Abstract
The alpine meadows of the Qinghai–Tibet Plateau have significant potential for storing soil carbon, which is important to global carbon sequestration. Grazing is a major threat to its potential for carbon sequestration. However, grazing poses a major threat to this potential by speeding up the breakdown of organic matter in the soil and releasing carbon, which may further lead to positive carbon-climate change feedback and threaten ecological security. Therefore, in order to accurately explore the driving mechanism and regulatory factors of soil organic matter decomposition in grazing alpine meadows on the Qinghai–Tibet Plateau, we took the grazing sample plots of typical alpine meadows as the research object and set up grazing intensities of different life cycles, aiming to explore the relationship and main regulatory factors of grazing on soil organic matter decomposition and soil microorganisms. The results show the following: (1) soil microorganisms, especially Acidobacteria and Acidobacteria, drove the decomposition of organic matter in the soil, thereby accelerating the release of soil carbon, which was not conducive to soil carbon sequestration in grassland; (2) the grazing triggering effect formed a positive feedback with soil microbial carbon release, accelerating the decomposition of organic matter and soil carbon loss; and (3) the grazing ban and light grazing were more conducive to slowing down soil organic matter decomposition and increasing soil carbon sequestration.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As a major ecological barrier to China and even Asia, the Qinghai‒Tibet Plateau (QTP) is a sensitive and ecologically fragile area of global change with great environmental significance and responsibility [2, 45]. Alpine meadows on the QTP have 1–5 times greater soil carbon sequestration capacity than other vegetation types [3, 15]. These meadows sequester 94% of the soil carbon across the QTP [15], playing a crucial role in the global soil carbon cycle [29, 55]. The QTP is also China’s largest alpine pastoral region, supporting nearly 5.3 million people through alpine livestock production by grazing. Additionally, the production value of the QTP is highly significant [15, 46, 48]. Uncontrolled grazing disrupts soil carbon balance in grasslands by altering soil microorganism communities, increasing soil CO2 release, and decreasing soil carbon content [23, 24, 40]. This increase in soil carbon emissions from alpine meadows due to grazing may lead to positive feedback and threaten ecological security. In recent years, 19–60% of the natural grasslands on the QTP have been degraded due to unscientifically managed grazing [46], and harmonizing production and ecological functions has become a priority for the regional development of the Tibetan Plateau. Therefore, it is necessary to accurately identify the regulatory factors of soil degradation in the alpine meadow of the Tibetan Plateau driven by grazing.
New evidence suggests that soil microbes strongly influence atmospheric CO2 concentrations and key ecosystem carbon processes by participating in and driving the turnover of soil organic matter [3]. Grazing alters vegetation and environmental conditions, soil water content, soil pH, and species interactions, leading to remodeling of soil microbial communities and imbalances in soil ecological functions [5, 21, 23, 24, 43, 53]. Soil microbial community structure and function changes can be characterized by changes in soil microbial biomass [13]. These are positively correlated with microbial function and associated with soil function [61]. Therefore, changes in soil microbial biomass nutrients are important indicators of the impact of grazing on soil [4]. The impact of grazing varied due to the diverse ecosystem environments at the observation sites, leading to inconsistent responses [15, 23, 24, 27]. However, the grazing response of soil microorganisms was also inconsistent due to the different ecosystem environments in the observation sites. Water is the main regulator in the desert steppe [56, 57, 60], while organic matter was observed to be the important factor of grazing microorganisms effect in typical grasslands and alpine meadows [9, 33, 56, 57]. In addition, grazing intensity and length of grazing time also contribute to the difference in grazing effects. Heavy grazing can disrupt the relative stability of grassland ecosystems [26], reduce soil microbial diversity [58], and lead to grassland degradation and soil carbon loss. Grazing bans and light grazing can increase soil microbial diversity and ecosystem stability [36]. The grazing effect is also related to the grazing duration, and short-term grazing is not enough to offset the resilience of ecosystems [28], and the grazing effect and its regulatory factors cannot be accurately observed [36]. Overall, the synergistic effects of vegetation, grazing intensity, grazing effect accumulation time, and soil microorganisms jointly regulated grazing responses in grazing land [7, 18, 39]. Although a large number of experiments and observational studies have been carried out, the main driving mechanisms and regulatory factors of soil microbial grazing effects in alpine meadows on the QTP are still uncertain. To investigate the microbial factors and regulatory mechanisms responsible for soil carbon release due to grazing in alpine meadows on the Qinghai–Tibet Plateau, we selected a representative alpine meadow with a 10-year history of grazing as our research subject. We used two different grazing intensities to better understand the impact of grazing. We proposed the following hypotheses: (1) The variation of soil organic matter content is dominated by one or several soil microorganisms. (2) Soil microbial biomass carbon is the main source of soil organic matter content. The aim of this study was to explore the main driving factors and regulatory mechanisms of grazing soil carbon release and to provide support for the coordinated development of ecology and economy in alpine pastoral areas.
Materials and Methods
Site Description and Experimental Design
This study was conducted at the Haibei National Field Research Station of the Alpine Grassland Ecosystem (37°37′ N, 101°19′ E), which is located in the northeastern part of the Qinghai‒Tibet Plateau. The area has a typical plateau continental climate with a long cold season from October to April and a short warm season from May to September. The mean annual temperature is − 1.2 °C, with a maximum of 9.9 °C in July and a minimum of − 15.2 °C in January [10]. The mean annual precipitation is 489 mm, with 80% concentrated in the growing season. The main vegetation type is alpine meadow, with 44% of the area accounting for alpine meadow. Its soil carbon storage accounts for 56% of the QTP [54]. Soils of the alpine meadow are the most widespread type, supporting a variety of vegetation [14]. The species in these soils are Kobresia Willd plants, and the vegetation density is low. The vegetation group at the sample site was Carex moorcroftii Falc. ex Boott and Carex alatauensis S. R. Zhang; common auxiliary species include Carex atrofusca Schkuhr subsp. minor (Boott) T. Koy, Poa crymophila Keng, and Deyeuxia flavens Keng [47].
The study focused on the alpine swamp meadow. The soil had high moisture content, and the vegetation thrived. Detailed information is provided in Table 1. The experimental site was used as an experimental site beginning in 2009. The experimental treatments were carried out continuously during that period. The plots included grazing prohibition (CK) and different intensity grazing groups. The feed intake, calculated from the livestock carrying capacity of the grassland, was used as the standard for setting grazing intensity. The maximum feed intake in the designated area was 30% for light grazing (LG) and 70% for heavy grazing [41, 51]. The geographical location and layout of the sample plots are shown in Fig. 1. The area of the closed grazing group was 3 × 15 m2, there was no human interference except for sampling throughout the year, and the area of the grazing treatment group was 9 × 9 m2. For the grazing group, warm season grazing experiments were carried out from May to September every year, the grazing livestock were Tibetan sheep, the grazing intensity treatment groups were grazed continuously for 48 h at the beginning of each month, and the standard and heavy grazing groups were grazed by 3 and 12 Tibetan sheep, respectively, with consistent ages and body conditions. The standard grazing group and the heavy grazing group grazed 3 and 12 Tibetan sheep of the same age and body condition, respectively. The sheep were allowed to feed and move freely and then released from the experimental area after 48 h, after which the grazing experiment was complete.
Study area profile map. a The geographical location and vegetation cover information of the experimental plots. b, c The geographical information and soil characteristics of the experimental sample, which was collected at the edge of the swamp wetland, and the vegetation was of the type of alpine swamp meadow. d–f The grazing intensity of the sample plots
Research Methodology
Soil samples were collected during different periods of the growing season: the greening period (May), the full-grass period (July), the yellow-grass period (September), and the dead-grass period (October). Soil samples were collected 15 days after the end of every grazing during the growing season. using the “S” sampling method [42, 44]. At the time of collection, vegetation and weathered material on the ground were removed first, and 1 cm of topsoil was removed from the surface layer [42, 44]. After removal, a soil perforator with a diameter of 3.5 cm was inserted into the soil. In order to better reflect the microbial characteristics, the surface layer of the 0–10 cm soil [35, 42, 44] as a soil sample. The soil samples for each grazing intensity treatment were taken 6 times, with 6 replications in total. The samples were temporarily stored at low temperatures and then transported to the laboratory for subsequent analysis. For the soil holes caused by sample collection, the soil of the natural grazing pasture at the periphery of the sample site was used to fill in to minimize human disturbance.
The soil samples collected from different grazing intensities were divided equally into three parts for each replicate, one for the analysis and determination of soil nutrient content and one for the determination of soil microbial biomass carbon(MBC) and soil microbial biomass nitrogen(MBN). The soil samples used for soil nutrient analysis were placed in a cool and dry environment and allowed to dry naturally. After air-drying, the soil samples were sieved to remove plant roots, stones, and other impurities, and after removing the impurities, the soil was milled for the determination of soil organic matter(SOM), total nitrogen(TN), and pH. The SOM content was determined via the potassium dichromate oxidation method (Qian [34]). The soil TN content was determined by the Kjeldahl method. The potentiometric method determined the soil pH [59]. For another fresh soil sample used for the MBC and MBN assays, the soil was sieved to remove impurities and then extracted using chloroform fumigation leaching [17, 37]. The soil solution was extracted by fumigating the soil with chloroform, and the difference between fumigated and unfumigated soil carbon and nitrogen was used to calculate the soil microbial MBC and MBN [4]. Finally, the remaining soil is reserved for crucial microflora testing.
Detection of Relative and Absolute Abundance of Soil Microbes
The Accu16S Assay is the precise method for assessing microorganisms' relative and absolute abundance. Accu16S (Accurate 16S absolute quantification sequencing) was meticulously performed by Genesky Biotechnologies Inc., Shanghai, 201,315 (China). First, total genomic DNA was extracted using the FastDNA SPIN Kit for Soil (MP Biomedicals, Santa Ana, CA) per the manufacturer’s instructions [6]. The integrity of genomic DNA was rigorously examined through agarose gel electrophoresis, and the concentration and purity were meticulously analyzed using the Nanodrop 2000 and Qubit 3.0 Spectrophotometer. Carefully designed spike-ins with conserved regions akin to natural 16S rRNA genes and variable regions replaced by random sequences with about 40% GC content were deliberately synthesized. A precisely calibrated gradient of spike-ins with varying copy numbers was methodically added to the sample DNA. Subsequently, the V4-V5 hypervariable regions of the 16S rRNA gene and spike-ins were amplified using the primers 515F (5′-GTGCCAGCMGCCGCGG-3′) and 907R (5′-CCGTCAATTCMTTTRAGTTT-3′) and then meticulously sequenced using the Illumina NovaSeq 6000 sequencer [8].
The raw reads were processed in QIIME2. Adaptor and primer sequences were trimmed using the cutadapt plugin. Quality control and identification of amplicon sequence variants (ASVs) were efficiently carried out with the DADA2 plugin. Taxonomic assignments were meticulously performed using a pre-trained Naive Bayes classifier. Spike-in sequences were accurately identified, and reads were meticulously counted. The absolute copy number of each ASV in each sample was precisely calculated based on the read counts. Additionally, the spike-in sequence must be removed in the subsequent analysis [19].
The quantitative results of the 16S rRNA gene qPCR were multiplied with the relative abundance of microorganisms to convert the absolute abundance [49].
Analysis of the Data
At the end of the experiment, the data were entered and organized in Microsoft Excel. The organized data were then imported into R and GraphPad Prism 8.4.3 for analysis and graphing. The R package “agricolae” was used for multiple comparisons with a significance level of p < 0.05. GraphPad Prism 8.4.3 was employed to verify the significance of the results and to create graphs. Significance levels are indicated by asterisks: *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001. Soil nutrient content and pH data were used to construct box-and-whisker plots and line plots using GraphPad Prism 8.4.3. Adobe Illustrator was used to combine the images, label the axes, and determine the significance of differences. Soil microbial MBC and MBN data were also used to create bar charts and line graphs with GraphPad Prism 8.4.3, with Adobe Illustrator used for image stitching and labeling. The significance of differences was assessed as described above. Finally, SPSS software version 24.0 (SOM, TN, PH, SMC, C/N, MBC, MBN, MBC/MBN) was utilized for statistical analysis, applying Duncan’s test for determining multiple comparisons of means. We calculated Spearman's correlation coefficient for pairwise comparison to determine the correlation between microbial composition, soil physicochemical properties, and microbial biomass carbon and nitrogen. We used Cytoscape v3.9.0 software for network analysis. We calculated Spearman's correlation coefficient for pairwise comparison to determine the correlation between microbial composition, soil physicochemical properties, and microbial biomass carbon and nitrogen.
Results
Changes in Soil Functional Traits in Response to Grazing
The data showed the following: (1) A significant decrease in SMC (soil moisture content) was found in both the CK and the grazing treatment groups (LG and HG) during the growing season (July and September). The decrease in SMC in this growing season was not regulated by grazing but grazing would reduce SMC (Fig. 2a), and the water content of grazing land was significantly lower than CK before (May) and after (October) the growing season. (2) Soil pH was significantly affected by grazing intensity and grazing reduced soil pH, which was the lowest value under heavy grazing (Fig. 2c). (3) SOM was significantly increased in the growing season due to the dual effects of season and grazing, and the highest content was found in the heavily grazed land (HG) (Fig. 2e, f). (4) The soil C/N and SOM trend was consistent and significantly regulated by growing season and grazing intensity (Fig. 2i, j).
Diagram of the response of soil functional traits to grazing intensity and its relationship with phenology. a, b The response of the soil moisture content (SMC) to grazing and its relationship with months; c, d the response of the soil pH to grazing and its relationship with months; e, f the response of the soil organic matter (SOM) content to grazing and its relationship with months; g, h the response of the soil total nitrogen (TN) content to grazing and its relationship with months
Changes in Soil Microbial Biomass Nutrients in Response to Grazing
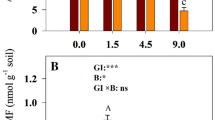
Through analysis and detection, the data results showed the following: (1) MBC increased significantly in the growing season, and the trend was consistent in CK and both LG and HG (Fig. 3a, b). (2) MBC and MBN showed opposite trends in response to grazing intensity, with grazing increasing MBN content while decreasing MBC content (Fig. 3c, d). (3) The stoichiometric analysis of MBC and MBN showed that MBCMBN had the same trend as MBC, significantly increasing in the growing season, but significantly decreasing in grazing land (Fig. 3e, f).
Response of soil microbial biomass carbon and nitrogen to grazing and its relationship with month. a, b The change in soil microbial biomass carbon (MBC) in response to grazing intensity and its relationship with month, and c, d the change in soil microbial biomass nitrogen (MBN) in response to grazing intensity and its relationship with month. e, f The change in MBC/MBN
Soil Microbial Community Composition Under Varying Levels of Grazing Intensity
After grazing was analyzed through high-throughput sequencing of soil microbial species composition, the results showed the following: (1) soil bacteria dominant species of Proteobacteria, at the gate level, are Acidobacteria, Actinobacteria, Planctomycetes, and Chloroflexi (Fig. 4a); the dominant species at the generic level are Gp6, Gp4, Gp17, Gp7, and Nitrospira (Fig. 4b). (2) Phylum and genus-level inundations were stable, and genera-level inundations of dominant species increased with the increase of grazing intensity (Fig. 4). (3) The advantage of soil Fungi in gate level for Ascomycota, Rozellomycota, Fungi_phy_Incertae_sedis, Basidiomycota, and Mortierellomycota (Fig. 5a) belongs to the level of dominant species to GS11_gen_Incertae_sedis, Fungi_phy_Incertae_sedis, Mortierella, Thelebolus, and Stagonospora (Fig. 5b). (4) The abundance of fungi at phylum level and genus level showed a decreasing trend with the increase of grazing intensity (Fig. 5).
The diversity of soil bacteria and fungi was analyzed. The results showed the following: (1) Grazing significantly increased the bacterial richness (Fig. 6a, f), and the highest richness was achieved under HG. (2) The diversity of beta in the grazing group was different from that in the CK group but overlapped between LG and HG (Fig. 7). (3) Grazing increased the alpha diversity of soil fungi, and the LG diversity index was the highest, while the HG diversity was significantly decreased (Fig. 8). (4) Grazing also caused the differentiation of soil fungi, and there were significant differences between the CK group and LG and HG groups (Fig. 9a, b).
Correlation Analysis of Soil Microbial Composition with Soil Physicochemical Properties and MBC and MBN
The Spearman correlation analysis revealed several significant findings: (1) Bacterial genera were strongly correlated with changes in MBC (soil microbial biomass carbon) and MBN (soil microbial biomass nitrogen) content. Notably, MBC and MBN exhibited a highly significant positive correlation (Fig. 10). (2) The bacterial genera associated with MBC and MBN were largely the same and showed a significant positive correlation. The impact of bacteria was markedly higher than that of fungi, as indicated in Table 2. (3) Specifically, the bacterium Gaiella displayed a notable positive correlation with MBC but demonstrated no correlation with MBN. Additionally, apart from the bacterial flora, SOM (soil organic matter) also exhibited a significant positive correlation with MBC but not with MBN (Table 2). (4) Upon analyzing the absolute abundance of bacteria, it was observed that the abundance of Gaiella in the LG (low grazing) and HG (high grazing) groups was higher than that in the CK (control) group (Fig. 11a). Furthermore, the abundance increased significantly with the rise in grazing intensity (Fig. 11b).
Spearman correlation analysis of soil bacteria and fungi with physical and chemical properties and soil microbial biomass carbon and nitrogen nodes with a correlation coefficient greater than 0.5 were selected. The red line represents a positive correlation, while the blue line represents a negative correlation. The size of the nodes and the yellow-to-red gradient indicate the number of correlations; nodes with more correlations are larger and redder, while those with fewer correlations are lighter and smaller
Distribution of dominant bacterial genera at different grazing intensities (a). Absolute abundances were converted by multiplying the 16S rRNA gene qPCR quantification results by the relative abundance of microorganisms. Absolute quantitative absolute abundance value of the dominant bacterium Gaiella (b)
Discussion
Growing Season Effects of Changes in SMC and SOM Contents
During this study, grazing experiments were carried out throughout the entire lifecycle of the alpine meadow. The lifecycle was divided into three phases: May for the regreening period, July to September for the grass-bearing period, and October for the yellowing period. It was observed that there were noticeable seasonal variations in soil moisture content (SMC) and soil organic matter (SOM) (Fig. 2b, f). The SMC content decreased significantly during the growing season (July–September). This decreasing trend was also observed in the control group (CK) and the two grazing groups, LG and HG. Similarly, SOM content increased significantly throughout the growing season and significantly correlated with grazing intensity. The highest SOM content was recorded in the HG group during the growing season. The seasonal changes in SMC and SOM were associated with the vigorous growth of plants and soil microorganisms during the growing season.
Based on a two-factor analysis of months and grazing intensity, seasons significantly impacted the SMC (Fig. 2b). On the other hand, SOM was substantially influenced by both the growing season and grazing. Factors related to vegetation growth, closely tied to SMC, play a crucial role. During the growing season, vegetation and soil microorganisms experience robust growth and absorb significant water from the soil for their own development and reproduction. Additionally, the plateau’s growing season for vegetation coincides with the highest temperature and strongest radiation, leading to surface evaporation and the decline of SMC (G. [23, 24]). The rise in SOM content due to the growing season and grazing is likely attributed to soil microorganisms. Consequently, the community composition and abundance of soil microorganisms were examined after grazing, and correlation analysis was carried out by selecting dominant strains. The results revealed that while microbial composition did not correlate significantly with SOM content, MBC demonstrated a significant positive correlation with SOM (Fig. 10). Consequently, the seasonal increase in SOM can be primarily attributed to soil microorganisms' increased activated carbon content.
In addition, correlation analysis also showed a significant negative correlation between SMC content and dominant flora and a significant negative correlation between MBC and MBN (Fig. 10, Table 2). In particular, seven microbial genera, Gaiella, Mortierella, GS11_gen_Incertae_sedis, Solicoccozyma, Stagonospora, Dactylonectria, and Clavaria, which were significantly positively correlated with MBC content, were significantly or extremely negatively correlated with SMC. The decrease of soil moisture content during the growing season promoted the proliferation of soil microorganisms, thereby increasing total soil organic matter.
These results support our initial hypothesis that the soil organic matter content variation is dominated by one or several soil microorganisms. The rise in soil organic matter during the growing season declined after the growing season. A C/N stoichiometric analysis also revealed a negative correlation between C/N and SOM, with a correlation coefficient of less than 0.5 (− 0.357). This suggests that the rate of soil carbon mineralization was high, and the increase in soil carbon content resulting from microbial proliferation came from the decomposition of soil carbon by microorganisms. This could lead to soil carbon loss and is not beneficial to soil carbon sequestration. Furthermore, grazing expedited this process.
Microbial-Dominated Soil Carbon Decomposition in Grazing Grassland
Grazing, as a kind of human disturbance, strongly affects the pattern of the grassland ecosystem and material cycle, and there is a synergistic effect [43, 50]. First, grazing livestock increased grassland litter and decreased aboveground biomass through feeding and trampling and caused changes in the soil environment, mainly manifested in decreased soil pH and SMC (Fig. 2) [12]. Second, the excretion of grazing livestock, as an important nutrient return, strongly influences the material cycle [1]. Soil microorganisms are important drivers of the geochemical cycle [25] and depend on the soil environment, so the environmental changes caused by grazing also act on microorganisms [61], resulting in grazing effects.
Grazing livestock directly damages the soil ecosystem environment through feeding and trampling, thereby affecting vegetation and soil microbial growth (Ma [27]). In this study, the SMC, SOM, and pH of grazing land were significantly different from those of CK soil (Fig. 2), and this environmental change was an important factor affecting the growth of vegetation and soil microorganisms. In addition, the early return of litter and manure input caused by grazing also provided a carbon source for the growth and reproduction of some soil microorganisms, and some studies have proved that grazing causes changes in the dominant species of soil microbial communities and the well-nutrient bacterial communities become the dominant species, thereby accelerating the decomposition of organic matter and the release of CO2 [52]. The results of this study showed that bacteria had stronger interaction with SOM in grazing soil (Fig. 10), and the dominant bacteria in Acidobacteria, Actinobacteria, and Nitrospirae had the greatest effect (Fig. 11a). Research showed that Gp6 in Acidobacteria, Gp7, and other bacteria have the effect of promoting litter decomposition [32]. The bacteria in Acidobacteria promote decomposition and nutrient release by producing enzymes to dissolve cellulose and hemicellulose in pasture litter [20] While dominant bacteria such as Gaiella in Actinobacteria are related to the absorption of phosphate and decomposition of organic matter [38], they appeared in the late stage of decomposition of organic matter [16], which can promote the decomposition of organic matter and produce antibiotics to inhibit the growth of other microorganisms such as fungi, thus forming advantages [31]. In this study, MBC and MBN have basically the same positive correlation flora. Compared with MBN, MBC has more positive interaction between Gaiella and SOM (Table 2), which is the reason for the increase of MBC content in the growing season. Another dominant bacterium, Nitrospirae, is a kind of autotrophic nitrifier whose increased abundance is conducive to nitrogen decomposition [11], providing nitrogen support for the growth of plants and microorganisms [30]. Therefore, the increase of soil SOM and MBC in grazing grassland during the growing season was due to the stimulating effect of grazing on microorganisms [22]. MBC and activated carbon produced by soil microorganisms accelerate carbon and nitrogen decomposition to release nutrients and accelerate reproduction are the main sources of SOM, driving the carbon decomposition of grassland soil in the growing season. Compared with CK, more nutrient input in grazing land provides more carbon sources for microorganisms, so the contents of SOM and MBC increase significantly. The results confirm our hypothesis that MBC is the main source of SOM content variation. Although SOM and MBC levels increase during the growing season, they return to their original levels, preventing effective soil carbon storage. The rapid release of soil carbon by microorganisms hinders carbon sequestration in grazing grasslands, with grazing further accelerating carbon release. A more moderate grazing approach is recommended to slow carbon release and enhance soil carbon fixation to address this. Finding ways to regulate and reduce the rate of carbon release by microorganisms could boost the carbon storage capacity of grassland soil, ultimately contributing to global carbon neutrality.
Conclusion
According to the results of this study, the following conclusions were drawn: (1) Soil microorganisms, especially Acidobacteria and Acidobacteria, drove the decomposition of organic matter in the soil, thereby accelerating the release of soil carbon, which was not conducive to soil carbon sequestration in grassland. (2) The grazing triggering effect formed a positive feedback with soil microbial carbon release, accelerating the decomposition of organic matter and soil carbon loss. (3) The grazing ban and light grazing were more conducive to slowing down soil organic matter decomposition and increasing soil carbon sequestration.
Data Availability
No datasets were generated or analysed during the current study.
References
Aarons S, O’Connor C, Hosseini H, Gourley C (2009) Dung pads increase pasture production, soil nutrients and microbial biomass carbon in grazed dairy systems. Nutr Cycl Agroecosyst 84(1):81–92. https://doi.org/10.1007/s10705-008-9228-5
Badgery W, Kemp D, Yingjun Z, Zhongwu W, Guodong H, Fujiang H, Nan L, Michalk D, Behrendt K (2020) Optimising grazing for livestock production and environmental benefits in Chinese grasslands. Rangel J 42(5):347–358. https://doi.org/10.1071/RJ20049
Bai Y, Cotrufo M (2022) Grassland soil carbon sequestration: current understanding, challenges, and solutions. Science 377(6606):603–608. https://doi.org/10.1126/science.abo2380
Bai Y, Zha X, Chen S (2020) Effects of the vegetation restoration years on soil microbial community composition and biomass in degraded lands in Changting County China. J For Res 31(4):1295–1308. https://doi.org/10.1007/s11676-019-00879-z
Banerjee S, Helgason B, Wang L, Winsley T, Ferrari B, Siciliano S (2016) Legacy effects of soil moisture on microbial community structure and N2O emissions. Soil Biol Biochem 95:40–50. https://doi.org/10.1016/j.soilbio.2015.12.004
Bolyen E, Rideout J, Dillon M, Bokulich N, Abnet C, Al-Ghalith G, Alexander H, Alm E, Arumugam M, Asnicar F, Bai Y, Bisanz J, Bittinger K, Brejnrod A, Brislawn C, Brown C, Callahan B, Caraballo-Rodríguez A, Chase J, … Caporaso J (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2 (vol 37, pg 852, 2019). Nat Biotechnol 37(9), 1091–1091. https://doi.org/10.1038/s41587-019-0252-6
Buchkowski R, Bradford M, Grandy A, Schmitz O, Wieder W (2017) Applying population and community ecology theory to advance understanding of belowground biogeochemistry. Ecol Lett 20(2):231–245. https://doi.org/10.1111/ele.12712
Callahan B, McMurdie P, Rosen M, Han A, Johnson A, Holmes S (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581. https://doi.org/10.1038/NMETH.3869
Chen D, Li Q, Huo L, Xu Q, Chen X, He F, Zhao L (2023) Soil nutrients directly drive soil microbial biomass and carbon metabolism in the Sanjiangyuan Alpine Grassland. J Soil Sci Plant Nutrit 23(3):3548–3560. https://doi.org/10.1007/s42729-023-01270-y
Chen Y, Han M, Yuan X, Hou Y, Qin W, Zhou H, Zhao X, Klein J, Zhu B (2022) Warming has a minor effect on surface soil organic carbon in alpine meadow ecosystems on the Qinghai-Tibetan Plateau. Glob Change Biol 28(4):1618–1629. https://doi.org/10.1111/gcb.15984
Daims H, Lebedeva E, Pjevac P, Han P, Herbold C, Albertsen M, Jehmlich N, Palatinszky M, Vierheilig J, Bulaev A, Kirkegaard R, von Bergen M, Rattei T, Bendinger B, Nielsen P, Wagner M (2015) Complete nitrification by Nitrospira bacteria. Nature 528(7583):504. https://doi.org/10.1038/nature16461
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15(10):579–590. https://doi.org/10.1038/nrmicro.2017.87
Hao Z, Zhao Y, Wang X, Wu J, Jiang S, Xiao J, Wang K, Zhou X, Liu H, Li J, & Sun Y (2021). Thresholds in aridity and soil carbon-to-nitrogen ratio govern the accumulation of soil microbial residues. Commun Earth Environ 2(1). https://doi.org/10.1038/s43247-021-00306-4
Hou Y, Chen Y, Chen X, He K, Zhu B (2019) Changes in soil organic matter stability with depth in two alpine ecosystems on the Tibetan Plateau. Geoderma 351:153–162. https://doi.org/10.1016/j.geoderma.2019.05.034
Huang Y, Xin Z, Liu J, & Liu Q. (2022). Divergences of soil carbon turnover and regulation in alpine steppes an meadows on the Tibetan Plateau. Sci Total Environ 814. https://doi.org/10.1016/j.scitotenv.2021.152687
Javed Z, Tripathi GD, Mishra M, Dashora K (2021) Actinomycetes – the microbial machinery for the organic-cycling, plant growth, and sustainable soil health. Biocatal Agric Biotechnol 31:101893. https://doi.org/10.1016/j.bcab.2020.101893
Jenkinson D, Brookes P, Powlson D (2004) Measuring soil microbial biomass. Soil Biol Biochem 36(1):5–7. https://doi.org/10.1016/j.soilbio.2003.10.002
Jing J, Minghua S (2010) Review of the roles of plants and soil microorganisms in regulating ecosystem nutrient cycling. Acta Phytoecologica Sinica 34(8):979–988
Jiang S, Yu Y, Gao R, Wang H, Zhang J, Li R, Long X, Shen Q, Chen W, Cai F (2019) High-throughput absolute quantification sequencing reveals the effect of different fertilizer applications on bacterial community in a tomato cultivated coastal saline soil. Sci Total Environ 687:601–609. https://doi.org/10.1016/j.scitotenv.2019.06.105
Kalam S, Basu A, Ahmad I, Sayyed R, El-Enshasy H, Dailin D, & Suriani N (2020). Recent understanding of soil acidobacteria and their ecological significance: a critical review. Front Microbiol 11. https://doi.org/10.3389/fmicb.2020.580024
Lau J, Lennon J (2011) Evolutionary ecology of plant-microbe interactions: soil microbial structure alters selection on plant traits. New Phytol 192(1):215–224. https://doi.org/10.1111/j.1469-8137.2011.03790.x
Li C, Xiao C, Li M, Xu L, He N (2023) The quality and quantity of SOM determines the mineralization of recently added labile C and priming of native SOM in grazed grasslands. Geoderma 432:116385. https://doi.org/10.1016/j.geoderma.2023.116385
Li G, Zhang Z, Shi L, Zhou Y, Yang M, Cao J, Wu S, & Lei G (2018). Effects of different grazing intensities on Soil C, N, and P in an alpine meadow on the QinghaiTibetan Plateau, China. Int J Environ Res Public Health, 15(11). https://doi.org/10.3390/ijerph15112584
Li L, Zhu-Barker X, Ye R, Doane T, Horwath W (2018) Soil microbial biomass size and soil carbon influence the priming effect from carbon inputs depending on nitrogen availability. Soil Biol Biochem 119:41–49. https://doi.org/10.1016/j.soilbio.2018.01.003
Liang C, Schimel J, & Jastrow J (2017). The importance of anabolism in microbial control over soil carbon storage. Nat Microbiol 2(8). https://doi.org/10.1038/nmicrobiol.2017.105
Liu Y, Duan D, Jiang F, Tian Z, Feng X, Wu N, Hou F, Kardol P, Nan Z, Chen T (2022) Long-term heavy grazing increases community-level foliar fungal diseases by shifting plant composition. J Appl Ecol 59(3):791–800. https://doi.org/10.1111/1365-2664.14093
Ying Ma, Xiaokui K, Jie L, Yunfeng Y (2021) Response of soil greenhouse gases emissions to grazing in alpine grasslands:a review. Microbiology China 48(10):3835–3846
Macheroum A, & Chenchouni H (2022). Short-term land degradation driven by livestock grazing does not affect soil properties in semiarid steppe rangelands. Front Environ Sci 10. https://doi.org/10.3389/fenvs.2022.846045
Maestre F, Le Bagousse-Pinguet Y, Delgado-Baquerizo M, Eldridge D, Saiz H, Berdugo M, Gozalo B, Ochoa V, Guirado E, García-Gómez M, Valencia E, Gaitán J, Asensio S, Mendoza B, Plaza C, Díaz-Martínez P, Rey A, Hu H, He J, … Gross N (2022). Grazing and ecosystem service delivery in global drylands. Science, 378(6622), 915–920. https://doi.org/10.1126/science.abq4062
Nunes-Alves C (2016). Microbial Ecology Do it yourself nitrification. Nat Rev Microbiol 14(2). https://doi.org/10.1038/nrmicro.2015.20
Ouchene R, Intertaglia L, Zaatout N, Kecha M, Suzuki MT (2022) Selective isolation, antimicrobial screening and phylogenetic diversity of marine actinomycetes derived from the Coast of Bejaia City (Algeria), a polluted and microbiologically unexplored environment. J Appl Microbiol 132(4):2870–2882. https://doi.org/10.1111/jam.15415
Pankratov T, Ivanova A, Dedysh S, Liesack W (2011) Bacterial populations and environmental factors controlling cellulose degradation in an acidic Sphagnum peat. Environ Microbiol 13(7):1800–1814. https://doi.org/10.1111/j.1462-2920.2011.02491.x
Xiaoqian P, Wei W (2016) Spatial pattern of soil microbial biomass carbon and its driver in temperate grasslands of Inner Mongolia. Microbiology China 43(9):1918–1930
Bao Q, Ling L, Xiao X (2011) Comparative tests on different methods for content of soil organic matter. J Hohai Univ Nat Sci 39(1):34–38
Ren C, Liu K, Dou P, Shao X, Zhang D, Wang K, Liu X, Li J, Wang K (2022) Soil nutrients drive microbial changes to alter surface soil aggregate stability in typical grasslands. J Soil Sci Plant Nutrit 22(4):4943–4959. https://doi.org/10.1007/s42729-022-00972-z
H Ren, J Xie, Y Ke, Q Yu, A Lkhagva, M Liu, 2024 Moderate grazing weakens legacy effects of grazing history on primary productivity and biodiversity in a meadow grassland. AgricultEcosyst Environ 362 https://doi.org/10.1016/j.agee.2023.108825
Rotbart N, Borisover M, Bukhanovsky N, Beriozkin A, Eshel G, Bar-Tal A, Oren A (2020) The assessment of microbial biomass C in subsoil samples using fumigation-extraction is negligibly affected by residual chloroform. Arid Land Res Manage 34(4):474–482. https://doi.org/10.1080/15324982.2020.1767226
Severino R, Froufe H, Barroso C, Albuquerque L, Lobo-da-Cunha A, da Costa M, & Egas C (2019). High-quality draft genome sequence of Gaiella occulta isolated from a 150 meter deep mineral water borehole and comparison with the genome sequences of other deep-branching lineages of the phylum Actinobacteria. Microbiologyopen, 8(9). https://doi.org/10.1002/mbo3.840
Sharrar A, Crits-Christoph A, Meheust R, Diamond S, Starr E, & Banfield J (2020). Bacterial secondary metabolite biosynthetic potential in soil varies with phylum, depth, and vegetation type. Mbio, 11(3). https://doi.org/10.1128/mBio.00416-20
Sokol N, Slessarev E, Marschmann G, Nicolas A, Blazewicz S, Brodie E, Firestone M, Foley M, Hestrin R, Hungate B, Koch B, Stone B, Sullivan M, Zablocki O, Pett-Ridge J, LLNL Soil Microbiome Consortium (2022) Life and death in the soil microbiome: how ecological processes influence biogeochemistry. Nat Rev Microbiol 20(7):415–430. https://doi.org/10.1038/s41579-022-00695-z
Sollenberger LE, Aiken GE, & Wallau MO (2020). Chapter 5—Managing grazing in forage–livestock systems. In M Rouquette & GE Aiken (Eds.), Management Strategies for Sustainable Cattle Production in Southern Pastures (pp. 77–100). Academic Press. https://doi.org/10.1016/B978-0-12-814474-9.00005-0
Wang B, Wu L, Chen D, Wu Y, Hu S, Li L, Bai Y (2020) Grazing simplifies soil micro-food webs and decouples their relationships with ecosystem functions in grasslands. Glob Change Biol 26(2):960–970. https://doi.org/10.1111/gcb.14841
Wang H, Liu S, Chang S, Wang J, Shi Z, Huang X, Wen Y, Lu L, Cai D (2015) Soil microbial community composition rather than litter quality is linked with soil organic carbon chemical composition in plantations in subtropical China. J Soils Sediments 15(5):1094–1103. https://doi.org/10.1007/s11368-015-1118-2
Wang J, Zhao C, Zhao L, Wen J, & Li Q (2020). Effects of grazing on the allocation of mass of soil aggregates and aggregate-associated organic carbon in an alpine meadow. PloS One, 15(6). https://doi.org/10.1371/journal.pone.0234477
Wang S, Fan J, Li Y, & Huang L (2019). Effects of grazing exclusion on biomass growth and species diversity among various grassland types of the Tibetan Plateau. Sustainability, 11(6). https://doi.org/10.3390/su11061705
Wang Y, Lv W, Xue K, Wang S, Zhang L, Hu R, Zeng H, Xu X, Li Y, Jiang L, Hao Y, Du J, Sun J, Dorji T, Piao S, Wang C, Luo C, Zhang Z, Chang X, … Niu H (2022). Grassland changes and adaptive management on the Qinghai-Tibetan Plateau. Nat Rev Earth Environ 3(10), 668–683. https://doi.org/10.1038/s43017-022-00330-8
Wang Y, Pei W, Cao G, Guo X, Zhou H, & Du Y (2021). Moderate grazing increases water use efficiency for environmental health in alpine meadows of the Tibetan Plateau. Front Ecol Evol 9. https://doi.org/10.3389/fevo.2021.684321
Wei Y, Lu H, Wang J, Wang X, & Sun J (2022). Dual influence of climate change and anthropogenic activities on the spatiotemporal vegetation dynamics over the Qinghai-Tibetan Plateau From 1981 to 2015. Earths Fut 10(5). https://doi.org/10.1029/2021EF002566
Wen T, Xie P, Liu H, Liu T, Zhao M, Yang S, Niu G, Hale L, Singh B, Kowalchuk G, Shen Q, & Yuan J (2023). Tapping the rhizosphere metabolites for the prebiotic control of soil-borne bacterial wilt disease. Nat Commun 14(1). https://doi.org/10.1038/s41467-023-40184-2
J Xiao, S Dong, Z Zhao, Y Han, S Li, H Shen, C Ding 2021 Stabilization of soil organic carbon in the alpine meadow is dependent on the nitrogen deposition level on the Qinghai-Tibetan Plateau. EcolEng 170 https://doi.org/10.1016/j.ecoleng.2021.106348
Minyun Xu, Lijie G, Yunqi Li (2014) A review on grassland carrying capacity (II): Parameters and calculation methods. Acta Pratacul Sin 23(4):311–321
Xun W, Yan R, Ren Y, Jin D, Xiong W, Zhang G, Cui Z, Xin X, & Zhang R (2018). Grazing-induced microbiome alterations drive soil organic carbon turnover and productivity in meadow steppe. Microbiome, 6. https://doi.org/10.1186/s40168-018-0544-y
Yan Y, Kuramae E, de Hollander M, Klinkhamer P, van Veen J (2017) Functional traits dominate the diversity-related selection of bacterial communities in the rhizosphere. ISME J 11(1):56–66. https://doi.org/10.1038/ismej.2016.108
Yang Y, Fang J, Tang Y, Ji C, Zheng C, He J, Zhu B (2008) Storage, patterns and controls of soil organic carbon in the Tibetan grasslands. Glob Change Biol 14(7):1592–1599. https://doi.org/10.1111/j.1365-2486.2008.01591.x
Yang Y, Shi Y, Sun W, Chang J, Zhu J, Chen L, Wang X, Guo Y, Zhang H, Yu L, Zhao S, Xu K, Zhu J, Shen H, Wang Y, Peng Y, Zhao X, Wang X, Hu H, … Fang J (2022). Terrestrial carbon sinks in China and around the world and their contribution to carbon neutrality. Scie China-Life Sci 65(5), 861–895. https://doi.org/10.1007/s11427-021-2045-5
Yang Y, Zhang H, Liu W, Sun J, Zhao M, Han G, & Pan Q (2023). Effects of grazing intensity on diversity and composition of rhizosphere and non-rhizosphere microbial communities in a desert grassland. Ecol Evol 13(7). https://doi.org/10.1002/ece3.10300
Yang Z, Miao P, Zheng Y, Guo J, Li Y, Liu T, & He X (2023). Impacts of Grazing on Vegetation and Soil Physicochemical Properties in Northern Yinshan Mountain Grasslands. Sustainability, 15(22). https://doi.org/10.3390/su152216028
Yao Z, Shi L, He Y, Peng C, Lin Z, Hu M, Yin N, Xu H, Zhang D, & Shao X (2023). Grazing intensity, duration, and grassland type determine the relationship between soil microbial diversity and ecosystem multifunctionality in Chinese grasslands: a meta-analysis. Ecol Indicat 154. https://doi.org/10.1016/j.ecolind.2023.110801
Zbiral J (2001) Comparison of methods for soil pH determination. Rostlinna Vyroba 47(10):463–466
Zhang M, Delgado-Baquerizo M, Li G, Isbell F, Wang Y, Hautier Y, Wang Y, Xiao Y, Cai J, Pan X, & Wang L (2023). Experimental impacts of grazing on grassland biodiversity and function are explained by aridity. Nat Commun 14(1). https://doi.org/10.1038/s41467-023-40809-6
Zhou Z, Wang C, & Luo Y (2020). Meta-analysis of the impacts of global change factors on soil microbial diversity and functionality. Nat Commun 11(1). https://doi.org/10.1038/s41467-020-16881-7
Funding
This study was supported by the Qinghai Province Applied Basic Research Program (2023-ZJ-767) and the Qinghai Province Talented Scientist program (E439051D05).
Author information
Authors and Affiliations
Contributions
All the authors of this research paper directly participated in the planning, execution, or analysis of the study. Lin Wei (Data organization, article writing), Na Zhao, Shixiao Xu, (Data collection and organization to provide guidance and assistance for the article) Yalin Wang, Na Li, (Various help with article writing). All the authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wei, L., Wang, Y., Li, N. et al. Bacteria-Like Gaiella Accelerate Soil Carbon Loss by Decomposing Organic Matter of Grazing Soils in Alpine Meadows on the Qinghai–Tibet Plateau. Microb Ecol 87, 104 (2024). https://doi.org/10.1007/s00248-024-02414-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02414-y