Abstract
The northern pig-tailed macaque (Macaca leonina) has been confirmed to be an independent species from the pig-tailed macaque group of Old World monkey. We have previously reported that the northern pig-tailed macaques were also susceptible to HIV-1. Here, to make this animal a potential HIV/AIDS model and to discover the mechanism of virus control, we attempted to assess the role of major histocompatibility complex (MHC) class I-restricted immune responses to HIV-1 infection, which was associated with viral replication and disease progression. As an initial step, we first cloned and characterized the classical MHC class I gene of northern pig-tailed macaques. In this study, we identified 39 MHC class I alleles including 17 MHC-A and 22 MHC-B alleles. Out of these identified alleles, 30 were novel and 9 were identical to alleles previously reported from other macaque species. The MHC-A and MHC-B loci were both duplicates as rhesus macaques and southern pig-tailed macaques. In addition, we also detected the patterns of positive selection in northern pig-tailed macaques and revealed the existence of balance selection with 20 positive selection sites in the peptide binding region. The analysis of B and F peptide binding pockets in northern and southern pig-tailed macaques and rhesus macaques suggested that they were likely to share a few common peptides to present. Thus, this study provides important MHC immunogenetics information and adds values to northern pig-tailed macaques as a promising HIV/AIDS model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-human primates have been widely used as essential models in biomedical research based on their phylogenetic, immunologic, and genetic proximity to humans. Particularly, macaque species, including rhesus macaques (Macaca mulatta), cynomolgus macaques (Macaca fascicularis), and pig-tailed macaques (Macaca nemestrina group), are commonly served as HIV/AIDS models, when infected with simian immunodeficiency virus (SIV) or chimeric SIV/HIV (SHIV) (Smith et al. 2005a). Unlike rhesus and cynomolgus macaques, pig-tailed macaques can be challenged with minimally modified HIV-1 strains and develop acute infection, due to a non-functional TRIM5-cyclophilin fusion that excludes a major barrier for HIV-1 infection in macaque species (Liao et al. 2007; Kuang et al. 2009). In addition, pig-tailed macaques can also be susceptible to influenza, chlamydia, and tuberculosis (Karl et al. 2014). These induce increasing utilization of this species in HIV/AIDS and other biomedical researches.
To further develop a pig-tailed macaque model in HIV/AIDS research, it is necessary to improve our knowledge about the T cell immunity and major histocompatibility complex (MHC) immunogenetics. MHC class I-restricted CD8+ T cell response plays a vital role in controlling HIV/SIV infection in macaque species. To characterize CD8+ T cell response, detailed knowledge of MHC class I alleles present in macaques is required. Besides, the class I MHC molecules are expressed on all nucleated cells and essential to present peptides as well as cytotoxic T lymphocyte (CTL) responses.
For pig-tailed macaques, little is known about MHC immunogenetics. As in rhesus macaques, pig-tailed macaques also present the classical MHC class I gene like A and B loci as well as other non-classical genes like E and I loci (Lafont et al. 2003). The A and B loci are duplicates. To date, sequences for about 447 MHC alleles of this species have been named and characterized (de Groot et al. 2012). The most studied pig-tailed macaque MHC class I allele, Mane-A1*084:01 (previously known as Mane-A*10) presenting the immunodominant SIV Gag epitope KP9, was reported to be associated with delayed progressive SIV disease and lower viral load (Smith et al. 2005a, b).
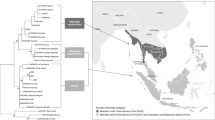
Recently, morphological characteristic and zoogeographical works have been studied extensively and the taxonomic status of macaques has also been re-evaluated. The pig-tailed macaque group has been divided into three independent species: southern pig-tailed macaque or Sunda pig-tailed macaque (M. nemestrina), northern pig-tailed macaque (Macaca leonina), and Mentawai macaque (Macaca pagensis) (Groves 2001; Malaivijitnond et al. 2012). The southern pig-tailed macaque ranges from about 7° 30′ N in Malay Peninsula, Sumatra, Bangka, and Borneo. The northern pig-tailed macaque is distributed in Peninsular Thailand, Burma, Bangladesh, India, and the southernmost Yunnan, China. The Mentawai macaque is located in the Mentawai islands (Kuang et al. 2009). So far, previously studied pig-tailed macaques were mainly M. nemestrina (Lei et al. 2013). The northern pig-tailed macaques were also susceptible to HIV-1 (Lei et al. 2014). In an effort to develop this local species to be a promising animal model, we have previously studied the reference values of blood chemistry, hematology, and basic immunological parameters of the northern pig-tailed macaque (Pang et al. 2013; Zhang et al. 2014; Zheng et al. 2014). However, to our knowledge, there are only 15 MHC alleles identified from this species according to the new nomenclature (Yan et al. 2013). Here, we analyzed MHC class I genes of the local species northern pig-tailed macaques in Yunnan using a cloning and sequencing method. Seventeen MHC-A and 22 MHC-B alleles were identified in our study. Exploring the frequencies of MHC class I alleles may contribute to further developing this potential AIDS model.
Materials and methods
Northern pig-tailed macaque samples
Whole blood samples from 12 unrelated northern pig-tailed macaques were generously provided by the Kunming Primate Research Center of Chinese Academy of Sciences, Kunming Institute of Zoology, Chinese Academy of Sciences. These animals were maintained in accordance with the institutional Animal Care and Use Committee of the Kunming Institute of Zoology, Chinese Academy of Sciences. Peripheral blood mononuclear cells (PBMCs) were obtained from whole blood using Ficoll-Paque Plus (Hao Yang, Tianjin, China) gradient centrifugation according to the manufacturer’s instruction.
RNA isolation and complementary DNA synthesis
Total RNA was isolated with a TRIzol reagent (TaKaRa, Dalian, China) from each PBMC sample. The integrity and concentration of RNA was determined with an Eppendorf BioPhotometer (Eppendorf, Germany). The complementary DNA (cDNA) was synthesized using the PrimeScript 1st Strand cDNA Synthesis Kit (TaKaRa) following the manufacturer’s instruction.
Amplification and cloning of northern pig-tailed macaque MHC class I cDNA
Polymerase chain reaction (PCR) was performed to amplify the MHC class I cDNA using a TransTaq DNA Polymerase High Fidelity (TransGen, Beijing, China) with locus-specific primer pairs. These primers were specific for untranslated regions around Mane-A (Mane5UA: GATTCTCCGCAGACGCCCA, Mane3UA: AAGTCAGGGTTCTTCAAGTCA) or Mane-B (Mane5UB1: AGAGTCTCCTCAGACGCCGA, Mane3UB: TGCCAGAGTGTCTTCAAAAGG) transcripts as described previously (Lafont et al. 2003). The following parameters were used for all cDNA amplifications: initial denaturation at 94 for 5 min; 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 90 s; and a final extension of 72 °C for 10 min to generate a 3′ dA overhang. Subsequently, PCR products were analyzed using 1 % agarose gel and purified using the DNA Gel Extraction Kit (Generay, Shanghai, China). Purified PCR products were cloned into the pMD-19T Simple Vector (TaKaRa), and the vectors were then transformed into Escherichia coli Trans5α Chemically Competent Cell (TransGen). For each macaque, a total of 30–50 independent cDNA clones were picked for each locus and then were sequenced on both strands using the BigDye terminator method using an ABI 3730xl automated sequence analyzer. Sequences were assembled by SeqMan (DNASTAR, Madison, WI, USA).
Sequences and phylogeny analysis
Sequence analysis was performed using DNAssist 2.2 and BLAST-2.2.30. A cDNA sequence was considered to be a real allele, when identified in at least three identical clones from independent PCR or from different individuals. The sequence was submitted to GenBank and the IMGT/MHC Non-human Primate Immuno Polymorphism Database-MHC (IPD-MHC) for official nomenclature assignments (Robinson et al. 2013; de Groot et al. 2012). Non-synonymous substitutions (d N) and synonymous substitutions (d S) were calculated using the modified Nei and Gojobori model with the Juke-Cantor correction for multiple hits (Nei and Kumar 2000). The ratio of d N/d S was tested for positive selection using the Z test with MEGA-6.06-mac (Tamura et al. 2013). DnaSP (Librado and Rozas 2009) was used to calculate the number of polymorphic sites and substitutions, nucleotide diversity, and average number of nucleotide differences. To infer the positive selected site (PPS) in MHC class I alleles identified from northern pig-tailed macaques, maximum likelihood-based random-site model analysis was used in CodeML within the PAML 4.8 (Yang 2007). Three null models of neutral evolution (M0, M1a, and M7) and three associated alternative models (M3, M2a, and M8) were used. Finally, the codons with w = d N/d S > 1 were identified using the empirical Bayes theorem method (Nielson and Yang 1998). Sequences were aligned using the ClustalW program of MEGA-6.06-mac, and a phylogenetic tree was generated according to the neighbor-joining method (Saitou and Nei 1987) of the same software. Genetic distances were computed using Kimura’s two-parameter method (Kimura 1980), and bootstrap analysis was based on 2000 replications. Values >50 % were shown on the tree.
Pocket analysis
To measure the peptide binding potential, amino acid composition of the B and P pockets was identified and compared. B and P pockets interacted with the most common peptide binding anchor, the second and the carboxyl-terminal resides of the peptides. The B and F pocket analysis was generated as described previously (Lafont et al. 2003). Briefly, the anchor residues involved in B pocket binding were 7, 9, 24, 25, 34, 45, 63, 66, 67, 70, and 99 and the residues involved in F pocket binding were 77, 80, 81, 84, 95, 116, 123, 143, 146, and 147. Additionally, numbering of residues of the class I heavy chain began with position 1 being the first residue of the mature chain where the leader sequence was excluded. Separated residues were aligned by MEGA-6.06-mac and Weblog 3.4 (Crooks et al. 2004).
GenBank accession numbers of MHC sequences
Northern pig-tailed macaque MHC Malo allele sequences have been deposited in GenBank under accession numbers KT214429–KT214467. The GenBank accession numbers for other allele sequences used in this manuscript were as follows: Mane-A1*003:02 (KF012913), Mane-A1*073:01 (KF012916), Mane-B*074:04:03 (KF012970), Mane-B*047:03 (KF012966), Mane-B*019:01 (KF012968), Mane-A2*05:01 (EF010510), Mane-B*5601 (FJ875237), Mane-B*058:01 (GQ153511), HLA-A*01:01 (BC003069), HLA-A*02:01 (AF055066), HLA-A*11:01 (AF030897), HLA-B*27:02 (JN703996), HLA-B*35:02 (GQ119000), HLA-B*53:01 (AJ311599), Mafa-A1*001:01 (AM295828), Mafa-A2*05:01 (AM295861), Mafa-A3*13:06 (AB447563), Mafa-B*031:01 (AY958152), Mamu-A1*001:01 (AJ539307), Mamu-A1*003:01 (U41379), Mamu-A2*24:01 (AJ542576), Mamu-A3*13:02 (AF157400), Mamu-B*013:02 (AB540185), Mamu-B*017:01 (AF199358), Mane-A1*084:01 (AY557348), Mane-A1*010:01 (EF010520), Mane-A2*05:01 (EF010513), Mane-A3*13:01 (EF010521), Mane-B*017:01 (AY204733), Mane-B*016:01 (AY557361), and Mane-B*131:01 (HQ992784).
Results and discussion
Summary of the identified MHC class I alleles
In our study, northern pig-tailed macaque (M. leonina) MHC class I alleles were designated Malo following the nomenclature proposed by Klein et al. (1990). We analyzed 853 MHC class I cDNA sequences from 12 northern pig-tailed macaques including 414 MHC-A sequences and 439 MHC-B sequences. Seventeen MHC-A and 22 MHC-B alleles were identified and submitted to GenBank and IPD-MHC databases. Of these 39 MHC class I sequences, only 2 MHC-A and 3 MHC-B alleles (Mane-A1*003:02, Mane-A1*073:01, Mane-B*074:04:03, Mane-B*047:03, and Mane-B*019:01) were previously reported from the northern pig-tailed macaques (Yan et al. 2013). We renamed these 5 alleles with northern pig-tailed macaque MHC four-letter code Malo as Malo-A1*003:01, Malo-A1*073:01, Malo-B*074:01, Malo-B*047:01, and Malo-B*019:01, respectively. The three other identified MHC alleles are identical to Mane-A2*05:01, Mane-B*56:01, and Mane-B*058:01, which were reported previously in southern pig-tailed macaques. Moreover, Malo-B*043:01 was identical to the rhesus macaque MHC Mamu-B*4301 and was first reported in pig-tailed macaques. The remaining 30 sequences obtained in this study were all novel. The accession number, IPD name, reference animals, and identities to previously reported alleles are listed in Table 1.
In addition, the complexity of the MHC gene in rhesus macaque, cynomolgus macaque, and southern pig-tailed macaque was reported to be higher than that in humans (Daza-Vamenta et al. 2004). Similar to these macaque species, we found that northern pig-tailed macaques also had duplication at MHC-A and MHC-B loci. For each animal, we identified 2–6 alleles at each MHC class I locus, indicating the duplication of Malo-A and Malo-B loci. Sequences like MHC-C genes were not detected in our study. This finding was consistent with the previous reports on southern pig-tailed macaque and rhesus macaque (Lafont et al. 2003). Among the 39 identified alleles, 23 were shared between two or more individuals. The Malo-A2*05 lineage and Malo-B*047 lineage were the most frequent MHC-A and MHC-B alleles, respectively. The summary of shared alleles is shown in Table 2, and the number of each allele clone detected from particular macaques was present in the cells. For instance, the combination of Malo-A1*040:01, Malo-B*120:01, Malo-B*078:01, and Malo-B*072:01 was detected in macaques 05211, 07246, and 05004, suggesting that this combination may be haplotypes. Haplotypes are localized groups of alleles inherited together on a chromosome. There have been several haplotypes reported from rhesus macaques, cynomolgus macaques, and southern pig-tailed macaques (Karl et al. 2008; Saito et al. 2012; Pratt et al. 2006). We hypothesized that haplotypes shared by northern pig-tailed macaques could also be present. Further studies would require more number of animals and more identified alleles.
Polymorphism and positive select analysis
The predicted amino acid sequences of northern pig-tailed macaque MHC class I open reading frame (ORF) were aligned by MegAlign software (DNASTAR). As shown in Fig. 1, the molecules were made up of leading peptide, two highly variable domains α1 and α2, and one conserved domain α3 containing interactions with CD8 and β2-microglobulin, followed by transmembrane and cytoplasmic domains as tail. The variability of the Malo-A and Malo-B gene was concentrated in α1 and α2 domains encoded by exon 2 and exon 3, as expected for classical MHC class I genes. These α1 and α2 domains contained peptides and T cell receptor (TCR) binding regions. As shown in Table 3, Malo-A and Malo-B ORF sequences contain 126 and 176 polymorphic sites. Forty-four and 35 % of these mutations were located in the peptide binding sites of α1 and α2 domains, respectively. The sequences of α1 domain exhibited 60 and 87 segregating sites with 76 and 112 mutations as well as a nucleotide diversity of 0.083 and 0.102 in Malo-A and Malo-B, respectively, while in α2 domain, there are 55 and 89 segregating sites with 73 and 111 mutations and a nucleotide diversity of 0.069 and 0.086, respectively. These findings suggested that Malo has a high level of genetic diversity and polymorphism. Most of the substitutions were located in α1 and α2 domains. In these domains, the amino acid substitutions caused by nucleotide mutation may directly change the ability to binding peptides and consequently affect the progression of the disease. Polymorphism in peptide binding region is significant to maintain a highly variable MHC pool and increase the potential range of epitope presented, which may attribute to host response to a larger selection of antigens or pathogens.
Alignment of predicted amino acid sequences identified from 12 northern pig-tailed macaque MHC class I ORFs. The sequences were compared with the Mane-A*084:01 molecule from southern pig-tailed macaques. Identity with Mane-A*084:01 is indicated with periods. Gaps are indicated by blank spaces. Residues in the α1 and α2 domains, interacting with peptide, TCR, or both, are indicated by p, t, or b, respectively, according to Lafont et al. (2003). The cysteine disulfide bridge is presented by asterisks
In addition, the sizes of the sequences we identified ranged from 1080 to 1101 bp. Therefore, the molecules encoded by these sequences were composed of 359–366 amino acids (aa). As shown in Fig. 1, Malo-B allele cytoplasmic domains were 24 aa with the exception of Malo-B*025:01, Malo-B*24:01, Malo-B*073:01, and Malo-B*047:02, which were of the same size as Malo-A in cytoplasmic domains containing 27 aa. In addition, there was much more polymorphism in view of either peptide length variance or amino acid mutations in the leading peptide region. Most of the cDNA sequences of Malo alleles had double start codons (ATGs) at the 5′ terminal of the leading peptide region. While Malo-B*025:01, Malo-B*24:01, Malo-B*073:01, Malo-B*047:02, Malo-B*017:01, Malo-B*061:01, and Malo-B*013:01 alleles could only use the second ATG as the start codon due to the deletion of a guanosine (G) right after the first ATG. This change resulted in 3-amino acid abridgment. This mutation was also reported in rhesus macaques, Tibetan macaques (Macaca thibetana), stump-tailed macaques (Macaca arctoides), and Assamese macaques (Macaca assamensis), indicating that the presence of single or double start codons is not restricted to certain macaque species (Daza-Vamenta et al. 2004; Ouyang et al. 2008; Yan et al. 2013). Interestingly, we found that two 1101-bp Malo-A alleles (Malo-A2*05:01 and Malo-A2*05:02) had 1-amino acid insertion (A) in the leader peptide domain (25 aa). In a macaque family, the variation and polymorphism in the leading peptide domain of MHC class I alleles could have immunological significance. In humans, the non-classical HLA-E molecules are important ligands of CD94/NKG2 receptors, which have been shown to participate in the activated or inhibitory reaction of natural killer (NK) cells (Lee et al. 1998). Recent studies have implied that HLA-A, HLA-B, and HLA-G may take part in the NK responses by a nonapeptide ligand (VMAPRTLLL). They provide the ligand for HLA-E from their leading peptides, which is essential for NK activity or inhibition (Lee et al. 1998; Miller et al. 2003). In our study, 28 of the 39 full-length MHC alleles had the typical (V)MAPRTLLL motif in their leading peptides, with the exception of Malo-A*003:01 (IMAPRTLL); Malo-A2*05:01 and Malo-A2*05:02 (VMGPARTLLL); Malo-A3*13:01 (VMAPRTFLL); Malo-B*047:01 (VMAPRNLLL); Malo-B*120:01, Malo-B*094:01, Malo-B*018:01, and Malo-B*013:01 (VMAPRILLL); Malo-B*017:01 (VMAPGTLLL); and Malo-B*091:01 (VVAPRTLLL) (Fig. 1). Although there is still no proof to confirm that the nonapeptide ligand (VMAPRTLLL) has a similar function in macaque species, the variation and different lengths of MHC leading peptides might have changed or enriched the non-peptide reservoir for MHC-E binding to cope with variant pathogens (Ouyang et al. 2008).
Based on the high level of polymorphism, we detected the patterns of positive selection in northern pig-tailed macaques by evaluating the ratio of non-synonymous substitution (d N)/synonymous substitution (d S). The ratio of d N/d S is commonly used to detect a balancing selection. The balancing selection is indicated by a ratio >1, while a ratio <1 is interpreted as an evidence of purifying selection and a ratio of 1 is considered neutral amino changes. In this study, we evaluated the ratio of d N/d S in α1 and α2 domains of Malo-A and Malo-B, with substantially elevated d N/d S ratios and a tendency towards positive selection. Especially for the peptide binding sites (PBSs) in the α1 domain of Malo-A and Malo-B, the number of non-synonymous substitutions (d N) was much higher than synonymous substitutions (d S) with ratios 5.727 (p = 0.003) and 5.220 (p = 0.002), respectively (Table 3) while, in the non-PBS region, the ratio <1 indicated that such sites were under a purifying selection and such highly conserved amino acid residues may also be essential for the proper function of Malo.
Then, we inferred 20 positive selected sites for the α1 and α2 domains of the northern pig-tailed macaque MHC class I molecule by CodeML in PAML 4.8 software. Three null models of neutral evolution (M0, M1a, and M7) and three associated alternative models (M3, M2a, and M8) were utilized. In models M2a and M8, 20 codons in α1 and α2 domains were identified being under a positive selection with an empirical Bayes theorem method. Most of these PPSs (80 %) were interacted with binding peptides, TCR, or both (labeled in Table 4), suggesting that the selective pressure mainly came from antigens or pathogens, while in this highly variant domain, several conserved sites were still responsible for stabilizing or folding the protein correctly. For example, a cysteine disulfide bridge is conserved in all macaque MHC sequences, which is important for folding protein (indicated by asterisks in Fig. 1). In essence, these conserved codons in human HLA and macaque MHC are conserved in the same position, indicating that these amino acids are under a strong purifying selection.
Phylogenetic analysis and allele share
To determine the relationship between northern pig-tailed macaque (Malo) and other macaque species, we collected typical southern pig-tailed macaques (Mane), rhesus macaques (Mamu), cynomolgus macaques (Mafa), and human (HLA) MHC alleles from GenBank and built a neighbor-joining non-root tree (Fig. 2). It was obvious that several macaque species clustered together but did not segregate based on their species of origin, suggesting their close evolution relationship with no separation of lineages among different macaque species. In contrast, HLA alleles grouped independently from the macaque MHC cluster. Two main branches contained MHC-A and MHC-B alleles. MHC-A branches were distributed into three clusters: A1, A2, and A3, which is in parallel with the previous studies on southern pig-tailed macaques and rhesus macaques (Lafont et al. 2003; Karl et al. 2008).
Neighbor-joining tree of northern pig-tailed macaques (Malo), southern pig-tailed macaques (Mane), rhesus macaques (Mamu), cynomolgus macaques (Mafa), and human (HLA) MHC alleles. Sequences were aligned using the ClustalW program of MEGA-6.6-mac software with minor manual adjustments. Phylogenetic trees were constructed based on the alignment using the neighbor-joining method of the same software. Genetic distances were estimated using a Kimura’s two-parameter method. Bootstrap analysis was performed (2000 replicates) to assign confidence to tree nodes. Values >50 % are shown on the tree. The Malo-A and Malo-B alleles identified in our study are indicated by filled circle and filled triangle, respectively
In addition, around 90 % of cDNA sequences identified in our study were unique to northern pig-tailed macaques when compared with other macaque species including southern pig-tailed macaques. Interestingly, although pig-tailed macaque group has been divided into three independent species (Groves 2001), the pig-tailed macaque is still under controversial taxonomical reclassification. Whether the northern and southern forms are two distinct species, subspecies, or just forms of the same species remains equivocal (Malaivijitnond et al. 2012). Here, based on the limited MHC immunogenetics information, the differences in MHC alleles between the two groups of pig-tailed macaques were greater than the differences found between different forms of rhesus macaques and cynomolgus macaques (Karl et al. 2008; Saito et al. 2012). The differences may be consequences of geographic separation caused by a marine transgression during the Pleistocene period. The separation facilitates the evolutionary divergence of the two forms of pig-tailed macaques (Malaivijitnond et al. 2012). As a result, the unique alleles of northern and southern pig-tailed macaques supported the recent classification that the two forms of pig-tailed macaques are actually two different species, M. leonina and M. nemestrina, based on different morphological characters and mtDNA analysis.
On the other hand, among the 39 alleles, 4 of the 9 previously reported sequences were identical to alleles described in other macaque species. Malo-B*043:01 was previously identified as Mamu-B*043:01 in rhesus macaque, while Malo-B*056:01, Malo-B*058:01, and Malo-A2*05:01 were identical to Mane-B*056:01, Mane-B*58:01(Mn-B*nov079), and Mane-A2*05:01(Mane-A*06) in southern pig-tailed macaque, respectively. Interestingly, the Mane-A2*05:01 and its orthologous genes Mamu-A2*05:01 and Mafa-A2*05:01 have been shown to have high frequency in southern pig-tailed macaque, rhesus macaque, and cynomolgus macaque, respectively (Wu et al. 2008). By cloning and sequence methodology, we additionally determined that Malo-A2*05:01 was also prevalent (50 %) among 12 northern pig-tailed macaques in our study, which demonstrates that the locus encoding Mane-A2*05:01/Mamu-A2*05:01 is highly prevalent in macaques. By analyzing full-length Malo-A transcripts from six Malo-A2*05:01 cloning positive individuals having different MHC genotypes, we found that the Malo-A2*05:01 is expressed at low level, similar to Mane-A2*05:01/Mamu-A2*05:01. As shown in Fig. 3, 23–46 Malo-A cDNA clones were sequenced and 2–4 Malo-A alleles were detected from each animal. In total, 203 clones were isolated and only 16 (8 %) of them were detected as Malo-A2*05:01 clones, the frequency of which was five- or tenfold lower than the most abundant MHC-A allele. Only 3–13 % of the Malo-A2*05:01 clones were identified in each monkey (Fig. 3). Moreover, such infrequent sharing of orthologous alleles between several different macaque species suggests that the certain MHC class I alleles may share a common ancestor and play a crucial role in immune responses.
Implications for biomedical research
Multiple lines of studies have demonstrated that CD8+ T cells and their strong and specific cytotoxic T lymphocyte (CTL) response are important for the control of SIV/HIV replication in macaques or humans (Gauduin et al. 1998; Martin and Carrington 2013). During the activation of CTL, the MHC molecule is essential to present peptides derived from virus protein to TCR of Ag-specific CD8+ T cells. To measure the peptide binding potential of the northern pig-tailed macaque MHC, we identified and compared the amino acid composition of the B and F pockets of the MHC molecule. These two pockets interacted with the most common peptide binding anchor, the second and the carboxyl-terminal resides of the peptides (Fig. 4). We compared the northern pig-tailed macaque MHC (Malo) in this study with all the reported southern pig-tailed macaque MHC (Mane) and rhesus macaque MHC (Mamu) from the IPD website (de Groot et al. 2012). There was no significant difference between these three macaque species, neither the amino acid diversity nor the site conservation, suggesting that they may share some common epitopes presented to T cells. MHC-B alleles seemed to have more amino acid diversity and variability, especially for the B pocket. There were several highly variable amino acid sites, three in B pocket and one in F pocket, containing more than five unique resides (resides 9, 70, 99, and 116). Such unstable sites may contribute to the binding of different peptides and the MHC molecular binding of a wider range of epitopes. Consistently, there were several Mamu-B alleles like Mamu-B*008:01, Mamu-B*017:01, Mamu-B*003:01:01, and Mamu-B*004:01 considered to be protective genes, possibly as a consequence of their extensive diversity. Similarly, in humans, HLA-B showed greater differential effects on HIV outcome and its protective role in HIV was also supported by functional data (Martin and Carrington 2013). Although relatively less MHC-A alleles have been reported, there were also some alleles like Mamu-A1*001:01 in rhesus macaques and Mane-A1*084:01 in southern pig-tailed macaques reported as protective genes (Pal et al. 2002; Smith et al. 2005a).
Comparison of B and F pocket residues from northern pig-tailed macaque (Malo), southern pig-tailed macaque (Mane), and rhesus macaque (Mamu) MHC allele sequence. The overall height of each stack indicates the sequence conservation at that position (measured in bits), whereas the height of symbols within the stack reflects the relative frequency of the corresponding amino or nucleic acid at that position. The maximum sequence conservation per site is log220 ≈ 4.32 bits for proteins according to the manuscript of Weblog 3.4 (Crooks et al. 2004)
In addition, Malo-B*017:01 and Malo-B*047:01 identified in our study were strikingly similar to Mamu-B*017:01 and Mamu-B*047:01, respectively, in the amino acid level. Many studies have shown that Mamu-B*017:01 was associated with the control of SIV in rhesus macaques and haplotypes containing Mamu-B*047:01 were correlated with slow disease progression (Yant et al. 2006; Sauermann et al. 2008). There was only one substitution, serine to proline, between Malo-B*017:01 and Mamu-B*017:01, and this change may not alert the peptide binding specificity since the substitution was not located at the peptide binding site or the binding pocket. Thus, these two alleles may share the same epitope as the previously characterized Mamu-B*017:01 SIV-derived peptides (Mothé et al. 2002). Therefore, it is possible that reagents like MHC tetramers of Mamu-B*017:01 are available to analyze Malo-B*017:01-restricted CD8+ T cell responses in pig-tailed macaques. Additionally, the predicted amino acid of Malo-B*017:01 was identical to that of Mane-B*017:01. Consequently, the Malo-B*017:01-positive animals are likely to present Nef IW9 and Env FW9 epitopes, which may shorten the research time.
As mentioned above, the pig-tailed macaque is a promising HIV/AIDS model and the knowledge of MHC-restricted CD8+ T cell responses will be useful in vaccine development and biomedical research. Because of the limited MHC immunogenetics information in this species, however, only Mane-A1*084:01 and its restricted CD8+ T cell responses to SIV have been well characterized (Smith et al. 2005a, b) with a few studies examining the CTL responses in SIV-infected pig-tailed macaques (Gooneratne et al. 2014). Although the protective role of CTL responses in HIV-infected pig-tailed macaques has also been demonstrated (Kent et al. 1997), the HIV epitopes and specific MHC molecules have not been identified yet. In the future, developing reagents to better understand the MHC-restricted CD8+ T cell responses and other functional immunological studies in SIV/HIV-infected pig-tailed macaques could address these questions, based on the characterization of the full-length MHC class I cDNA sequences in pig-tailed macaques. The MHC cDNA sequences identified in our study provide a significant beginning and add more information to the limited MHC immunogenetics knowledge available for northern pig-tailed macaques. Based on more identification of Malo alleles associated with disease progression and development of reagents as well as better techniques for immunological studies, the northern pig-tailed macaque will become a potential non-human primate animal model for SIV/HIV pathogenesis and vaccine research. Moreover, the identification of shared and unique MHC sequences may also contribute to the non-human primate’s biogeographic study.
References
Crooks GE, Hon G, Chandonia JM et al (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Daza-Vamenta R, Glusman G, Rowen L et al (2004) Genetic divergence of the rhesus macaque major histocompatibility complex. Genome Res 14(8):1501–1515
de Groot NG, Otting N, Robinson J et al (2012) Nomenclature report on the major histocompatibility complex genes and alleles of great ape, Old and New World monkey species. Immunogenetics 64:615–631
Gauduin MC, Glickman RL, Means R et al (1998) Inhibition of simian immunodeficiency virus (SIV) replication by CD8+ T lymphocytes from macaques immunized with live attenuated SIV. J Virol 72:6315–6324
Gooneratne SL, Alinejad-Rokny H, Ebrahimi D et al (2014) Linking pig-tailed macaque major histocompatibility complex class I haplotypes and cytotoxic T lymphocyte escape mutations in simian immunodeficiency virus infection. J Virol 88:14310–14325
Groves C (2001) Primate taxonomy. Smithsonian Institution, Washington DC
Karl JA, Wiseman RW, Campbell KJ et al (2008) Identification of MHC class I sequences in Chinese-origin rhesus macaques. Immunogenetics 60:37–46
Karl JA, Heimbruch KE, Vriezen CE et al (2014) Survey of major histocompatibility complex class II diversity in pig-tailed macaques. Immunogenetics 66:613–623
Kent SJ, Woodward A, Zhao A (1997) Human immunodeficiency virus type 1 (HIV-1)-specific T cell responses correlate with control of acute HIV-1 infection in macaques. J Infect Dis 176:1188–1197
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Klein J, Bontrop RE, Dawkins RL et al (1990) Nomenclature for the major histocompatibility complexes of different species: a proposal. Immunogenetics 31:217–219
Kuang YQ, Tang X, Liu FL et al (2009) Genotyping of TRIM5 locus in northern pig-tailed macaques (Macaca leonina), a primate species susceptible to human immunodeficiency virus type 1 infection. Retrovirology 6:58
Lafont BA, Buckler-White A, Plishka R et al (2003) Characterization of pig-tailed macaque classical MHC class I genes: implications for MHC evolution and antigen presentation in macaques. J Immunol 171:875–885
Lafont BA, McGraw CM, Stukes SA et al (2007) The locus encoding an oligomorphic family of MHC-A alleles (Mane-A*06/Mamu-A*05) is present at high frequency in several macaque species. Immunogenetics 59:211–223
Lee N, Llano M, Carretero M et al (1998) HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proc Natl Acad Sci U S A 95:5199–5204
Lei AH, Pang W, Zhang GH et al (2013) Use and research of pigtailed macaques in nonhuman primate HIV/AIDS models. Zool Res 34:77–78
Lei AH, Zhang GH, Tian RR et al (2014) Replication potentials of HIV-1/HSIV in PBMCs from northern pig-tailed macaque (Macaca leonina). Zool Res 35:186–195
Liao CH, Kuang YQ, Liu HL et al (2007) A novel fusion gene, TRIM5-cyclophilin A in the pig-tailed macaque determines its susceptibility to HIV-1 infection. AIDS 21:S19–S26
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Malaivijitnond S, Arsaithamkul V, Tanaka H et al (2012) Boundary zone between northern and southern pig-tailed macaques and their morphological differences. Primates 53:377–389
Martin MP, Carrington M (2013) Immunogenetics of HIV disease. Immunol Rev 254:245–264
Miller JD, Weber DA, Ibegbu C et al (2003) Analysis of HLA-E peptide-binding specificity and contact residues in bound peptide required for recognition by CD94/NKG2. J Immunol 171:1369–1375
Mothé BR, Sidney J, Dzuris JL et al (2002) Characterization of the peptide-binding specificity of Mamu-B*17 and identification of Mamu-B*17-restricted epitopes derived from simian immunodeficiency virus proteins. J Immunol 169:210–219
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press
Nielson R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Ouyang D, Xu L, Dai Z et al (2008) Identification of major histocompatibility complex class I alleles in Chinese rhesus macaques. Acta Biochim Biophys Sin 40:919–927
Pal R, Venzon D, Letvin NL et al (2002) ALVAC-SIV-gag-pol-env-based vaccination and macaque major histocompatibility complex class I (A* 01) delay simian immunodeficiency virus SIVmac-induced immunodeficiency. J Virol 76(1):292–302
Pang W, Lv LB, Wang Y et al (2013) Measurement and analysis of hematology and blood chemistry parameters in northern pig-tailed macaques (Macaca leonina). Zool Res 34:89–96
Pratt BF, O’Connor DH, Lafont BAP et al (2006) MHC class I allele frequencies in pigtail macaques of diverse origin. Immunogenetics 58:995–1001
Robinson J, Halliwell JA, McWilliam H et al (2013) IPD—the immuno polymorphism database. Nucleic Acids Res 41:D1234–D1240
Saito Y, Naruse TK, Akari H et al (2012) Diversity of MHC class I haplotypes in cynomolgus macaques. Immunogenetics 64:131–141
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sauermann U, Siddiqui R, Suh YS et al (2008) Mhc class I haplotypes associated with survival time in simian immunodeficiency virus (SIV)-infected rhesus macaques. Genes Immun 9:69–80
Smith MZ, Dale CJ, De Rose R et al (2005a) Analysis of pigtail macaque major histocompatibility complex class I molecules presenting immunodominant simian immunodeficiency virus epitopes. J Virol 79:684–695
Smith MZ, Fernandez CS, Chung A et al (2005b) The pigtail macaque MHC class I allele Mane-A*10 presents an immunodominant SIV Gag epitope: identification, tetramer development and implications of immune escape and reversion. J Med Primatol 34:282–293
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30:2725–2729
Wu J, Bassinger S, Montoya GD et al (2008) Allelic diversity within the high frequency Mamu-A2*05/Mane-A2*05 (Mane-A*06)/Mafa-A2*05 family of macaque MHC-A loci. Tissue Antigens 72:29–38
Yan X, Li A, Zeng L et al (2013) Identification of MHC class I sequences in four species of Macaca of China. Immunogenetics 65:851–859
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yant LJ, Friedrich TC, Johnson RC et al (2006) The high-frequency major histocompatibility complex class I allele Mamu-B*17 is associated with control of simian immunodeficiency virus SIVmac239 replication. J Virol 80:5074–5077
Zhang XL, Wei P, De-Yao D et al (2014) Analysis of immunoglobulin, complements and CRP levels in serum of captive northern pig-tailed macaques (Macaca leonina). Zool Res 35:196–203
Zheng HY, Zhang MX, Zhang LT et al (2014) Flow cytometric characterizations of leukocyte subpopulations in the peripheral blood of northern pig-tailed macaques (Macaca leonina). Zool Res 35:465–473
Acknowledgments
The authors would like to thank Muhammad Shahzad for editing the manuscript and his helpful discussions and Natasja de Groot, Nel Otting, and the IMGT Non-human Primate Nomenclature Committee for naming the Malo-A and Malo-B sequences. This work was supported by the National Basic Research Program of China (2012CBA01305), the National Natural Science Foundation of China (81172876, 81373124, 81471620, 81571606), the Knowledge Innovation Program of CAS (KSCX2-EW-R-13, KJZD-EW-L10-02), the Natural Science Foundation of Yunnan Province (2014FB181), and the Collaborative Innovation Center for Natural Products and Biological Drugs of Yunnan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Lian, XD., Zhang, XH., Dai, ZX. et al. Cloning, sequencing, and polymorphism analysis of novel classical MHC class I alleles in northern pig-tailed macaques (Macaca leonina). Immunogenetics 68, 261–274 (2016). https://doi.org/10.1007/s00251-015-0897-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-015-0897-3