Abstract
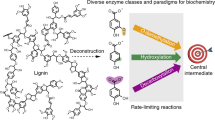
Lignin is the major aromatic biopolymer in nature, and it is considered a valuable feedstock for the future supply of aromatics. Hence, its valorisation in biorefineries is of high importance, and various chemical and enzymatic approaches for lignin depolymerisation have been reported. Among the enzymes known to act on lignin, β-etherases offer the possibility for a selective cleavage of the β-O-4 aryl ether bonds present in lignin. These enzymes, together with glutathione lyases, catalyse a reductive, glutathione-dependent ether bond cleavage displaying high stereospecificity. β-Etherases and glutathione lyases both belong to the superfamily of glutathione transferases, and several structures have been solved recently. Additionally, different approaches for their application in lignin valorisation have been reported in the last years. This review gives an overview on the current knowledge on β-etherases and glutathione lyases, their biochemical and structural features, and critically discusses their potential for application in biorefineries.








Similar content being viewed by others
References
Abdel-Hamid AM, Solbiati JO, Cann IK (2013) Insights into lignin degradation and its potential industrial applications. Adv Appl Microbiol 82:1–28. https://doi.org/10.1016/B978-0-12-407679-2.00001-6
Adler E (1977) Lignin chemistry—past, present and future. Wood Sci Technol 11:169–218
Ahmad M, Roberts JN, Hardiman EM, Singh R, Eltis LD, Bugg TDH (2011) Identification of DypB from Rhodococcus jostii RHA1 as a lignin peroxidase. Biochemistry 50:5096–5107. https://doi.org/10.1021/bi101892z
Ahmed AAQ, McKay TJM (2017) Potential of Bacillus sp. LG7 as a promising source of ligninolytic enzymes for industrial and biotechnological applications. Proc Natl Acad Sci India B Biol Sci. https://doi.org/10.1007/s40011-017-0957-6
Allocati N, Federici L, Masulli M, Di Ilio C (2009) Glutathione transferases in bacteria. FEBS J 276:58–75. https://doi.org/10.1111/j.1742-4658.2008.06743.x
Bandounas L, Wierckx NJP, de Winde JH, Ruijssenaars (2011) Isolation and characterization of novel bacterial strains exhibiting ligninolytic potential. BMC Biotechnol 11:94. https://doi.org/10.1186/1472-6750-11-94
Brown ME, Chang MCY (2014) Exploring bacterial lignin degradation. Curr Opin Chem Biol 19:1–7. https://doi.org/10.1016/j.cbpa.2013.11.015
Brown ME, Barros T, Chang MCY (2012) Identification and characterization of a multifunctional dye peroxidase from a lignin-reactive bacterium. ACS Chem Biol 7:2074–2081. https://doi.org/10.1021/cb300383y
Bugg TDH, Rahmanpour R (2015) Enzymatic conversion of lignin into renewable chemicals. Curr Opin Chem Biol 29:10–17. https://doi.org/10.1016/j.cbpa.2015.06.009
Bugg TDH, Ahmad M, Hardiman EM, Singh R (2011) The emerging role for bacteria in lignin degradation and bio-product formation. Curr Opin Biotechnol 22:394–400. https://doi.org/10.1016/j.copbio.2010.10.009
Chen YR, Sarkanen S, Wang YY (2012) Lignin-degrading enzyme activities. In: Himmel M (ed) Biomass conversion. Methods in molecular biology (methods and protocols), vol 908. Humana Press, Totowa, pp 251–268. https://doi.org/10.1007/978-1-61779-956-3_21
Crestini C, Melone F, Sette M, Saladino R (2011) Milled wood lignin: a linear oligomer. Biomacromolecules 12:3928–3935. https://doi.org/10.1021/bm200948r
de Gonzalo G, Colpa DI, Habib MHM, Fraaije MW (2016) Bacterial enzymes involved in lignin degradation. J Biotechnol 236:110–119. https://doi.org/10.1016/j.jbiotec.2016.08.011
Faix O (1991) Classification of lignins from different botanical origins by FT-IR spectroscopy. Holzforschung 45:21–27. https://doi.org/10.1515/hfsg.1991.45.s1.21
FritzPatrick M, Champagne P, Cunningham MF, Whitney RA (2010) A biorefinery processing perspective: treatment of lignocellulosic materials for the production of value-added products. Bioresour Technol 101:8915–8922. https://doi.org/10.1016/j.biortech.2010.06.125
Gall DL, Kim H, Lu F, Donohue TJ, Noguera DR, Ralph J (2014a) Stereochemical features of glutathione-dependent enzymes in the Sphingobium sp. strain SYK-6 β-aryl etherase pathway. J Biol Chem 289:8656–8667. https://doi.org/10.1074/jbc.M113.536250
Gall DL, Ralph J, Donohue TJ, Noguera DR (2014b) A group of sequence-related sphingomonad enzymes catalyzes cleavage of β-aryl ether linkages in lignin β-guaiacyl and β-syringyl ether dimers. Environ Sci Technol 48:12454–12463. https://doi.org/10.1021/es503886d
Gall DL, Kontur WS, Lan W, Kim H, Li Y, Ralph J, Donohue TJ, Noguera DR (2018) In vitro enzymatic depolymerization of lignin with release of syringyl, guaiacyl, and tricin units. Appl Environ Microbiol. https://doi.org/10.1128/AEM.02076-17
Grande PM, Viell J, Theyssen N, Marquardt W, Domínguez de María P, Leitner W (2015) Fractionation of lignocellulosic biomass using the OrganoCat process. Green Chem 17:3533–3539. https://doi.org/10.1039/C4GC02534B
Helmich KE, Pereira JH, Gall DL, Heins RA, McAndrew RP, Bingman C, Deng K, Holland KC, Noguera DR, Simmons BA, Sale KL, Ralph J, Donohue TJ, Adams PD, Phillips GN (2016) Structural basis of stereospecificity in the bacterial enzymatic cleavage of β-aryl ether bonds in lignin. J Biol Chem 291:5234–5246. https://doi.org/10.1074/jbc.M115.694307
Hendriks ATWM, Zeeman G (2009) Pretreatments to enhance the digestibility of lignocellulosic biomass. Bioresour Technol 100:10–18. https://doi.org/10.1016/j.biortech.2008.05.027
Isikgor FH, Becer R (2015) Lignocellulosic biomass: a sustainable platform for the production of bio-based chemicals and polymers. Polym Chem 6:4497–4559. https://doi.org/10.1039/c5py00263j
Kontur WS, Bingman CA, Olmsted CN, Wassarman DR, Ulbrich A, Gall DL, Smith RW, Yusko LM, Fox BG, Noguera DR, Coon JJ, Donohue TJ (2018) Novosphingobium aromaticivorans uses a Nu-class glutathione-S-transferase as a glutathione lyase in breaking the β-aryl ether bond of lignin. J Biol Chem 293:4955–4968. https://doi.org/10.1074/jbc.RA117.001268
Lancefield CS, Ojo OS, Tran F, Westwood NJ (2015) Isolation of functionalized phenolic monomers through selective oxidation and C-O bond cleavage of the β-O-4 linkages in lignin. Angew Chem Int Ed Engl 54:258–262. https://doi.org/10.1002/ange.201409408
Masai E, Katayama Y, Nishikawa S, Yamasaki M, Morohoshi N, Haraguchi T (1989) Detection and localization of a new enzyme catalyzing the β-aryl ether cleavage in the soil bacterium (Pseudomonas paucimobilis SYK-6). FEBS Lett 249:348–352. https://doi.org/10.1016/0014-5793(89)80656-8
Masai E, Katayama Y, Kubota S, Kawai S, Yamasaki M, Morohoshi N (1993a) A bacterial enzyme degrading the model lignin compound β-etherase is a member of the glutathione-S-transferase superfamily. FEBS Lett 323:135–140. https://doi.org/10.1016/0014-5793(93)81465-C
Masai E, Kubota S, Katayama Y, Kawai S, Yamasaki M, Morohoshi N (1993b) Characterization of the Cα-dehydrogenase gene involved in the cleavage of β-aryl ether by Pseudomonas paucimobilis. Biotechnol Biochem 57:1655–1659. https://doi.org/10.1271/bbb.57.1655
Masai E, Katayama Y, Nishikawa S, Fukuda M (1999) Characterization of Sphingomonas paucimobilis SYK-6 genes involved in degradation of lignin-related compounds. J Ind Microbiol Biotechnol 23:364–373. https://doi.org/10.1038/sj.jim.2900747
Masai E, Ichimura A, Sato Y, Miyauchi K, Katayama Y, Fukuda M (2003) Roles of the enantioselective glutathione-S-transferases in cleavage of β-aryl ether. J Bacteriol 185:1768–1775. https://doi.org/10.1128/JB.185.6.1768-1775.2003
Masai E, Katayama Y, Fukuda M (2007) Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci Biotechnol Biochem 71:1–15. https://doi.org/10.1271/bbb.60437
Meux E, Prosper P, Masai E, Mulliert G, Dumarçay S, Morel M, Didierjean C, Gelhaye E, Favier F (2012) Sphingobium sp. SYK-6 LigG involved in lignin degradation is structurally and biochemically related to the glutathione transferase ω class. FEBS Lett 586:3944–3950. https://doi.org/10.1016/j.febslet.2012.09.036
Ohta Y, Nishi S, Hasegawa R, Hatada Y (2015) Combination of six enzymes of a marine Novosphingobium converts the stereoisomers of β-O-4 lignin model dimers into the respective monomers. Sci Rep 5:15105–15118. https://doi.org/10.1038/srep15105
Ohta Y, Hasegawa R, Kurosawa K, Maeda AH, Koizumi T, Nishimura H, Okada H, Qu C, Saito K, Watanabe T, Hatada Y (2016) Enzymatic specific production and chemical functionalization of phenylpropanone platform monomers from lignin. ChemSusChem 10:425–433. https://doi.org/10.1002/cssc.201601235
Otsuka Y, Sonoki T, Ikeda S, Kajita S, Nakamura M, Katayama Y (2003) Detection and characterization of a novel extracellular fungal enzyme that catalyzes the specific and hydrolytic cleavage of lignin guaiacylglycerol β-aryl ether linkages. Eur J Biochem 270:2353–2362. https://doi.org/10.1046/j.1432-1033.2003.03545.x
Pandey MP, Kim CS (2011) Lignin depolymerization and conversion: a review of thermochemical methods. Chem Eng Technol 34:29–41. https://doi.org/10.1002/ceat.201000270
Pereira JH, Heins RA, Gall DL, McAndrew RP, Deng K, Holland KC, Donohue TJ, Noguera DR, Simmons BA, Sale KL, Ralph J, Adams PD (2016) Structural and biochemical characterization of the early and late enzymes in the lignin β-aryl ether cleavage pathway from Sphingobium sp. SYK-6. J Biol Chem 291:10228–10238. https://doi.org/10.1074/jbc.M115.700427
Picart P, Müller C, Mottweiler J, Wiermans L, Bolm C, Domínguez de María P, Schallmey A (2014) From gene towards selective biomass valorization: bacterial β-etherases with catalytic activity on lignin-like polymers. ChemSusChem 7:3164–3171. https://doi.org/10.1002/cssc.201402465
Picart P, Domínguez de María P, Schallmey A (2015a) From gene to biorefinery: microbial β-etherases as promising biocatalysts for lignin valorization. Front Microbiol 6:916. https://doi.org/10.3389/fmicb.2015.00916
Picart P, Sevenich M, Domínguez de María P, Schallmey A (2015b) Exploring glutathione lyases as biocatalysts: paving the way for enzymatic lignin depolymerization and future stereoselective applications. Green Chem 17:4931–4940. https://doi.org/10.1039/C5GC01078K
Picart P, Wiermans L, Pérez-Sánchez M, Grande PM, Schallmey A, Domínguez de María P (2016) Assessing lignin types to screen novel biomass-degrading microbial strains: synthetic lignin as useful carbon source. ACS Sustain Chem Eng 4:651–655. https://doi.org/10.1021/acssuschemeng.5b00961
Picart P, Liu H, Grande PM, Anders N, Zhu L, Klankermayer J, Leitner W, Domínguez de María P, Schwaneberg U, Schallmey A (2017) Multi-step biocatalytic depolymerization of lignin. Appl Microbiol Biotechnol 101:6277–6287. https://doi.org/10.1007/s00253-017-8360-z
Pollegioni L, Tonin F, Rosini E (2015) Lignin-degrading enzymes. FEBS J 282:1190–1213. https://doi.org/10.1111/febs.13224
Ragauskas AJ, Beckham GT, Biddy MT, Chandra R, Chen F, Davis MF, Davison BH, Nixon RA, Glina P, Keller M, Langan P, Naskar AK, Saddler JN, Tschaplinski TJ, Tuskan GA, Wyman CE (2014) Lignin valorization: improving lignin processing in the biorefinery. Science 344:1246843. https://doi.org/10.1126/science.1246843
Rahimi A, Azarpira A, Kim H, Ralph J, Stahl SS (2013) Chemoselective metal-free aerobic alcohol oxidation in lignin. J Am Chem Soc 135:6415–6418. https://doi.org/10.1021/ja401793n
Rahimi A, Ulbrich A, Coon JJ, Stahl SS (2014) Formic-acid induced depolymerization of oxidized lignin to aromatics. Nature 13:249–252. https://doi.org/10.1038/nature13867
Rahmanpour R, Bugg TDH (2015) Characterisation of Dyp-type peroxidases from Pseudomonas fluorescens Pf-5: oxidation of Mn(II) and polymeric lignin by Dyp1B. Arch Biochem Biophys 574:93–98. https://doi.org/10.1016/j.abb.2014.12.022
Ralph J, Lundquist K, Brunow G, Lu F, Kim H, Schatz PF, Marita JM, Hatfield RD, Ralph SA, Christensen JH, Boerjan W (2004) Lignins: natural polymers from oxidative coupling of 4-hydroxyphenylpropanoids. Phytochem Rev 3:29–60. https://doi.org/10.1023/B:PHYT.0000047809.65444.a4
Reiter J, Strittmatter H, Wiemann LO, Schieder D, Sieber V (2013) Enzymatic cleavage of lignin β-O-4 aryl ether bonds via net internal hydrogen transfer. Green Chem 15:1373–1381. https://doi.org/10.1039/C3GC40295A
Rinaldi R, Jastrzebski R, Clough MT, Ralph J, Kennema M, Bruijnincx PC, Weckhuysen BM (2016) Paving the way for lignin valorisation: recent advances in bioengineering, biorefining and catalysis. Angew Chem Int Ed Engl 55:8164–8215. https://doi.org/10.1002/anie.201510351
Sato Y, Moriuchi H, Hishiyama S, Otsuka Y, Oshima K, Kasai D, Nakamura M, Ohara S, Katayama Y, Fukuda M, Masai E (2009) Identification of three alcohol dehydrogenase genes involved in the stereospecific catabolism of arylglycerol-β-aryl ether by Sphingobium sp. strain SYK-6. Appl Environ Microbiol 75:5195–5201. https://doi.org/10.1128/AEM.00880-09
Tanamura K, Abe T, Kamimura N, Kasai D, Hishiyama S, Otsuka Y, Nakamura M, Kajita S, Katayama Y, Fukuda M, Masai E (2011) Characterization of the third glutathione S-transferase gene involved in enantioselective cleavage of the β-aryl ether by Sphingobium sp. strain SYK-6. Biosci Biotechnol Biochem 75:2404–2407. https://doi.org/10.1271/bbb.110525
vom Stein T, Grande PM, Kayser H, Sibilla F, Leitner W, Domínguez de María P (2011) From biomass to feedstock: one-step fractionation of lignocellulose components by the selective organic acid-catalyzed depolymerization of hemicellulose in a biphasic system. Green Chem 13:1772–1777. https://doi.org/10.1039/C1GC00002K
Wang C, Quyang X, Su S, Liang X, Zhang C, Wang W, Yuan Q, Li Q (2016) Effect of sulfonated lignin on enzymatic activity of the ligninolytic enzymes Cα-dehydrogenase LigD and β-etherase LigF. Enzym Microb Technol 93:59–69. https://doi.org/10.1016/j.enzmictec.2016.07.008
Zakzeski J, Bruijnincx PCA, Jongerius AL, Weckhuysen BM (2010) The catalytic valorization of lignin for the production of renewable chemicals. Chem Rev 110:3552–3599. https://doi.org/10.1021/cr900354u
Funding
This work was financially supported by the Ministry for Science and Culture of Lower Saxony, Germany.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Husarcíková, J., Voß, H., Domínguez de María, P. et al. Microbial β-etherases and glutathione lyases for lignin valorisation in biorefineries: current state and future perspectives. Appl Microbiol Biotechnol 102, 5391–5401 (2018). https://doi.org/10.1007/s00253-018-9040-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9040-3