Abstract
Plastic pollution poses a worldwide environmental challenge, affecting wildlife and human health. Assessing the biodegradation capabilities of natural microbiomes in environments contaminated with microplastics is crucial for mitigating the effects of plastic pollution. In this work, we evaluated the potential of landfill leachate (LL) and estuarine sediments (ES) to biodegrade polyethylene (PE), polyethylene terephthalate (PET), and polycaprolactone (PCL), under aerobic, anaerobic, thermophilic, and mesophilic conditions. PCL underwent extensive aerobic biodegradation with LL (99 ± 7%) and ES (78 ± 3%) within 50–60 days. Under anaerobic conditions, LL degraded 87 ± 19% of PCL in 60 days, whereas ES showed minimal biodegradation (3 ± 0.3%). PE and PET showed no notable degradation. Metataxonomics results (16S rRNA sequencing) revealed the presence of highly abundant thermophilic microorganisms assigned to Coprothermobacter sp. (6.8% and 28% relative abundance in anaerobic and aerobic incubations, respectively). Coprothermobacter spp. contain genes encoding two enzymes, an esterase and a thermostable monoacylglycerol lipase, that can potentially catalyze PCL hydrolysis. These results suggest that Coprothermobacter sp. may be pivotal in landfill leachate microbiomes for thermophilic PCL biodegradation across varying conditions. The anaerobic microbial community was dominated by hydrogenotrophic methanogens assigned to Methanothermobacter sp. (21%), pointing at possible syntrophic interactions with Coprothermobacter sp. (a H2-producer) during PCL biodegradation. In the aerobic experiments, fungi dominated the eukaryotic microbial community (e.g., Exophiala (41%), Penicillium (17%), and Mucor (18%)), suggesting that aerobic PCL biodegradation by LL involves collaboration between fungi and bacteria. Our findings bring insights on the microbial communities and microbial interactions mediating plastic biodegradation, offering valuable perspectives for plastic pollution mitigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plastics have become indispensable in modern society, playing a crucial role in various economic sectors such as agriculture and industry [1, 2]. However, their exponential production, insufficient recycling, and plastics’ slow degradation under natural conditions have resulted in the accumulation of plastic waste in ecosystems [3, 4]. Plastic waste represents 80–85% of the total marine litter [5], and a significant portion of plastic waste collected in the European Union is incinerated or buried in landfills [6]. New eco-friendly solutions for plastic waste treatment are necessary, and improved biodegradation strategies can have an important role in reducing plastic pollution.
Microplastics represent a significant environmental and health threat due to their small size and pervasive nature. Often invisible to the naked eye, microplastics are harder to detect and manage compared to larger plastic debris. They can spread more easily through the environment, reaching even the most remote areas. Moreover, microplastics can infiltrate various ecosystems, including oceans, rivers, soil, and the atmosphere, leading to widespread environmental and biological contamination. They can accumulate in the tissues of organisms, moving up the food chain and potentially impacting a wide range of species, including humans [7, 8].
Conventional plastics are synthetic and semi-synthetic polymers primarily derived from fossil carbon sources [9]. These materials are generally classified as non-biodegradable [10] and are extremely bio-inert, with a highly hydrophobic chain, which makes their biodegradation extremely difficult [1, 11, 12]. Amongst synthetic fossil-based plastics, polyethylene (PE) is the most widely produced [13], finding extensive use in the manufacture of bags, bottles, food packaging, and other products [11]. Due to the saturation of its chain with ethylene bonds, PE is a very hydrophobic polymer, and one of the most recalcitrant [13]. Although PE is considered non-biodegradable, evidence of its biodegradation was shown by complex microbial communities (e.g., waxworm gut microbiome) and pure cultures of bacteria and fungi [14,15,16]. However, most often, PE biodegradation occurs only partially and at very slow rate.
Polyethylene terephthalate (PET) is also a versatile polymer widely used in various everyday products [11]. The prevalence of PET in single-use plastic products makes it a significant contributor to plastic pollution [17]. Some microorganisms and enzymes were reported to biodegrade PET, but the crystalline regions are extremely resistant to biological attack [18, 19].
Due to the challenges in achieving an efficient biodegradation of recalcitrant plastics, it is worthy to replace PE and PET by biodegradable alternatives. Biodegradable plastics are polymers that undergo more easily biodegradation by microorganisms and are generally broken down into smaller molecules, such as carbon dioxide and water, with minimal production of toxic compounds [5, 20]. Poly(ε-caprolactone) (PCL) is a synthetic, fossil-based polyester and widely used biodegradable plastic [21]. PCL presents numerous benefits compared to other biodegradable plastics, leading to a rise in its applications. For example, it is resistant to water, oil, solvents, and chlorine [22]. Due to its high biocompatibility, blend compatibility, and low biodegradability rates, it is frequently used in long-term biomedical and tissue applications [22,23,24]. Among biodegradable plastics, PCL can have extended biodegradation times, leading to its accumulation in the environment where it is discarded [22, 25]. Total PCL biodegradation may take few months to several years, depending on the environmental conditions, as well as with variations in PCL properties (e.g., molecular weight and crystallinity) [24, 26]. Nevertheless, biodegradation has been reported to occur both in natural environments and in engineering environments (e.g., sewage sludge and compost) [24, 27], either under anaerobic or aerobic conditions [1, 28]. Recently, a PCL-degrading bacterium was isolated from a plastic-contaminated landfill and presented high biodegradation rates, showing that landfill microbiomes can efficiently biodegrade PCL [24,25,26].
Environmental microbiomes have the ability to adapt to external stimuli, such as plastic pollution, and may harbor microbes with the ability to biodegrade plastics more efficiently than the ones currently known. Therefore, in this work, we tested two different environmental microbiomes (landfill leachate and estuarine sediments), for their ability to biodegrade PCL, PE, and PET microplastics. These microbiomes were chosen because they originate from environments typically contaminated with plastic. In fact, despite significant efforts to recycle plastic, a substantial proportion of these materials still ends up in municipal landfills. Additionally, plastics and microplastics are abundant in marine and estuarine environments, where they often accumulate in sediments. Although PE and PET are considered recalcitrant, their microbial biodegradation has been reported, and thus, given the unexplored potential of the inocula used in our study, it was important to test their biodegradation capabilities. This work is aimed at screening microplastic biodegradation (under different incubation conditions, including aerobic, anaerobic, mesophilic, and thermophilic conditions) and identifying potential microplastic-degrading microorganisms. The temperature conditions were selected considering the origin of each inoculum, i.e., mesophilic environment for the estuarine sediment and thermophilic environment for the landfill leachate. These environments are poorly explored regarding microplastic biodegradation, and thus, this work is aimed at broadening current knowledge on the microbiology of microplastic biodegradation, opening new perspectives for the development of efficient biotechnological solutions for plastic waste treatment.
Materials and Methods
Plastic Materials
Pellets of PE, PET, and PCL were synthesized in the Institute for Polymers and Composites, University of Minho. Pellets were mechanically grinded to obtain particles with a 1 mm diameter.
Inocula
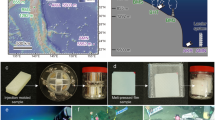
Biodegradation assays were conducted using inocula leachate from the municipal landfill (Resulima) in Viana do Castelo, Portugal, and estuarine sediments collected in the Cávado river estuary near Esposende, Portugal.
The leachate was transported to the laboratory and stored at 4 °C until further analysis. Before setting up the biodegradation assays, the leachate was concentrated by decantation and centrifugation (8000 g, 10 min). Volatile solids (VS) were determined as described elsewhere [30]. The leachate was incubated overnight at 55 °C to consume the residual substrate.
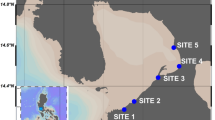
Estuarine sediments were sampled using a 6-mm diameter PVC tube, at the layer between 2 and 12 cm depth. Sediment was transported to the laboratory at 4 °C, homogenized, and sieved (5 mm). The salinity of the seawater was 20 g L−1. Before starting the assays, the sediment was incubated at 37 °C under saline conditions (20 g L−1 of NaCl in the aerobic assay; 10 g L−1 in the anaerobic assays), for 2 days, for consumption of residual substrate.
Biodegradation Assays
For both inocula, microbial degradation of plastics was investigated under anaerobic and aerobic conditions. Incubations were performed at 55–37 °C, in the assays inoculated with the leachate or the sediment, respectively. The good activity of the inocula was confirmed in control assays with microcrystalline cellulose (62.5 mg, average particle size 50 μm, Sigma-Aldrich, St. Louis, MO, USA). Methane production (in the anaerobic assays) and oxygen consumption (in the aerobic assays) started immediately, and cellulose biodegradation reached 95 ± 17% and 71 ± 11%, respectively. For the anaerobic assays, this is in agreement with the validation criterium defined by Fruteau De Laclos et al. and Holliger et al [29, 30], which states that microcrystalline cellulose conversion to methane should be 82–95% of the theoretical value.
Anaerobic Biodegradation Assays
Anaerobic biodegradation assays were performed in 120 mL bottles containing 45 mL of bicarbonate-buffered basal medium (BM), supplemented with salts and vitamins, as described by Stams et al. [31]. The assays with the sediment were also supplemented with NaCl (10 g L−1). This value is lower than the salinity measured in the seawater but was chosen considering that methanogenic microorganisms generally do not tolerate high salt concentrations in the medium [32, 33].
The bottles were loaded with 62.5 mg of each plastic in powder (PE, PET, and PCL) and inoculated with 1.5 mL of leachate (corresponding to a VS concentration of 1.4 g L−1) or 4.5 g of sediment (VS concentration of 2.4 g L−1). Bottles were closed with butyl rubber septa and aluminum crimp caps, and the headspace was flushed and pressurized with N2/CO2 (80:20%, v/v, 1.7 × 105 Pa). A reducing agent, Na2S•9H2O (1 mmol L−1), was added. These assays had no electron acceptors other than bicarbonate (methanogenic conditions). For the sediment, assays under sulfate-reducing (SR) conditions were also prepared, by adding sodium sulfate (20 mmol L−1). Blank tests, without the addition of plastics or any other carbon source, were also performed. All assays were conducted in triplicate, in the dark, with agitation at 150 rpm.
Methane was measured periodically. In the SR assays, sulfide was also measured to indirectly assess sulfate reduction. At the end of the incubation period, samples were collected from the assays with leachate and PCL, for VFA and microbial community composition analysis.
Aerobic Biodegradation Assays
The aerobic experiments were performed in closed bottles, to allow for the monitoring of oxygen consumption. Basal medium was used, composed of the following stock solutions (composition of the solutions in g L−1): 40 mL L−1 of solution (A) containing KH2PO4, 28.25 and K2HPO4, 146.08; 30 mL L−1 of solution (B) containing CaCl2•2H2O, 3.36 and NH4Cl, 28.64; and 30 mL L−1 of solution (C) containing MgSO4•7H2O, 3.06; FeSO4, 0.7; and ZnSO4, 0.4 [34]. NaCl (20 g L−1) was also added to the medium in the assays with the sediment. A volume of 50 mL of medium was added to each 120 mL bottle, along with the plastics and inocula (as described for the anaerobic biodegradation assays). Blank assays (without plastics or other added carbon source) were also performed. The bottles were sealed with butyl rubber septa and aluminum crimp caps and pressurized with atmospheric air. Oxygen content in the headspace of the bottles was immediately measured. All assays were conducted in triplicate, in the dark, with agitation (150 rpm). Oxygen measurements were performed twice a week. When the oxygen levels became low, the headspace was flushed with air, and fresh air was injected using a syringe. At the end of the incubation period, samples were collected from the assays with leachate and PCL, for analysis of the microbial community composition.
Analytical Methods
Methane (in the anaerobic assays) or oxygen (in the aerobic assays) was determined by gas chromatography (GC) using a GC BRUKER SCION 456 (Billerica, MA, USA), with a Molsieve packed column (13 × 80/100, 2 m length, 2.1 mm internal diameter) connected to a thermal conductivity detector. Argon was the carrier gas at a flow rate of 30 mL min−1, with temperatures of 100 °C, 35 °C, and 130 °C for injector, column, and detector, respectively.
For total dissolved sulfide analysis, samples were collected, immediately transferred to a zinc acetate solution (2% w/v), and analyzed with Hach cuvette tests (LCK653) and a DR 2800 spectrophotometer (Hach, Düsseldorf, Germany [35].
VFA analyses were carried out using HPLC equipment (Jasco, Tokyo, Japan) with a Rezex ROA-Organic Acid H+ (8%) LC Column (300 × 7.8 mm) at 60 °C. The mobile phase comprised a solution of sulfuric acid (5 mmol L−1), and crotonic acid was utilized as the internal standard. Elution was performed at a flow rate of 0.6 mL min−1, and compound detection occurred at 210 nm [36].
Data Analysis
Under anaerobic conditions, the biodegradability of the microplastics was determined considering the experimentally measured values of CH4 (BMPexp) and the theoretical biochemical CH4 production (BMPtheoric), according to Eq. 1.
The BMPtheoric was obtained from the element composition of the microplastics (C, H, N, O, and S), according to Reaction 1 and Eq. 2 [37]. BMPtheoric was expressed in volume of methane (L) at standard temperature and pressure (STP) per mass unit of plastic (g). The BMPtheoric values calculated for the different plastics in the study are shown in Table 1.
Reaction 1
Under aerobic conditions, the microplastics biodegradation (%) was calculated according to Eq. 3, where the O2,sample and O2,blank are the experimentally measured values of O2 consumed in sample and blank assays, respectively.
The theoretical O2 demand (ThOD) for complete aerobic mineralization of the polymer CcHhOo (expressed as mass of O2 per mass of polymer) was calculated according to Eq. 4, where Mr corresponds to the relative molecular mass of the polymer. The ThOD values determined for the different plastics in the study are shown in Table 2.
Microbial Community Composition
Microbial community composition was studied at the end of the PCL biodegradability assays inoculated with the landfill leachate, as well as in the inoculum leachate. Both prokaryotic and eukaryotic communities were investigated. Total DNA was isolated, and 16S rRNA and 18S rRNA genes were sequenced by Illumina MiSeq [38]. The primers used were the following: 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and (806R: 5′-GGACTACHVGGGTWTCTAAT-3′) [39], targeting the prokaryotic community, and EUK1391F (5′-GTACACACCGCCCGTC-3′) and EUKBR (5′-TGATCCTTCTGCAGGTTCACCTAC3′) [40] targeting the eukaryotic community. Sequencing and bioinformatics analyses were performed by RTL Genomics (Lubbock, TX, USA), and the detailed methodology is described by Salvador et al. (2019) [38]. The database used for taxonomic assignment consisted of high-quality sequences derived from NCBI that are maintained in the Research and Testing Laboratory (http://www.medicalbiofilm.org).
The FASTQ files were submitted to the European Nucleotide Archive (ENA) under the accession number PRJEB73311.
Mining Coprothermobacter Species Genomes for PCL Degrading Enzymes
The M-party tool (https://anaconda.org/bioconda/m-party) was harnessed to conduct an in-depth exploration of proteins potentially homologous to carboxylesterases, lipases, and cutinases, previously reported as capable to hydrolyze PCL [41,42,43]. The search was performed in the genomes of Coprothermobacter species, the ones which have their genomes sequenced (taxonomy_id:55,509, comprising the genome of Coprothermobacter proteolyticus (strain ATCC 35245and Coprothermobacter sp.), which are the closer relatives of the Coprothermobacter microorganisms found in our leachate sample. M-party compiled multiple enzymes obtained from the KEGG database corresponding to the EC numbers 3.1.1.1 (4466 enzymes), 3.1.1.3 (11,425 enzymes), and 3.1.1.74 (1350 enzymes). Clustering of sequences was performed with CD-HIT [44], using default parameters and an identity threshold of 70%. Multiple sequence alignment for each cluster was done with T-Coffee [45]. Finally, hidden Markov models (HMMs) were constructed with the multiple sequencing alignments by utilizing HMMER3 [46] using default settings.
Results and Discussion
Anaerobic Biodegradation Assays
Under anaerobic conditions, landfill leachate extensively biodegraded PCL with concomitant methane production. Cumulative methane production is represented in Fig. 1. After 60 days of incubation, the methane produced accounted for 87 ± 19% of the value that could be expected from complete PCL degradation (Table 1). These results are in accordance with the literature, with other authors reporting 80–92% PCL biodegradation in similar time periods [47,48,49]. A lag phase of approximately 10 days preceded the onset of methane production from PCL, probably due to microbial adaptation to the polymer and/or incubation conditions. The methane produced during the first 10 days of incubation in these assays closely followed the results of the blanks, pointing out that it was derived from the consumption of residual substrate still present in the inoculum. No VFA was detected at the end of the experiments.
For the estuarine sediment, PCL degradation started after 20 days of incubation (data not shown) and after 60 days corresponded only to 3.3 ± 0.3% biodegradation (considering the BMPtheoric in Table 1) (data not shown). This residual methane production can be attributed to the low methanogenic activity of the sediment. Indeed, estuarine environments are intermittently exposed to air and present relatively high salinity which does not favor the growth of strict anaerobes like methanogens [32, 33]. Low PCL biodegradation (2%) was also previously reported in batch assays, in this case using anaerobic sludge as inoculum, after 56 days of incubation [50]. In the assays with the sediment and sulfate, no sulfide production was observed throughout the experiment, pointing to the absence of sulfate-reducing microorganisms capable of degrading PCL.
Contrary to what was observed with PCL, in the assays with PE or PET cumulative methane production closely followed that of the blanks over all the experiment (Fig. 1 for leachate; data not shown for sediment). These results suggest that the methane produced was most likely derived from the consumption of residual substrates rather than from the biodegradation of the polymers. Similar results were obtained by other authors [51, 52]—for example, Selke et al. [51] reported that after 500 days of incubation, biogas production in assays containing these polymers did not exhibit a significant difference compared to the blank. When the sediment was incubated under sulfate-reducing conditions, no polymer biodegradation coupled with sulfate reduction was observed as well.
The disparity observed between PCL and the other tested polymers can be attributed to its higher susceptibility to microbial hydrolysis. Additionally, it presents a lower melting point (around 60 °C) [28], while PET is more susceptible to microbial attack within the temperature range of 75 to 80 °C [53]. These physical-chemical properties are a distinctive factor, resulting in different degradation profiles [24, 26].
Aerobic Biodegradation Assays
In the aerobic incubations, similarly to the anaerobic assays, PCL was the only microplastic biodegraded. PCL was consumed by the two inocula, as shown by the cumulative oxygen consumption curves obtained for the landfill leachate (Fig. 2a) and the estuarine sediment (Fig. 2b). Considering the ThOD values (Table 2), 99 ± 7% (50 days) and 78 ± 3% (63 days) of PCL mineralization were achieved by the leachate and the sediment, respectively.
Because PCL is a biodegradable plastic, it is expected that it should undergo biodegradation by environmental microbiomes and also under composting conditions. However, the efficiency of PCL biodegradation varies depending on the conditions applied and probably on the microbial composition of the microbiomes. Results similar to the ones obtained in our study were reported by other authors [24, 54], but, on the other hand, under aerobic composting conditions, Pradhan et al. [55] achieved only 60% biodegradation after 180 days. Variability in the results has also been reported across different studies regarding PCL biodegradation in marine environments, which were attributed to variations in experimental conditions, such as the use of artificial seawater and different sediment types (coastal or fluvial) [56].
Regarding PE and PET assays, both for inocula, the results obtained were similar to those observed in the blanks assays, revealing the lack of polymers’ biodegradation under aerobic conditions. This is in line with literature results, where mineralization was not observed with powdered PE during simulated composting at 58 °C under aerobic conditions [51].
PCL-Degrading Thermophilic Microbiomes
More extensive biodegradation of PCL was achieved in the assays inoculated with landfill leachate than in the ones with estuarine sediment, for a similar period. This fact might be related to the origin of each inoculum, and therefore, the composition of the microbial communities, as well as with the different experimental conditions applied (e.g., mesophile or thermophile temperature). Estuarine sediment is a natural inoculum, eventually contaminated by microplastics [57]. Landfill leachate results from a myriad of physical-chemical and biological processes during the decomposition of municipal wastes and is characterized by a miscellany of organic and xenobiotic compounds that induce a microbial community with the potential to present high diversity and metabolic functions [58].
Based on the promising results obtained with the landfill leachate, and considering that it originates from an understudied environment, with significant potential for harboring novel plastic-degrading microorganisms, this microbial community was further studied. Moreover, it is a thermophilic community, and not much is known about thermophilic microorganisms that can degrade plastic, which further reinforces the interest of this study. Utilizing thermophiles in biotechnological processes offers several advantages, namely, decreased viscosity of the medium and enhanced bioavailability and solubility of organic compounds, which results in higher reaction rates [59]. The use of stable enzymes at high temperatures prolongs hydrolysis of the polymers’ backbone [59, 60]. There are some known thermophilic microorganisms able to biodegrade PCL [60, 61], and leachate may be a source of microbes yet unknown with this capability.
PCL biodegradation involves several steps, starting with the hydrolysis of the polymer, which is usually the most difficult step. After hydrolysis, PCL monomers (6-hydroxycaproic acid) may undergo a cascade of reactions which finalizes in the TCA cycle [62, 63] (Fig. 3). It is still unknown whether this pathway occurs under thermophilic conditions and if it is performed by the majority of PCL biodegraders.
Anaerobic Community
Figure 4A displays the microbial community composition and the relative abundance of the microorganisms identified in the anaerobic assays with landfill leachate and PCL. Bacteria and Archaea accounted for 77% and 23% of the classified organisms, respectively. In a significant proportion of the retrieved sequences (~31%), taxonomic identification was only possible at the kingdom level (bacteria), possibly due to the scarce knowledge of plastic-biodegrading microorganisms and leachate microbiomes. Microorganisms from the Methanothermobacter (20.9% relative abundance), Caloramator (11.0%), Coprothermobacter (6.8%), Defluviitoga (6.1%), Acetomicrobium (2.3%), and Leucobacter (1.5%) genera predominated in the community, accounting for a total of 49% of the retrieved sequences.
Taxonomic characterization and relative abundance of the A prokaryotic microorganisms, given by 16S rRNA gene sequencing, of landfill leachate before (inoculum) and after the anaerobic and aerobic PCL biodegradation assays; and of the B eukaryotic microorganisms assigned to the fungi kingdom, obtained by 18S rRNA amplicon sequencing, at the end of the aerobic PCL biodegradability assays. Only OTU's with relative abundance higher than 1% were considered
Microorganisms assigned to the Methanothermobacter genus are thermophilic and produce methane by utilizing hydrogen as an energy source [64, 65]. Its high abundance in the anaerobic assays suggests that hydrogenotrophic methanogenesis has an important role in PCL conversion to methane. Still, Methanosaeta sp. (1.1% relative abundance, Fig. 4A) and Methanosarcina (0.5%) were also present in the community, showing that both hydrogenotrophic and acetoclatic pathways were occurring during PCL conversion to methane.
Coprothermobacter spp. are anaerobic thermophilic bacteria that possess substantial enzymatic activity, both intracellularly and extracellularly, particularly in proteolytic capabilities [66]. Moreover, these microbes are hydrogen producers and have been reported to facilitate interspecies hydrogen transfer and accelerate protein degradation when in co-culture with Methanothermobacter sp [66]. Jin et al. [49] reported a positive correlation between the relative abundance of Methanothermobacter sp. and Coprothermobacter sp. during the degradation of biodegradable plastics, including PCL. Similar syntrophic interactions can possibly be occurring in the anaerobic assays performed in this work with the landfill leachate and PCL, revealing novel microbial interactions in microplastic biodegradation.
Bacteria from the genus Caloramator (11% relative abundance) have not been associated with PCL biodegradation before [67, 68] but are able to ferment a wide range of substrates, including several carbohydrates derived from plant biomass. Therefore, their possible role in the hydrolysis of PCL may be hypothesized.
Lactobacillus spp. and Pseudomonas spp. were also identified in the anaerobic community, with 1.3% and 0.6% relative abundance, respectively (Fig. 4A). Bacteria from these two genera have been reported to secrete extracellular enzymes, such as lipases and esterases, which can break down the chemical bonds of PCL [72,73,74]. Furthermore, Pseudomonas spp. can also degrade PCL via intracellular enzymes [53]. Although being present with low relative abundance in the anaerobic community, these bacteria may also have a role in PCL degradation. Although preferring aerobic metabolism, bacteria from the Pseudomonas genus are capable to grow under anaerobic conditions, using nitrate or nitrite as electron acceptor [70]. Pseudomonas are also able to ferment arginine and pyruvate anaerobically [71, 72]. Arginine fermentation leads to very slow growth [73], while pyruvate fermentation has been associated with long-term survival and does not seem to contribute to anaerobic cell growth [72]. Substrate level phosphorylation coupled with the reduction of electron shuttles, such as phenazines, is another strategy described to promote energy conservation pathways that facilitate Pseudomonas’ anaerobic survival [74, 75]. Bearing in mind this highly versatile energy metabolism of Pseudomonas, it seems adequate not to rule out any possible role for Pseudomonas in PCL biodegradation under anaerobic conditions. Still, it is important to notice that Pseudomonas was the predominant genus identified in the leachate (42.9% relative abundance, Fig. 4A), but when incubated with PCL under strict anaerobic conditions, its relative abundance sharply decreased.
Aerobic Assays
The prokaryotes identified in the aerobic microbial community degrading PCL, as well as in the inoculum leachate, and their relative abundances, are displayed in Fig. 4A. A predominance of microorganisms from the bacteria domain was observed, accounting for 99.8% of the total diversity. Among the bacteria present in the samples, Coprothermobacter sp. was the most abundant, comprising 28% of the total identified community. Additionally, several groups belonging to the phylum Firmicutes constituted 62.6% of the total community, and other genera of bacteria were also present in relevant proportions, including Bacillus (8.9%), Symbiobacterium (6%), Ureibacillus (4.3%), Brevibacillus (2.3%), and Geobacillus (2%).
The hydrolytic function of Coprothermobacter on PCL has been previously hypothesized by Jin et al. [49] although conclusive evidence is still lacking. However, considering the significantly high relative abundances of Coprothermobacter sp. in the incubations with PCL (6.8% in anaerobic assays and 28% in aerobic assays), it strongly suggests that these microorganisms indeed play a significant role in the PCL biodegradation process. Coprothermobacter has been found to be one of the predominant bacteria in petroleum reservoirs [49], suggesting its capability to deal with recalcitrant compounds.
Inspection of the genomes of Coprothermobacter species revealed the presence of two enzymes that can potentially catalyze the hydrolysis of PCL. A total of 5374 HMMs were generated from enzyme sequences of carboxylesterases, lipases, and cutinases. The enzymes found were an esterase (Uniprot ID: A0A922ZSW3) from Coprothermobacter sp. and a thermostable monoacylglycerol lipase (Uniprot ID: B5Y789) from Coprothermobacter proteolyticus (strain ATCC 35,245), matching 52 and 71 HMMs derived from carboxylesterases (EC 3.1.1.1), respectively, with E-values ranging from 1.4E−88 to 1.1E−53.
Care should be taken with conclusions relying on the analysis of genomes of closely related species, since functional and physiological heterogeneity among species of the same genus may occur. This means that the Coprothermobacter present in leachate samples may have different hydrolytic enzymes in its genome or may have none. Also, despite the high homology between the hydrolytic enzymes found in the genomes of Coprothermobacter species to those previously associated with plastic biodegradation, enzyme activities and affinities to PCL as substrate need to be tested to unequivocal conclude about their function.
Nevertheless, these data taken together, i.e., the high abundance of Coprothermobacter species in aerobic and anaerobic assays where PCL was the only carbon and energy source, and the existence of hydrolytic enzymes (very close to carboxylesterases, lipases, and cutinases, previously reported as capable to hydrolyze PCL [41,42,43]), in the genomes of two Coprothermobacter species, suggests a role on PCL degradation and motivates further studies targeting the isolation of the species and testing its biodegradation activity towards PCL and other plastics.
Despite Coprothermobacter spp. are considered strictly anaerobic microorganisms, this bacterium was present in high abundance in the aerobic assays. Although it was never reported, Coprothermobacter spp. may be tolerant to oxygen, which might explain their capacity to endure in this community, where other strict anaerobic microorganisms could not (e.g., Methanothermobacter sp.).
Among the different bacterial genera identified in the community (Fig. 4A), the genus Bacillus (9% relative abundance) [76, 77] and Geobacillus (2%) [78] hav been previously associated with PCL biodegradation.
Besides bacteria, fungi were also present in this PCL-degrading community. Several eukaryotic microorganisms assigned to the fungi kingdom were identified by 18 S amplification (Fig. 4B).
The results indicate that the phylum Ascomycota dominates, accounting for 76.9% of the identified eukaryotic community (Fig. 4B). Within this phylum, the genera Exophiala (40.8%), Penicillium (16.9%), Aspergillus (2.7%), and Monascus (1.6%) are the most prevalent. Exophiala species were reported to degrade polyurethane (PU), a non-biodegradable polymer [79, 80]. On the other hand, Penicillium is well-known for its ability to degrade various plastics, particularly biodegradable polymers like PCL [81].
Mucor racemosus (18.4%), from the phylum Mucoromycota, is the second most abundant fungi identified in this community. Although no evidence was found in the literature regarding its ability to biodegrade PCL, some studies report its ability to degrade other polymers such as polyvinyl chloride (PVC) [82], polybutylene succinate (PBS) [83], and even crude petroleum by-products [84].
Several genera belonging to the order Saccharomycetales have also been identified, including Geotrichum (6.7%), Pichia (2.7%), Thelebolus (2.7%), and Kasachstania (2%). While there is limited literature available on the role of Geotrichum on PCL biodegradation, it has been reported to possess degrading properties towards other plastics, such as polycarbonate (PC) [84] and polyhexamethyleneguanidine (PHMB) [85].
Among the identified microorganisms, fungi are widely recognized as significant lipase-producers. Nakajima-Kambe et al. [86] reported that purified native and recombinant lipases from Aspergillus niger (2.7% relative abundance in the community) could degrade PCL.
All these results indicated that fungi and bacteria, especially Coprothermobacter sp., may be involved in PCL degradation in the aerobic assays, working individually or interacting within this complex microbial network.
Conclusion
In conclusion, this study demonstrates the potential of a landfill leachate and estuarine sediment as sources for biodegrading PCL. PCL biodegradation under both anaerobic and aerobic thermophilic conditions within approximately 50 days, using leachate as inoculum, as well as estuarine sediment in mesophilic aerobic conditions. However, the more recalcitrant polymers, PE and PET, did not show significant biodegradation. Taxonomic analysis of samples from the aerobic and anaerobic assays with PCL revealed the prominence of Coprothermobacter, and we found that Coprothermobacter species contain genes coding for enzymes that can potentially hydrolyze PCL. Also, given the high predominance of Methanothermobacter, it is likely that these two species collaborate in PCL biodegradation processes under methanogenic conditions.
These findings contribute to our understanding of the biodegradation potential of PCL and shed light on the microbial communities involved in the degradation process. The insights gained from this study could inform the development of more sustainable approaches for plastic waste management and highlight the importance of considering specific environmental conditions and microbial interactions in biodegradation studies.
Data Availability
Nucleotide sequencing data have been submitted to the European Nucleotide Archive (ENA) under the study accession number PRJEB73311.
References
Rana KI (2019) Usage of potential micro-organisms for degradation of plastics. Open J Environ Biol 007–015. https://doi.org/10.17352/ojeb.000010
Taghavi N, Udugama IA, Zhuang W-Q, Baroutian S (2021) Challenges in biodegradation of non-degradable thermoplastic waste: from environmental impact to operational readiness. Biotechnol Adv 49:107731. https://doi.org/10.1016/j.biotechadv.2021.107731
Rochman CM, Hoh E, Hentschel BT, Kaye S (2013) Long-term field measurement of sorption of organic contaminants to five types of plastic pellets: implications for plastic marine debris. Environ Sci Technol 47:1646–1654. https://doi.org/10.1021/es303700s
Yan F, Wei R, Cui Q et al (2021) Thermophilic whole-cell degradation of polyethylene terephthalate using engineered Clostridium thermocellum. Microb Biotechnol 14:374–385. https://doi.org/10.1111/1751-7915.13580
Battista F, Frison N, Bolzonella D (2021) Can bioplastics be treated in conventional anaerobic digesters for food waste treatment? Environ Technol Innov 22:101393. https://doi.org/10.1016/j.eti.2021.101393
Yu C, Dongsu B, Tao Z et al (2023) Anaerobic co-digestion of three commercial bio-plastic bags with food waste: effects on methane production and microbial community structure. Sci Total Environ 859:159967. https://doi.org/10.1016/j.scitotenv.2022.159967
Li Y, Tao L, Wang Q et al (2023) Potential health impact of microplastics: a review of environmental distribution, human exposure, and toxic effects. Environ Health 1:249–257. https://doi.org/10.1021/envhealth.3c00052
Zhu L, Kang Y, Ma M et al (2024) Tissue accumulation of microplastics and potential health risks in human. Sci Total Environ 915:170004. https://doi.org/10.1016/j.scitotenv.2024.170004
Gómez EF, Michel FC (2013) Biodegradability of conventional and bio-based plastics and natural fiber composites during composting, anaerobic digestion and long-term soil incubation. Polym Degrad Stab 98:2583–2591. https://doi.org/10.1016/j.polymdegradstab.2013.09.018
Fernandes M, Salvador A, Alves MM, Vicente AA (2020) Factors affecting polyhydroxyalkanoates biodegradation in soil. Polym Degrad Stab 182:109408. https://doi.org/10.1016/j.polymdegradstab.2020.109408
Ahmed T, Shahid M, Azeem F et al (2018) Biodegradation of plastics: current scenario and future prospects for environmental safety. Environ Sci Pollut Res 25:7287–7298. https://doi.org/10.1007/s11356-018-1234-9
Attallah OA, Mojicevic M, Garcia EL et al (2021) Macro and micro routes to high performance bioplastics: bioplastic biodegradability and mechanical and barrier properties. Polym (Basel) 13:2155. https://doi.org/10.3390/polym13132155
Wilkes RA, Aristilde L (2017) Degradation and metabolism of synthetic plastics and associated products by Pseudomonas sp.: capabilities and challenges. J Appl Microbiol 123:582–593. https://doi.org/10.1111/jam.13472
Tribedi P, Sil AK (2013) Low-density polyethylene degradation by Pseudomonas sp. AKS2 biofilm. Environ Sci Pollut Res 20:4146–4153. https://doi.org/10.1007/s11356-012-1378-y
Skariyachan S, Setlur AS, Naik SY et al (2017) Enhanced biodegradation of low and high-density polyethylene by novel bacterial consortia formulated from plastic-contaminated cow dung under thermophilic conditions. Environ Sci Pollut Res 24:8443–8457. https://doi.org/10.1007/s11356-017-8537-0
Ghatge S, Yang Y, Ahn JH, Hur HG (2020) Biodegradation of polyethylene: a brief review. Appl Biol Chem 63:27. https://doi.org/10.1186/s13765-020-00511-3
Kaushal J, Khatri M, Arya SK (2021) Recent insight into enzymatic degradation of plastics prevalent in the environment: a mini - review. Clean Eng Technol 2:100083. https://doi.org/10.1016/j.clet.2021.100083
Wei R, Zimmermann W (2017) Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: how far are we? Microb Biotechnol 10:1308–1322. https://doi.org/10.1111/1751-7915.12710
Urbanek AK, Kosiorowska KE, Mirończuk AM (2021) Current knowledge on polyethylene terephthalate degradation by genetically modified microorganisms. Front Bioeng Biotechnol 9. https://doi.org/10.3389/fbioe.2021.771133
Alshehrei F (2017) Biodegradation of synthetic and natural plastic by microorganisms. J Appl Environ Microbiol 5:8–19. https://doi.org/10.12691/jaem-5-1-2
Din MI, Ghaffar T, Najeeb J et al (2020) Potential perspectives of biodegradable plastics for food packaging application-review of properties and recent developments. Food Addit Contam: Part A 37:665–680. https://doi.org/10.1080/19440049.2020.1718219
Ilyas R, Zuhri M, Norrrahim M et al (2022) Natural fiber-reinforced polycaprolactone green and hybrid biocomposites for various advanced applications. Polym (Basel) 14:182. https://doi.org/10.3390/polym14010182
Blackwell CJ, Haernvall K, Guebitz GM et al (2018) Enzymatic degradation of star poly(ε-caprolactone) with different central units. Polym (Basel) 10. https://doi.org/10.3390/polym10111266
Nevoralová M, Koutný M, Ujčić A et al (2020) Structure characterization and biodegradation rate of poly(ε-caprolactone)/starch blends. Front Mater 7. https://doi.org/10.3389/fmats.2020.00141
Yoon Y, Park H, An S et al (2023) Bacterial degradation kinetics of poly(Ɛ-caprolactone) (PCL) film by Aquabacterium sp. CY2-9 isolated from plastic-contaminated landfill. J Environ Manage 335:117493. https://doi.org/10.1016/j.jenvman.2023.117493
Thakur M, Majid I, Hussain S, Nanda V (2021) Poly(ε-caprolactone): a potential polymer for biodegradable food packaging applications. Packaging Technol Sci 34:449–461. https://doi.org/10.1002/pts.2572
Nawaz A, Hasan F, Shah AA (2015) Degradation of poly(ɛ-caprolactone) (PCL) by a newly isolated Brevundimonas sp. strain MRL-AN1 from soil. FEMS Microbiol Lett 362:1–7. https://doi.org/10.1093/femsle/fnu004
Borghesi DC, Molina MF, Guerra MA, Campos MGN (2016) Biodegradation study of a novel poly-caprolactone-coffee husk composite film. Mater Res 19:752–758. https://doi.org/10.1590/1980-5373-MR-2015-0586
Fruteau De Laclos H, Hafner S, Holliger C (2018) Report on international inter-laboratory study on BMP tests. IOP Publishing PhysicsWeb. https://www.ktbl.de/fileadmin/user_upload/Allgemeines/Download/Ringversuch-Biogas/Report_Interlab-study_BMP-tests_February2018_korr.pdf. Accessed 31 Jan 2024
Holliger C, Astals S, de Laclos HF et al (2021) Towards a standardization of biomethane potential tests: a commentary. Water Sci Technol 83:247–250. https://doi.org/10.2166/wst.2020.569
Stams AJM, Van Dijk JB, Dijkema C, Plugge CM (1993) Growth of syntrophic propionate-oxidizing bacteria with fumarate in the absence of methanogenic bacteria. Appl Environ Microbiol 59:1114–1119. https://doi.org/10.1128/aem.59.4.1114-1119.1993
Riffat R, Krongthamchat K (2006) Specific methanogenic activity of halophilic and mixed cultures in saline wastewater. Int J Environ Sci Technol 2:291–299. https://doi.org/10.1007/BF03325889
Wang S, Hou X, Su H (2017) Exploration of the relationship between biogas production and microbial community under high salinity conditions. Sci Rep 7. https://doi.org/10.1038/s41598-017-01298-y
Moura I, Machado AV, Duarte FM, Nogueira R (2011) Biodegradability assessment of aliphatic polyesters-based blends using standard methods. J Appl Polym Sci 119:3338–3346. https://doi.org/10.1002/app.32966
Alves JI, Salvador AF, Castro AR et al (2020) Long-chain fatty acids degradation by desulfomonile species and proposal of “Candidatus desulfomonile palmitatoxidans.” Front Microbiol 11. https://doi.org/10.3389/fmicb.2020.539604
Silva AR, Duarte MS, Alves MM, Pereira L (2022) Bioremediation of perfluoroalkyl substances (PFAS) by anaerobic digestion: effect of PFAS on different trophic groups and methane production accelerated by carbon materials. Molecules 27. https://doi.org/10.3390/molecules27061895
Achinas S, Euverink GJW (2016) Theoretical analysis of biogas potential prediction from agricultural waste. Resource-Efficient Technol 2:143–147. https://doi.org/10.1016/j.reffit.2016.08.001
Salvador AF, Cavaleiro AJ, Paulo AMS et al (2019) Inhibition studies with 2-bromoethanesulfonate reveal a novel syntrophic relationship in anaerobic oleate degradation. Appl Environ Microbiol 85:e01733–e01718. https://doi.org/10.1128/AEM.01733-18
Caporaso JG, Lauber CL, Walters WA et al (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Stoeck T, Bass D, Nebel M et al (2010) Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol Ecol 19:21–31. https://doi.org/10.1111/j.1365-294X.2009.04480.x
Li L, Lin X, Bao J et al (2022) Two extracellular poly(ε-caprolactone)-degrading enzymes from Pseudomonas hydrolytica sp. DSWY01T: purification, characterization, and Gene Analysis. Front Bioeng Biotechnol 10. https://doi.org/10.3389/fbioe.2022.835847
Almeida BC, Figueiredo P, Carvalho ATP (2019) Polycaprolactone enzymatic hydrolysis: a mechanistic study. ACS Omega 4:6769–6774. https://doi.org/10.1021/acsomega.9b00345
Zampolli J, Vezzini D, Brocca S, Di Gennaro P (2024) Insights into the biodegradation of polycaprolactone through genomic analysis of two plastic-degrading Rhodococcus bacteria. Front Microbiol 14. https://doi.org/10.3389/fmicb.2023.1284956
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659. https://doi.org/10.1093/bioinformatics/btl158
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment 1 1Edited by J. Thornton. J Mol Biol 302:205–217. https://doi.org/10.1006/jmbi.2000.4042
Eddy SR (2011) Accelerated Profile HMM searches. PLoS Comput Biol 7:e1002195
Yagi H, Ninomiya F, Funabashi M, Kunioka M (2009) Anaerobic biodegradation tests of poly(lactic acid) and polycaprolactone using new evaluation system for methane fermentation in anaerobic sludge. Polym Degrad Stab 94:1397–1404. https://doi.org/10.1016/j.polymdegradstab.2009.05.012
Yagi H, Ninomiya F, Funabashi M, Kunioka M (2013) Thermophilic anaerobic biodegradation test and analysis of eubacteria involved in anaerobic biodegradation of four specified biodegradable polyesters. Polym Degrad Stab 98:1182–1187. https://doi.org/10.1016/j.polymdegradstab.2013.03.010
Jin Y, Cai F, Song C et al (2022) Degradation of biodegradable plastics by anaerobic digestion: morphological, micro-structural changes and microbial community dynamics. Sci Total Environ 834:155167. https://doi.org/10.1016/j.scitotenv.2022.155167
Narancic T, Verstichel S, Reddy Chaganti S et al (2018) Biodegradable plastic blends create new possibilities for end-of-life management of plastics but they are not a panacea for plastic pollution. Environ Sci Technol 52:10441–10452. https://doi.org/10.1021/acs.est.8b02963
Selke S, Auras R, Nguyen TA et al (2015) Evaluation of biodegradation-promoting additives for plastics. Environ Sci Technol 49:3769–3777. https://doi.org/10.1021/es504258u
Hermanová S, Šmejkalová P, Merna J, Zarevúcka M (2015) Biodegradation of waste PET based copolyesters in thermophilic anaerobic sludge. Polym Degrad Stab 111:176–184. https://doi.org/10.1016/j.polymdegradstab.2014.11.007
Mohanan N, Montazer Z, Sharma PK, Levin DB (2020) Microbial and enzymatic degradation of synthetic plastics. Front Microbiol 11:. https://doi.org/10.3389/fmicb.2020.580709
Al Hosni AS, Pittman JK, Robson GD (2019) Microbial degradation of four biodegradable polymers in soil and compost demonstrating polycaprolactone as an ideal compostable plastic. Waste Manag 97:105–114. https://doi.org/10.1016/J.WASMAN.2019.07.042
Pradhan R, Reddy M, Diebel W et al (2010) Comparative compostability and biodegradation studies of various components of green composites and their blends in simulated aerobic composting bioreactor. Int J Plast Technol 14. https://doi.org/10.1007/s12588-010-0009-z
Wang G, Huang D, Ji J et al (2021) Seawater-degradable polymers—fighting the marine plastic pollution. Adv Sci 8. https://doi.org/10.1002/advs.202001121
Wang W, Tao J, Yu K et al (2021) Vertical stratification of dissolved organic matter linked to distinct microbial communities in subtropic estuarine sediments. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.697860
Zhao R, Liu J, Feng J et al (2021) Microbial community composition and metabolic functions in landfill leachate from different landfills of China. Sci Total Environ 767:144861. https://doi.org/10.1016/j.scitotenv.2020.144861
Gomes E, de Souza AR, Orjuela GL et al (2016) Applications and benefits of thermophilic microorganisms and their enzymes for industrial biotechnology. In: Schmoll M, Dattenböck C (eds) Gene expression systems in Fungi: advancements and applications. Springer International Publishing, Cham, pp 459–492
Atanasova N, Stoitsova S, Paunova-Krasteva T, Kambourova M (2021) Plastic degradation by extremophilic bacteria. Int J Mol Sci 22:5610. https://doi.org/10.3390/ijms22115610
Tokiwa Y, Calabia B, Ugwu C, Aiba S (2009) Biodegradability of plastics. Int J Mol Sci 10:3722–3742. https://doi.org/10.3390/ijms10093722
Heimowska A, Morawska M, Bocho-Janiszewska A (2017) Biodegradation of poly(ϵ-caprolactone) in natural water environments. Pol J Chem Technol 19:120–126. https://doi.org/10.1515/pjct-2017-0017
Woodruff MA, Hutmacher DW (2010) The return of a forgotten polymer—polycaprolactone in the 21st century. Prog Polym Sci 35:1217–1256. https://doi.org/10.1016/J.PROGPOLYMSCI.2010.04.002
Kato S, Kosaka T, Watanabe K (2008) Comparative transcriptome analysis of responses of Methanothermobacter thermautotrophicus to different environmental stimuli. Environ Microbiol 10:893–905. https://doi.org/10.1111/j.1462-2920.2007.01508.x
Liczbiński P, Borowski S, Nowak A (2022) Isolation and use of Coprothermobacter spp. to improve anaerobic thermophilic digestion of grass. Molecules 27:4338. https://doi.org/10.3390/molecules27144338
Gagliano MC, Braguglia CM, Petruccioli M, Rossetti S (2015) Ecology and biotechnological potential of the thermophilic fermentative Coprothermobacter spp. FEMS Microbiol Ecol 91:fiv018. https://doi.org/10.1093/femsec/fiv018
Anne HE, M KD, K MR, et al (2019) Complete genome sequence of Caloramator sp. strain E03, a novel ethanologenic, thermophilic, obligately anaerobic bacterium. Microbiol Resour Announc 8. https://doi.org/10.1128/mra.00708-19
Ahlert S, Zimmermann R, Ebling J, König H (2016) Analysis of propionate-degrading consortia from agricultural biogas plants. Microbiologyopen 5:1027–1037. https://doi.org/10.1002/mbo3.386
Oh Y-R, Jang Y-A, Song JK, Eom GT (2022) Efficient enzymatic depolymerization of polycaprolactone into 6-hydroxyhexanoic acid by optimizing reaction conditions and microbial conversion of 6-hydroxyhexanoic acid into adipic acid for eco-friendly upcycling of polycaprolactone. Biochem Eng J 185:108504. https://doi.org/10.1016/j.bej.2022.108504
Carlson CA, Ingraham JL (1983) Comparison of denitrification by Pseudomonas stutzeri, Pseudomonas aeruginosa, and Paracoccus denitrificans. Appl Environ Microbiol 45:1247–1253. https://doi.org/10.1128/aem.45.4.1247-1253.1983
Vander Wauven C, Piérard A, Kley-Raymann M, Haas D (1984) Pseudomonas aeruginosa mutants affected in anaerobic growth on arginine: evidence for a four-gene cluster encoding the arginine deiminase pathway. J Bacteriol 160:928–934. https://doi.org/10.1128/jb.160.3.928-934.1984
Eschbach M, Schreiber K, Trunk K et al (2004) Long-term anaerobic survival of the opportunistic pathogen Pseudomonas aeruginosa via pyruvate fermentation. J Bacteriol 186:4596–4604. https://doi.org/10.1128/JB.186.14.4596-4604.2004
Arai H (2011) Regulation and function of versatile aerobic and anaerobic respiratory metabolism in Pseudomonas aeruginosa. Front Microbiol 2. https://doi.org/10.3389/fmicb.2011.00103
Glasser NR, Kern SE, Newman DK (2014) Phenazine redox cycling enhances anaerobic survival in Pseudomonas aeruginosa by facilitating generation of ATP and a proton-motive force. Mol Microbiol 92:399–412. https://doi.org/10.1111/mmi.12566
Ciemniecki JA, Newman DK (2020) The potential for redox-active metabolites to enhance or unlock anaerobic survival metabolisms in aerobes. J Bacteriol 202. https://doi.org/10.1128/JB.00797-19
Wu K-J, Wu C-S, Chang J-S (2007) Biodegradability and mechanical properties of polycaprolactone composites encapsulating phosphate-solubilizing bacterium Bacillus sp. PG01. Process Biochem 42:669–675. https://doi.org/10.1016/j.procbio.2006.12.009
Tiago I, Teixeira I, Silva S et al (2004) Metabolic and genetic diversity of mesophilic and thermophilic bacteria isolated from composted municipal sludge on poly-ε-caprolactones. Curr Microbiol 49:407–414. https://doi.org/10.1007/s00284-004-4353-0
Malunavicius V, Padaiga A, Stankeviciute J et al (2023) Engineered Geobacillus lipolytic enzymes – attractive polyesterases that degrade polycaprolactones and simultaneously produce esters. Int J Biol Macromol 253:127656. https://doi.org/10.1016/j.ijbiomac.2023.127656
Liu J, He J, Xue R et al (2021) Biodegradation and up-cycling of polyurethanes: progress, challenges, and prospects. Biotechnol Adv 48:107730. https://doi.org/10.1016/j.biotechadv.2021.107730
Magnin A, Pollet E, Phalip V, Avérous L (2020) Evaluation of biological degradation of polyurethanes. Biotechnol Adv 39:107457. https://doi.org/10.1016/j.biotechadv.2019.107457
Antipova TV, Zhelifonova VP, Zaitsev KV et al (2018) Biodegradation of poly-ε-caprolactones and poly-l-lactides by fungi. J Polym Environ 26:4350–4359. https://doi.org/10.1007/s10924-018-1307-3
Pardo-Rodríguez ML, Zorro-Mateus PJP (2021) Biodegradation of polyvinyl chloride by Mucor s.p. and Penicillium s.p. isolated from soil. Revista De Investigación Desarrollo E Innovación 11:387–400. https://doi.org/10.19053/20278306.v11.n2.2021.12763
Al Hosni AS (2019) Biodegradation of polycaprolactone bioplastic in comparision with other bioplastics and its impact on biota. Dissertation, University of Manchester
Adekunle AA, Oluyode TF (2005) Biodegradation of crude petroleum and petroleum products by fungi isolated from two oil seeds (melon and soybean). J Environ Biol 26:37–42
Brzezinska MS, Walczak M, Burkowska-But A et al (2019) Antifungal activity of polyhexamethyleneguanidine derivatives introduced into biodegradable polymers. J Polym Environ 27:1760–1769. https://doi.org/10.1007/s10924-019-01472-5
Nakajima-Kambe T, Edwinoliver NG, Maeda H et al (2012) Purification, cloning and expression of an Aspergillus niger lipase for degradation of poly(lactic acid) and poly(ε-caprolactone). Polym Degrad Stab 97:139–144. https://doi.org/10.1016/j.polymdegradstab.2011.11.009
Funding
Open access funding provided by FCT|FCCN (b-on). This work was supported by the Portuguese Foundation for Science and Technology (FCT) under the scope of the strategic funding of UIDB/04469/2020 unit, with DOI https://doi.org/10.54499/UIDB/04469/2020, and by the European Regional Development Fund under the scope of Norte2020—Programa Operacional Regional do Norte - MarPlas project (NORTE-01-0145-FEDER-000080). Authors LC and JCS received research support from FCT through the fellowships SFRH/BD/147271/2019 and 2023.01617.BD, respectively.
Author information
Authors and Affiliations
Contributions
AFS and AJC contributed to the study conception and design. AFS and SGB provided guidance to CSP, LC, JCS, DG, and JPF. Research was performed by CSP, LC, JCS, SGB, DG, and JPF. CSP, LC, and SGB drafted the manuscript with the support of AJC and AFS. AVM and AJC secured funding. All authors participated in data interpretation and scientific discussion as well as revisions of the final manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pires, C.S., Costa, L., Barbosa, S.G. et al. Microplastics Biodegradation by Estuarine and Landfill Microbiomes. Microb Ecol 87, 88 (2024). https://doi.org/10.1007/s00248-024-02399-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02399-8