Abstract
Bacterial azoreductases are enzymes that catalyze the reduction of ingested or industrial azo dyes. Although azoreductase genes have been well identified and characterized, the regulation of their expression has not been systematically investigated. To determine how different factors affect the expression of azoR, we extracted and analyzed transcriptional data from the Gene Expression Omnibus (GEO) resource, then confirmed computational predictions by quantitative reverse transcription polymerase chain reaction (qRT-PCR). Results showed that azoR expression was lower with higher glucose concentration, agitation speed, and incubation temperature, but higher at higher culture densities. Co-expression and clustering analysis indicated ten genes with similar expression patterns to azoR: melA, tpx, yhbW, yciK, fdnG, fpr, nfsA, nfsB, rutF, and chrR (yieF). In parallel, constructing a random transposon library in E. coli K-12 and screening 4320 of its colonies for altered methyl red (MR)-decolorizing activity identified another set of seven genes potentially involved in azoR regulation. Among these genes, arsC, relA, plsY, and trmM were confirmed as potential azoR regulators based on the phenotypic decolorization activity of their transposon mutants, and the expression of arsC and relA was confirmed, by qRT-PCR, to significantly increase in E. coli K-12 in response to different MR concentrations. Finally, the significant decrease in azoR transcription upon transposon insertion in arsC and relA (as compared to its expression in wild-type E. coli) suggests their probable involvement in azoR regulation. In conclusion, combining in silico analysis and random transposon mutagenesis suggested a set of potential regulators of azoR in E. coli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Azo dyes are among the most commonly used synthetic dyes, with extensive applications in food, pharmaceuticals, cosmetics, and textile industries, among others [1]. They are characterized by the presence of one or more azo groups (– N = N –) that account for their recalcitrant xenobiotic nature [2]. Approximately 15% of the azo dyes used in the textile industry are released into the environment as industrial effluents, which contaminate agricultural crops, decrease oxygen levels and pH in rivers, and disturb the aquatic ecosystem, evoking a significant environmental concern [3]. Additionally, many azo dyes are toxic, and their biotransformation products can be carcinogenic and mutagenic; therefore, they are disapproved for use as food components [4]. Because of this problem, ecofriendly, cost-effective biological systems have been thoroughly studied for their potential application as decolorizing agents in industry [5].
Bacteria and other microorganisms, including yeast and filamentous fungi, exert their degradative effect by secreting a number of enzymes, such as azoreductases, laccases, oxidases, and peroxidases [6]. A number of azoreductases were identified in Escherichia coli, two of which, AzoRI and AzoRII, are NAD(P)H dependent azoreductases that degrade Ponceau SX most efficiently, besides Tartrazine, Amaranth, and Orange II [7]. However, the most common azoreductase in E. coli is AzoR, an FMN-dependent-NADH azoreductase that follows a ping pong Bi-Bi mechanism of action in two cycles [8]. Other studies showed that some azoreductases, including the E. coli AzoR, act as quinone reductases, thus providing resistance to thiol-specific stress [9,10,11]. In Pseudomonas aeruginosa, azoreductases were predicted to be potentially involved in bacterial pathogenicity, and both azoR1 and azoR2 genes were required for systemic infection in mice [12].
Although azoreductase activity has been thoroughly studied in several bacteria, only limited research has been conducted on the regulation of its gene expression, particularly in E. coli. A pilot study used whole-genome microarrays in E. coli K-12 to determine the changes in gene expression during acid red 18 (AR18) decolorization, and found upregulated genes to be related to stress response, metabolism, and cell components. Upon deletion of five upregulated genes, the decolorization rate of AR18 by E. coli K-12 decreased but did not completely stop, suggesting the presence of alternative decolorization pathway [13].
The previous data suggest that bacterial azoreductases may have more than one physiological role that is yet to be elucidated to reveal how their homologs in human-associated and environmental isolates react to xenobiotics [10]. Here, we aimed to use two complementary approaches to investigate, on a genome-scale, the regulation of azoR in E. coli—as a model organism: (i) a computational approach that integrates data from public databases, and (ii) an experimental loss-of-function approach through random transposon mutagenesis, followed by screening and transcriptional analysis. Using those two complementary approaches, we shortlisted a set of four candidate regulators of azoR expression and experimentally confirmed the involvement of two of them, arsC and relA, in the decolorization process of the model dye methyl red.
Methods
Bacterial Strains and Culture Conditions
The wild-type (WT) E. coli K-12 MG1655 was used for the construction of the transposon library. ∆azoR, an E. coli K-12 strain with the azoR gene knocked out, from the Keio collection [14] was used for optimization of the 96-well plate decolorization assay.
E. coli K-12 was regularly grown on Luria Bertani (LB) broth or agar (Conda, Spain) at 37 °C with shaking at 250 rpm unless otherwise specified. For decolorization experiments, E. coli WT and mutants were cultivated in mineral salts medium (MSM) containing 4 g K2HPO4; 4 g KH2PO4; 2 g (NH4)2SO4; 0.5 g MgSO4·7H2O; 0.01 g CaCl2; 0.01 g FeSO4·7H2O per liter of distilled water (pH 7 ± 0.2) supplemented with 0.1% yeast. Methyl red (MR) (Alfa-Aesar, Germany) was prepared as a 500 mg/L stock then filter sterilized through a 0.22-µm sterile cellulose acetate membrane filter. Fifty microgram per milliliter kanamycin (Bio Basic, Canada) was used to positively select transposon mutants when necessary.
Systems Analysis of Transcriptomic Data
In Silico azoR Expression Analysis
As a first approach to study the regulation of published azoreductase genes in the organisms of interest, microarray data for the E. coli azoreductases were mined at the National Center for Biotechnology Information (NCBI)’s GEO database (https://www.ncbi.nlm.nih.gov/geo/) and analyzed.
Briefly, when we searched GEO for “azoR and E. coli,” 462 samples from 50 datasets of high-throughput gene expression data, recorded as percentile ranks, were collected, and azoR transcriptional data was analyzed using R statistical package (R version 4.0.2) to determine conditions under which it is significantly up or downregulated across various research studies not exclusively related to azoreduction. Any sample missing azoR expression value was filtered out. The selected 462 samples were those that had expression percentile ranks for the azoR gene (among all other measured E. coli genes), in addition to at least one experimental condition listed. The studied conditions included the effect of culture media, growth phase, glucose concentration, culture density, incubation temperature, and agitation speed on azoR expression.
In Vitro Validation of the Conditions Affecting azoR Expression in E. coli
To confirm the reliability of the analyzed public transcriptome data, we quantified azoR expression in E. coli K-12 grown under different growth conditions. To test the effect of different glucose concentrations while using it as a sole source of carbon, we used the RNeasy Mini Kit (Qiagen, Germany) to extract RNA from E. coli samples grown to mid-log phase in MSM, supplemented with different glucose concentrations of 0.02%, 0.2%, and 0.5%. To test the effects of different OD600, incubation temperatures, and agitation speeds, we extracted RNA from E. coli samples grown in LB broth to OD600 of 0.4, 0.8, and 1.2, as well as E. coli samples incubated at two different temperatures (30 °C and 37 °C), and finally, E. coli samples incubated at different agitation speeds of 60 and 250 rpm, respectively.
The purity and concentration of RNA were checked by Nanodrop (Quawell, China), then the extracted RNA was treated with DNaseI (New England Biolabs, USA) to ensure the absence of any genomic DNA contamination. Subsequently, we used RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher, USA) for cDNA synthesis, according to the manufacturer’s protocol. The purity and concentration of the prepared cDNA were measured by Nanodrop (Quawell, China). All qRT-PCR experiments were carried out in the StepOnePlus, Real-Time PCR System (Applied Biosystems, USA) with the Maxima Sybr Green qPCR Master Mix (Thermo Scientific, USA). The azoR primer pair (Supplementary Table S1) was designed by primer3plus (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) and synthesized by Macrogen (Korea). The results were normalized to the ihfB reference gene [15]. All assays were performed in triplicates.
The data were analyzed by the delta delta cycle threshold (∆∆Ct) relative quantification method. The data were then presented as fold increase or decrease in terms of the transcriptional level of each sample compared to the control. Statistical analysis between the Cts for each sample was conducted by one-way ANOVA followed by post hoc t-test with Tukey’s adjustment for multiple variables (i.e., glucose concentrations and OD600). For comparisons involving two variables (i.e., temperature and agitation speed), a paired t-test was used. GraphPad Prism 8 (GraphPad Software Tools, Inc., La Jolla, CA, USA) was used for visualizing the data and calculating the p values.
Co-expression Profile and Analysis
Profile neighbors are the top 200 genes with similar expression patterns to the gene of interest in this specific dataset. Profile neighbors’ data for each of the collected 50 datasets were retrieved from the GEO to determine co-expressed genes with azoR. Statistical analysis was carried out to determine the most repeated genes, in terms of expression patterns, to azoR within all datasets and thus help predict functionally related genes. The maximum number of repetitions was identified, and then a cutoff of genes showing at least half of this number of similar expression pattern repetitions with azoR were further processed. Genes (in the form of UniProt accession numbers) were then clustered according to their functional annotations using the Database for Annotation, Visualization and Integrated Discovery (DAVID) [16] (https://david.ncifcrf.gov/).
Transposon Library Construction and Screening
Random Transposon Mutagenesis in E. coli and Colony Selection
To determine the genetic elements involved in azoR regulation, we constructed an E. coli K-12 transposon library as follows. An overnight E. coli K-12 culture, grown to an OD600 of 0.5, was harvested and made electrocompetent by three rounds of 10% glycerol wash and subsequent centrifugation for 10 min at 1700 ×g and 4 °C. Fifty microliters of the prepared electrocompetent cells were transformed with 1 µl of EZ-Tn5™<KAN-2 > Tnp Transposome (Epicenter, Illumina, USA) by electroporation (Bio-Rad Micropulser, USA). Bacterial cells were recovered for 60 min using 950 µl super optimal broth with catabolite repression (SOC) medium at 37 °C with shaking at 200 rpm. The library was plated on LB agar, supplemented with 50 µg/ml kanamycin, for positive selection of successfully transposed cells. Kanamycin-resistant colonies were picked and arrayed into 96-well plates, and 25% glycerol stocks were prepared in duplicates and stored at −80 °C.
Decolorization Activity Screening of the Transposon Library
Decolorization activity of the mutant library was tested by a high-throughput 96-well plate assay as previously described [17], with some modifications that were based on our specific assay optimization. Tests for assay optimization were carried out to identify the conditions that maximize the differences in decolorization activity between the WT and the mutants in terms of wavelength, incubation time and culture media (Supplementary Methods and Supplementary Fig. S1 and S2).
Briefly, sterile, flat-bottomed 96-well plates (SPL, Life Sciences, Korea) were filled with 135 µl MSM supplemented with 0.1% yeast, 50 mg/L dye, and 50 µg/ml kanamycin. Fifteen microliters of an overnight culture of each of the mutants and the WT E. coli were added to the wells. Plates were incubated overnight at 30 °C with shaking at 150 rpm. Subsequently, decolorization was measured in a microplate reader (ELx800, Biotek, USA) set with a 450-nm filter. The decolorization ability of each strain, expressed as a percentage, was calculated by the following equation:
\(\mathrm{Decolorization}\;\mathrm{percentage}\;(\%)=\frac{\mathrm{Initial}\;\mathrm{absorbance}-\mathrm{Final}\;\mathrm{absorbance}}{\mathrm{Initial}\;\mathrm{absorbance}}\times100\)
Initial and final absorbance correspond to the absorbance measured at 0 and 16 h, respectively.
For ease of determining the cluster or deviation of the mutants, in terms of decolorization activity, from those of the WT strain, the relative mutant decolorization was calculated as follows:
\(\mathrm{Normalized}\;\mathrm{mutant}\;\mathrm{decolorization}=\frac{\mathrm{Mutant}'\mathrm s\;\mathrm{decolorization}\%}{\mathrm{WT}\;\mathrm{decolorization}\;\%}\times100\)
High-throughput screening of the decolorization activity of the library facilitated setting a cutoff for mutants showing a normalized mutant decolorization ≤ 65 or ≥ 110 for further testing. Such mutants were accordingly retested in triplicates by the same 96-well plate assay, before further confirmation in large volumes in 250-ml Erlenmeyer flasks, as previously described [18]. A two-tailed t-test was applied to determine the statistical significance between the decolorization percentages of the mutants and the WT.
Transposon Insertion Site Determination
Transposon insertion positions for the selected mutants were identified by the previously developed rapid amplification of transposon ends (RATE) PCR which is a single transposon-specific primer-PCR protocol that comprises three rounds of amplification in a single PCR run followed by sequencing the PCR products with a nested primer [19]. GeneAmp® High Fidelity PCR System was used for amplification of the extracted DNA from the chosen mutants in compliance with the manufacturer’s protocol.
The first round of PCR amplification was as follows: initial denaturation of 95 °C for 5 min, followed by 30 cycles of 95 °C/30 s., 55 °C/30 s., and 72 °C/3 min each. The second round comprised 30 cycles of 95 °C/30 s., 30 °C/30 s., and 72 °C/2 min. each. The final round, comprised 33 cycles of 95 °C/30 s., 55 °C/30 s., and 72 °C/2 min each. DNA from the gel bands was subsequently purified by the QIAquick Gel Extraction Kit (Qiagen®, Germany). DNA’s purity and concentration were further checked by the Nanodrop device (Quawell, China). Finally, a single nested kanamycin specific primer KAN2-RP1 (Table S1) was used for unidirectional Sanger sequencing of the PCR product. The samples were sequenced by Macrogen, Korea, via Green Tech, Egypt, in a high-throughput Applied Biosystems 3730XL sequencer.
A nucleotide BLAST (BLASTn) [20] of the query sequences against the E. coli K-12 substr. MG1655 genome, with default settings, was performed after transposon ends removal and ambiguous base removal by the sequence editor Chromas (software version 2.6.6).
Quantifying Gene Expression of Candidate azoR Regulators by qRT-PCR
To further investigate the role of four of the sequenced mutants (∆arsC, ∆relA, ∆plsY, and ∆trmM) as candidate azoR regulators, first, azoR expression was compared with that of the four genes of interest in the WT. An overnight culture of E. coli K-12 was normalized to a starting OD600 of 0.05 in (i) fresh MSM supplemented with 0.1% yeast—as a control, (ii) fresh MSM supplemented with 0.1% yeast and 5 mg/L MR, and (iii) Fresh MSM supplemented with 0.1% yeast and 50 mg/L MR. Triplicates of all samples were grown at 30 °C with shaking at 150 rpm until an OD600 of 0.15 was reached; then the cells were harvested. qRT-PCR experiments were carried out as previously explained.
Second, the difference in azoR gene expression between the WT E. coli K-12, ∆arsC, and ∆relA under the three conditions tested in the first experiment was investigated, and the data were analyzed by the ∆∆Ct method as detailed above.
Results
Systems Analysis of Transcriptomic Data
In Silico Analysis of azoR Expression Under Different Growth Conditions
Expression data for azoR, from 462 samples in 50 microarray datasets of different E. coli strains, were retrieved from the GEO (Supplementary Dataset 1) and compared in relation to different growth conditions (Supplementary Fig. S3).
For the rest of the study, transcriptomic data from E. coli K-12 substrains (411 samples) were filtered out (Supplementary Dataset 2) and were plotted against different growth conditions to determine the conditions modulating its transcription positively or negatively. We focused on K-12 to reduce variability and because K12 was the strain used in in vitro studies. The analysis showed azoR to be highly expressed at the following conditions: increase in the culture optical density (OD600) from 0.4 to 0.7 (Fig. 1a); growth at the log phase (Fig. 1b); growth in minimal media (Fig. 1c); supplementation with high glucose concentration of 0.5% (Fig. 1d); growth at low agitation speed of 60 rpm (Fig. 1e); and at lower temperatures of 25 °C or 30 °C (Fig. 1f). On the other hand, azoR expression was low at OD600 under 0.4 and above 0.7; in stationary phase; upon growth in rich media, e.g., trypticase soy broth (TSB); at glucose concentrations of 0.1 to 0.4%; at agitation speeds of 200 to 300 rpm and at incubation temperature of 37 °C.
Effect of different growth conditions on the transcripts level of azoR in E. coli K-12 according to GEO expression data. Boxplots showing the computed effects of a culture density expressed as OD600, b growth phases, c culture media, d glucose concentrations, e agitation speed, and f incubation temperatures on azoR expression. The X-axis displays the tested conditions used in different datasets, while the Y-axis represents the percentile rank of the GEO expression data as statistically analyzed by R (version 4.0.2). The box spans the central 50% of the data. Significantly different categories (in comparison the median percentile rank for all values for each condition) are shown with asterisks. ns = non significant.
In Vitro Expression Analysis of azoR Under Different Growth Conditions
The effects of OD600, glucose concentration, agitation speed, and incubation temperature on the expression of azoR were tested by qRT-PCR to validate the predictions inferred from the computational analysis. The relative abundance of azoR mRNA increased as culture density increased, partially agreeing with the computational prediction since computational predictions showed a decrease in azoR expression over OD600 of 0.7 (Fig. 2a). However, azoR expression significantly decreased (p < 0.01 at 0.2% glucose and p < 0.001 at 0.5% glucose) at higher glucose concentrations, unlike computational predictions (Fig. 2b). On the other hand, the results of the two other tested conditions conformed to the computational predictions, as follows: a significant decrease in azoR expression level (p < 0.001) at higher agitation speed was observed (Fig. 2c). Finally, a significant increase (p < 0.0001) in azoR RNA levels was observed at 30 °C, as compared to 37 °C (Fig. 2d).
Effect of different growth conditions on azoR expression. Bar chart showing qRT-PCR (in vitro) results, expressed as expression fold change of azoR, of the effects of different a culture densities, b glucose concentrations, c agitation speeds, and d incubation temperatures on azoR expression. The X-axis represents the variables of the tested growth condition, and the Y-axis represents the expression fold change of azoR. Statistical comparisons between Ct values were tested for significance by one-way ANOVA followed by post hoc t-test with Tukey’s adjustment (a and b) or just t-tests (c and d). **p value < 0.01, ***p value < 0.001, and ****p value < 0.0001. Data presented are the mean of three replicates, and error bars represent the standard deviation
Co-expression Profile and Analysis
Identification of Nine Functionally Related Co-expressed Genes with azoR
Data about azoR transcriptional “profile neighbors” were extracted from the GEO and analyzed in R software, in terms of the repetition of co-expressed genes in the different datasets, which indicates a likelihood of functional relationship to azoR. This analysis showed that 4345 genes have a similar expression pattern to that of the azoR only in a single dataset, while 1280 genes were azoR profile neighbors twice, 553 genes were repeated three times, 229 genes were repeated four times, 77 genes were repeated five times, 28 genes were repeated six times, five genes were repeated seven times, only one gene (pspE) was repeated eight times, and finally, one gene (tesA) was repeated ten times (Supplementary Dataset 3).
We used the DAVID database to identify, categorize, and classify the functional annotations of 112 genes that were profile neighbors of azoR at least five times (Table 1). The analysis elucidated that 34% of the genes are involved in various metabolic processes; 18% in several cellular responses, including responses to DNA damage, antibiotic resistance, stress, and others; 10% of the genes are involved in transcription regulation processes; 9% in transport, while others (with lower representation) were shown to be involved in other processes, such as DNA recombination and repair, biofilm formation, lysozyme inhibition, cell division, and proteolysis. Meanwhile, 12% of the genes remained uncharacterized. The functional clustering feature of the DAVID database was applied to the same set of genes, resulting in 13 clusters, which were filtered to eight clusters with at least two members each (Fig. S4). AzoR was clustered with ten functionally related co-expressed genes, which were annotated as oxidoreductases, flavoproteins, and reducing equivalents (FAD, FMN, NADH, or NADPH)-binders: nfsA, nfsB, yieF, fdnG, yciK, fpr, rutF, yhbW, tpx, and melA (Fig. 3).
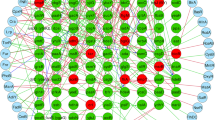
Functional clustering of azoR co-expressed genes. Heatmap displaying the clustering of genes co-expressed with azoR within at least five different GEO datasets. Clustering was generated by the DAVID software, according to the functional annotations. The horizontal axis displays the functional categories, while the vertical axis displays the genes
Transposon Mutagenesis and Screening Identify Seven Genes Potentially Involved in azoR Regulation
Transposon Mutagenesis and Screening
A genome-wide random insertion library was constructed for E. coli K-12, substrain MG1655, with the Tn5 transposome, EZ-Tn5™ <KAN-2>, which confers kanamycin resistance to recipient colonies, thus allowing selection of mutants on LB agar supplemented with kanamycin. Kanamycin-resistant colonies (n = 4320) were picked and arrayed into 45 96-well plates.
The mutant library was screened for the ability of each clone to decolorize MR in 96-well plates. After the preliminary screening of the library, 428 mutants showed relative decolorization ≤ 65%, and 56 mutants showed relative decolorization ≥ 110%. Repetition of the test for the selected mutants, in at least two experiments with three replicates per experiment, allowed the exclusion of outliers or inconsistent results. This resulted in the selection of 15 mutants with reduced decolorization activity and five mutants with increased decolorization activity relative to the WT E. coli, with the same cutoffs used in the preliminary screening experiment (Table 2). The mutant decolorization percentages were statistically significantly different from that of the WT (two-tailed t-test p values < 0.05).
The transposon insertion sites for the chosen strains were determined by RATE PCR, followed by sequencing of PCR products. Seven isolates showed more than one band, and in those cases, DNA from all bands was extracted. Sequences with poor quality or with no hit to the E. coli genome were discarded, while good-quality sequences were trimmed then aligned against the E. coli K-12 substrain MG1655 genome, and the first hit in each case was retrieved (Table 2).
Confirmation of the Involvement of arsC and relA in the Decolorization Process
The expression levels of the candidate regulatory genes of interest (arsC, relA, plsY, and trmM) versus azoR in the WT E. coli were determined after the bacteria were exposed to MR concentrations of 5 and 50 mg/L. These levels were compared to their expression levels in the absence of the dye. azoR, arsC, relA, and trmM were differentially expressed at one or both MR concentrations. The expression of azoR significantly increased (p < 0.0001) in presence of both MR concentrations (Fig. 4a), whereas arsC and relA expression significantly increased (p < 0.05 and p < 0.01, respectively) only in response to 5 mg/L dye, despite their genetic upregulation at both concentrations (Fig. 4b and c). Expression of plsY did not significantly increase in response to 5 mg/L dye (Fig. 4d). Finally, the expression of trmM increased (p < 0.05) only in response to 5 mg/L dye (Fig. 4e).
Effect of different MR concentrations on the expression of candidate genes. Bar charts showing the differences in gene expression of a azoR, b arsC, c relA, d plsY, and e trmM upon exposure to different MR concentrations. The X-axis represents the samples exposed to different MR concentrations, and the Y-axis represents the expression fold change of the gene of interest. Comparisons between Ct values were tested for statistical significance by one-way ANOVA followed by post hoc t-test with Tukey’s adjustment. *p value < 0.05, **p value < 0.01, ***p value < 0.001, and ****p value < 0.0001. The data are the mean of three replicates, and error bars represent the standard deviation
∆arsC and ∆relA were further studied based on their promising involvement in the decolorization process and possible feedback or compensatory activity on azoR, as elucidated in the previous phenotypic and genotypic experiments. In the second RT-qPCR assay, the gene expression levels of azoR in the WT E. coli and in both arsC and relA mutants were compared upon stimulation with MR concentrations of 5 and 50 mg/L, with the expression level of azoR in WT E. coli (unexposed to the dye) used as control.
The transcriptional activity of azoR in both ∆arsC and ∆relA, compared to that in the WT E. coli, significantly decreased (p < 0.001) upon induction by MR. This significant decrease may explain the decreased phenotypic activity of these mutants (Fig. 5).
Difference in azoR expression in WT E. coli, and mutants of interest. Bar charts showing the difference in azoR expression levels between WT E. coli, ∆arsC, and ∆relA upon stimulation with 5 mg/L and 50 mg/L MR as compared to control experiment with no dye added. The X-axis represents the different strains exposed to different MR concentrations. Each strain was exposed to three MR concentrations (0, 5, and 50 mg/L). The Y-axis represents the expression fold change for the gene of interest. Comparisons between Ct values were tested for statistical significance by one-way ANOVA followed by post hoc t-test with Tukey’s adjustment for differences from the “no-dye” WT control (indicated by °). ***p value < 0.001 and ****p value < 0.0001. The data are the mean of three replicates and error bars represent the standard deviation
Discussion
In this study, we investigated the regulation of azoR expression in E. coli K-12 to better understand the physiological role these enzymes might play. This role is particularly interesting to fully uncover because, although azo dyes are mostly connected to human activity, the presence of bacterial azoreductases that predate humans suggests that these enzymes may have different physiological roles in bacteria [9].
Combining a systems approach, involving computational transcriptome analysis, with a reductionist qRT- PCR-based approach demonstrated the following:
First, azoR expression was higher at 30 °C than at 37 °C, suggesting that 30 °C is the optimal temperature for decolorization activity and is consistent with previous results that showed an enhanced in vitro activity of azoreductases at 30 °C rather than 37 °C [21].
Second, increasing agitation was predicted to decrease azoR expression, conforming with previous phenotypic results [22]. This could be a result of increased oxygen transfer, which decreases enzymatic activities [23], or competition between oxygen and azoR over electron donors [24].
Third, increasing culture density from 0.4 to 0.8 to 1.2 was found to consistently increase azoR expression, partially conforming to computational analyses which predicted that initially increasing OD increases azoR expression to a certain extent after which the activity decreases, which is consistent with a previous study [25].
Finally, increasing glucose concentration decreased azoR transcription, unlike the results of computational analysis, in which azoR expression values tended to increase with increasing glucose concentration; however, the GEO-retrieved data were quite heterogeneous. For example, for glucose concentration of 0.2%, there were two subpopulations of azoR expression values (Supplementary Fig. S5).
The experimental effect of glucose concentration (Fig. 2b), which was observed under controlled conditions (MSM minimal media and OD = 0.5), could be attributed to the abiotic stress the bacteria encounter at a decreased glucose concentration while surviving in a carbon-depleted minimal media as opposed to higher glucose concentrations. Using glucose as a carbon source was found to phenotypically enhance azo-reducing activity in several studies, until a certain threshold after which the decolorization activity decreases, probably due to preferential consumption of the dye rather than the external carbon source [26, 27].
The observed discrepancy between in vitro experiments and in silico analysis of some factors, especially glucose %, is most likely due to the controlled in vitro experimental setting—unlike the heterogeneity of GEO datasets. For example, the culture density effect was consistent in vitro (Fig. 2a), while the computational analysis showed a large spread in expression values at OD 0.4 and 0.5 (Fig. 1a). This is because the in vitro experiment was conducted in one culture medium, at a fixed agitation speed (250 rpm) and fixed culture conditions, while the computational analysis was conducted on data from all available conditions (Fig. 1 and Supplementary Fig. S3). Likewise, the discrepancy in measured azoR expression under different glucose % might be due to the wide variation between other confounding experimental conditions. For example, out of 190 GEO experiments for which glucose concentration was stated, 142 experiments (74.7%) were conducted at a glucose concentration of 0.2%, but a wide variation of agitation speed, OD, and type of media, and these variations had significant effects on azoR expression (Fig. 6).
Variations of azoR expression levels in E. coli K-12 within the 142 samples that had been grown in 0.2% glucose, according to GEO expression data. Significant variations of a growth phase, b agitation speed, c culture media, and d type of media within. Significantly different categories (in comparison the median percentile rank for all values for each condition) are shown with asterisks. ns = non significant.
Using the GEO datasets for co-expression analysis to determine genes with similar expression patterns to that of azoR, followed by functional categorization, allowed us to functionally cluster azoR with ten co-expressed genes, all of them encoding oxidoreductases, namely melA, tpx, yhbW, yciK, fdnG, fpr, nfsA, nfsB, rutF, and chrR (yieF). Three of the clustered genes were identified as quinone reductase genes: nfsA, nfsB, and yieF. They are also clustered based on the required cofactor as energy donor for the oxidoreduction reaction. In this context, rutF, azoR, melA, and fdnG encode NAD-binding enzymes, nfsA and fpr encode NADP-binding enzymes, while nfsB and yieF encode both NAD- and NADP-binding enzymes. Finally, rutF, azoR, nfsA, nfsB, fpr, and yieF encode flavoprotein (FMN/FAD)-binding enzymes. Details about those genes, encoded functions, and their interrelations are provided in Supplementary Material.
As previously mentioned, azoreduction is an electron transport process that utilizes electron donors for subsequent reduction of azo dyes [28]. Among the components involved in the electron transport chain (ETC) are dehydrogenases, such as the formate dehydrogenase N alpha subunit (FdnG), which is able to transfer electrons to terminal reductases using quinones as redox mediators [29]. This co-occurrence in the ETC could explain the correlation between fdnG and azoR with probable involvement in the azo reduction process, and agrees with the upregulation of another dehydrogenase gene, fdhF, up to 4.5 folds upon exposure to AR18, and its subsequent downregulation upon enzyme deletion [13]. Likewise, yciK, which encodes a putative yet unannotated oxidoreductase, was among azoR profile neighbors in five datasets in our study, and was also upregulated in response to the azo dye AR18 [13]. This suggests its probable role as an azoreductase, a hypothesis that requires further investigation.
In addition to computational systems analysis, another system approach to comprehensively study azoR regulation was to construct a Tn5 transposon insertion mutant library of 4320 kanamycin-resistant mutants in E. coli K-12 MG1655. When screened for MR decolorization ability, 20 mutants were found to have significantly different decolorizing abilities from the WT strain, suggesting that the transposon interrupted key genes in the reduction process. Transposon insertions were identified in seven different genes, namely arsC, relA, plsY, trmM, gshA, glnE, and the gene encoding the hypothetical “YqcC family protein.”
Arsenate reductase (arsC) is a part of the arsenic resistance operon (ars) in E. coli, which provides resistance against As5+ [30,31,32]. Other ars operon products include the protein ArsH, which is an FMN reductase with a demonstrated ability to reduce chromate [33], ferric iron [33, 34], quinones [35] and, interestingly, azo dyes [36,37,38] as well as a quencher of oxidative stress [39]. This is particularly interesting because, although the E. coli K-12 genome lacks an arsH gene [39] in its ars operon, the redox arsC gene product showed in our study a similar azo dye-reducing ability in both phenotypic and genetic assays. We therefore suggest that the arsenic resistance operon is also involved in MR decolorization and acts as a stress quencher brought about by exposure to azo dyes.
The second transposon insertion was detected in the relA gene, encoding a GDP/GTP pyrophosphokinase. RelA is a stringent factor that plays an important role in bacterial adaptation to amino acid starvation through the production of guanosine tetraphosphate (ppGpp) and guanosine pentaphosphate (pppGpp) [40]. Besides, glutamate excretion as a result of amino acid starvation is a relA-dependent process [41]. Another pathway for the stimulation of relA transcription is a result of nitrogen starvation or glutamine limitation by the σ54-dependent response regulator NtrC through the GlnD pathway [42]. Therefore, an insertion in the relA gene is expected to cause drainage of the glutamine pool as well as accumulation of glutamate, under nitrogen and amino acid starvation, respectively.
To the best of our knowledge, no prior studies correlated the products of relA and azoR. Here, the threefold upregulation of relA in response to MR exposure, and subsequent decrease in azoR activity and transcription upon insertion in relA, suggest they are correlated. We propose this upregulation as a stringent response by which the cells adapt to their exposure to azo dyes.
An insertion was identified in the putative glycerol-3-phosphate acyltransferase gene (plsY or ygiH) which plays a role in phospholipid biosynthesis [43]. plsY showed a non-significant upregulation in response to MR exposure, and its mutant showed a defected azoreductase activity in vitro. Surprisingly, this coincides with the upregulation of another acyl-glyerol-3-phosphate acyltransferase in Staphylococcus aureus upon exposure to Sudan III [44].
Seven transposon insertions were identified in tRNA (adenosine(37)-N6)-methyltransferase (trmM or yfiC). This gene’s product methylates adenosine 37 in tRNA1Val at the N6 position, which helps stabilize the tRNA structure. YfiC was also found to resist osmotic and oxidative stress [45].
Another insertion was identified in the glutamine synthetase adenylyl transferase gene (glnE). Glutamine synthetase (GS) catalyzes glutamine synthesis which, along with glutamate and other metabolites, is required for nitrogen assimilation. GlnE activates and reduces GS activity under low and high nitrogen levels, respectively. A reduction in GS activity is regulated by cellular glutamine concentration [46], and therefore an insertion in the glnE gene might result in a depletion of glutamate levels due to the continuous GS activity. This hypothesis is supported by a reported transcriptional upregulation of glnR, the glutamine synthetase repressor in S. aureus, upon exposure to the azo dye, Sudan III [44].
An insertion was observed in gamma-glutamyl cysteine synthase (gshA), which is involved in glutathione (GSH) biosynthesis, a thiol that plays a crucial role in protection against stress, as an antioxidant and a redox buffer [47]. An insertion in gshA had no effect on E. coli’s resistance to oxidative damage and gamma radiation; however, it contributed to thiol-specific stress [48]. Moreover, the lack of GSH, as a result of gshA inactivation, increased the sensitivity of E. coli to both mercury and arsenite [49]. azoR expression increased in E. coli in response to quinones to prevent thiol-specific stress, which might have led to GSH depletion [9], but the effect of GSH absence on azoR transcripts in E. coli was not studied further. We, therefore, propose that the decrease in azo reduction in this clone resulted from the absence of glutathione and therefore increased oxidative stress. In other terms, this mutation presumably leads to the accumulation of glutamate.
The phenotypic decrease in azo reduction upon insertions in gshA or glnE suggests that azoR might be involved in ammonium assimilation. Our computational analysis supports this suggestion as it indicated the co-expression between azoR and glnD in five different datasets. GlnD also plays an important role in nitrogen assimilation regulation and metabolism [50].
Finally, an ambiguous mutation in a 21-bp segment of the gene encoding the hypothetical protein YqcC was identified in a phenotype that showed higher azo-reducing activity than the WT E. coli. The exact function of the protein is unknown; nonetheless, it was reported to have a biofilm-related function [51].
To validate some of the above predictions, we measured the transcripts of arsC, relA, plsY, and trmM, as well as that of the well-known azoR gene in WT E. coli K-12 upon exposure to two different MR concentrations. While the transcription of the four genes increased in response to the lower MR concentration, only arsC and relA transcriptionally increased at both MR concentrations. This suggests a regulatory or feedback mechanism, resulting in a compensatory action through the genetic upregulation of these genes—probably in response to the oxidative stress imposed on the bacterial cells by exposure to the dye. The significant decrease in azoR transcriptional activity in ∆arsC and ∆relA mutants suggests a role these two genes might play in regulating azoreductase activity and justifies the observed phenotypic decrease in decolorization activity by both strains. It also suggests a potential relation between arsC and arsH, as both exhibit similar azo dye-reducing activity and play a part in the arsenical resistance operon.
Conclusion and Outlook
In conclusion, this work followed a systems approach to study azoR gene expression in E. coli K-12 combining computational and experimental genome-wide mutagenesis tools. Based on the computationally analyzed transcriptomic data, we hypothesize the possible regulation of azoR by either the SoxRS or MarRAB regulons or both; however, this hypothesis needs further investigation. Our work also suggests that yciK is a probable azoreductase and that the arsenic resistance operon is involved in azo dye decolorization.
Using a loss-of-function approach by transposon mutagenesis coupled with qRT-PCR, we identified arsC and relA as probable azoR regulators based on the significant transcriptional upregulation in E. coli K-12 upon exposure to MR. This hypothesis is backed by the consequent decrease in azoR transcriptional activity in ∆arsC and ∆relA strains as compared to the WT E. coli.
This work highlights the importance of better understanding the physiological role azoreductases might play. Future work includes precisely deleting these genes and defining their phenotypes along with subsequent complementation assays.
Data Availability
No datasets were generated or analyzed during the current study.
References
Benkhaya S, M’rabet S, El Harfi A (2020) Classifications, properties, recent synthesis and applications of azo dyes. Heliyon 6:e03271. https://doi.org/10.1016/J.HELIYON.2020.E03271
Misal SA, Gawai KR (2018) Azoreductase: a key player of xenobiotic metabolism. Bioresour Bioprocess 5:17. https://doi.org/10.1186/s40643-018-0206-8
Srinivasan S, Sadasivam SK (2021) Biodegradation of textile azo dyes by textile effluent non-adapted and adapted Aeromonas hydrophila. Environ Res 194:110643. https://doi.org/10.1016/J.ENVRES.2020.110643
Cui D, Li G, Zhao D, Gu X, Wang C, Zhao M (2012) Purification and characterization of an azoreductase from Escherichia coli CD-2 possessing quinone reductase activity. Process Biochem 47:544–549. https://doi.org/10.1016/j.procbio.2011.12.013
Chen H (2006) Recent advances in azo dye degrading enzyme research. Curr Protein Pept Sci 7:101–111
Senthil Rathi B, Senthil Kumar P (2022) Sustainable approach on the biodegradation of azo dyes: a short review. Curr Opin Green Sustain Chem 33:100578. https://doi.org/10.1016/J.COGSC.2021.100578
Ghosh DK, Ghosh S, Sadhukhan P, Mandal A, Chaudhuri J (1993) Purification of two azoreductases from Escherichia coli K12. Indian J Exp Biol 31:951–954
Nakanishi M, Yatome C, Ishida N, Kitade Y (2001) Putative ACP phosphodiesterase gene (acpD) encodes an azoreductase. J Biol Chem 276:46394–46399. https://doi.org/10.1074/jbc.M104483200
Liu G, Zhou J, Fu QS, Wang J (2009) The Escherichia coli azoreductase AzoR is involved in resistance to thiol-specific stress caused by electrophilic quinones. J Bacteriol 191:6394–6400. https://doi.org/10.1128/JB.00552-09
Ryan A, Wang C, Laurieri N, Westwood I, Sim E (2010) Reaction mechanism of azoreductases suggests convergent evolution with quinone oxidoreductases. Protein Cell 1:780–790. https://doi.org/10.1007/s13238-010-0090-2
Töwe S, Leelakriangsak M, Kobayashi K et al (2007) The MarR-type repressor MhqR (YkvE) regulates multiple dioxygenases/glyoxalases and an azoreductase which confer resistance to 2-methylhydroquinone and catechol in Bacillus subtilis. Mol Microbiol 66:40–54. https://doi.org/10.1111/j.1365-2958.2007.05891.x
Skurnik D, Roux D, Aschard H et al (2013) A comprehensive analysis of in vitro and in vivo genetic fitness of Pseudomonas aeruginosa using high-throughput sequencing of transposon libraries. PLoS Pathog 9:e1003582. https://doi.org/10.1371/journal.ppat.1003582
Zhang H, Lu H, Wang J, Zhang T, Liu G, Zhou J (2014) Transcriptional analysis of Escherichia coli during Acid Red 18 decolorization. Process Biochem 49:1260–1265. https://doi.org/10.1016/j.procbio.2014.04.007
Baba T, Ara T, Hasegawa M et al (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:20060008. https://doi.org/10.1038/msb4100050
Zhou K, Zhou L, Lim Q, Zou R, Stephanopoulos G, Too HP (2011) Novel reference genes for quantifying transcriptional responses of Escherichia coli to protein overexpression by quantitative PCR. BMC Mol Biol 12:18. https://doi.org/10.1186/1471-2199-12-18
Sherman BT, Hao M, Qiu J et al (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50:W216–W221. https://doi.org/10.1093/NAR/GKAC194
Lucas M, Mertens V, Corbisier A, Vanhulle S (2008) Synthetic dyes decolourisation by white-rot fungi: Development of original microtitre plate method and screening. 42:97–106. https://doi.org/10.1016/j.enzmictec.2007.07.023
Hashem RA, Samir R, Essam TM, Ali AE, Amin MA (2018) Optimization and enhancement of textile reactive Remazol black B decolorization and detoxification by environmentally isolated pH tolerant Pseudomonas aeruginosa KY284155. AMB Express 8:83. https://doi.org/10.1186/s13568-018-0616-1
Karlyshev AV, Pallen MJ, Wren BW (2000) Single-primer PCR procedure for rapid identification of transposon insertion sites. Biotechniques 28:1078–1082. https://doi.org/10.2144/00286bm05
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Sari IP, Simarani K (2019) Decolorization of selected azo dye by Lysinibacillus fusiformis W1B6: biodegradation optimization, isotherm, and kinetic study biosorption mechanism. Adsorpt Sci Technol 37:492–508. https://doi.org/10.1177/0263617419848897
Al-Tohamy R, Kenawy ER, Sun J, Ali SS (2020) Performance of a newly isolated salt-tolerant yeast strain Sterigmatomyces halophilus SSA-1575 for azo dye decolorization and detoxification. Front Microbiol 11:1163. https://doi.org/10.3389/FMICB.2020.01163/BIBTEX
Kaushik P, Malik A (2009) Fungal dye decolourization: recent advances and future potential. Environ Int 35:127–141. https://doi.org/10.1016/j.envint.2008.05.010
Kumar N, Sinha S, Mehrotra T, Singh R, Tandon S, Thakur IS (2019) Biodecolorization of azo dye Acid Black 24 by Bacillus pseudomycoides: process optimization using Box Behnken design model and toxicity assessment. Bioresour Technol Rep 8:100311. https://doi.org/10.1016/J.BITEB.2019.100311
Agrawal S, Tipre D, Patel B, Dave S (2014) Optimization of triazo Acid Black 210 dye degradation by Providencia sp SRS82 and elucidation of degradation pathway. Process Biochem 49:110–119. https://doi.org/10.1016/j.procbio.2013.10.006
Dong H, Guo T, Zhang W et al (2019) Biochemical characterization of a novel azoreductase from Streptomyces sp: application in eco-friendly decolorization of azo dye wastewater. Int J Biol Macromol 140:1037–1046. https://doi.org/10.1016/J.IJBIOMAC.2019.08.196
Jafari N, Kasra-Kermanshahi R, Reaz Soudi M (2013) Screening, identification and optimization of a yeast strain, Candida palmioleophila JKS4, capable of azo dye decolorization. Iran J Microbiol 5:434
Pearce CI, Lloyd JR, Guthrie JT (2003) The removal of colour from textile wastewater using whole bacterial cells: a review. Dye Pigment 58:179–196. https://doi.org/10.1016/S0143-7208(03)00064-0
Unden G, Bongaerts J (1997) Alternative respiratory pathways of Escherichia coli: energetics and transcriptional regulation in response to electron acceptors. Biochim Biophys Acta - Bioenerg 1320:217–234. https://doi.org/10.1016/S0005-2728(97)00034-0
Carlin A, Shi W, Dey S, Rosen BP (1995) The ars operon of Escherichia coli confers arsenical and antimonial resistance. J Bacteriol 177:981–986. https://doi.org/10.1128/jb.177.4.981-986.1995
Suzuki K, Wakao N, Kimura T, Sakka K, Ohmiya K (1998) Expression and regulation of the arsenic resistance operon of Acidiphilium multivorum AIU 301 plasmid pKW301 in Escherichia coli. Appl Environ Microbiol 64:411–418
Mukhopadhyay R, Rosen BP (2002) Arsenate reductases in prokaryotes and eukaryotes. Environ Health Perspect 110:745–748. https://doi.org/10.1289/ehp.02110s5745
Xue X-M, Yan Y, Xu H-J, Wang N, Zhang X, Ye J (2014) ArsH from Synechocystis sp. PCC 6803 reduces chromate and ferric iron. FEMS Microbiol Lett 356:105–112. https://doi.org/10.1111/1574-6968.12481
Mo H, Chen Q, Du J et al (2011) Ferric reductase activity of the ArsH protein from Acidithiobacillus ferrooxidans. J Microbiol Biotechnol 21:464–469. https://doi.org/10.4014/jmb.1101.01020
Hervás M, López-Maury L, León P, Sánchez-Riego AM, Florencio FJ, Navarro JA (2012) ArsH from the cyanobacterium Synechocystis sp. PCC 6803 is an efficient NADPH-dependent quinone reductase. Biochemistry 51:1178–1187. https://doi.org/10.1021/bi201904p
Crescente V, Holland SM, Kashyap S, Polycarpou E, Sim E, Ryan A (2016) Identification of novel members of the bacterial azoreductase family in Pseudomonas aeruginosa. Biochem J 473:549–558. https://doi.org/10.1042/BJ20150856
Vorontsov II, Minasov G, Brunzelle JS et al (2007) Crystal structure of an apo form of Shigella flexneri ArsH protein with an NADPH-dependent FMN reductase activity. Protein Sci 16:2483–2490. https://doi.org/10.1110/ps.073029607
Ye J, Yang HC, Rosen BP, Bhattacharjee H (2007) Crystal structure of the flavoprotein ArsH from Sinorhizobium meliloti. FEBS Lett 581:3996–4000. https://doi.org/10.1016/j.febslet.2007.07.039
Páez-Espino AD, Nikel PI, Chavarría M, Lorenzo V (2020) ArsH protects Pseudomonas putida from oxidative damage caused by exposure to arsenic. Environ Microbiol 22:2230–2242. https://doi.org/10.1111/1462-2920.14991
Arenz S, Abdelshahid M, Sohmen D et al (2016) The stringent factor RelA adopts an open conformation on the ribosome to stimulate ppGpp synthesis. Nucleic Acids Res 44:6471–6481. https://doi.org/10.1093/nar/gkw470
Burkovski A, Weil B, Krämer R (1995) Glutamate excretion in Escherichia coli: dependency on the reIA and spoT genotype. Arch Microbiol 164:24–28. https://doi.org/10.1007/BF02568730
Ronneau S, Hallez R (2019) Make and break the alarmone: regulation of (p)ppGpp synthetase/hydrolase enzymes in bacteria. FEMS Microbiol Rev 43:389–400. https://doi.org/10.1093/femsre/fuz009
Lu YJ, Zhang YM, Grimes KD, Qi J, Lee RE, Rock CO (2006) Acyl-phosphates initiate membrane phospholipid synthesis in Gram-positive pathogens. Mol Cell 23:765–772. https://doi.org/10.1016/j.molcel.2006.06.030
Pan H, Xu J, Gew O et al (2015) Differential gene expression in Staphylococcus aureus exposed to Orange II and Sudan III azo dyes. 745–757. https://doi.org/10.1007/s10295-015-1599-4
Golovina AY, Sergiev PV, Golovin AV et al (2009) The yfiC gene of E. coli encodes an adenine-N6 methyltransferase that specifically modifies A37 of tRNA1Val(cmo5UAC). RNA 15:1134–1141. https://doi.org/10.1261/rna.1494409
Okano H, Hwa T, Lenz P, Yan D (2010) Reversible adenylylation of glutamine synthetase is dynamically counterbalanced during steady-state growth of Escherichia coli. J Mol Biol 404:522–536. https://doi.org/10.1016/j.jmb.2010.09.046
Meister A, Anderson ME (1983) Glutathione. Annu Rev Biochem 52:711–760. https://doi.org/10.1146/annurev.bi.52.070183.003431
Greenberg JT, Demple B (1986) Glutathione in Escherichia coli is dispensable for resistance to H2O2 and gamma radiation. J Bacteriol 168:1026–1029. https://doi.org/10.1128/jb.168.2.1026-1029.1986
Latinwo LM, Donald C, Ikediobi C, Silver S (1998) Effects of intracellular glutathione on sensitivity of Escherichia coli to mercury and arsenite. Biochem Biophys Res Commun 242:67–70. https://doi.org/10.1006/bbrc.1997.7911
Zhang Y, Pohlmann EL, Serate J, Conrad MC, Roberts GP (2010) Mutagenesis and functional characterization of the four domains of GlnD, a bifunctional nitrogen sensor protein. J Bacteriol 192:2711–2721. https://doi.org/10.1128/JB.01674-09
Beloin C, Valle J, Latour-Lambert P et al (2004) Global impact of mature biofilm lifestyle on Escherichia coli K-12 gene expression. Mol Microbiol 51:659–674. https://doi.org/10.1046/j.1365-2958.2003.03865.x
Acknowledgements
The authors thank Dr. Jonathan Monk, from the University of California, San Diego, for providing the ∆azoR deletion clone from the Keio collection library. MAS thanks Yomna A. Hashem, PhD, for useful advice and training on using the real-time PCR instrument.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was not funded by any grants from public or private agencies. MAS received limited funding from the British University in Egypt (BUE) graduate program to cover the consumables of her thesis work, including the experimental work. The BUE also supported the purchase of the electroporator used in creating the transposon library.
Author information
Authors and Affiliations
Contributions
A.M.H. and R.K.A. conceived the study. A.M.H. and R.K.A. supervised M.A.S. throughout the entire project. M.A.S., A.M.H., and R.K.A. designed experiments. M.A.S. performed all laboratory experiments and analyzed the data. M.A.S. performed bioinformatics and genomic analyses. M.A.S. and R.K.A. performed statistical analysis. M.A.S., H.T.N., and R.K.A. generated the figures. H.T.N. and A.M.H. provided resources to support experimental analysis. M.A.S. and H.T.N. drafted the manuscript, and R.K.A. revised different drafts. All authors revised and agreed with the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salem, M.A., Nour El-Din, H.T., Hashem, A.M. et al. Genome-Scale Investigation of the Regulation of azoR Expression in Escherichia coli Using Computational Analysis and Transposon Mutagenesis. Microb Ecol 87, 63 (2024). https://doi.org/10.1007/s00248-024-02380-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02380-5