Abstract
In the small intestine of celiac disease patients, dietary wheat gluten and similar proteins in barley and rye trigger an inflammatory response. While strict adherence to a gluten-free diet induces full recovery in most patients, a small percentage of patients fail to recover. In a subset of these refractory celiac disease patients, an (aberrant) oligoclonal intraepithelial lymphocyte population develops into overt lymphoma. Celiac disease is strongly associated with HLA-DQ2 and/or HLA-DQ8, as both genotypes predispose for disease development. This association can be explained by the fact that gluten peptides can be presented in HLA-DQ2 and HLA-DQ8 molecules on antigen presenting cells. Gluten-specific CD4+ T cells in the lamina propria respond to these peptides, and this likely enhances cytotoxicity of intraepithelial lymphocytes against the intestinal epithelium. We propose a threshold model for the development of celiac disease, in which the efficiency of gluten presentation to CD4+ T cells determines the likelihood of developing celiac disease and its complications. Key factors that influence the efficiency of gluten presentation include: (1) the level of gluten intake, (2) the enzyme tissue transglutaminase 2 which modifies gluten into high affinity binding peptides for HLA-DQ2 and HLA-DQ8, (3) the HLA-DQ type, as HLA-DQ2 binds a wider range of gluten peptides than HLA-DQ8, (4) the gene dose of HLA-DQ2 and HLA-DQ8, and finally,(5) additional genetic polymorphisms that may influence T cell reactivity. This threshold model might also help to understand the development of refractory celiac disease and lymphoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
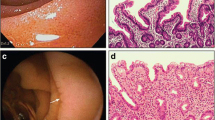
With a prevalence of 1% in western populations, celiac disease (CD) is one of the most common inflammatory disorders of the small intestine (Green and Cellier 2007). CD is often assumed to have its onset in childhood, but it has recently been suggested that adults can also develop CD (Vilppula et al. 2009). Clinical manifestations vary according to age group: infants and young children present with diarrhea, abdominal distention, and failure to thrive, whereas adults that develop CD not only present with diarrhea, but also with silent manifestations such as anemia, osteoporosis, or neurological symptoms (Green and Cellier 2007). Immunohistochemistry of the small intestine of patients shows villous atrophy, crypt hyperplasia, and elevated levels of intraepithelial lymphocytes (IELs). The only therapy until now is a gluten-free diet, which will normalize the clinical and histological manifestations and allows the patients to live an otherwise normal life.
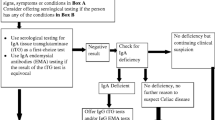
A small percentage of adult-onset CD patients develop a primary or secondary resistance to a gluten-free diet (Fig. 1). This condition is called refractory celiac disease (RCD) and is characterized by persisting villous atrophy and elevated levels of IELs. Currently, RCD is subdivided into two subtypes: RCD type I (RCD I) and RCD type II (RCD II) that both display clinical and histological resistance to a gluten-free diet (Daum et al. 2005). RCD II, however, is associated with the presence of an aberrant IEL population that lacks surface T cell receptor (TCR)-CD3 expression, but contains intracellular CD3ε and has clonal TCRγ-gene rearrangements (Cellier et al. 1998). These aberrant IELs can gain chromosomal abnormalities and develop into surface TCR-CD3- lymphoma cells (Deleeuw et al. 2007; Verkarre et al. 2003). RCD II is therefore considered a premalignant condition, and roughly 50% of the RCD II patients develop overt lymphoma within 5 years of diagnosis (Al-Toma et al. 2007; Malamut et al. 2009). In summary, the majority of CD patients has an uncomplicated disease course and can be treated with a gluten-free diet (Fig. 1). RCD II and RCD-associated lymphoma, however, are difficult to treat and have therefore poor 5-year survival rates of <44% and <20%, respectively (Al-Toma et al. 2007).
Prevalence of CD and complicated CD in the Caucasian population. Approximately 25% of the general Caucasian population is HLA-DQ2+. From these genetically susceptible individuals, only 4% develop CD. In majority of the CD patients, the disease course is uncomplicated. Roughly 3% of the CD patients will not respond to a gluten-free diet and develop RCD. A subset of RCD patients develop RCD II of which approximately 50% develop RCD-associated lymphoma (not shown)
The development of CD is determined by both environmental and genetic factors. In the 1950s, ingestion of wheat products was described to cause malabsorption symptoms in patients (Dicke et al. 1953). Later on, it was established that gluten, the storage proteins in wheat, barley and rye, caused a cell-mediated immune response in the small intestine (Ferguson et al. 1975). In addition to this environmental factor, CD development involves genetic predisposition, as the vast majority of the CD patients possess human leukocyte antigen (HLA)-DQ2 and/or HLA-DQ8 (Spurkland et al. 1997). HLA-DQ2 is a genotype that is present in roughly 25% of the European population (Bourgey et al. 2007; Fig. 1). Yet, only ~4% of all HLA-DQ2+ individuals develop CD (Fig. 1) (Wolters and Wijmenga 2008). CD development in HLA-DQ2- and HLA-DQ8-negative individuals is extremely rare (Karell et al. 2003). These findings came together with the observation that CD4+ T cell lines from duodenal biopsies of CD patients specifically respond to gluten peptides presented by HLA-DQ2 and/or HLA-DQ8 (Lundin et al. 1993, 1994). The etiology of RCD is much less clear, but seems to be associated with HLA-DQ2 (Al-Toma et al. 2006).
In short, the complex interplay of multiple genetic and environmental factors will determine the development of CD and its complications. This review describes current knowledge on gluten, HLA-DQ, and the immunopathogenesis of CD and its complications. In addition, we present a likely sequence of events in the development of CD and discuss the factors that may influence the risk of CD development.
Gluten: the disease-inducing environmental trigger
The visco-elastic properties of gluten are essential for dough formation of wheat flour and give bread its unique texture and taste. Because of its unique properties, gluten is widely used in the food industry: not only in products that are readily associated with wheat, like bread, cookies and pasta, but also as a hidden ingredient in sauces, instant soups, and even medication. Consequently, the daily gluten intake on a gluten-containing diet in Western Europe and the United States is high, between 15 and 20 g per day. The omnipresence of gluten makes adherence to a gluten-free diet challenging for CD patients.
Gluten is a heterogeneous mixture of gliadins and glutenins in wheat or similar proteins in barley and rye. Each wheat variety expresses multiple α-, γ-, and ω-gliadins in addition to low- and high-molecular weight glutenins. Gluten has a very high content of the amino acids glutamine (30%) and proline (15%). By virtue of its high glutamine content, gluten is rich in nitrogen, an essential factor for seed germination. The high proline content renders gluten highly resistant to degradation by gastrointestinal enzymes, making it possible for large immunogenic gluten peptides to reach the mucosal surface (Shan et al. 2002, 2005). As modern wheat varieties contain three complete genomes encoding gliadins and glutenins, up to 100 different gluten proteins may be present in a single wheat variety, and many of these are implicated in the pathogenesis of CD.
HLA-DQ: the strongest disease-associated gene locus by far
The strong genetic influence in CD is apparent, as the concordance between monozygotic twins is 80%, whereas in dizygotic twins, this is only 11% (Nistico et al. 2006), which is approximately the same as the risk for first-degree relatives (Dube et al. 2005). The main genetic influence in CD is HLA, which was first indicated by studies describing the predominance of HLA-B8 and HLA-DR3 serotypes in CD patients (Falchuk et al. 1972; Keuning et al. 1976). Later studies established that the strongest association is with HLA-DQ2 (DQA*0501, DQB*0201, termed HLA-DQ2.5 hereafter) (Sollid et al. 1989), which is encoded together with HLA-B8 and HLA-DR3 on the highly conserved ancestral haplotype 8.1 (Price et al. 1999). CD is associated, to a lesser extent, with HLA-DQ8 (DQA*03, DQB*0302) (Spurkland et al. 1997). The strong association between HLA-DQ2.5 and CD is further illustrated by the observation that individuals homozygous for HLA-DQ2.5 have a fivefold increased risk for development of CD compared to individuals heterozygous for HLA-DQ2.5 (Mearin et al. 1983). Similarly, HLA-DQ 2.5 homozygosity is associated with the development of RCD II and RCD-associated lymphoma, whereas this association is less clear for HLA-DQ2.5 heterozygosity and HLA-DQ8 (Al-Toma et al. 2006). Another HLA-DQ2 variant exists: HLA-DQ2.2 (DQA*0201, DQB*0202), which has a peptide-binding motif that is almost identical to that of HLA-DQ2.5 (van de Wal et al. 1997). Whereas HLA-DQ2.5 predisposes to CD, HLA-DQ2.2 does not. This difference is related to the peptide-binding properties of these HLA-DQ2 variants (see below). The estimated risk effect of HLA-DQ2 and HLA-DQ8 on CD development is estimated to be ~35% (Hunt et al. 2008).
Not all HLA-DQ2.5+ and HLA-DQ8+ individuals develop CD, indicating that these HLA genotypes are necessary, but not sufficient for CD development. Recent large-scale genetic association studies identified many additional genetic loci that all make a small contribution to the risk to develop CD (Box 1 and Table 1). Importantly, most of these genes encode proteins involved in immunity, which supports the notion that CD is an immune-related disorder and possibly provides clues on the immunopathogenesis of CD.
The adaptive immune response: gluten, HLA-DQ, and CD4+ T cells
As noted above, CD develops almost exclusively in HLA-DQ2.5+ or HLA-DQ8+ individuals. It is also well established that in CD patients, gluten-derived peptides presented by either HLA-DQ2.5 or HLA-DQ8 induce a CD4+ T cell response. Both HLA-DQ2.5 and HLA-DQ8 prefer to bind peptides with negatively charged amino acids at anchor residues. Gluten peptides, however, are virtually devoid of negative charges, and native gluten peptides thus bind poorly to HLA-DQ2.5 or HLA-DQ8. Consequently, CD4+ T cells specific for native gluten peptides are rare. It has become clear that the enzyme tissue transglutaminase 2 (TG2) can modify gluten peptides to fit the requirements for high affinity binding to HLA-DQ2 and HLA-DQ8 (Molberg et al. 1998; van de Wal et al. 1998a). TG2 can convert non-charged glutamine into negatively charged glutamic acid, a process called deamidation. Because gluten has a high content of glutamine (Q) and proline (P), the sequences QP, QXP and QXXP (in which X can represent any amino acid) are frequently found in gluten peptides. Strikingly, only in the sequence QXP that the glutamine is converted, which results in highly selective introduction of negative charges in gluten peptides (Vader et al. 2002). This specific deamidation process introduces the negative charges at the positions favored by HLA-DQ2 and HLA-DQ8, thereby expanding the presentable gluten peptide repertoire (Fig. 2). As a result, the gluten-specific CD4+ T cell repertoire is substantially expanded, which enhances the inflammation and disease development (Fig. 2).
Self-amplifying loops in the development of CD. HLA-DQ2+ or HLADQ8+ individuals can present native gluten peptides on their APC to CD4+ T cells. The IFN-γ released in this process can in turn boost HLA-DQ2/8 expression. This low-grade inflammation might eventually lead to tissue damage with TG2 release. TG2 is able to expand the presentable gluten peptide repertoire by deamidation. As a result, the CD4+ T cell response is strongly enhanced, leading to more IFN-γ, tissue damage, and increased release of TG2. The end result is full-blown CD
TG2 is mostly retained intracellularly in an inactive form and is activated upon its release during tissue damage (Lorand and Graham 2003; Siegel et al. 2008). Therefore, something should trigger tissue damage which initiates TG2 release, allowing the modification of gluten peptides. Whereas CD4+ T cell responses against native gluten peptides are relatively rare, they could represent the first breach in oral tolerance to gluten. The presentation of native gluten peptides by HLA-DQ2 or HLA-DQ8 to CD4+ T cells will lead to the production of IFN-γ (Fig. 2). IFN-γ will in turn lead to higher expression of the HLA-DQ molecules and thereby, to increased gluten peptide presentation (Fig. 2). In the presence of gluten, this could become a self-amplifying loop that could cause limited tissue damage locally. This tissue damage would lead to the release of TG2 that will modify native gluten peptides into high affinity ligands for HLA-DQ2 and/or HLA-DQ8, thereby expanding the gluten-specific CD4+ T cell responses and leading to additional tissue damage: the initiation of a second self-amplifying loop (Fig. 2). Alternatively, infections occurring in the gastrointestinal tract would generate a pro-inflammatory milieu that might lead to loss of tolerance to native gluten peptides and generate tissue damage simultaneously and thus, initiate deamidation by TG2.
Activated intraepithelial lymphocytes damage the intestinal epithelium
Intraepithelial lymphocytes are localized between intestinal epithelial cells at the basolateral side of the epithelium and are thought to play an important role in immunosurveillance of the epithelium. The IEL population in the small intestine is a mixed population of TCRαβ+ T cells, TCRγδ+ T cells, and NK cells, although the vast majority of the IELs are CD8+ TCRαβ+ T cells (Jabri and Ebert 2007). Furthermore, most of these TCR+ IELs express a variety of NK cell receptors that is distinct from the NK cell receptors expressed on blood T cells (Jabri et al. 2000). The NK cell receptors are thought to act mainly as T cell co-stimulators, lowering the threshold for T cell activation in stressful times (Bauer et al. 1999).
In active CD, the number of CD8+ TCRαβ+ and TCRγδ+ IELs is markedly increased. It is unclear whether this is a response to changes in the homeostasis of the epithelium, or a consequence of the pro-inflammatory milieu created by the CD4+ T cell response in the lamina propria (Fig. 3). Similar to IELs of normal controls, IELs from active CD express NK cell receptors. When compared to normal controls, however, IELs from CD patients have acquired a more activating NK cell receptor repertoire (Jabri et al. 2000). In the healthy small intestine, IELs predominantly express the inhibitory CD94/NKG2A receptor. In contrast, IELs in CD express high levels of activating receptors like CD94/NKG2C and NKG2D (Meresse et al. 2004, 2006). Simultaneously, intestinal epithelial cells in CD upregulate MIC and HLA-E, the ligands for NKG2D and CD94/NKG2C, respectively. Interaction of NKG2D and CD94/NKG2C with their ligands will enhance IFN-γ production and cytolysis, leading to tissue damage (Fig. 3). An important factor in acquiring an activating NK cell receptor repertoire is interleukin 15 (IL-15; Box 2), which has been shown to upregulate both NKG2D and CD94/NKG2C on IELs of active CD patients and boost their ability to lyse enterocytes (Meresse et al. 2004, 2006). In addition, IL-15 can alter the NK cell receptor function, leading to NK cell receptor-mediated cytotoxicity independent of TCR specificity (Meresse et al. 2004, 2006). In conclusion, while gluten-specific CD4+ T cells elicit an inflammatory response in the lamina propria, IELs in the epithelium acquire activating NK receptors and the ability to lyse stressed epithelial cells independent of T cell receptor signaling, which likely contributes to the typical tissue damage in CD.
Crosstalk between the CD4+ Tcell response and IEL cytotoxicity. The CD4+ T cell response to gluten may lead to IFN-γ production and potentially to upregulation of IL-15, which in turn boosts IEL-mediated cytotoxicity. Activated IELs lyse the epithelium, which leads to TG2 release and subsequent deamidation of gluten peptides. This may constitute yet another self-amplifying feedback loop, as deamidation of gluten peptides will enhance the CD4+ T cell response (see Fig. 2)
In RCD, the survival, expansion, and acquisition of an NK cell-like phenotype by IELs is even more pronounced than in CD, possibly as a result of the presence of larger amounts of IL-15. RCD II patients have an aberrant clonal IEL population that lacks surface TCR-CD3 expression. Studies on aberrant TCR-CD3- IEL lines from RCD II patients showed that, upon stimulation with IL-15, these cell lines express granzyme B and lyse the intestinal epithelial cell line HT29, suggesting a role for aberrant IELs in perpetuating epithelial damage in RCD II (Mention et al. 2003). Therefore, IL-15-dependent NK cell-like transformation of IELs may be an essential step in the immunopathology of RCD.
A threshold model for the risk of CD development
The expansion of the presentable gluten peptide repertoire due to the release and activity of TG2 is a critical step in the pathogenesis of full-blown CD. Several lines of evidence support the notion that the level of gluten presentation to T cells critically influences the risk of disease development.
First, HLA-DQ2.5 homozygous individuals have a fivefold higher risk of CD development than HLA-DQ2.5 heterozygous individuals (Mearin et al. 1983). This gene dose effect directly correlates with the magnitude of the CD4+ T cell response: antigen presenting cells (APC) from HLA-DQ2.5 homozygous individuals induce very strong proliferative T cell responses and IFN-γ production, while APC from HLA-DQ2.5/DQX heterozygous individuals induce much weaker responses (Vader et al. 2003b). These data indicate that the number of HLA-DQ2.5 molecules capable of presenting gluten peptides on the surface of APC will define the magnitude of the CD4+ T cell response.
Second, whereas HLA-DQ2.5 is associated with CD development, the homologous HLA-DQ2.2 is not. Although these two variants have almost identical peptide-binding motifs, HLA-DQ2.2 can only bind a subset of the gluten peptides that can bind to HLA-DQ2.5. This difference is explained by the fact that a proline at position 3 in peptides has an adverse effect on peptide binding to HLA-DQ2.2 (van de Wal et al. 1997). As gluten epitopes cluster in proline-rich regions (Arentz-Hansen et al. 2002), many gluten peptides have a proline at position 3 and do not bind to HLA-DQ2.2 (Vader et al. 2003b). Consequently, HLA-DQ2.5 is able to present a much broader repertoire of gluten peptides than HLA-DQ2.2. In addition, HLA-DQ2.5 is better at retaining gluten peptides in its binding groove compared to HLA-DQ2.2 (Fallang et al. 2009). As a result, gluten peptide presentation by HLA-DQ2.5 is protracted compared to presentation by HLA-DQ2.2, which will increase the chance for productive CD4+ T cell stimulation.
Third, CD is associated mainly with HLA-DQ2.5 and, to a lesser extent, with HLA-DQ8. Although a variety of gluten peptides has been identified that can stimulate HLA-DQ8 restricted T cells from CD patients, one α-gliadin peptide in particular appears to be immunodominant, as this peptide invariably induces specific T cell responses in HLA-DQ8+ CD patients (Henderson et al. 2007; Tollefsen et al. 2006; van de Wal et al. 1998b, 1999; Kooy et al. unpublished data). In contrast to the HLA-DQ2.5 restricted α-gliadin peptides, the HLA-DQ8 peptide is not derived from a proline-rich region of the α-gliadin protein, and therefore likely susceptible to degradation in the gastrointestinal tract. Furthermore, whereas for HLA-DQ2 a single deamidation in a gluten peptide is sufficient to evoke a CD4+ T cell response, for HLA-DQ8, deamidation at two positions is preferred (Henderson et al. 2007), which may limit the generation of strong antigenic gluten peptides. The fact that the immunodominant HLA-DQ8 peptide is more readily degraded and requires more deamidation steps, limits the availability for antigen presentation and may therefore limit the risk to develop CD.
Fourth, further evidence that the level of gluten presentation is a critical parameter comes from a totally different angle: most CD patients tolerate oat even though it has been shown that the gluten-like molecules in oat can elicit CD4+ T cell responses in CD patients (Arentz-Hansen et al. 2004; Vader et al. 2003a) There are two striking differences between the relatively safe oat and the disease-inducing cereals wheat, barley, and rye: (1) while the gluten-like molecules in oat contain only two antigenic sequences, dozens are found in gluten and the gluten-like molecules of barley and rye, (2) the “gluten” content of oat is much lower compared to the other cereals. Consumption of oat thus results in a much lower exposure to antigenic peptides, in comparison with the other cereals, and this is apparently tolerated, as it does not lead to disease in the majority of patients.
Collectively, these data indicate the presence of a threshold to develop CD. Initiation of CD becomes more likely with increased T cell exposure to gluten antigens. This exposure is influenced by the type and amount of HLA-DQ, as this determines the efficiency of gluten peptide presentation to CD4+ T cells. For HLA-DQ2.5 homozygous individuals, the threshold to develop CD is most easily exceeded, whereas for HLA-DQ2.2+ and HLA-DQ8+ individuals, the threshold is much higher.
CD development: a series of unfortunate events
The past two decades have witnessed the identification of several critical immunological factors in CD, from which a likely sequence of events in the development of this disease can be deduced. It is well known that healthy individuals can have antibodies against native gluten peptides. As an antibody response is controlled by CD4+ T cell help, such individuals most likely have CD4+ T cells specific for native gluten peptides, indicating that the mere presence of such T cells is, in general, not sufficient to exceed the threshold to develop CD (Fig. 4). This indicates that in the majority of individuals, tolerogenic and regulatory processes in the intestine keep gluten-specific T cell responses in check. This steady state can be breached by frequent episodes of enteroviral infections, as this leads to the secretion of inflammatory cytokines and differentiation of Th1 cells (Stene et al. 2006), thereby enhancing the response to gluten (Fig. 4). Subsequently, the combined effect of low-level gluten reactivity and pathogen-induced inflammation could lead to tissue damage and the release of TG2. In turn, the activity of TG2 would generate a large repertoire of deamidated gluten peptides with high affinity for HLA-DQ, thereby boosting the gluten-specific T cell response (Fig. 4). At this point in CD development, self-amplifying loops are in action: CD4+ T cell responses against native gluten peptides lead to IFN-γ production to upregulation of HLA, and further amplification of the gluten-specific T cell response. TG2 released upon tissue damage expands the presentable gluten peptide repertoire, which will ultimately lead to more tissue damage (Fig. 2). Due to the massive expansion of the gluten-specific T cell pool, regulatory processes are no longer able to contain the T cell responses, and exposure to gluten suffices to perpetuate inflammation. Eliminating gluten from the diet is the only way to stop this process. At all stages in this scenario, the risk to develop CD is influenced by the HLA-DQ genotype, as HLA-DQ2.5 homozygous individuals will present more HLA-gluten complexes than HLA-DQ2.5 heterozygous individuals (Fig. 4). Simultaneously, an infiltrate of IELs is formed in the epithelium, which is potentially driven by the inflammatory milieu created by CD4+ T cells in the lamina propria. These IELs upregulate activating NK cell receptors and acquire the ability to lyse enterocytes independent of the TCR.
Threshold model for CD development. As HLA-DQ2.5 homozygous individuals can present more gluten peptides on their APC than HLA-DQ2.5 heterozygous individuals, HLA-DQ2.5 homozygotes have a higher risk of CD development a priori. In steady state conditions, presentation of native gluten peptides is unlikely to induce disease. This steady state can be breached by frequent viral infections and low-grade T cell responses to native gluten peptides that lead to release of TG2 upon tissue damage. TG2 activity will expand the presentable gluten peptide repertoire extensively, thereby increasing the risk to develop CD. The recently uncovered non-HLA genes associated with CD likely lower the threshold to develop CD
The development of full-blown CD is most likely the result of an unfortunate series of events which, in isolation, would not lead to disease, but, combined, have a detrimental outcome. It is important to note that exposure to gluten, frequent enteroviral infections, and occasional TG2 activation likely occur in every individual, but usually do not result in CD development, even in HLA-DQ2+ and/or HLA-DQ8+ individuals (Fig. 1). It has now become clear that the presence of a higher number of additional non-HLA risk alleles (Table 1) is directly correlated with an increase in the risk to develop CD (Romanos et al. 2009). This suggests that the influence of non-HLA genes lowers the threshold to develop CD and could skew the balance towards disease development (Fig. 4). It is plausible that non-HLA genes also increase the risk to develop complicated CD, although this hypothesis will be difficult to test as this patient population is very small. Thus, even though key pieces of the celiac puzzle have been collected and assembled, the picture is not yet complete.
Open questions
Although the molecular basis for the involvement of HLA-DQ in CD is now well established, a number of issues remain unclear.
First, it is still controversial how immunogenic gluten peptides from the intestinal lumen reach the lamina propria where they can prime gluten-specific T cells. It has been suggested that gluten peptides can be transported during transient increased intestinal permeability during enteroviral infections (Stene et al. 2006) or by IgA-mediated retrotranscytosis (Matysiak-Budnik et al. 2008). Yet, this issue is far from resolved.
Second, a direct effect of gluten on the intestinal mucosa has been attributed to a peptide from α-gliadin, p31-49 (Box 3). It remains unclear, however, whether and how this peptide would exert its activity and contribute to the disease development (Box 3).
Third, TG2 is a crucial factor in expanding the presentable gluten peptide repertoire. In steady state conditions, TG2 is present in an inactive form intracellular and on the cell surface. An intriguing question is, therefore, how TG2 is activated and released in CD. We propose that TG2 is released upon tissue damage induced by the initial CD4+ T cell response to native gluten peptides (Figs. 2 and 3). Alternatively, a recent study proposed a role for TLR3 ligands released during enteroviral infections that, upon ligation with TLR3, could result in TG2 activation (Siegel et al. 2008). These two possibilities are not mutually exclusive.
Fourth, in active CD, disrupted IL-15 regulation results in massive overexpression of IL-15. It remains unclear what causes this disruption. As on a gluten-free diet, the adaptive CD4+ T cell response and IL-15 expression both decrease (Mention et al. 2003); it is possible that the adaptive CD4+ T cell response has a direct effect on IL-15 expression (Fig. 3). Alternatively, innate signals delivered through TLRs may be responsible for elevated IL-15 levels.
Finally, the events leading from uncomplicated CD to RCD II and subsequent lymphoma development are still poorly understood. Lymphoma cells develop from the aberrant IELs in RCD II. One view is that aberrant IELs derive from mature TCR+ IELs that have undergone oligoclonal expansion and lost surface TCR-CD3 expression due to overstimulation (Cellier et al. 1998). Alternatively, aberrant IELs could derive from a distinct population of CD3− CD7+ precursor cells that can develop into T cells and NK cells (Gunther et al. 2005). In favor of the first hypothesis: although aberrant IELs lack surface TCR-CD3 expression, they do express CD3 intracellularly and display TCR-γ-gene rearrangements (Malamut et al. 2009). Furthermore, microarray analysis in one study on TCR-CD3+ IEL lines from CD patients revealed a significant decrease in the transcript levels of TCRα- and TCRβ-chains (Meresse et al. 2006), indicating that IELs may lower TCR expression in CD. We found that aberrant IELs not only express CD3ε intracellularly, but also have intracellular expression of the CD3γ, CD3δ, and ζ-chains (Tjon et al. 2008). In favor of the second hypothesis: the TCR chains were not always present (Tjon et al. 2008), and TCR rearrangements were often incomplete (Tjon et al. unpublished data). Furthermore, the full complement of CD3 chains and incomplete TCR rearrangements have also been observed in NK cell precursors, and even mature NK cells can carry partially rearranged TCRs. A recent study indicated that extrathymic TCR-gene rearrangement is an ongoing event in the human small intestine throughout life (Bas et al. 2009). This raises the possibility that aberrant IELs derive from cells in an early stage of extrathymic lymphocyte development.
Concluding remarks
Life used to be simple: CD was a rare disease, diagnosed in 1 in 1,000 individuals. Patients were HLA-DQ2+ of HLA-DQ8+ and could be treated effectively with a gluten-free diet. That was about it.
Now we know that CD affects ~1% of the population in Western Europe and the USA, most of which remain undiagnosed. Although good insight has been gained on the immunopathology of CD—inflammation in both lamina propria and epithelium—it remains unclear what triggers the development of CD and why not every patient is equally affected. In addition, with the recognition of RCD and RCD-associated lymphoma that do not respond to a gluten-free diet, CD has become a far more complicated disease.
References
Al-Toma A, Goerres MS, Meijer JW, Pena AS, Crusius JB, Mulder CJ (2006) Human leukocyte antigen-DQ2 homozygosity and the development of refractory celiac disease and enteropathy-associated T-cell lymphoma. Clin Gastroenterol Hepatol 4:315–319
Al-Toma A, Verbeek WH, Hadithi M, von Blomberg BM, Mulder CJ (2007) Survival in refractory coeliac disease and enteropathy-associated T-cell lymphoma: retrospective evaluation of single-centre experience. Gut 56:1373–1378
Arentz-Hansen H, McAdam SN, Molberg O, Fleckenstein B, Lundin KE, Jorgensen TJ, Jung G, Roepstorff P, Sollid LM (2002) Celiac lesion T cells recognize epitopes that cluster in regions of gliadins rich in proline residues. Gastroenterology 123:803–809
Arentz-Hansen H, Fleckenstein B, Molberg O, Scott H, Koning F, Jung G, Roepstorff P, Lundin KE, Sollid LM (2004) The molecular basis for oat intolerance in patients with celiac disease. PLoS Med 1:e1
Bas A, Forsberg G, Sjoberg V, Hammarstrom S, Hernell O, Hammarstrom ML (2009) Aberrant extrathymic T cell receptor gene rearrangement in the small intestinal mucosa: a risk factor for coeliac disease? Gut 58:189–195
Bauer S, Groh V, Wu J, Steinle A, Phillips JH, Lanier LL, Spies T (1999) Activation of NK cells and T cells by NKG2D, a receptor for stress-inducible MICA. Science 285:727–729
Benahmed M, Meresse B, Arnulf B, Barbe U, Mention JJ, Verkarre V, Allez M, Cellier C, Hermine O, Cerf-Bensussan N (2007) Inhibition of TGF-beta signaling by IL-15: a new role for IL-15 in the loss of immune homeostasis in celiac disease. Gastroenterology 132:994–1008
Bourgey M, Calcagno G, Tinto N, Gennarelli D, Margaritte-Jeannin P, Greco L, Limongelli MG, Esposito O, Marano C, Troncone R, Spampanato A, Clerget-Darpoux F, Sacchetti L (2007) HLA related genetic risk for coeliac disease. Gut 56:1054–1059
Budagian V, Bulanova E, Paus R, Bulfone-Paus S (2006) IL-15/IL-15 receptor biology: a guided tour through an expanding universe. Cytokine Growth Factor Rev 17:259–280
Cellier C, Patey N, Mauvieux L, Jabri B, Delabesse E, Cervoni JP, Burtin ML, Guy-Grand D, Bouhnik Y, Modigliani R, Barbier JP, Macintyre E, Brousse N, Cerf-Bensussan N (1998) Abnormal intestinal intraepithelial lymphocytes in refractory sprue. Gastroenterology 114:471–481
Ciclitira PJ, Evans DJ, Fagg NL, Lennox ES, Dowling RH (1984) Clinical testing of gliadin fractions in coeliac patients. Clin Sci Lond 66:357–364
Daum S, Cellier C, Mulder CJ (2005) Refractory coeliac disease. Best Pract Res Clin Gastroenterol 19:413–424
Deleeuw RJ, Zettl A, Klinker E, Haralambieva E, Trottier M, Chari R, Ge Y, Gascoyne RD, Chott A, Muller-Hermelink HK, Lam WL (2007) Whole-genome analysis and HLA genotyping of enteropathy-type T-cell lymphoma reveals 2 distinct lymphoma subtypes. Gastroenterology 132:1902–1911
Di Sabatino A, Ciccocioppo R, Cupelli F, Cinque B, Millimaggi D, Clarkson MM, Paulli M, Cifone MG, Corazza GR (2006) Epithelium derived interleukin 15 regulates intraepithelial lymphocyte Th1 cytokine production, cytotoxicity, and survival in coeliac disease. Gut 55:469–477
Dicke WK, Weijers HA, van de Kamer JH (1953) Coeliac disease II. The presence in wheat of a factor having a deleterious effect in cases of coeliac disease. Acta Paediatr 42:34–42
Dube C, Rostom A, Sy R, Cranney A, Saloojee N, Garritty C, Sampson M, Zhang L, Yazdi F, Mamaladze V, Pan I, Macneil J, Mack D, Patel D, Moher D (2005) The prevalence of celiac disease in average-risk and at-risk Western European populations: a systematic review. Gastroenterology 128:S57–S67
Dubois PC, Trynka G, Franke L, Hunt KA, Romanos J, Curtotti A, Zhernakova A, Heap GA, Adany R, Aromaa A, Bardella MT, van den Berg LH, Bockett NA, de la Concha EG, Dema B, Fehrmann RS, Fernandez-Arquero M, Fiatal S, Grandone E, Green PM, Groen HJ, Gwilliam R, Houwen RH, Hunt SE, Kaukinen K, Kelleher D, Korponay-Szabo I, Kurppa K, Macmathuna P, Maki M, Mazzilli MC, McCann OT, Mearin ML, Mein CA, Mirza MM, Mistry V, Mora B, Morley KI, Mulder CJ, Murray JA, Nunez C, Oosterom E, Ophoff RA, Polanco I, Peltonen L, Platteel M, Rybak A, Salomaa V, Schweizer JJ, Sperandeo MP, Tack GJ, Turner G, Veldink JH, Verbeek WH, Weersma RK, Wolters VM, Urcelay E, Cukrowska B, Greco L, Neuhausen SL, McManus R, Barisani D, Deloukas P, Barrett JC, Saavalainen P, Wijmenga C, van Heel DA (2010) Multiple common variants for celiac disease influencing immune gene expression. Nat Genet 42:295–302
Ebert EC (1998) Interleukin 15 is a potent stimulant of intraepithelial lymphocytes. Gastroenterology 115:1439–1445
Ebert EC (2009) Interleukin 21 up-regulates perforin-mediated cytotoxic activity of human intra-epithelial lymphocytes. Immunology 127:206–215
Falchuk ZM, Rogentine GN, Strober W (1972) Predominance of histocompatibility antigen HL-A8 in patients with gluten-sensitive enteropathy. J Clin Invest 51:1602–1605
Fallang LE, Bergseng E, Hotta K, Berg-Larsen A, Kim CY, Sollid LM (2009) Differences in the risk of celiac disease associated with HLA-DQ2.5 or HLA-DQ2.2 are related to sustained gluten antigen presentation. Nat Immunol 10:1096–1101
Fehniger TA, Caligiuri MA (2001) Interleukin 15: biology and relevance to human disease. Blood 97:14–32
Ferguson A, Macdonald TT, McClure JP, Holden RJ (1975) Cell-mediated immunity to gliadin within the small-intestinal mucosa in coeliac disease. Lancet 1:895–897
Greco L, Babron MC, Corazza GR, Percopo S, Sica R, Clot F, Fulchignoni-Lataud MC, Zavattari P, Momigliano-Richiardi P, Casari G, Gasparini P, Tosi R, Mantovani V, De VS, Iacono G, D'Alfonso A, Selinger-Leneman H, Lemainque A, Serre JL, Clerget-Darpoux F (2001) Existence of a genetic risk factor on chromosome 5q in Italian coeliac disease families. Ann Hum Genet 65:35–41
Green PH, Cellier C (2007) Celiac disease. N Engl J Med 357:1731–1743
Gunther U, Holloway JA, Gordon JN, Knight A, Chance V, Hanley NA, Wilson DI, French R, Spencer J, Steer H, Anderson G, Macdonald TT (2005) Phenotypic characterization of CD3-7+ cells in developing human intestine and an analysis of their ability to differentiate into T cells. J Immunol 174:5414–5422
Henderson KN, Tye-Din JA, Reid HH, Chen Z, Borg NA, Beissbarth T, Tatham A, Mannering SI, Purcell AW, Dudek NL, van Heel DA, McCluskey J, Rossjohn J, Anderson RP (2007) A structural and immunological basis for the role of human leukocyte antigen DQ8 in celiac disease. Immunity 27:23–34
Hunt KA, Zhernakova A, Turner G, Heap GA, Franke L, Bruinenberg M, Romanos J, Dinesen LC, Ryan AW, Panesar D, Gwilliam R, Takeuchi F, McLaren WM, Holmes GK, Howdle PD, Walters JR, Sanders DS, Playford RJ, Trynka G, Mulder CJ, Mearin ML, Verbeek WH, Trimble V, Stevens FM, O'Morain C, Kennedy NP, Kelleher D, Pennington DJ, Strachan DP, McArdle WL, Mein CA, Wapenaar MC, Deloukas P, McGinnis R, McManus R, Wijmenga C, van Heel DA (2008) Newly identified genetic risk variants for celiac disease related to the immune response. Nat Genet 40:395–402
Jabri B, Ebert E (2007) Human CD8+ intraepithelial lymphocytes: a unique model to study the regulation of effector cytotoxic T lymphocytes in tissue. Immunol Rev 215:202–214
Jabri B, de Serre NP, Cellier C, Evans K, Gache C, Carvalho C, Mougenot JF, Allez M, Jian R, Desreumaux P, Colombel JF, Matuchansky C, Cugnenc H, Lopez-Botet M, Vivier E, Moretta A, Roberts AI, Ebert EC, Guy-Grand D, Brousse N, Schmitz J, Cerf-Bensussan N (2000) Selective expansion of intraepithelial lymphocytes expressing the HLA-E-specific natural killer receptor CD94 in celiac disease. Gastroenterology 118:867–879
Karell K, Louka AS, Moodie SJ, Ascher H, Clot F, Greco L, Ciclitira PJ, Sollid LM, Partanen J (2003) HLA types in celiac disease patients not carrying the DQA1*05-DQB1*02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease. Hum Immunol 64:469–477
Keuning JJ, Pena AS, van LA, van Hooff JP, Va Rood JJ (1976) HLA-DW3 associated with coeliac disease. Lancet 1:506–508
Lorand L, Graham RM (2003) Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat Rev Mol Cell Biol 4:140–156
Lundin KE, Scott H, Hansen T, Paulsen G, Halstensen TS, Fausa O, Thorsby E, Sollid LM (1993) Gliadin-specific, HLA-DQ(alpha 1*0501, beta 1*0201) restricted T cells isolated from the small intestinal mucosa of celiac disease patients. J Exp Med 178:187–196
Lundin KE, Scott H, Fausa O, Thorsby E, Sollid LM (1994) T cells from the small intestinal mucosa of a DR4, DQ7/DR4, DQ8 celiac disease patient preferentially recognize gliadin when presented by DQ8. Hum Immunol 41:285–291
Maiuri L, Ciacci C, Ricciardelli I, Vacca L, Raia V, Auricchio S, Picard J, Osman M, Quaratino S, Londei M (2003) Association between innate response to gliadin and activation of pathogenic T cells in coeliac disease. Lancet 362:30–37
Malamut G, Afchain P, Verkarre V, Lecomte T, Amiot A, Damotte D, Bouhnik Y, Colombel JF, Delchier JC, Allez M, Cosnes J, Lavergne-Slove A, Meresse B, Trinquart L, Macintyre E, Radford-Weiss I, Hermine O, Brousse N, Cerf-Bensussan N, Cellier C (2009) Presentation and long-term follow-up of refractory celiac disease: comparison of type I with type II. Gastroenterology 136:81–90
Matysiak-Budnik T, Moura IC, Arcos-Fajardo M, Lebreton C, Menard S, Candalh C, Ben-Khalifa K, Dugave C, Tamouza H, van NG, Bouhnik Y, Lamarque D, Chaussade S, Malamut G, Cellier C, Cerf-Bensussan N, Monteiro RC, Heyman M (2008) Secretory IgA mediates retrotranscytosis of intact gliadin peptides via the transferrin receptor in celiac disease. J Exp Med 205:143–54
McGowan KE, Lyon ME, Butzner JD (2008) Celiac disease and IgA deficiency: complications of serological testing approaches encountered in the clinic. Clin Chem 54:1203–1209
Mearin ML, Biemond I, Pena AS, Polanco I, Vazquez C, Schreuder GT, de Vries RR, van Rood JJ (1983) HLA-DR phenotypes in Spanish coeliac children: their contribution to the understanding of the genetics of the disease. Gut 24:532–537
Mention JJ, Ben AM, Begue B, Barbe U, Verkarre V, Asnafi V, Colombel JF, Cugnenc PH, Ruemmele FM, McIntyre E, Brousse N, Cellier C, Cerf-Bensussan N (2003) Interleukin 15: a key to disrupted intraepithelial lymphocyte homeostasis and lymphomagenesis in celiac disease. Gastroenterology 125:730–745
Meresse B, Chen Z, Ciszewski C, Tretiakova M, Bhagat G, Krausz TN, Raulet DH, Lanier LL, Groh V, Spies T, Ebert EC, Green PH, Jabri B (2004) Coordinated induction by IL15 of a TCR-independent NKG2D signaling pathway converts CTL into lymphokine-activated killer cells in celiac disease. Immunity 21:357–366
Meresse B, Curran SA, Ciszewski C, Orbelyan G, Setty M, Bhagat G, Lee L, Tretiakova M, Semrad C, Kistner E, Winchester RJ, Braud V, Lanier LL, Geraghty DE, Green PH, Guandalini S, Jabri B (2006) Reprogramming of CTLs into natural killer-like cells in celiac disease. J Exp Med 203:1343–1355
Molberg O, McAdam SN, Korner R, Quarsten H, Kristiansen C, Madsen L, Fugger L, Scott H, Noren O, Roepstorff P, Lundin KE, Sjostrom H, Sollid LM (1998) Tissue transglutaminase selectively modifies gliadin peptides that are recognized by gut-derived T cells in celiac disease. Nat Med 4:713–717
Nistico L, Fagnani C, Coto I, Percopo S, Cotichini R, Limongelli MG, Paparo F, D'Alfonso S, Giordano M, Sferlazzas C, Magazzu G, Momigliano-Richiardi P, Greco L, Stazi MA (2006) Concordance, disease progression, and heritability of coeliac disease in Italian twins. Gut 55:803–808
Parrish-Novak J, Dillon SR, Nelson A, Hammond A, Sprecher C, Gross JA, Johnston J, Madden K, Xu W, West J, Schrader S, Burkhead S, Heipel M, Brandt C, Kuijper JL, Kramer J, Conklin D, Presnell SR, Berry J, Shiota F, Bort S, Hambly K, Mudri S, Clegg C, Moore M, Grant FJ, Lofton-Day C, Gilbert T, Rayond F, Ching A, Yao L, Smith D, Webster P, Whitmore T, Maurer M, Kaushansky K, Holly RD, Foster D (2000) Interleukin 21 and its receptor are involved in NK cell expansion and regulation of lymphocyte function. Nature 408:57–63
Price P, Witt C, Allcock R, Sayer D, Garlepp M, Kok CC, French M, Mallal S, Christiansen F (1999) The genetic basis for the association of the 8.1 ancestral haplotype (A1, B8, DR3) with multiple immunopathological diseases. Immunol Rev 167:257–274
Romanos J, van Diemen CC, Nolte IM, Trynka G, Zhernakova A, Fu J, Bardella MT, Barisani D, McManus R, van Heel DA, Wijmenga C (2009) Analysis of HLA and non-HLA alleles can identify individuals at high risk for celiac disease. Gastroenterology 137(834–40):840
Shan L, Molberg O, Parrot I, Hausch F, Filiz F, Gray GM, Sollid LM, Khosla C (2002) Structural basis for gluten intolerance in celiac sprue. Science 297:2275–2279
Shan L, Qiao SW, Arentz-Hansen H, Molberg O, Gray GM, Sollid LM, Khosla C (2005) Identification and analysis of multivalent proteolytically resistant peptides from gluten: implications for celiac sprue. J Proteome Res 4:1732–1741
Siegel M, Strnad P, Watts RE, Choi K, Jabri B, Omary MB, Khosla C (2008) Extracellular transglutaminase 2 is catalytically inactive, but is transiently activated upon tissue injury. PLoS ONE 3:e1861
Sollid LM, Markussen G, Ek J, Gjerde H, Vartdal F, Thorsby E (1989) Evidence for a primary association of celiac disease to a particular HLA-DQ alpha/beta heterodimer. J Exp Med 169:345–350
Spurkland A, Ingvarsson G, Falk ES, Knutsen I, Sollid LM, Thorsby E (1997) Dermatitis herpetiformis and celiac disease are both primarily associated with the HLA-DQ (alpha 1*0501, beta 1*02) or the HLA-DQ (alpha 1*03, beta 1*0302) heterodimers. Tissue Antigens 49:29–34
Stene LC, Honeyman MC, Hoffenberg EJ, Haas JE, Sokol RJ, Emery L, Taki I, Norris JM, Erlich HA, Eisenbarth GS, Rewers M (2006) Rotavirus infection frequency and risk of celiac disease autoimmunity in early childhood: a longitudinal study. Am J Gastroenterol 101:2333–2340
Sturgess R, Day P, Ellis HJ, Lundin KE, Gjertsen HA, Kontakou M, Ciclitira PJ (1994) Wheat peptide challenge in coeliac disease. Lancet 343:758–761
Tjon JM, Verbeek WH, Kooy-Winkelaar YM, Nguyen BH, van der Slik AR, Thompson A, Heemskerk MH, Schreurs MW, Dekking LH, Mulder CJ, van Bergen J, Koning F (2008) Defective synthesis or association of T-cell receptor chains underlies loss of surface T-cell receptor-CD3 expression in enteropathy-associated T-cell lymphoma. Blood 112:5103–5110
Tollefsen S, Arentz-Hansen H, Fleckenstein B, Molberg O, Raki M, Kwok WW, Jung G, Lundin KE, Sollid LM (2006) HLA-DQ2 and -DQ8 signatures of gluten T cell epitopes in celiac disease. J Clin Invest 116:2226–2236
Trynka G, Zhernakova A, Romanos J, Franke L, Hunt KA, Turner G, Bruinenberg M, Heap GA, Platteel M, Ryan AW, de KC H, GK HPD, Walters JR, Sanders DS, Mulder CJ, Mearin ML, Verbeek WH, Trimble V, Stevens FM, Kelleher D, Barisani D, Bardella MT, McManus R, van Heel DA, Wijmenga C (2009) Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappaB signalling. Gut 58:1078–1083
Vader LW, De RA, van der Wal Y, Kooy YM, Benckhuijsen W, Mearin ML, Drijfhout JW, van VP, Koning F (2002) Specificity of tissue transglutaminase explains cereal toxicity in celiac disease. J Exp Med 195:643–649
Vader LW, Stepniak DT, Bunnik EM, Kooy YM, De HW, Drijfhout JW, Van Veelen PA, Koning F (2003a) Characterization of cereal toxicity for celiac disease patients based on protein homology in grains. Gastroenterology 125:1105–1113
Vader W, Stepniak D, Kooy Y, Mearin L, Thompson A, van Rood JJ, Spaenij L, Koning F (2003b) The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc Natl Acad Sci USA 100:12390–12395
van Belzen MJ, Meijer JW, Sandkuijl LA, Bardoel AF, Mulder CJ, Pearson PL, Houwen RH, Wijmenga C (2003) A major non-HLA locus in celiac disease maps to chromosome 19. Gastroenterology 125:1032–1041
van de Wal Y, Kooy Y, van VP, Pena S, Mearin L, Papadopoulos G, Koning F (1998a) Selective deamidation by tissue transglutaminase strongly enhances gliadin-specific T cell reactivity. J Immunol 161:1585–1588
van de Wal Y, Kooy YM, Drijfhout JW, Amons R, Papadopoulos GK, Koning F (1997) Unique peptide binding characteristics of the disease-associated DQ(alpha 1*0501, beta 1*0201) vs the non-disease-associated DQ(alpha 1*0201, beta 1*0202) molecule. Immunogenetics 46:484–492
van de Wal Y, Kooy YM, van Veelen PA, Pena SA, Mearin LM, Molberg O, Lundin KE, Sollid LM, Mutis T, Benckhuijsen WE, Drijfhout JW, Koning F (1998b) Small intestinal T cells of celiac disease patients recognize a natural pepsin fragment of gliadin. Proc Natl Acad Sci USA 95:10050–10054
van de Wal Y, Kooy YM, van VP, Vader W, Vader W, August SA, Drijfhout JW, Pena SA, Koning F (1999) Glutenin is involved in the gluten-driven mucosal T cell response. Eur J Immunol 29:3133–3139
van Heel DA, Hunt K, Greco L, Wijmenga C (2005) Genetics in coeliac disease. Best Pract Res Clin Gastroenterol 19:323–339
van Heel DA, Franke L, Hunt KA, Gwilliam R, Zhernakova A, Inouye M, Wapenaar MC, Barnardo MC, Bethel G, Holmes GK, Feighery C, Jewell D, Kelleher D, Kumar P, Travis S, Walters JR, Sanders DS, Howdle P, Swift J, Playford RJ, McLaren WM, Mearin ML, Mulder CJ, McManus R, McGinnis R, Cardon LR, Deloukas P, Wijmenga C (2007) A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat Genet 39:827–829
Verkarre V, Romana SP, Cellier C, Asnafi V, Mention JJ, Barbe U, Nusbaum S, Hermine O, Macintyre E, Brousse N, Cerf-Bensussan N, Radford-Weiss I (2003) Recurrent partial trisomy 1q22-q44 in clonal intraepithelial lymphocytes in refractory celiac sprue. Gastroenterology 125:40–46
Vilppula A, Kaukinen K, Luostarinen L, Krekela I, Patrikainen H, Valve R, Maki M, Collin P (2009) Increasing prevalence and high incidence of celiac disease in elderly people: a population-based study. BMC Gastroenterol 9:49
Wolters VM, Wijmenga C (2008) Genetic background of celiac disease and its clinical implications. Am J Gastroenterol 103:190–195
Zhernakova A, Elbers CC, Ferwerda B, Romanos J, Trynka G, Dubois PC, de Kovel CG, Franke L, Oosting M, Barisani D, Bardella MT, Joosten LA, Saavalainen P, van Heel DA, Catassi C, Netea MG, Wijmenga C (2010) Evolutionary and functional analysis of celiac risk loci reveals SH2B3 as a protective factor against bacterial infection. Am J Hum Genet 86:970–977
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Tjon, J.ML., van Bergen, J. & Koning, F. Celiac disease: how complicated can it get?. Immunogenetics 62, 641–651 (2010). https://doi.org/10.1007/s00251-010-0465-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-010-0465-9