Abstract
Since the discovery of human leukocyte antigens (HLAs), the function of major histocompatibility complex (MHC) gene families in a wide range of diseases have been the subject of research for decades. In particular, the associations of autoimmune disorders to allelic variants and candidate genes encoding the MHC are well documented. However, despite decades of research, the knowledge of MHC associations with human disease susceptibility have been predominantly studied in European origin, with limited understanding in different populations and ethnic groups. This is particularly evident in countries and ethnic populations of the Arabian Peninsula. Human MHC haplotypes, and its association with diseases, of the variable ethnic groups of this region are poorly studied. This review compiled published manuscripts that have reported a list of autoimmune diseases (insulin-dependent diabetes mellitus, systemic lupus erythematosus, myasthenia gravis, rheumatoid arthritis, psoriasis vulgaris, and multiple sclerosis) associated with MHC class I and class II in the populations of the Arabian Peninsula, specifically Bahrain, Kuwait, Oman, Qatar, Saudi Arabia, the United Arab Emirates, and Yemen. Data available was compared with other three ethnic groups, namely Caucasians, Asians, and Africans. The limited data available in the public domain on the association between MHC gene and autoimmune diseases highlight the challenges in the Middle Eastern region.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the 1950s, work described by Jean Dausset, Jon van Rood, and Rose Payne laid the foundation for what is now known as the human major histocompatibility complex (MHC). The MHC region is one of the most gene-dense regions of the genome that spans 3.6 megabase contiguous sequence (The MHC sequencing consortium 1999) to an expanded 7.6 megabases (Mungall et al. 2003), on chromosome 6p21.3. Members of these MHC gene families are involved in self recognition (human leukocyte antigens (HLAs)), inflammation (tumor necrosis factor (TNF)), and components of the complement cascade (C4, C2 and complement factor B or Bf). Human MHC haplotypes may be compared among individuals and populations in the form of single nucleotide polymorphisms (SNPs), insertion-deletions (indel), as well as gross large-scale differences involving gene copy number differences. An example of these three categories of genetic variation is evident in the complement C4 gene, located in the central region of the MHC. Despite sharing 99% sequence homology, the two forms of this gene, C4A and C4B, are defined by 5 important nucleotide differences in exon 26, which contribute to isotype-specific motifs (Yu 1991; Yu et al. 1986). A HERV-K(C4) insertion in intron 9 of some, but not all, C4 genes gives rise to a long (21 kb C4L) and a short (14.6 kb C4S) form (Dangel et al. 1994; Yu et al. 1986). The complement C4 genes are segmentally duplicated as part of one to four modular RCCX cassettes, such that two to eight copies of the C4 gene may be present in a diploid genome, where each chromosome 6 consisting of one to four copies of a single C4 gene (Fernando et al. 2010).
HLA gene variants were first associated with human diseases in the late 1960s, with the discovery of the susceptibility effects of HLA-B in Hodgkin’s lymphoma (Amiel 1967). Subsequently, many other associations have been established, such as the association between HLA-A2 and acute lymphocytic leukemia (Walford et al. 1970), the classical HLA-B27 association with ankylosing spondylitis (Brewerton et al. 1973; Schlosstein et al. 1973), among others. Since then, associations with other non-HLA-genes have been implicated. For example, the presence of C4 null alleles has been associated with systemic lupus erythematosus (SLE) (Christiansen et al. 1983; Fielder et al. 1983), systemic sclerosis (Briggs et al. 1993), and myasthenia gravis (Franciotta et al. 2001). By comparing conserved extended or ancestral haplotypes of the MHC, some have argued that this C4 null association accounts for only part of the association, and have implicated the possible role of other MHC genes (Christiansen et al. 1991). Over time, the MHC has been recognized as the region of the genome with arguably the greatest number of human disease associations (Trowsdale and Knight 2013). Interestingly, the influence of MHC genes is not merely confined to immune conditions, but also includes susceptibility to neuropathologies, malignancies, and infectious diseases (Howell 2014; Trowsdale 2011). With recent advances in high-throughput DNA-based technologies and increased knowledge of the complexity and extreme polymorphism of the MHC, HLA disease associations have subsequently been reported.
Population Definition for this Review: the Arabian Peninsula
This review focuses on the populations of a region in the Middle East, specifically, the Arabian Peninsula. The land mass of the Arabian Peninsula comprises of seven countries: Bahrain, Kuwait, Oman, Qatar, Saudi Arabia, the United Arab Emirates (UAE), and Yemen. The peninsula is surrounded by the Red and Arabian Seas, as well as the Gulfs of Arden, Oman, and Persia. It has land borders with Egypt to the west, Jordan to the northwest, and Iraq to the northeast. These connect the peninsula to Africa and the northern Levantine region, which leads to Europe and Asia. The population of the Arabian Peninsula, which is often referred to as the population of Arabian Gulf, consists of diverse ethnic groups, with influences from neighboring regions. The first inhabitants were the Bedouins, a group that dates back to at least 1200 BC. This group travelled widely throughout the Arabian Peninsula in search of water and grazing pastures, and their caravans were used by travelers from surrounding regions to traverse through the region. Through trade, populations from North and sub-Saharan Africa, the Levant, and Southern or Central Asia intermingled with the locals, with some remaining within the gulf region. A genetic study of the present-day population revealed that genetic variants of ethnic groups can be traced to Central Asia (Al-Jenaidi et al.), Southern Asia (Baluchistan), and North Africa (Iberia and Egypt) (Hajjej et al. 2018; Tay et al. 2020).
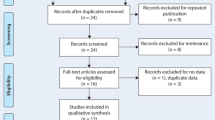
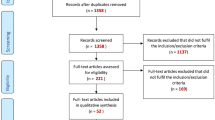
A genome-wide association analyses, based on systematically dispersed SNPs across the genomes of 1000 citizens of the UAE, has provided some insight into the admixture that has occurred over history in the south eastern corner of this gulf region, with genetic influences from the Central, Southern, and sub-Saharan Africa (Fig. 1) (Tay et al. 2020). Trade and exchange throughout history has been well documented between these three regions, and as expected, there are contributions from populations from North Africa, Central Asia, and Europe through migration of people overland. A secondary route from central Africa across the Bab al-Mandab Strait has also influenced the composition of the population in this region. There has been minimal to no influence from the East Asia, Oceania, and the Americas. Although a generalization, it is not unexpected that the unique composition of the seven countries of the Arabian Peninsula has arisen from the gradual admixture of populations that have inhabited the region through interactions with those from neighboring areas (Fig. 2).
Proportions of ancestral genomes that have accumulated in the contemporary population of the UAE. Data was obtained from the genome wide analysis study of 1000 UAE citizens which provide insights into the admixture of the contemporary population. The main ancestral contribution can be traced back to South Asia (27%) which can be explained by a history of trade and exchange between the regions (Tay et al. 2020)
Histogram showing the distribution of variants number at chromosome 6 with troughs (low variability) and peaks (high variability) (Tay et al. 2020). Data were obtained from a genome wide association studies on 1,000 UAE citizens. Unsurprisingly, there is a significant peak in the number of variants on the short arm of chromosome 6, specifically around the MHC region. The MHC contains gene families with variable copy numbers. The size of MHC haplotypes can vary according to the gene copy number around the HLA class II and C4/CYP21 regions. The table shows the different gene copy numbers in different MHC ancestral haplotypes (Zhang et al. 1990)
Motivation for this review
The genetic architecture and disease susceptibility among the Arabian Peninsula region, including its MHC, remain poorly characterized (Popejoy and Fullerton 2016). The Clinical Genome Resource Ancestry and Diversity Working Group continues to highlight the importance of developing guidance for incorporating race, ethnicity, and ancestry data for applications in clinical genomics. Nevertheless, there is substantial disparity across populations in the amount of information available on clinically relevant variants in the Genome Aggregation Database (gnomAD) (Popejoy et al. 2018). This highlights the importance of implementing programs to increase the diversity of genome sequencing and clinical genomics, and reduce uncertainty of variant interpretation, especially in those that have been poorly characterized.
Scope of this review
In keeping with the inclusion of race, ethnicity, and ancestry, this review summarizes the association between HLA class I/II genes and autoimmune diseases in populations of the Arabian Peninsula, contrasting available data with other major ethnic groups, including the Caucasian, Asian, and African populations. This review primarily focuses on a list of major autoimmune diseases, specifically insulin-dependent diabetes mellitus, systemic lupus erythematosus, myasthenia gravis, rheumatoid arthritis, psoriasis vulgaris, and multiple sclerosis. It is well established that predispositions to autoimmunity is a result of a certain disease-specific variation at HLA and non-HLA genes, in addition to epigenetic and environmental elements, which lead to the pathophysiology of the disease and its clinical features (Ramos et al. 2015). Furthermore, the already limited data on autoimmune diseases presented challenges in finding literature on associations with non-immune-related diseases.
A comprehensive search was performed on PubMed, Medline, ScienceDirect, and Google Scholar publications from database inception until April 2020. Search terms included a combination of free-text terms or controlled vocabulary terms for variations of “Arab(ian)” or countries within the region of interest (Bahrain, Kuwait, Oman, Qatar, Saudi Arabia, the United Arab Emirates (including UAE) and Yemen) and “disease association(s)” and “MHC and/or HLA”. Studies using different HLA typing methods such as serological, cellular and molecular methods were considered. Only full text articles published in English were included.
The MHC
General structure
The human MHC is an important region of the human genome located on chromosome 6. The boundary of the region can be fairly arbitrary and is dependent on the genes of interest at the exterminates of the region. In 1999, the MHC sequencing consortium quoted a figure of 3.6-megabases (Mb) (The MHC sequencing consortium 1999) from the HSET loci at the centromeric end to the P5-15 pseudogene in the telomeric end. Ten years later, a region of approximately 4 Mb was referred to with the MHC, containing approximately 0.5% (> 150) of the estimated 32,000 known protein coding genes in the genome (Shiina et al. 2009). Depending on the haplotype, it is reasonable to suggest that the MHC spans a 4 megabase stretch on the short arm of chromosome 6 and contains genes that are involved in numerous immune-related process (Shiina et al. 2009), as well as transplantation (Ketheesan et al. 1999; Nakamura et al. 2019; Tay et al. 1995a, b; Zou and Stastny 2009).
The detailed organization of the human MHC published in 2009 show a region that is densely packed and highly polymorphic, with multigene families (Shiina et al. 2009). One example is the HLA class II gene cluster, where each haplotype contains one copy of the HLA-DRA gene but the number of HLA-DRB genes are variable. Specifically, 9 different HLA-DRB genes have been identified in the HLA class II region, some are expressed (HLA-DRB1, HLA-DRB3, HLA-DRB4, and HLA-DRB5) and others are pseudogenes (Shiina et al. 2009). Not all HLA-DRB genes are found on a haplotype. Within the organizational structure around the C4/CYP21 region, a module known as RCCX, containing sequences of CYP21, TNX, and C4 is duplicated. However, a haplotype can contain up to four RCCX modules (Fernando et al. 2010; Traherne 2008).
These multigene families are involved in an array of different functions. Some are involved in signaling, whereas others are transcription factors of the inflammatory response. Other loci in this region encode for proteins involved in antigen processing and presentation during the adaptive immune response. Further, the region carries genes that encode for components that interact with Natural Killer (NK) cells and cytokines, as well as components of the complement cascade (Shiina et al. 2009).
Genes of the MHC
Genes across the MHC region encode proteins that are important for self-recognition (e.g., HLA), inflammation (e.g., TNF) and the complement cascade (e.g., C4, C2, and Bf). Many of these genes tend to be highly polymorphic, some of which are duplicated and present in multiple copies (Dawkins et al. 1999).
The highly polymorphic classical class I genes (HLA-A, HLA-B, HLA-C) encode peptide binding receptors on the cell surface of all nucleated cells. The class I heterodimeric cell surface receptor platform contains two polypeptide heavy α chains and a β-2 microglobulin, where a peptide-binding groove is located between the α1 and α2 domains. These receptors are responsible for presenting peptide fragments of non-self-antigens to cytotoxic CD8+ T cell receptors (TCRs). The activation of the T cells could lead to tolerance, if the presented peptides are identified as “self,” initiation of cell-mediated immunity in case of “non-self” antigens, or to a maladaptive autoimmune response in the presented “self” peptide was misidentified.
On the other hand, the non-classical class I genes such as HLA-F, HLA-G, and HLA-E are characterized by limited levels of polymorphisms and lower cell surface expression compared to the classical class I genes (Shiina et al. 2009). Interspersed between the classical HLA class I genes are loci that encode for the MHC class I chain related (MIC) proteins, MICA and MICB (Bahram et al. 1994; Leelayuwat et al. 1994). These proteins do not appear to have a role in antigen presentation but act as a stress-induced self-antigen that is recognized by γδ T cells, natural killer (NK), and CD8+ T cells (Ghadially et al. 2017).
The classical HLA class II multigene family (HLA-DR, HLA-DQ, and HLA-DP) are expressed on the surface of antigen-presenting cells (e.g., dendritic cells, macrophages, and B cells), involved in presenting peptides to CD4+ T cells (Wieczorek et al. 2017). The HLA class II molecules include two polypeptide chains, an α and a β chain (Alfonso and Karlsson 2000). There are a number of non-classical HLA class II loci, which include HLA-DM and HLA-DO. HLA-DM is involved in peptide loading of class II molecules in the endosomal/lysosomal system, whereas HLA-DO acts as a chaperon for antigenic peptides (Alfonso and Karlsson 2000).
Flanked by the HLA class I and HLA class II regions, the central MHC also carries a number of multigene families including complement factor genes (C2, C4, and CFB), cytokine genes (TNF, LTA and LTB) (Shiina et al. 2009), and other genes that are not linked to the immune function or inflammation. Many of these genes are duplicated and polymorphic (Zhang et al. 1990). The TNF is a multifunctional cytokine, secreted by macrophages and T lymphocytes, involved in systematic inflammation. It regulates immune cells and protects against tumors. The three complement genes (C2, C4, and Bf) in the central portion of the MHC encode for proteins involved in the early stages of the complement cascade. As mentioned previously, the actual number of C4 genes can vary from one to four per haplotype or two to eight per diploid genome (Bánlaki et al. 2012; Castley and Martinez 2012; Fernando et al. 2008) and it is dependent on the ancestral haplotype (AH) (Zhang et al. 1990). In this regard, Zhang et al. (1990) has suggested that relative deficiencies of the products of the central MHC genes could impact the susceptibility to autoimmune diseases (Zhang et al. 1990).
The extensive degree of polymorphism of the MHC genes found in humans at the population level is thought to contribute to the success of displaying a broad range of non-self peptides to the T cell antigen receptor. These variants are partly caused by point mutations or gene conversion. Variation in HLA genes that produce amino acid changes, including those in the peptide-binding groove of the MHC molecules, can alter the binding preferences, leading to a different peptide-binding repertoire (Dendrou et al. 2018). If a change in the HLA molecule provides fitness advantage to its host, it will be maintained in the population. Hence, it is not surprising that many HLA genes are associated with hundreds of diseases, owing to their polymorphic nature and role in immune response initiation (van Deutekom and Keşmir 2015).
Interestingly, an HLA molecule could increase the likelihood of developing a particular disease, while a closely related HLA molecule cannot. For instance, HLA-B*27:05 has been found to cause ankylosing spondylitis, while HLA-B*27:09 does not have an effect in the disease predisposition (Fiorillo et al. 1998). Even though the difference between both alleles is minimal, and mainly at the peptide-binding groove, it produces an entirely different outcome (van Deutekom and Keşmir 2015). HLA-B is known to have the highest number of alleles, but HLA-DR/DQ subregion has been associated with the highest number of diseases (Kishore and Petrek 2018).
The reporting and naming of HLA alleles and haplotypes in this review follows the HLA nomenclature system described by Marsh (Marsh et al. 2010). The asterisk “*” sign indicates that the typing has been performed by a molecular method. The digits before the first colon (field 1) define the type or the allele group. The next set of digits (field 2) denotes the subtype, and the third set of digits is used to show synonymous variants. Many studies reported in this review describe HLA allelic associations to 2-digit (first field) or 4-digit (second field) resolution.
HLA allele diversity and MHC haplotypes
According to the international HLA nomenclature committee, there were 19,031 and 7183 HLA class I and class II alleles, respectively, in December 2019. The most frequent haplotypes observed were the AH 8.1 haplotype (HLA-A1-B8-DR3) in Caucasians; the HLA-A*33:03-B*58:01-DRB1*03:01 combination in Asians, and the HLA-A*30:01-B*42:01-DRB1*03:02 haplotype in Africans (Gourraud et al. 2014). To date, no large-scale study has characterized ancestral haplotypes in the ethnic populations of Arabs. Based on recent studies of HLA frequencies in the population of the UAE, the most frequent HLA-A/HLA-B combinations are HLA-A*02-B*50, HLA-A*02-B*51, HLA-A*26-B*08, HLA-A*11-B*40, and HLA-A*03-B*50 (Kulski et al. 2019). Only two haplotypes from the list of the most frequent haplotypes observed in Caucasians, Asians, and Africans from the 1000-genome project were observed in the top 10 HLA-A-B haplotypes in UAE. In Saudi Arabia, HLA-A*02, HLA-B*50, and HLA-B*51 were the most frequent HLA alleles described in the population; whereas in neighboring Oman, HLA-A*02, HLA-B*08, and HLA-B*50 were the most frequent alleles (Hajeer et al. 2009).
Disease associations and the Arabian MHC
Insulin-dependent diabetes mellitus
Insulin-dependent diabetes mellitus (IDDM), also known as type 1 diabetes (T1D), is an autoimmune disease where pancreatic islet β cells are destroyed by the immune system, resulting in low production of insulin and insulin deficiency. Consequently, the control of blood sugar level is disrupted, leading to complications that causes serious health issues and shortened life expectancy (Polychronakos and Li 2011).
IDDM can affect individuals at any age, but usually develops at a young age making it a common childhood disease. The incidence of the IDDM among children and adolescents in Saudi Arabia, Algeria, and Morocco rank in top 10 countries worldwide. The Middle East and North African (MENA) region ranks third region in terms of number of children and adolescents with IDDM (Nam Han et al. 2017). Epidemiological studies in Arab countries show that there are between 31.4 and 37.1 new cases per 100,000 population (Al-Herbish et al. 2008).
Genome-wide association studies (GWAS) have highlighted the complexity of IDDM and have implicated the involvement of numerous genes in the development of the disease (Noble and Valdes 2011). The insulin (INS), inositol 1-, 4-, 5-trisphosphate receptor type 3 (ITPR3), cytotoxic T-lymphocyte-associated protein 4 (CTLA4), protein tyrosine phosphatase, non-receptor type 22 (PTPN22), and interleukin 2 receptor alpha (IL2RA) genes are non-HLA risk loci that have been confirmed (Steck and Rewers 2011).
The HLA genes represent almost half of the genes found to be associated with IDDM. The association of the disease with HLA class II (HLA-DR, HLA-DQ, and HLA-DP) have been found to be linked to the geographic location or racial background of individuals (Table 1) (Ikegami et al. 2006). Alleles that have a strong predisposition to the disease include HLA-DPB1*04:02, HLA-DPB1*03:01, and HLA-DPB1*02:02 (Steck and Rewers 2011). The strong linkage disequilibrium between the genes within the MHC region makes it difficult to evaluate the direct effect of each gene separately. Interestingly, studies on Caucasian families showed that a number of HLA class I alleles that contributes to the onset of IDDM are also associated with type 2 diabetes (T2D) (Jahromi and Al-Ozairi 2019).
The increasing incidence of IDDM among the populations of the Arabian Peninsula is thought to be due unhealthy lifestyle choices, changes in breastfeeding practices, higher hygiene standards, and factors that contribute to vitamin D deficiency. Moreover, the high rates of endogamous and consanguineous marriages are also believed to have contributed to the higher prevalence of the IDDM and other autoimmune diseases (Jahromi and Al-Ozairi 2019).
A study in Saudi Arabia that was conducted on IDDM patients revealed a significant correlation between the disease and the haplotypes HLA-DRB1*04-DQB1*02:01/02:02. The same haplotype association, known as HLA-DR4, was also found in Caucasian (Noble and Valdes 2011) and Japanese patients (Ikegami et al. 2007). Further, HLA-DPB1*04:01 was found to be significantly protective in the Saudi Arabian population (Al-Hussein et al. 2003).
Studies on the population of Bahrain reported a high frequency (70.1%) of haplotypes HLA-DRB1*03:01:01-DQB1*02:01 and HLA-DRB1*03:01:01-DQB1*03:02 in IDDM patients (Al-Jenaidi et al. 2005). The HLA-DRB1*03:01:01-DQB1*02:01 haplotype (also known as HLA-DR3) was also found to have a great susceptibility to the disease in Caucasians (Noble and Valdes 2011). In contrast, the MHC haplotypes HLA-DRB1*10:01:01-DQB1*05:01:01, HLA-DRB1*11:01:01-DQB1*03:01:01, as well as HLA-DRB1*11:01:01-DQB1*05:01:01 were found to be protective in the patients from Bahrain (Table 1) (Al-Harbi et al. 2004; Al-Jenaidi et al. 2005).
Systemic lupus erythematosus
Systemic lupus erythematosus (SLE) is a chronic multisystem autoimmune disease that exhibits variable phenotypes and severity depending on ethnicity (Adwan 2018). Women are predominantly affected (Feldman et al. 2013). Clinical features include hematological abnormalities, skin, joint diseases, renal disease, and neuropsychiatric complications (Feldman et al. 2013). Although clinical expression studies of SLE in Arabian populations are limited, similar phenotypic features and disease factors have generally been observed when compared to other populations (Adwan 2018).
The prevalence of SLE among populations of the Arabian Gulf region has been shown to be higher than most other ethnicities. Specifically, a recent epidemiological study of the population of the UAE revealed an age standardized rate of 103 per 100,000 population, which is higher than most Caucasian populations (Al Dhanhani et al. 2017).
SLE is a multifactorial disease where genetics, hormonal, and environmental factors are involved (Hachicha et al. 2018). GWAS studies have highlighted over 80 loci with variants that contribute in a cumulative way (Morris et al. 2012; Yeoh et al. 2018) The exact weight of each genetic factor has yet to be determined, but genes of the MHC region seem to be heavily involved. HLA class II genes, particularly HLA-DRB1, HLA-DQB1, and HLA-DQA1, have been reported as the strongest risk factors, and some alleles have been associated with particular clinical phenotypes (Graham et al. 2007; Wadi et al. 2014).
Some of the alleles are not directly linked to the SLE disease but associated to clinical phenotypes of SLE. For example, HLA-DQB1*03 seems to be related with skin manifestations, HLA-DRB1*15 with nephritis, DRB1*10 with hematological manifestations, and HLA-DRB1*11 with neurological manifestations (Ke et al. 2018; Wadi et al. 2014).
A worldwide recurrent association has been observed for MHC class II alleles HLA-DRB1*03:01, HLA-DRB1*15:01, and HLA-DQB1*06:01 (Table 2) (Al-Motwee et al. 2013; Fernando et al. 2008). Additionally, a protective effect has been reported for HLA-DRB1*14:01 (Graham et al. 2007).
Other MHC genes have also been associated to SLE, including genes of the complement cascade (C4A and C4B) and TNF. However, LD between these genes and some HLA haplotypes make the interpretation of their role in SLE difficult (Fernando et al. 2008).
HLA-DRB1*15-DQB1*06 has been confirmed as a risk haplotype in Saudi Arabian SLE patients (Table 2). However, the HLA-DQB1*06 allele frequency (40%) was higher than that of HLA-DRB1*15 (20%), suggesting that HLA-DQB1 might represent an independent association with the disease. Moreover, the study found that alleles HLA-DRB1*16 and HLA-DRB1*04 are protective against the disease (Table 2) (Al-Motwee et al. 2013). Other associations with SLE have also been found within MHC class I, including HLA-A*29 and HLA-B*51 (Al-Motwee et al. 2013).
Some contrasting results have been observed depending on ethnicity. For example, HLA-DRB3 was found to be a protective allele among Saudi Arabians (Wadi et al. 2014), contradicting the studies from other populations including Malaysians (Chai et al. 2012) and Jamaicans (Smikle et al. 2002).
Myasthenia gravis
Myasthenia gravis (MG) is a rare, chronic autoimmune neuromuscular disease that causes weakness in the skeletal muscles. The disease is a result of defects in the transmission of nerve impulses to muscles due to autoantibodies that target molecules at the neuromuscular junction. This causes impairment for the neuromuscular transmission in 80% of the patients (Avidan et al. 2014). The autoantibodies can act opposing to the acetylcholine receptor (AChR), the muscle-specific kinase (MuSK), or the lipoprotein-related protein 4 (LRP4). Moreover, the disease can be categorized based on the age of onset, thymic abnormalities, ocular involvement and type of autoantibodies (Avidan et al. 2014).
The disease can be further classified into early-onset MG (EOMG), where patients are less than 50 years old, and late-onset MG (LOMG), where patients are more than 50 years old. Both EOMG and LOMG have different HLA allele(s) associations and disease characteristics. For example, EOMG is more common in females and LOMG in males (Feng et al. 2019).
The prevalence of MG among European societies has been estimated to be around 1 to 5000 (Berrih-Aknin 2014). On the other hand, there has been no data available on the prevalence of the disease in the population of the Arabian Peninsula to date.
The relatively low prevalence of the disease has made it difficult for GWAS efforts to cover all probable heritability factors and confirm the specific genetic susceptibility factor(s). In addition, the LD between the MHC genes, including HLA, has also made it difficult to predict the exact susceptibility gene.
Other non-HLA genes have been shown to be linked to EOMC, including interleukin-10 (IL-10) and cellular tyrosine phosphatase 22 (PTPN22) on chromosome 1, cytotoxic T cell late antigen 4 (CTLA4) and muscle nicotinic acetylcholine receptor α-subunit (CHRNA-1) on chromosome 2, TNF-α on chromosome 6, and TNFAIP3-interacting protein 1 (TNIP1) on chromosome 5 (Avidan et al. 2014).
In a study by Hajeer et al. (2009) involving patients from Saudi Arabia, the haplotype comprising of HLA-B*08 and HLA-DRB1*03 was found to be frequent in MG patients. Further, HLA-B*08 showed the strongest susceptibility effect and was linked to young age at onset, and with the female gender. Similarly, a recent study performed on Swedish patients with AH8.1 showed that HLA-B*08:01 is the unique risk factor that marked EOMG (Table 3) (Varade et al. 2018).
Interestingly, HLA-B*51 was found to be another risk allele in the population of Saudi Arabia, with an allele frequency of 19.3% in the general population (Hajjej et al. 2018). Another observation was the protective effect of HLA-B*50, a common allele in Saudi Arabians, but rarely found in other ethnicities (Hajeer et al. 2009).
Rheumatoid arthritis
Rheumatoid arthritis (RA) is an autoimmune disease that affects the joints. This long-term disease results in swollen and painful joints, most commonly the wrist and the hands (McInnes and Schett 2017). The annual incidence of RA worldwide is around 3 cases per 10,000 (Saxena et al. 2017). However, the prevalence of RA in populations of the Arabian Peninsula remains uncertain.
Genetic factors, when combined with environmental events, can define the pathogenesis of the disease. Multiple pathways, including environmental factors (e.g., smoking, obesity, and vitamin D) and mucosal microbiome, seem to be involved, resulting in the same clinical phenotype. Low concordance has been observed between twins, suggesting a greater contribution from environmental factors (Firestein and McInnes 2017).
Despite more than 100 non-MHC candidates identified, genes within the MHC have long been regarded as the major genetic risk factor of the disease (Kunz and Ibrahim 2011). The individual contributions of non-MHC genes seem to be low, but when combined with others, have been suggested to increase the risk by interaction (Kurkó et al. 2013; Okada et al. 2014). Among these gene variants, PTPN22 (1p13.2), PAD14 (1p36.13), and CDK6 (7q21.2) are known to play an important role in increasing the risk of the disease (Snir et al. 2014).
Gene(s) within the MHC class II region are considered to be the main genetic contributor, accounting for more than 40% of the genetic risk (Firestein and McInnes 2017). The significant genetic risk of RA has been primarily explained by the shared epitope (SE) hypothesis proposed by Gregersen et al. (1987). SE refers to an amino acid sequence motif with residues 67 to 74 overrepresented by a number of HLA-DRB1 alleles in RA patients. The SE is thought to act as a “signal transduction ligand that interacts with an evolutionarily-conserved receptors” (de Almeida et al. 2011). Their shared amino acid sequences (QKRAA, QRRAA, and RRRAA) are located at positions 70 to 74 of the third hypervariable region of the HLA-DRB1 antigen-binding groove (Madeleine et al. 2010). The amino acid motif with the strongest association is QKRAA, encoded by HLA-DRB1*04:01, followed by QRRAA encoded by HLA-DRB1*04:04, HLA-DRB1*01:01 and HLA-DRB1*04:05. Finally, the rarest of the 3 SE motifs, RRRAA, is encoded by DRB1*10:01 (de Almeida et al. 2011). It has been found that more than 90% of RA patients express at least one of these SE alleles (Firestein and McInnes 2017). Other alleles that exhibit risk to RA include HLA-DRB1*01:02, HLA-DRB1*04:08, HLA-DRB1*04:10, HLA-DRB1*10:01 (Table 4). These alleles share one of the three SE homologous amino acid sequence variants that are associated with increased severity of RA in Caucasians (Madeleine et al. 2010).
The role of these SEs is not fully understood but recent studies have highlighted an increase of efficiency in peptide presentation to T cells for altered citrullinated peptides. This results in production of interleukin 17 (IL-17), interferon gamma (IFN-γ), and anti-citrullinated peptide antibodies (ACPA), which are seen in 80 to 90% of the patients (Firestein and McInnes 2017).
Furthermore, a list of diseases has also been associated to the same SEs including polymyalgia rheumatica, giant cell arteries, IDDM, SLE, autoimmune hepatitis, chronic lymphoid leukemia, and psoriatic arthritis (de Almeida et al. 2011).
A study conducted on RA patients in Saudi Arabia using a polymerase chain reaction sequence specific primers (PCR-SSP) method revealed that the strongest risk allele is HLA-DRB1*04 followed by HLA-DRB1*08 and HLA-DRB1*10, when compared with healthy controls (Al-Swailem et al. 2006). Further subtyping of the HLA-DRB1*04 locus of the patients has showed strong association with 11 subtypes, specifically HLA-DRB1*04:01, HLA-DRB1*04:02, HLA-DRB1*04:03:01, HLA-DRB1*04:03:02, HLA-DRB1*04:04, HLA-DRB1*04:05, HLA-DRB1*04:06, HLA-DRB1*04:08, HLA-DRB1*04:10, HLA-DRB1*04:14, and HLA-DRB1*04:24.
Further comparison has indicated a significantly positive association of the disease to HLA-DRB1*04:05, which is one of the shared epitopes. A negative association of HLA-DRB1*04:03 with RA has also been reported (Table 4) (Al-Swailem et al. 2006). Surprisingly, a significant protective effect of RA has been observed in HLA-DRB1*06 indicating that HLA-DRB1 could have both negative and postive correlation with the disease (Al-Swailem et al. 2006).
A study on 69 children in Kuwait with juvenile Rheumatoid arthritis using PCR-SSP showed a high incidence of HLA-DR3 in patients, when compared to the control group. The study revealed that this was due to the significant contribution of HLA-DRB1*03:07 and HLA-DRB1*03:08 alleles (Table 4) with no other HLA-DR alleles found in this study (Alsaeid et al. 2002).
A recent pan-Arabian RA genetics GWAS study of patients from the UAE, Saudi Arabia, Qatar, Lebanon, and Jordan reported replication of associated HLA alleles to those of European ancestry. Imputation of the HLA alleles and the HLA amino acid polymorphisims have confirmed the significant effect of HLA-DRB1 amino acid position 11 to RA predisposition in Arabs. Moreover, two novel Arab-specific loci reached genome-wide significance in association analyses of RA (Saxena et al. 2017).
Psoriasis vulgaris
Psoriasis vulgaris (PV) is a common hyper proliferative dermatosis that causes the skin to flake particularly around joints. It is characterized by abnormal symmetrical epidermal proliferation accompanied by an inflammatory response. The disease affects around 1% of the world population, but its prevalence in Arabian populations has not been accurately defined (Capon 2017). Potential triggers of PV are believed to involve a combination of genetic and environmental factors including physical trauma, infections, stress, alcohol, smoking, and obesity (Capon 2017).
Genetic studies have revealed more than 30 SNPs associated with PV, affecting both the skin and the immune system. Nevertheless, only two SNPs have been found to cause the disease independently. GWAS have revealed that the MHC region accounts for the highest association signal observed in patients with threshold of P < 5.0 × 10–8 (Ayala-Fontanez et al. 2016). The most significantly associated alleles, being HLA-C*06 and HLA-B*57, have been described by Schmitt-Egenolf et al. (1993). The alleles appear to mark MHC haplotypes with the greatest disease risk. In particular, the association with HLA-C*06 allele is very strong and has been found in more than 60% of psoriasis patients (Table 5) (Ayala-Fontanez et al. 2016). Moreover, the ancestral haplotype (AH), specifically AH57.1 (HLA-C*06-B*57-DRB1*07:01-DQA1*02:01-DQB1*03:03), was the first to be observed in PV with 26 times higher frequency in patients than in controls (Szczerkowska-Dobosz 2005). Further, differential expression of the MICA and MICB in the class I region of the MHC have been found to be potentially implicated in the etiology of the disease (Tay et al. 2000).
A study in Oman conducted on 55 local PV patients showed contradicting disease association results. The HLA-C6 antigen was reported in 27.8% of the patients and 16% in controls, but showed no significant contribution to disease risk (Al-Mamari et al. 2009). Moreover, HLA-B57 was reduced in both groups and no increase was observed for HLA-DR7 in patients. A novel association, with HLA-B52, was reported as a potential risk allele for PV in the Arabian population of Oman. The study also reported a negative association between a number of HLA alleles and the disease including HLA-A68, HLA-DR13, HLA-DR52, and HLA-DQ2 (Table 5) (Al-Mamari et al. 2009). Although the result of this study suggests that there are selective racial and environmental factors contributing to the etiology of the disease the sample size is too small and larger regional efforts will be required.
Multiple sclerosis
Multiple sclerosis (MS) is demyelinating disease of the central nervous system where the immune system attacks the myelin sheath of the neurons. This causes disruptions in sensory, motor, autonomic and neurocognitive functions. The incidence of the disease is approximately tenfold higher risk in the Caucasian population, when compared to other populations from around the world, with a prevalence of approximately 108 per 100,000. Women and young adults with ancestry from Northern and Central Europe were found to be particularly at a higher risk of developing MS (Mohammed 2016). There has been an increase in the number of MS diagnosed in the Arabian Gulf region, rising from 31 to 55 MS cases per 100,000 individuals in a 2016 report (Mohammed 2016).
Around 50 genome-wide screens have been performed to investigate the role of MHC and non-MHC genes in the risk to MS. However, as suggested by Schmidt et al. (2007), MS is not a homogenous disease with a single-locus model, but a collection of diseases with different genetic etiologies (Schmidt et al. 2007).
Possible non-MHC genetic risk of the disease includes the myelin oligonucleotide glycoprotein (MOG), which lies in the MHC class I region (Wadi et al. 2014). It represents a component of the myelin expressed on the outer surface of the sheath that wraps around the neurons and exons. It is believed that when disrupted, it could influence the action potential electrical conduction that might lead to MS. However, its function in the progression of the disease has been questioned in recent years (Bronge et al. 2019).
Case–control and family-based research have revealed that haplotype DRB1*15:01-DQA1*01:02-DQB1*06:02 is strongly associated to MS in different ethnic groups (Schmidt et al. 2007). Interestingly, this haplotype has also been linked to other diseases, including narcolepsy and SLE (Mohammed 2016). In Saudi Arabian patients, HLA-DRB1*15:01, HLA-DQB1*06:02, and HLA-DQB1*06:01 have been shown to confer the highest risk to MS, with HLA-DRB1*15:01 showing an increased risk of previous relapses. The HLA-DRB1*15:01 allele seems to present the largest risk to MS in populations around the world including Caucasians (Patsopoulos et al. 2013), the Chinese (Qiu et al. 2011), and some populations in the Middle East (Galehdari et al. 2018) (Table 6).
On the other hand, the HLA-DQB1*02:03, HLA-DQB1*05:03, and HLA-DQB1*06:14 alleles were not found in the Saudi Arabian MS group (Table 6) (Al Jumah et al. 2018). Further, the same study showed that patients that carried HLA-DRB1*15:01 and HLA-DQB1*06:02 had lower levels of vitamin D in serum when compared to those without these alleles (Al Jumah et al. 2018). The vitamin D effect is of a particular interest as it has been linked to MS susceptibility and the alteration of MHC class II gene expression (Handunnetthi et al. 2010). Specifically, a previous study by Ramagopalan et al. (2009) has revealed that the expression of HLA-DRB1*15:01 may be controlled by vitamin D. More research is required to investigate the interaction between HLA allele expression and vitamin D, especially in the populations of the Arabian Gulf where vitamin D deficiency is common (Singh et al. 2019).
A study conducted on MS patients in Bahrain using tissue typing methods by polymerase chain reaction (PCR) reported a number of HLA associations, suggesting that there might be population-specific genetic factors that contribute to susceptibility or protection to MS among the populations of the Arabian Gulf. The study did not find any significance between HLA-DRB1*15:01 allele and patients from Bahrain. A number of HLA class I alleles, specifically HLA-A19, HLA-A2, HLA-A9, HLA-B35, HLA-B40, and HLA-B5, where found elevated in MS patients, while HLA-A10 showed a protective effect against the disease. Some HLA class II antigens, specifically HLA-DR3, HLA-DR4, and HLA-DR5, were also found to be higher in patients (Table 6) (Al-Nashmi et al. 2018). The associations found in the study differed to those reported in Caucasians but similar to a study on patients in another gulf country, Kuwait (Al-Shammri et al. 2004).
Other MHC associated diseases with no data available in the Arabian Gulf population
Although there are a number of studies demonstrating the involvement of MHC genes in susceptibility to autoimmune diseases for the Arabian Gulf populations, a clear picture is still far from available. There is progress in elucidating the genetic elements of major autoimmune diseases, but many have not been adequately studied in the populations of the Arabian Peninsula region. These include the deficiency in information on Hoshimoto thyroiditis, Addison’s disease, celiac disease, polymyalgia rheumaticia, autoimmune hepatitis, polyglandulat, and Behçet’s disease (Table 7). Some of diseases, such as celiac disease, are very common among the populations of the Arabian Peninsula (Saeed et al. 2017). However, due to the lack of organized national and regional epidemiological studies, the exact levels of prevalence of many of these diseases remain unknown. More effort is needed to improve our understanding of the MHC region and its association with autoimmune diseases in the populations of the Arabian Peninsula, and the surrounding Middle East region.
Conclusion
Published manuscripts on Arabian HLA-disease associations show inconsistent results from one population to another (e.g., IDDM, RA, and MS). The significance of studies involving Arabian populations have been compromised by methodological heterogeneity, due to low sample size and participant selection, effecting the robustness of the results. The studies have also tended not to adjust for co-morbidities, and did not provide information on whether the participants were diagnosed with any other autoimmune diseases, leading to possible statistical heterogeneity and confounding results. Hence, the sample size and study design of the included manuscripts, limited meaningful interpretation of the data.
It is evident that HLA allele and MHC haplotype frequencies vary according to ethnicity and geographic origin. Variation among closely related ethnic groups can be classified by haplotype distribution, LD analysis and different allele frequencies (Hajjej et al. 2018). Manuscripts on the MHC region of the Arabian Peninsula have presented allele or haplotype associations. Yet, the majority of the reports have been based on serological methods and low resolution DNA-based HLA typing. Hence, the accuracy of HLA genotyping is particularly important to confirm disease associations.
Challenges faced by HLA-disease association studies include incomplete penetrance, interactions between genes, epigenetic factors, and environmental influences which all contribute to the clinical manifestation and phenotype severity of the disease, making it difficult to reach a conclusive genetic association (Dendrou et al. 2018). Disease association studies are negatively impacted by the rising number of HLA alleles and the ambiguity from the use of traditional methods (e.g., SSOP), compared to sequencing methods. To overcome these limitations, Next Generation Sequencing (NGS) methods may be used to provide consistent levels of resolution of HLA typing (Brown et al. 2016), and collaborative workshop activities focused on Middle Eastern populations may encourage further research in the region.
As suggested, typing based on NGS should provide the appropriate level of analytical specificity and sensitivity, reduce ambiguity, and provide more accurate allelic information for disease association studies (Profaizer et al. 2020). Publications describing DNA-based HLA typing are increasing, and with the decreasing cost of next generation sequencing technology (Klasberg et al. 2019), there is no doubt that the pool of data and our understanding of MHC disease associations will improve. NGS technology has proven its ability in identifying novel HLA alleles undetected by the clinical Sanger sequencing-based typing (SBT) (Brown et al. 2016). Particularly, NGS has successfully utilized to detect 33 novel alleles in Saudi Arabian potential bone marrow donors (Bishara et al. 2020), despite allelic definitions of HLA in public databases remain limited.
It has been well established that HLA genes are in strong long-range linkage disequilibrium leading to the existence of “conserved extended” (CEH) (Yunis et al. 2003) or “ancestral” (AH) (Degli-Esposti et al. 1992) haplotypes. These MHC haplotypes are not transmitted randomly through generations. Rather, there is transmission of specific MHC genomic subregions comprising hundreds of kilobases in length (Dawkins et al. 1999). Therefore, the susceptibility for diseases should be looked at from the perspective of MHC ancestral haplotypes and their recombinants (Dawkins and Lloyd 2019).
The MHC ancestral haplotype and the genetic architecture of the Arabian Peninsula population is not well documented, if at all. Studying Arabian MHC ancestral haplotypes, is important to provide a reference for the presence and prevalence of HLA haplotypes and alleles in the population (Kishore and Petrek 2018). This will augment research on HLA-disease susceptibility and improve histocompatibility matching for hematopoietic stem cells and bone marrow (Ameen et al. 2020) for patients of this region.
Construction of MHC ancestral haplotypes in the region could be possible through family studies as they improve the statistical power of analyses (Jahromi and Al-Ozairi 2019), and provide the opportunity to dissect the specific MHC region of interest (Dawkins et al. 1999). Finally, this review highlights the need for greater effort into characterizing the MHC region and associated diseases for the Arabian population. This is needed to assist in early diagnosis, future prevention and treatment of the disease, in addition to optimizing the matching of bone marrow donor and recipient pairs, as well as the organ distribution for transplant purposes in the region.
Availability of data and material
This review discuses published data
Change history
05 March 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00251-021-01210-z
References
Adwan M (2018) Clinical and serologic characteristics of systemic lupus erythematosus in the Arab world: a pooled analysis of 3,273 patients. Arch Rheumatol 33:455–463
Al Dhanhani A, Agarwal M, Othman Y, Bakoush O (2017) Incidence and prevalence of systemic lupus erythematosus among the native Arab population in UAE. Lupus 26:664–669
Al Jumah M, Kojan S, Al Shehri AM, Al Balwi M, Al Abdulkarim I, Masuadi EM, Alhaidan Y, Alabdulrahman A, Fakhoury HM, Hajeer AH (2018) HLA class II polymorphism in Saudi patients with multiple sclerosis. HLA 91:17–22
Al-Harbi EM, Abbassi AJ, Tamim H, al-Jenaidi F, Kooheji M, Kamal M, al-Mahroos S, al-Nasir F, Motala AA, Almawi WY (2004) Specific HLA-DRB and -DQB alleles and haplotypes confer disease susceptibility or resistance in Bahraini type 1 diabetes patients. Clin Diagn Lab Immunol 11:292–296
Al-Herbish A, El-Mouzan MI, Al-Salloum AA, Al-Qurachi MM, Al-Omar AA (2008) Prevalence of type 1 diabetes mellitus in Saudi Arabian children and adolescents. Saudi Med J 9:1285–1288
Al-Hussein KA, Rama NR, Ahmad M, Rozemuller E, Tilanus MG (2003) HLA-DPB1*0401 is associated with dominant protection against type 1 diabetes in the general Saudi population and in subjects with a high-risk DR/DQ haplotype. Eur J Immunogenet 30:115–119
Al-Jenaidi FA, Wakim-Ghorayeb SF, Al-Abbasi A, Arekat MR, Irani-Hakime N, Najm P, Al-Ola K, Motala AA, Almawi WY (2005) Contribution of selective HLA-DRB1/DQB1 alleles and haplotypes to the genetic susceptibility of type 1 diabetes among Lebanese and Bahraini Arabs. J Clin Endocrinol Metab 90:5104–5109
Al-Mamari F, Al-Shirawi A, Banodkar D, Al-Hashmi S, Al-Yahyaae F, Varghese M, Raeburn J (2009) HLA antigens in Omani psoriasis vulgaris patients. Oman Med J 24
Al-Motwee S, Jawdat D, Jehani GS, Anazi H, Shubaili A, Sutton P, Uyar AF, Hajeer AH (2013) Association of HLA-DRB1*15 and HLADQB1*06 with SLE in Saudis. Ann Saudi Med 33:229–234
Al-Nashmi M, Taha S, Salem AH, Alsharoqi I, Bakhiet M (2018) Distinct HLA class I and II genotypes and haplotypes are associated with multiple sclerosis in Bahrain. Biomed Rep 9:531–539
Al-Shammri S, Nelson R, Al-Muzairi I, Akanji A (2004) HLA determinants of susceptibility to multiple sclerosis in an Arabian Gulf population. Multiple Sclerosis 10
Al-Swailem R, Al-Rayes H, Sobki S, Arfin M, Tariq M (2006) HLA-DRB1 association in Saudi rheumatoid arthritis patients. Rheumatol Int 26:1019–1024
Alfonso C, Karlsson L (2000) Nonclassical MHC class II molecules. Annu Rev Immunol 18:113–142
Alsaeid K, Haider Z, Kamal H, Srivastva BS, Ayoub EM (2002) Prevalence of human leukocyte antigen (HLA) DRB1 alleles in Kuwaiti children with juvenile rheumatoid arthritis. Eur J Immunogenet 29:1–5
Ameen R, Al Shemmari S, Marsh S (2020) HLA haplotype frequencies and genetic profiles of the Kuwaiti population. Med Princ Pract 29:39–45
Amiel J (1967) Study of the leukocyte phenotypes in Hodgkin’s disease in histocompatibility testing. In: Teraski PI, editor. Histocompatibility testing Munksgaard; Copenhagen 79–81
Avidan N, Le Panse R, Berrih-Aknin S, Miller A (2014) Genetic basis of myasthenia gravis - a comprehensive review. J Autoimmun 52:146–153
Ayala-Fontanez N, Soler DC, McCormick TS (2016) Current knowledge on psoriasis and autoimmune diseases. Psoriasis (Auckl) 6:7–32
Bahram S, Bresnahan M, Geraghty DE, Spies T (1994) A second lineage of mammalian major histocompatibility complex class I genes. Proc Natl Acad Sci U S A 91:6259–6263
Bánlaki Z, Doleschall M, Rajczy K, Fust G, Szilágyi A (2012) Fine-tuned characterization of RCCX copy number variants and their relationship with extended MHC haplotypes. Genes Immun 13:530–535
Berrih-Aknin S (2014) Orphanet: Myasthenia gravis
Bettencourt A, Carvalho C, Leal B, Bras S, Lopes D, Martins da Silva A, Santos E, Torres T, Almeida I, Farinha F, Barbosa P, Marinho A, Selores M, Correia J, Vasconcelos C, Costa PP, da Silva BM (2015) The protective role of HLA-DRB1( *)13 in autoimmune diseases. J Immunol Res 2015:948723
Bishara A, Brautbar C, Cereb N, Israel S (2020) Identification of 33 novel HLA alleles in Arab potential bone marrow donors. HLA 95:128–130
Brewerton D, Hart F, Nicholls A, Caffrey M, James D, Sturrock R (1973) Ankylosing spondylitis and HL-A 27. Lancet 1:904–907
Briggs D, Stephens C, Vaughan R, Welsh K, Black C (1993) A molecular and serologic analysis of the major histocompatibility complex and complement component C4 in systemic sclerosis. Arthritis Rheum 36:943–954
Bronge M, Ruhrmann S, Carvalho-Queiroz C, Nilsson O, Kaiser A, Holmgren E, Macrini C, Winklmeier S, Meinl E, Brundin L, Khademi M, Olsson T, Gafvelin G, Grönlund H (2019) Myelin oligodendrocyte glycoprotein revisited-sensitive detection of MOG-specific T-cells in multiple sclerosis. J Autoimmun 102:38–49
Brown N, Kheradmand T, Wang J, Marino S (2016) Identification and characterization of novel HLA alleles: utility of next-generation sequencing methods. Hum Immunol 77:313–316
Bugawan TL, Klitz W, Alejandrino M, Ching J, Panelo A, Solfelix CM, Petrone A, Buzzetti R, Pozzilli P, Erlich H (2002) The association of specific HLA class I and II alleles with type 1 diabetes among Filipinos. Tissue Antigens 59:452–469
Capon F (2017) The Genetic Basis of Psoriasis. Int J Mol Sci 18
Castley AS, Martinez OP (2012) Molecular analysis of complement component C4 gene copy number. Methods Mol Biol 882:159–171
Chai H, Phipps M, Chua K (2012) Genetic risk factors of systemic lupus erythematosus in the Malaysian population: a mini review. Clin Dev Immunol 2012
Christiansen FT, Dawkins RL, Uko G, McCluskey J, Kay PH, Zilko PJ (1983) Complement allotyping in SLE: association with C4A null. Aust N Z J Med 13:483–488
Christiansen FT, Zhang WJ, Griffiths M, Mallal SA, Dawkins RL (1991) Major histocompatibility complex (MHC) complement deficiency, ancestral haplotypes and systemic lupus erythematosus (SLE): C4 deficiency explains some but not all of the influence of the MHC. J Rheumatol 18:1350–1358
Cid MC, Ercilla G, Vilaseca J, Sanmarti R, Villalta J, Ingelmo M, Urbano‐Marquez A (1988) Polymyalgia rheumatica: a syndrome associated with hla–dr4 antigen. Arthritis Rheum 31
Dangel AW, Mendoza AR, Baker BJ, Daniel CM, Carroll MC, Wu LC, Yu CY (1994) The dichotomous size variation of human complement C4 genes is mediated by a novel family of endogenous retroviruses, which also establishes species-specific genomic patterns among Old World primates. Immunogenetics 40:425–436
Dawkins R, Leelayuwat C, Gaudieri S, Tay G, Hui J, Cattley S, Martinez P, Kulski J (1999) Genomics of the major histocompatibility complex: haplotypes, duplication, retroviruses and disease. Immunol Rev 167:275–304
Dawkins R, Lloyd S (2019) MHC genomics and disease: looking back to go forward. Cells 8:944
de Almeida DE, Ling S, Holoshitz J (2011) New insights into the functional role of the rheumatoid arthritis shared epitope. FEBS Lett 585:3619–3626
Degli-Esposti MA, Leaver AL, Christiansen FT, Witt CS, Abraham LJ, Dawkins RL (1992) Ancestral haplotypes: conserved population MHC haplotypes. Hum Immunol 34:242–252
Dendrou C, Petersen J, Rossjohn J, Fugger L (2018) HLA variation and disease. Nat Rev Immunol 18:325–339
Erlich H, Valdes AM, Noble J, Carlson JA, Varney M, Concannon P, Mychaleckyj JC, Todd JA, Bonella P, Fear AL, Lavant E, Louey A, Moonsamy P (2008) Type 1 Diabetes Genetics C, HLA DR-DQ haplotypes and genotypes and type 1 diabetes risk: analysis of the type 1 diabetes genetics consortium families Diabetes 57 1084 1092
Feldman C, Hiraki L, Liu J, Fischer M, Solomon D, Alarcón G, Winkelmayer W, Costenbader K (2013) Epidemiology and sociodemographics of systemic lupus erythematosus and lupus nephritis among US adults with Medicaid coverage, 2000–2004. Arthritis Rheum 65:753–763
Feng X, Li W, Song J, Liu X, Gu Y, Yan C, Wu H, Xi J, Zhou S, Zhao C (2019) HLA typing using next-generation sequencing for Chinese juvenile- and adult-onset myasthenia gravis patients. J Clin Neurosci 59:179–184
Fernando MM, Boteva L, Morris DL, Zhou B, Wu YL, Lokki ML, Yu CY, Rioux JD, Hollox EJ, Vyse TJ (2010) Assessment of complement C4 gene copy number using the paralog ratio test. Hum Mutat 31:866–874
Fernando MM, Stevens CR, Walsh EC, De Jager PL, Goyette P, Plenge RM, Vyse TJ, Rioux JD (2008) Defining the role of the MHC in autoimmunity: a review and pooled analysis. PLoS Genet 4:e1000024
Fielder AH, Walport MJ, Batchelor JR, Rynes RI, Black CM, Dodi IA, Hughes GR (1983) Family study of the major histocompatibility complex in patients with systemic lupus erythematosus: importance of null alleles of C4A and C4B in determining disease susceptibility. Br Med J (Clin Res Ed) 286:425–428
Fiorillo M, Greco G, Maragno M, Potolicchio I, Monizio A, Dupuis M, Sorrentino R (1998) The naturally occurring polymorphism Asp116–>His116, differentiating the ankylosing spondylitis-associated HLA-B*2705 from the non-associated HLA-B*2709 subtype, influences peptide-specific CD8 T cell recognition. Eur J Immunol 28:2508–2516
Firestein GS, McInnes IB (2017) Immunopathogenesis of rheumatoid arthritis. Immunity 46:183–196
Franciotta D, Cuccia M, Dondi E, Piccolo G, Cosi V (2001) Polymorphic markers in MHC class II/III region: a study on Italian patients with myasthenia gravis. J Neurol Sci 190:11–16
Galehdari H, Mohaghegh M, Majdinasab N, Khatami SR, Hosseini Behbahani M (2018) Analysis of HLA-A*03 in multiple sclerosis patients in Khuzestan Province. Gene, Cell and Tissue In Press, Iran
Ghadially H, Brown L, Lloyd C, Lewis L, Lewis A, Dillon J, Sainson R, Jovanovic J, Tigue NJ, Bannister D, Bamber L, Valge-Archer V, Wilkinson RW (2017) MHC class I chain-related protein A and B (MICA and MICB) are predominantly expressed intracellularly in tumour and normal tissue. Br J Cancer 116:1208–1217
Gough S, Simmonds M (2007) The HLA region and autoimmune disease: associations and mechanisms of action. Curr Genomics 8:453–465
Gourraud P-A, Khankhanian P, Cereb N, Yang SY, Feolo M, Maiers M, Rioux JD, Hauser S, Oksenberg J (2014) HLADiversity in the 1000 Genomes Dataset. PloS One 8
Graham RR, Ortmann W, Rodine P, Espe K, Langefeld C, Lange E, Williams A, Beck S, Kyogoku C, Moser K, Gaffney P, Gregersen PK, Criswell LA, Harley JB, Behrens TW (2007) Specific combinations of HLA-DR2 and DR3 class II haplotypes contribute graded risk for disease susceptibility and autoantibodies in human SLE. Eur J Hum Genet 15:823–830
Gregersen P, Silver J, Winchester R (1987) The shared epitope hypothesis. An approach to understanding the molecular genetics of susceptibility to rheumatoid arthritis. Arthritis Rheum 30:1205–1213
Hachicha H, Kammoun A, Mahfoudh N, Marzouk S, Sawsan F, Fakhfakh R, Fourati H, Haddouk S, Frikha F, Gaddour L, Hakim F, Bahloul Z, Makni H, Masmoudi H (2018) Human leukocyte antigens-DRB1*03 is associated with systemic lupus erythematosus and anti-SSB production in South Tunisia. Int J Health Sci (Qassim) 12(1):27
Hajeer AH, Sawidan FA, Bohlega S, Saleh S, Sutton P, Shubaili A, Tahan AA, Al Jumah M (2009) HLA class I and class II polymorphisms in Saudi patients with myasthenia gravis. Int J Immunogenet 36:169–172
Hajjej A, Almawi WY, Arnaiz-Villena A, Hattab L, Hmida S (2018) The genetic heterogeneity of Arab populations as inferred from HLA genes. PLoS ONE 13:e0192269
Handunnetthi L, Ramagopalan SV, Ebers GC (2010) Multiple sclerosis, vitamin D, and HLA-DRB1*15. Neurology 74:1905–1910
Howell WM (2014) HLA and disease: guilt by association. Int J Immunogenet 41:1–12
Ikegami H, Fujisawa T, Kawabata Y, Noso S, Ogihara T (2006) Genetics of type 1 diabetes: similarities and differences between Asian and Caucasian populations. Ann N Y Acad Sci 1079:51–59
Ikegami H, Kawabata Y, Noso S, Fujisawa T, Ogihara T (2007) Genetics of type 1 diabetes in Asian and Caucasian populations. Diabetes Res Clin Pract 77(Suppl 1):S116–S121
Jahromi M, Al-Ozairi E (2019) Human leukocyte antigen (HLA) and islet autoantibodies are tools to characterize type 1 diabetes in Arab countries: emphasis on Kuwait. Disease Markers 2019
Jun KR, Choi SE, Cha CH, Oh HB, Heo YS, Ahn HY, Lee KJ (2007) Meta-analysis of the association between HLA-DRB1 allele and rheumatoid arthritis susceptibility in Asian populations. J Korean Med Sci 22:973–980
Ke X, Wen‐Quan N, Yong C (2018) Association of HLA‐DR3 and HLA‐DR15 polymorphisms with risk of systemic lupus erythematosus. CN Med J 131
Ketheesan N, Tay GK, Witt CS, Christiansen FT, Taylor RR, Dawkins RL (1999) The significance of HLA matching in cardiac transplantation. J Heart Lung Transplant 18:226–230
Kishore A, Petrek M (2018) Next-generation sequencing based HLA typing: deciphering immunogenetic aspects of sarcoidosis. Front Gen 9
Klasberg S, Surendranath V, Lange V, Schöfl G (2019) Bioinformatics strategies, challenges, and opportunities for next generation sequencing-based HLA genotyping. Transfusion Medicine and Hemotherapy 46:312–325
Kulski J, AlSafar HS, Mawart A, Henschel A, Tay G (2019) HLA class I allele lineages and haplotype frequencies in Arabs of the United Arab Emirates International. J Immunogenetics 46
Kunz M, Ibrahim S (2011) Non-major histocompatibility complex rheumatoid arthritis susceptibility genes. Crit Rev Immunol 31:99–114
Kurkó J, Besenyei T, Laki J, Glant T, Mikecz K, Szekanecz Z (2013) Genetics of rheumatoid arthritis - a comprehensive review. Clin Rev Allergy Immunol 45:170–179
Leelayuwat C, Townend DC, Degli-Esposti MA, Abraham LJ, Dawkins RL (1994) A new polymorphic and multicopy MHC gene family related to nonmammalian class I. Immunogenetics 40:339–351
Madeleine S-N, Axel F, Sylvette B, Jean-Marie T, Cem G (2010) Diagnostic value of anti-cyclic citrullinated peptides and association with HLA-DRB1 shared epitope alleles in African rheumatoid arthritis patients. Arthritis Research & Therapy 12
Marsh S, Albert E, Bodmer W, Bontrop R, Dupont B, Erlich H, Fernández-Viña M, Geraghty D, Holdsworth R, Hurley C, Lau M, Lee K, Mach B, Maiers M, Mayr W, Müller C, Parham P, Petersdorf E, Sasazuki T, Strominger J, Svejgaard A, Terasaki P, Tiercy J, Trowsdale J (2010) Nomenclature for factors of the HLA system, 2010. Tissue Antigens 75:291–455
McInnes IB, Schett G (2017) Pathogenetic insights from the treatment of rheumatoid arthritis. Lancet 389:2328
Mohammed EMA (2016) Multiple sclerosis is prominent in the Gulf states: Review. Pathogenesis 3:19–38
Morris DL, Taylor KE, Fernando MM, Nititham J, Alarcon-Riquelme ME, Barcellos LF, Behrens TW, Cotsapas C, Gaffney PM, Graham RR, Pons-Estel BA, Gregersen PK, Harley JB, Hauser SL, Hom G, International MHC, Autoimmunity Genetics N, Langefeld CD, Noble JA, Rioux JD, Seldin MF, Systemic Lupus Erythematosus Genetics C, Criswell LA, Vyse TJ (2012) Unraveling multiple MHC gene associations with systemic lupus erythematosus: model choice indicates a role for HLA alleles and non-HLA genes in Europeans. Am J Hum Genet 91:778–793
Mungall AJ, Palmer SA, Sims SK, Edwards CA, Ashurst JL, Wilming L, Jones MC, Horton R, Hunt SE, Scott CE, Gilbert JG, Clamp ME, Bethel G, Milne S, Ainscough R, Almeida JP, Ambrose KD, Andrews TD, Ashwell RI, Babbage AK, Bagguley CL, Bailey J, Banerjee R, Barker DJ, Barlow KF, Bates K, Beare DM, Beasley H, Beasley O, Bird CP, Blakey S, Bray-Allen S, Brook J, Brown AJ, Brown JY, Burford DC, Burrill W, Burton J, Carder C, Carter NP, Chapman JC, Clark SY, Clark G, Clee CM, Clegg S, Cobley V, Collier RE, Collins JE, Colman LK, Corby NR, Coville GJ, Culley KM, Dhami P, Davies J, Dunn M, Earthrowl ME, Ellington AE, Evans KA, Faulkner L, Francis MD, Frankish A, Frankland J, French L, Garner P, Garnett J, Ghori MJ, Gilby LM, Gillson CJ, Glithero RJ, Grafham DV, Grant M, Gribble S, Griffiths C, Griffiths M, Hall R, Halls KS, Hammond S, Harley JL, Hart EA, Heath PD, Heathcott R, Holmes SJ, Howden PJ, Howe KL, Howell GR, Huckle E, Humphray SJ, Humphries MD, Hunt AR, Johnson CM, Joy AA, Kay M, Keenan SJ, Kimberley AM, King A, Laird GK, Langford C, Lawlor S, Leongamornlert DA, Leversha M et al (2003) The DNA sequence and analysis of human chromosome 6. Nature 425:805–811
Nakamura T, Shirouzu T, Nakata K, Yoshimura N, Ushigome H (2019) The role of major histocompatibility complex in organ transplantation- donor specific anti-major histocompatibility complex antibodies analysis goes to the next stage. Int J Mol Sci 20
Nam Han C, Joses K, Jean Claude M, Katherine O, Leonor G, Wolfgang R, Gojka R, Nita F, Rana D, Alireza E, Edward B, Ian H, Otaliba Ln, de Morais N, Pablo Aschner M, Joshi S, Juliana C, Jonathan S, Samuels TA, Pavkov M, Reja A (2017) IDF DIABETES ATLAS. 8 edn. International Diabetes Federation
Noble JA, Valdes AM (2011) Genetics of the HLA region in the prediction of type 1 diabetes. Curr Diab Rep 11:533–542
Okada Y, Wu D, Trynka G, Raj T, Terao C, Ikari K, Kochi Y, Ohmura K, Suzuki A, Yoshida S, Graham RR, Manoharan A, Ortmann W, Bhangale T, Denny JC, Carroll RJ, Eyler AE, Greenberg JD, Kremer JM, Pappas DA, Jiang L, Yin J, Ye L, Su DF, Yang J, Xie G, Keystone E, Westra HJ, Esko T, Metspalu A, Zhou X, Gupta N, Mirel D, Stahl EA, Diogo D, Cui J, Liao K, Guo MH, Myouzen K, Kawaguchi T, Coenen MJ, van Riel PL, van de Laar MA, Guchelaar HJ, Huizinga TW, Dieude P, Mariette X, Bridges SL Jr, Zhernakova A, Toes RE, Tak PP, Miceli-Richard C, Bang SY, Lee HS, Martin J, Gonzalez-Gay MA, Rodriguez-Rodriguez L, Rantapaa-Dahlqvist S, Arlestig L, Choi HK, Kamatani Y, Galan P, Lathrop M, consortium R, consortium G, Eyre S, Bowes J, Barton A, de Vries N, Moreland LW, Criswell LA, Karlson EW, Taniguchi A, Yamada R, Kubo M, Liu JS, Bae SC, Worthington J, Padyukov L, Klareskog L, Gregersen PK, Raychaudhuri S, Stranger BE, De Jager PL, Franke L, Visscher PM, Brown MA, Yamanaka H, Mimori T, Takahashi A, Xu H, Behrens TW, Siminovitch KA, Momohara S, Matsuda F, Yamamoto K, Plenge RM (2014) Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506:376–381
Oliveira LC, Porta G, Marin ML, Bittencourt PL, Kalil J, Goldberg AC (2011) Autoimmune hepatitis, HLA and extended haplotypes. Autoimmun Rev 10:189–193
Patsopoulos NA, Barcellos LF, Hintzen RQ, Schaefer C, van Duijn CM, Noble JA, Raj T, Imsgc A, Gourraud PA, Stranger BE, Oksenberg J, Olsson T, Taylor BV, Sawcer S, Hafler DA, Carrington M, De Jager PL, de Bakker PI (2013) Fine-mapping the genetic association of the major histocompatibility complex in multiple sclerosis: HLA and non-HLA effects. PLoS Genet 9:e1003926
Pirie FJ, Hammond MG, Motala AA, Omar MAK (2001) HLA class II antigens in South African Blacks with type I diabetes. Tissue Antigens 57:348–352
Polychronakos C, Li Q (2011) Understanding type 1 diabetes through genetics: advances and prospects. Nat Rev Genet 12:781–792
Popejoy AB, Fullerton SM (2016) Genomics is failing on diversity. Nature 538:161–164
Popejoy AB, Ritter DI, Crooks K, Currey E, Fullerton SM, Hindorff LA, Koenig B, Ramos EM, Sorokin EP, Wand H, Wright MW, Zou J, Gignoux CR, Bonham VL, Plon SE, Bustamante CD, Clinical Genome Resource A, Diversity Working G (2018) The clinical imperative for inclusivity: race, ethnicity, and ancestry (REA) in genomics. Hum Mutat 39:1713–1720
Prinz J (2018) Human leukocyte antigen-class I alleles and the autoreactive T cell response in psoriasis pathogenesis. Front Immunol 9:954
Profaizer T, Pole A, Monds C, Delgado J, Lázár-Molnár E (2020) Clinical utility of next generation sequencing based HLA typing for disease association and pharmacogenetic testing. Hum Immunol 81:354–360
Qiu W, James I, Carroll WM, Mastaglia FL, Kermode AG (2011) HLA-DR allele polymorphism and multiple sclerosis in Chinese populations: a meta-analysis. Mult Scler 17:382–388
Ramagopalan SV, Maugeri NJ, Handunnetthi L, Lincoln MR, Orton SM, Dyment DA, Deluca GC, Herrera BM, Chao MJ, Sadovnick AD, Ebers GC, Knight JC (2009) Expression of the multiple sclerosis-associated MHC class II Allele HLA-DRB1*1501 is regulated by vitamin D. PLoS Genet 5:e1000369
Ramos P, Shedlock A, Langefeld C (2015) Genetics of autoimmune diseases: insights from population genetics. J Hum Genet 60:657–664
Saeed A, Assiri A, Assiri H, Ullah A, Rashid M (2017) Celiac disease in Saudi children. Evaluation of clinical features and diagnosis. Saudi Med J 38:895–899
Saxena R, Plenge RM, Bjonnes AC, Dashti HS, Okada Y, Gad El Haq W, Hammoudeh M, Al Emadi S, Masri BK, Halabi H, Badsha H, Uthman IW, Margolin L, Gupta N, Mahfoud ZR, Kapiri M, Dargham SR, Aranki G, Kazkaz LA, Arayssi T (2017) A multinational Arab genome-wide association study identifies new genetic associations for rheumatoid arthritis. Arthritis Rheumatol 69:976–985
Schlosstein L, Terasaki P, Bluestone R, Pearson C (1973) High association of an HL-A antigen, W27, with ankylosing spondylitis. N Engl J Med 288:704–706
Schmidt H, Williamson D, Ashley-Koch A (2007) HLA-DR15 haplotype and multiple sclerosis: a HuGE review. Am J Epidemiol 165:1097–1109
Schmitt-Egenolf M, Boehncke H, Stander M, Eiermann T, Sterry W (1993) Oligonucleotide typing reveals associa- tion of type I psoriasis with the HLA- DRB1*0701/2-DQA1*0201, -DQB1*0303 extended haplotype. J Invest Dermatol 100:749–752
Shiina T, Hosomichi K, Inoko H, Kulski JK (2009) The HLA genomic loci map: expression, interaction, diversity and disease. J Hum Genet 54:15–39
Singh P, Kumar M, Al Khodor S (2019) Vitamin D deficiency in the Gulf Cooperation Council: exploring the triad of genetic predisposition, the gut microbiome and the immune system. Front Immunol 10:1042
Sirikong M, Tsuchiya H, Chandanayingyong D, Bejrachandra S, Suthipinittharm P, Luangtrakool K, Srinak D, Thongpradit R, Siriboonrit U, Tokunaga K (2002) Association of HLA-DRB1*1502 – DQB1*0501 haplotype with susceptibility to systemic lupus erythematosus in Thais. Tissue Antigens 59:113–117
Smikle M, Christian N, DeCeulaer K, Bartom E, Roye-Green K, Dowe G, Anderson N, Nicholson G (2002) HLA-DRB alleles and systemic lupus erythematosus in Jamaicans. South Med J 95:717–719
Snir O, Gomez-Cabrero D, Montes A, Perez-Pampin E, Gómez-Reino J, Seddighzadeh M, Klich K, Israelsson L, Ding B, Catrina A, Holmdahl R, Alfredsson L, Klareskog L, Tegnér J, Gonzalez A, Malmström V, Padyukov L (2014) Non-HLA genes PTPN22, CDK6 and PADI4 are associated with specific autoantibodies in HLA-defined subgroups of rheumatoid arthritis. Arthritis Res Ther 16:414
Steck AK, Rewers MJ (2011) Genetics of type 1 diabetes. Clin Chem 57:176–185
Szczerkowska−Dobosz A, (2005) Human leukocyte antigens as psoriasis inheritance and susceptibility markers. Arch Immunol Ther Exp 53:428–433
Tay GK, Henschel A, Daw Elbait G, Al Safar HS (2020) Genetic Diversity and Low Stratification of the Population of the United Arab Emirates. Frontiers in Genetics 11
Tay GK, Hui J, Gaudieri S, Schmitt-Egenolf M, Martinez OP, Leelayuwat C, Williamson JF, Eiermann TH, Dawkins RL (2000) PERB11 (MIC): a polymorphic MHC gene is expressed in skin and single nucleotide polymorphisms are associated with psoriasis. Clin Exp Immunol 119:553–558
Tay GK, Witt CS, Christiansen FT, Charron D, Baker D, Herrmann R, Smith LK, Diepeveen D, Mallal S, McCluskey J et al (1995) Matching for MHC haplotypes results in improved survival following unrelated bone marrow transplantation. Bone Marrow Transplant 15:381–385
Tay GK, Witt CS, Christiansen FT, Corbett JM, Dawkins RL (1995) The identification of MHC identical siblings without HLA typing. Exp Hematol 23:1655–1660
The MHC sequencing consortium (1999) Complete sequence and gene map of a human major histocompatibility complex. The MHC sequencing consortium Nature 401:921–923
Traherne JA (2008) Human MHC architecture and evolution: implications for disease association studies. Int J Immunogenet 35:179–192
Trowsdale J (2011) The MHC, disease and selection. Immunol Lett 137:1–8
Trowsdale J, Knight JC (2013) Major histocompatibility complex genomics and human disease. Annu Rev Genomics Hum Genet 14:301–323
van Deutekom H, Keşmir C (2015) Zooming into the binding groove of HLA molecules: which positions and which substitutions change peptide binding most? Immunogenetics 67:425–436
van Drongelen V, Holoshitz J (2017) Human leukocyte antigen-disease associations in rheumatoid arthritis. Rheum Dis Clin North Am 43:363–376
Varade J, Wang N, Lim CK, Zhang T, Zhang Y, Liu X, Piehl F, Matell R, Cao H, Xu X, Hammarstrom L (2018) Novel genetic loci associated HLA-B*08:01 positive myasthenia gravis. J Autoimmun 88:43–49
Wadi W, Noor eldeen A, Essam M, Adel A, Khaled H, Hamad A, Gasim G (2014) Relation between HLA typing and clinical presentations in systemic lupus erythematosus patients in Al-Qassim region, Saudi Arabia. Int J Health Sci 8
Walford R, Finkelstein S, Neerhout R, Konrad P, Shanbrom E (1970) Acute childhood leukaemia in relation to the HL–A human transplantation genes. Nature 225:461–462
Wallace GR (2014) HLA-B*51 the primary risk in Behcet disease. Proc Natl Acad Sci U S A 111:8706–8707
Weinstock C, Matheis N, Barkia S, Haager MC, Janson A, Markovic A, Bux J, Kahaly GJ (2011) Autoimmune polyglandular syndrome type 2 shows the same HLA class II pattern as type 1 diabetes. Tissue Antigens 77:317–324
Wieczorek M, Abualrous ET, Sticht J, Álvaro-Benito M, Stolzenberg S, Noé F, Freund C (2017) Major Histocompatibility Complex (MHC) Class I and MHC Class II Proteins: Conformational Plasticity in Antigen Presentation. Frontiers in Immunology 8
Xie YC, Qu Y, Sun L, Li HF, Zhang H, Shi HJ, Jiang B, Zhao Y, Qiao SS, Wang SH, Wang DX (2011) Association between HLA-DRB1 and myasthenia gravis in a northern Han Chinese population. J Clin Neurosci 18:1524–1527
Yeoh SA, Dias SS, Isenberg DA (2018) Advances in systemic lupus erythematosus. Medicine 46:84–92
Yu CY (1991) The complete exon-intron structure of a human complement component C4A gene. DNA sequences, polymorphism, and linkage to the 21-hydroxylase gene. J Immunol 146:1057–1066
Yu CY, Belt KT, Giles CM, Campbell RD, Porter RR (1986) Structural basis of the polymorphism of human complement components C4A and C4B: gene size, reactivity and antigenicity. EMBO J 5:2873–2881
Yunis EJ, Larsen CE, Fernandez-Viña M, Awdeh ZL, Romero T, Hansen JA, Alper CA (2003) Inheritable variable sizes of DNA stretches in the human MHC: conserved extended haplotypes and their fragments or blocks. Tissue Antigens 62:1–20
Zayed H (2016) Genetic epidemiology of type 1 diabetes in the 22 Arab countries. Curr Diab Rep 16:37
Zhang WJ, Degli-Esposti MA, Cobain TJ, Cameron PU, Christiansen FT, Dawkins RL (1990) Differences in gene copy number carried by different MHC ancestral haplotypes. Quantitation after physical separation of haplotypes by pulsed field gel electrophoresis. J Exp Med 171:2101–2114
Zhang X-J, Zhang A-P, Yang S, Gao M, Wei S-C, He P-P, Wang H-Y, Song Y-X, Cui Y, Chen J-J (2003) Association of HLA class I alleles with psoriasis vulgaris in southeastern Chinese Hans. J Dermatol Sci 33:1–6
Zou Y, Stastny P (2009) The role of major histocompatibility complex class I chain-related gene A antibodies in organ transplantation. Curr Opin Organ Transplant 14:414–418
Acknowledgement
We thank Dr Andres Henschel and Dr Gihan El Bait for their assistance in preparing Figures 1 and 2. We are grateful to Mira Mousa for proof-reading the manuscript.
Funding
This research is supported by a research grant from Khalifa University that was jointly funded by Sandooq Al Watan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Halima Al Naqbi and Aurélie Mawart contributed as first authors.
The original online version of this article was revised: The Figure 2 image was incorrectly presented with unconverted data; Tables' presentation has been updated and the abbreviated sections has been expanded and reference citation (Popejoy and Fullerton 2016) cited under "Motivation for this review" section has been deleted. These has been corrected in the original article.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Al Naqbi, H., Mawart, A., Alshamsi, J. et al. Major histocompatibility complex (MHC) associations with diseases in ethnic groups of the Arabian Peninsula. Immunogenetics 73, 131–152 (2021). https://doi.org/10.1007/s00251-021-01204-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-021-01204-x