Abstract
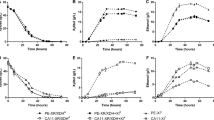
Engineered S. cerevisiae employing the xylose reductase pathway enables efficient xylose valorization to fuels and chemicals. However, toxicity of thermochemically pretreated biomass hydrolysate on S. cerevisiae is one of the key technical challenges to upgrade biomass-derived sugars including xylose and glucose into high-value products. We investigated the effect of glycolaldehyde, one of the biomass-derived highly toxic aldehyde compounds, and its combinatorial inhibitory effect with other major fermentation inhibitors commonly found in plant hydrolysate such as methylglyoxal, 5-HMF, furfural, vanillin, and acetic acid on engineered xylose-fermenting S. cerevisiae in xylose and/or glucose media. We elucidated that glycolaldehyde and methylglyoxal are the key inhibitory short-aliphatic aldehydes on engineered xylose-fermenting S. cerevisiae in xylose-containing medium. Indeed, the degree of toxicity of these tested fermentation inhibitors varies with the sole carbon source of the medium. We demonstrate that genome integration of an extra copy of autologous GRE2 with its native promotor substantially improved the toxic tolerance of engineered xylose-fermenting S. cerevisiae to major inhibitory compounds including glycolaldehyde in the xylose-containing medium, and xylose-rich, lignocellulosic hydrolysate derived from Miscanthus giganteus, and concurrently improved the ethanol fermentation profile. Outcomes of this study will aid the development of next-generation robust S. cerevisiae strains for efficient fermentation of hexose and pentose sugars found in biomass hydrolysate.




Similar content being viewed by others
References
Almeida JR, Modig T, Petersson A, Hähn-Hägerdal B, Lidén G, Gorwa-Grauslund MF (2007) Increased tolerance and conversion of inhibitors in lignocellulosic hydrolysates by Saccharomyces cerevisiae. J Chem Technol Biotechnol 82(4):340–349
Ask M, Bettiga M, Duraiswamy VR, Olsson L (2013a) Pulsed addition of HMF and furfural to batch-grown xylose-utilizing Saccharomyces cerevisiae results in different physiological responses in glucose and xylose consumption phase. Biotechnology for Biofuels 6(1):181
Ask M, Bettiga M, Mapelli V, Olsson L (2013b) The influence of HMF and furfural on redox-balance and energy-state of xylose-utilizing Saccharomyces cerevisiae. Biotechnology for Biofuels 6(1):22
Becuwe M, Léon S (2014) Integrated control of transporter endocytosis and recycling by the arrestin-related protein Rod1 and the ubiquitin ligase Rsp5. Elife 3:e03307
Cavka A, Stagge S, Jönsson LJ (2015) Identification of small aliphatic aldehydes in pretreated lignocellulosic feedstocks and evaluation of their inhibitory effects on yeast. J Agric Food Chem 63(44):9747–9754
Celton M, Goelzer A, Camarasa C, Fromion V, Dequin S (2012) A constraint-based model analysis of the metabolic consequences of increased NADPH oxidation in Saccharomyces cerevisiae. Metab Eng 14(4):366–379
Cherubini F (2010) The biorefinery concept: using biomass instead of oil for producing energy and chemicals. Energy Convers Manag 51(7):1412–1421
Chiaramonti D, Prussi M, Ferrero S, Oriani L, Ottonello P, Torre P, Cherchi F (2012) Review of pretreatment processes for lignocellulosic ethanol production, and development of an innovative method. Biomass Bioenergy 46:25–35
Delgenes J, Moletta R, Navarro J (1996) Effects of lignocellulose degradation products on ethanol fermentations of glucose and xylose by Saccharomyces cerevisiae, Zymomonas mobilis, Pichia stipitis, and Candida shehatae. Enzym Microb Technol 19(3):220–225
Dolz-Edo L, Rienzo A, Poveda-Huertes D, Pascual-Ahuir A, Proft M (2013) Deciphering dynamic dose responses of natural promoters and single cis elements upon osmotic and oxidative stress in yeast. Mol Cell Biol 33(11):2228–2240
dos Santos Cordeiro Perna M, Bastos RG, Ceccato-Antonini SR (2018) Single and combined effects of acetic acid, furfural, and sugars on the growth of the pentose-fermenting yeast Meyerozyma guilliermondii. 3 Biotech 8(2):119
Garay-Arroyo A, Covarrubias AA (1999) Three genes whose expression is induced by stress in Saccharomyces cerevisiae. Yeast 15(10a):879–892
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Glomb MA, Monnier VM (1995) Mechanism of protein modification by glyoxal and glycolaldehyde, reactive intermediates of the Maillard reaction. J Biol Chem 270(17):10017–10026
Guo P-C, Bao Z-Z, Ma X-X, Xia Q, Li W-F (2014) Structural insights into the cofactor-assisted substrate recognition of yeast methylglyoxal/isovaleraldehyde reductase Gre2. Biochim Biophys Acta (BBA) - Proteins Proteomics 1844(9):1486–1492
Hasunuma T, Sanda T, Yamada R, Yoshimura K, Ishii J, Kondo A (2011) Metabolic pathway engineering based on metabolomics confers acetic and formic acid tolerance to a recombinant xylose-fermenting strain of Saccharomyces cerevisiae. Microb Cell Factories 10(1):2
Jayakody LN, Horie K, Hayashi N, Kitagaki H (2012) Improvement of tolerance of Saccharomyces cerevisiae to hot-compressed water-treated cellulose by expression of ADH1. Appl Microbiol Biotechnol 94(1):273–283
Jayakody LN, Hayashi N, Kitagaki H (2013a) Molecular mechanisms for detoxification of major aldehyde inhibitors for production of bioethanol by Saccharomyces cerevisiae from hot-compressed water-treated lignocelluloses. In: Vilas AM (ed) Material and process for energy: communicating current research and technology development. Formatex Research Center, Badajoz, pp 302–311
Jayakody LN, Horie K, Hayashi N, Kitagaki H (2013b) Engineering redox cofactor utilization for detoxification of glycolaldehyde, a key inhibitor of bioethanol production, in yeast Saccharomyces cerevisiae. Appl Microbiol Biotechnol 97(14):6589–6600
Jayakody LN, Kadowaki M, Tsuge K, Horie K, Suzuki A, Hayashi N, Kitagaki H (2015) SUMO expression shortens the lag phase of Saccharomyces cerevisiae yeast growth caused by complex interactive effects of major mixed fermentation inhibitors found in hot-compressed water-treated lignocellulosic hydrolysate. Appl Microbiol Biotechnol 99(1):501–515
Jayakody LN, Ferdouse J, Hayashi N, Kitagaki H (2017) Identification and detoxification of glycolaldehyde, an unattended bioethanol fermentation inhibitor. Crit Rev Biotechnol 37(2):177–189
Jayakody LN, Johnson CW, Whitham JM, Giannone RJ, Black BA, Cleveland NS, Klingeman DM, Michener WE, Olstad JL, Vardon DR (2018) Thermochemical wastewater valorization via enhanced microbial toxicity tolerance. Energy Environ Sci 11:1625–1638
Kim SR, Kwee NR, Kim H, Jin YS (2013a) Feasibility of xylose fermentation by engineered Saccharomyces cerevisiae overexpressing endogenous aldose reductase (GRE3), xylitol dehydrogenase (XYL2), and xylulokinase (XYL3) from Scheffersomyces stipitis. FEMS Yeast Res 13(3):312–321
Kim SR, Skerker JM, Kang W, Lesmana A, Wei N, Arkin AP, Jin Y-S (2013b) Rational and evolutionary engineering approaches uncover a small set of genetic changes efficient for rapid xylose fermentation in Saccharomyces cerevisiae. PLoS One 8(2):e57048
Kimura A, Inoue Y (1993) Glyoxalase-I in microorganisms-molecular characteristics, genetics and biochemical regulation. Biochem Soc Trans 21:518–522
Kua J, Galloway MM, Millage KD, Avila JE, De Haan DO (2013) Glycolaldehyde monomer and oligomer equilibria in aqueous solution: comparing computational chemistry and NMR data. J Phys Chem A 117(14):2997–3008
Li Y-C, Gou Z-X, Zhang Y, Xia Z-Y, Tang Y-Q, Kida K (2017) Inhibitor tolerance of a recombinant flocculating industrial Saccharomyces cerevisiae strain during glucose and xylose co-fermentation. Braz J Microbiol 48(4):791–800
Liu ZL (2011) Molecular mechanisms of yeast tolerance and in situ detoxification of lignocellulose hydrolysates. Appl Microbiol Biotechnol 90(3):809–825
Liu C, Wyman CE (2005) Partial flow of compressed-hot water through corn stover to enhance hemicellulose sugar recovery and enzymatic digestibility of cellulose. Bioresour Technol 96(18):1978–1985
LoPachin RM, Gavin T (2014) Molecular mechanisms of aldehyde toxicity: a chemical perspective. Chem Res Toxicol 27(7):1081–1091
LoPachin RM, Gavin T, DeCaprio A, Barber DS (2011) Application of the hard and soft, acids and bases (HSAB) theory to toxicant–target interactions. Chem Res Toxicol 25(2):239–251
Lu X, Yamauchi K, Phaiboonsilpa N, Saka S (2009) Two-step hydrolysis of Japanese beech as treated by semi-flow hot-compressed water. J Wood Sci 55(5):367–375
Ma M, Liu ZL (2010) Comparative transcriptome profiling analyses during the lag phase uncover YAP1, PDR1, PDR3, RPN4, and HSF1 as key regulatory genes in genomic adaptation to the lignocellulose derived inhibitor HMF for Saccharomyces cerevisiae. BMC Genomics 11(1):660
Mitraki A, Haase-Pettingell C, King J (1991) Mechanisms of inclusion body formation. ACS Publications
Moon J, Liu ZL (2012) Engineered NADH-dependent GRE2 from Saccharomyces cerevisiae by directed enzyme evolution enhances HMF reduction using additional cofactor NADPH. Enzym Microb Technol 50(2):115–120
Nanda S, Kozinski JA, Dalai AK (2016) Lignocellulosic biomass: a review of conversion technologies and fuel products. Curr Biochem Eng 3(1):24–36
Nguyen TTM, Ishida Y, Kato S, Iwaki A, Izawa S (2018) The VFH1 (YLL056C) promoter is vanillin-inducible and enables mRNA translation despite pronounced translation repression caused by severe vanillin stress in Saccharomyces cerevisiae. Yeast 35:465–475
Roy A, Hashmi S, Li Z, Dement AD, Cho KH, Kim J-H (2016) The glucose metabolite methylglyoxal inhibits expression of the glucose transporter genes by inactivating the cell surface glucose sensors Rgt2 and Snf3 in yeast. Mol Biol Cell 27(5):862–871
Salusjärvi L, Kankainen M, Soliymani R, Pitkänen J-P, Penttilä M, Ruohonen L (2008) Regulation of xylose metabolism in recombinant Saccharomyces cerevisiae. Microb Cell Factories 7(1):18
Skerker JM, Leon D, Price MN, Mar JS, Tarjan DR, Wetmore KM, Deutschbauer AM, Baumohl JK, Bauer S, Ibánez AB (2013) Dissecting a complex chemical stress: chemogenomic profiling of plant hydrolysates. Mol Syst Biol 9(1):674
Thomas JG, Baneyx F (1996) Protein misfolding and inclusion body formation in recombinant Escherichia coli cells overexpressing heat-shock proteins. J Biol Chem 271(19):11141–11147
Wang X, Jin M, Balan V, Jones AD, Li X, Li BZ, Dale BE, Yuan YJ (2014) Comparative metabolic profiling revealed limitations in xylose-fermenting yeast during co-fermentation of glucose and xylose in the presence of inhibitors. Biotechnol Bioeng 111(1):152–164
Wang X, Ma M, Liu ZL, Xiang Q, Li X, Liu N, Zhang X (2016) GRE2 from Scheffersomyces stipitis as an aldehyde reductase contributes tolerance to aldehyde inhibitors derived from lignocellulosic biomass. Appl Microbiol Biotechnol 100(15):6671–6682
Warringer J, Blomberg A (2006) Involvement of yeast YOL151W/GRE2 in ergosterol metabolism. Yeast 23(5):389–398
Webb SR, Lee H (1992) Characterization of xylose reductase from the yeast Pichia stipitis: evidence for functional thiol and histidyl groups. Microbiology 138(9):1857–1863
Yu Y, Lou X, Wu H (2007) Some recent advances in hydrolysis of biomass in hot-compressed water and its comparisons with other hydrolysis methods. Energy Fuel 22(1):46–60
Zhang G, Kong II, Kim H, Liu J, Cate JH, Jin YS (2014) Construction of a quadruple auxotrophic mutant of an industrial polyploidy Saccharomyces cerevisiae using RNA-guided Cas9 nuclease. Appl Environ Microbiol 80(24):7694–701
Zhang GC, Kong II, Wei N, Peng D, Turner TL, Sung BH, Sohn JH, Jin YS (2016) Optimization of an acetate reduction pathway for producing cellulosic ethanol by engineered yeast. Biotechnol Bioeng 113(12):2587–2596
Funding
This study was funded by the Energy Bioscience Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 457 kb)
Rights and permissions
About this article
Cite this article
Jayakody, L.N., Turner, T.L., Yun, E.J. et al. Expression of Gre2p improves tolerance of engineered xylose-fermenting Saccharomyces cerevisiae to glycolaldehyde under xylose metabolism. Appl Microbiol Biotechnol 102, 8121–8133 (2018). https://doi.org/10.1007/s00253-018-9216-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9216-x