, Sugino T (2019) Evaluating the rumen-protected lysine stability in forage-based total mixed rations in vitro and determining the lysine Brix value. Anim Sci J 90:932–938
Jami E, Israel A, Kotser A, Mizrahi I (2013) Exploring the bovine rumen bacterial community from birth to adulthood. ISME J 7:1069–1079
Ji P, Tucker HA, Clark RE, Miura M, Ballard CS (2016) Short communication: effect of on-farm feeding practices on rumen protected lysine products. J Dairy Sci 99:1242–1246
Kumar S, Treloar BP, Teh KH, McKenzie CM, Henderson G, Attwood GT, Waters SM, Patchett ML, Janssen PH (2018) Sharpea and Kandleria are lactic acid producing rumen bacteria that do not change their fermentation products when co-cultured with a methanogen. Anaerobe 54:31–38
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Thurber RLV, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(814):821
Lee C, Hristov AN, Cassidy TW, Heyler KS, Lapierre H, Varga GA, de Veth MJ, Patton RA, Parys C (2012) Rumen-protected lysine, methionine, and histidine increase milk protein yield in dairy cows fed a metabolizable protein-deficient diet. J Dairy Sci 95:6042–6056
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Mardinoglu A, Shoaie S, Bergentall M, Ghaffari P, Zhang C, Larsson E, Backhed F, Nielsen J (2015) The gut microbiota modulates host amino acid and glutathione metabolism in mice. Mol Syst Biol 11:834
Meehan CJ, Beiko RG (2014) A phylogenomic view of ecological specialization in the Lachnospiraceae, a family of digestive tract-associated bacteria. Genome Biol Evol 6:703–713
Nakajima M, Tanaka N, Furukawa N, Nihira T, Kodutsumi Y, Takahashi Y, Sugimoto N, Miyanaga A, Fushinobu S, Taguchi H, Nakai H (2017) Mechanistic insight into the substrate specificity of 1,2-β-oligoglucan phosphorylase from Lachnoclostridium phytofermentans. Sci Rep 7:42671
Neubauer V, Petri R, Humer E, Kroger I, Mann E, Reisinger N, Wagner M, Zebeli Q (2018) High-grain diets supplemented with phytogenic compounds or autolyzed yeast modulate ruminal bacterial community and fermentation in dry cows. J Dairy Sci 101:2335–2349
Nichols JR, Schingoethe DJ, Maiga HA, Brouk MJ, Piepenbrink MS (1998) Evaluation of corn distillers grains and ruminally protected lysine and methionine for lactating dairy cows. J Dairy Sci 81:482–491
Noftsger S, St-Pierre NR (2003) Supplementation of methionine and selection of highly digestible rumen undegradable protein to improve nitrogen efficiency for milk production. J Dairy Sci 86:958–969
NRC (2001) Nutrient requirements of dairy cattle, 10th edn. National Academies Press, Washington D.C
Onodera R (1993) Methionine and lysine metabolism in the rumen and the possible effects of their metabolites on the nutrition and physiology of ruminants. Amino Acids 5:217–232
Ørskov ER, McDonald I (1979) The estimation of protein degradability in the rumen from incubation measurements weighted according to rate of passage. J Agric Sci 92:499–503
Papas AM, Sniffen CJ, Muscato TV (1984) Effectiveness of rumen protected methionine for delivering methionine postruminally in dairy cows. J Dairy Sci 67:545–552
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124
Rist VT, Weiss E, Eklund M, Mosenthin R (2013) Impact of dietary protein on microbiota composition and activity in the gastrointestinal tract of piglets in relation to gut health: a review. Animal 7:1067–1078
Robinson PH, DePeters EJ, Shinzato I, Sato H (2005) Influence of feeding free lysine to early lactation dairy cows on ruminal lysine escape, rumen fermentation and productivity. Anim Feed Sci Technol 118:201–214
Robinson PH, DePeters EJ, Shinzato I, Sato H (2006) Rumen lysine escape, rumen fermentation, and productivity of early lactation dairy cows fed free lysine. Anim Feed Sci Technol 128:31–41
Rossi F, Maurizio M, Francesco M, Giovanna C, Gianfranco P (2003) Rumen degradation and intestinal digestibility of rumen protected amino acids: comparison between in situ and in vitro data. Anim Feed Sci Technol 108:223–229
Sakkers M, Erasmus LJ, Robinson PH, Meeske R, Garrett JE (2013) Determining in vivo ruminal stability of three ruminally protected nutrients in lactating Jersey cows. Anim Feed Sci Technol 185:133–139
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Shang Q, Shi J, Song G, Zhang M, Cai C, Hao J, Li G, Yu G (2016) Structural modulation of gut microbiota by chondroitin sulfate and its oligosaccharide. Int J Biol Macromol 89:489–498
Storm E, Orskov ER, Smart R (1984) The nutritive value of rumen micro-organisms in ruminants. 4. The limiting amino acids of microbial protein in growing sheep determined by a new approach. Br J Nutr 52:613–620
Südekum KH, Wolffram S, Ader P, Robert J-C (2004) Bioavailability of three ruminally protected methionine sources in cattle. Anim Feed Sci Technol 113:17–25
Sun ZH, Tan ZL, Liu SM, Tayo GO, Lin B, Teng B, Tang SX, Wang WJ, Liao YP, Pan YF, Wang JR, Zhao XG, Hu Y (2007) Effects of dietary methionine and lysine sources on nutrient digestion, nitrogen utilization, and duodenal amino acid flow in growing goats. J Anim Sci 85:3340–3347
Van Soest PJ, Robertson JB, Lewis BA (1991) Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 74:3583–3597
Vandecasteele JP, Hermann M (1972) Regulation of a catabolic pathway: lysine degradation in Pseudomonas putida. Eur J Biochem 31:80–85
Velle W, Kanui TI, Aulie A, Sjaastad OV (1999) Ruminal escape and apparent degradation of amino acids administered intraruminally in mixtures to cows. J Dairy Sci 81:3231–3238
Venema K (2010) Role of gut microbiota in the control of energy and carbohydrate metabolism. Curr Opin Clin Nutr Metab Care 13:432–438
Wang J, Diao Q, Tu Y, Zhang N, Xu X (2012) The limiting sequence and proper ratio of lysine, methionine and threonine for calves fed milk replacers containing soy protein. Asian Australas J Anim Sci 25:224–233
Watanabe M, Kaku N, Ueki K, Ueki A (2016) Falcatimonas natans gen. nov., sp nov., a strictly anaerobic, amino-acid-decomposing bacterium isolated from a methanogenic reactor of cattle waste. Int J Syst Evol Microbiol 66:4639–4644
Wei T, Simko V (2013) corrplot: visualization of a correlation matrix. MMWR Morb Mortal Wkly Rep 52:145–151
Xu N, Tan GC, Wang HY, Gai XP (2016) Effect of biochar additions to soil on nitrogen leaching, microbial biomass and bacterial community structure. Eur J Soil Biol 74:1–8
Yin J, Li Y, Han H, Liu Z, Zeng X, Li T, Yin Y (2018) Long-term effects of lysine concentration on growth performance, intestinal microbiome, and metabolic profiles in a pig model. Food Funct 9:4153–4163
Zhu W, Tang CH, Sun XP, Liu JX, Wu YM, Yuan YM, Zhang XK (2013) Rumen microbial protein synthesis and milk performance in lactating dairy cows fed the fortified corn stover diet in comparison with alfalfa diet. J Anim Vet Adv 12:633–639
Zinn RA, Shen Y (1998) An evaluation of ruminally degradable intake protein and metabolizable amino acid requirements of feedlot calves
Funding
This work was supported, in part, by the Earmarked Fund for Beijing Dairy Industry Innovation Consortium of Agriculture Research System (BAIC06), Chinese Academy of Agricultural Science and Technology Innovation Project (CAAS-XTCX-2016011-01), Fundamental Research Funds for Central Non-profit Scientific Institution (Y2019CG08), Agricultural Science and Technology Innovation Program (CAAS-ASTIP-2017-FRI-04), Science and Technology Open Cooperation Project of Henan Province (182106000035)-Study on the Pattern of Diet Amino Acid for Different Physiological Stages of Heifers, and Key Research and Development Program of Hebei Province (19226621D).
Author information
Authors and Affiliations
Contributions
Fanlin Kong and Mengqi Tang carried out both experiments and then did the sampling and laboratory works. Yanliang Bi drafted the manuscript and critically reviewed the manuscript. Yanxia Gao provided the materials needed in in situ experiment and conceived the study. Tong Fu provided the materials needed in experiment 2 and conceived the study. Yan Tu and Qiyu Diao designed and approved the study plan. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Experimental procedures were approved by the Animal Ethics Committee of the Chinese academy of agricultural sciences (committee permit number: FRI-CAAS20180110), and humane animal care and handling procedures were followed throughout the experiment.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Code availability
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 134 kb)
Rights and permissions
About this article
Cite this article
Kong, F., Gao, Y., Tang, M. et al. Effects of dietary rumen–protected Lys levels on rumen fermentation and bacterial community composition in Holstein heifers. Appl Microbiol Biotechnol 104, 6623–6634 (2020). https://doi.org/10.1007/s00253-020-10684-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10684-y