Abstract
Aeromonas hydrophila is an opportunistic motile pathogen with a broad host range, infecting both terrestrial and aquatic animals. Environmental and geographical conditions exert selective pressure on both geno- and phenotypes of pathogens. Flagellin, directly exposed to external environments and containing important immunogenic epitopes, may display significant variability in response to external conditions. In this study, we conducted a comparative analysis of ~ 150 A. hydrophila genomes, leading to the identification of six subunits of the flagellin gene (fla-1 to fla-4, flaA, and flaB). Individual strains harbored different composition of flagellin subunits and copies. The composition of subunits showed distinct patterns depending on environmental sources. Strains from aquatic environments were mainly comprised of fla-1 to fla-4 subunits, while terrestrial strains predominated in groups harboring flaA and flaB subunits. Each flagellin showed varying levels of expression, with flaA and flaB demonstrating significantly higher expression compared to others. One of the chemotaxis pathways that control flagellin movement through a two-component system was significantly upregulated in flaA(+ 1)/flaB(+ 1) group, whereas flaA and flaB showed different transcriptomic expressions. The genes positively correlated with flaA expression were relevant to biofilm formation and bacterial chemotaxis, but flaB showed a negative correlation with the genes in ABC transporters and quorum sensing pathway. However, the expression patterns of fla-2 to fla-4 were identical. This suggests various types of flagellin subunits may have different biological functions. The composition and expression levels of flagellin subunits could provide valuable insights into the adaptation of A. hydrophila and the differences among strains in response to various external environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aeromonas hydrophila is a gram-negative facultative anaerobic bacterium. It has been isolated from diverse environmental habitats (e.g., drinking water, wastewater, sewage, food, milk, soil) and has a broad host range from fish to humans [1,2,3,4]. A. hydrophila causes infectious disease with severe clinical signs such as gastroenteritis, necrotizing fasciitis, exophthalmia, fin rot, dropsy, hemorrhage, and septicemia as opportunistic bacterial pathogens in aquatic to terrestrial species [5,6,7]. In terms of fish diseases, A. hydrophila can infect various ranges of species [8,9,10]. Among all the genomes publicly available on GenBank until 2022, A. hydrophila has been reported to have been isolated from dozens of fish species, such as goldfish, carp, snakehead, eel, tilapia, catfish, bluegill, large yellow croaker, sturgeon, and salmon as summarized in Table S1. A. hydrophila can be both a primary and secondary bacterial pathogen in fish, depending on the fish’s health conditions [8, 11]. It is well known as an opportunistic pathogen that can infect individuals under poor and stressed conditions. Due to these characteristics, outbreaks of A. hydrophila are used as indicators of poor water quality management and polluted environments. In addition, A. hydrophila can survive in a wide range of water temperatures (0–45 °C) [8, 12, 13]. A. hydrophila naturally confronts a variety of environments that require adaptation. These attributes have the potential to act as evolutionary drivers, allowing bacteria to endure prolonged exposure to various challenging environments and adapt accordingly. A. hydrophila harbor numerous extracellular proteins which are known to be associated with environmental adaptability [14]. In response to varying environmental conditions, bacteria undergo adaptations by modulating gene expression and composition, leading to the emergence of diverse phenotypic characteristics, including virulence [4, 15, 16]. Not only Aeromonas but there are many cases that the genotypic differences affected significantly phenotypic characteristics such as pathogenicity and pH resistance in other fish bacterial pathogens [17, 18]. Alterations in the bacterial genome can exert profound effects on the intricate transcriptional network, subsequently influencing the associated phenotypic traits [19]. This suggests an increasing importance in acquiring comprehensive insights into the genotypic characteristics of A. hydrophila that are associated with its transcriptomic and/or phenotypic traits. Despite its significance, the integration of genomic and transcriptomic data in A. hydrophila across diverse evolutionary contexts has remained insufficiently addressed up until now.
In recent years, massive genomic and transcriptomic results have been released, and all these works contribute to the understanding of the physiological characteristics of fish bacterial pathogens [20,21,22,23,24,25,26,27]. The significance of interpreting and analyzing massive and deep sequencing data is increasingly recognized in the exploration of novel insights into bacterial evolution and the dynamics of genetic content alteration [28,29,30]. The integration of omics data derived from independent studies holds significant potential in providing novel insights and generating substantial impact through the application of big data analysis in microbiological research [31]. However, each study employs different bioinformatics pipelines, strategies, and tools for individual specific purposes, thus posing a challenge in intuitively comparing results across different datasets.
Flagella are widely recognized for their crucial roles in motility, adhesion, and biofilm formation, which are directly associated with the growth and adaptation of bacteria in viscous environments or spaces [32,33,34]. The diversity of flagella subunits can influence the assembly, shape, size, and movement of flagella, which results in variations in functionality [35,36,37]. Given the fact that a single polar flagellum allows A. hydrophila to swim efficiently and move in diverse environments, facilitating its invasion of host cells, the understanding of genotypic and transcriptomic characteristics in the flagellin subunits can reveal the microbiological significance of flagellin in A. hydrophila [38, 39]. During the pan-genome and comparative genomic study, we have noticed that strains from terrestrial and aquatic environments tend to harbor different compositional patterns of flagellin subunits, known to possess unique and typical/atypical expression patterns and/or functionalities. Accordingly, this study manually aligned and processed A. hydrophila genome and/or transcriptome data using a standardized in-house pipeline. The main objective was to elucidate the genotypic and transcriptomic characteristics of A. hydrophila flagellin subunits that might greatly impact their phenotypic traits.
Materials and Methods
Strains and Genomic Data
In this study, A. hydrophila genomes with available sample histories, including information on isolation site, date, and source from the BioProject database, were utilized. Table S1 provides comprehensive information about the strains used in this study, including isolated year, country, host, and accession numbers in GenBank, RefSeq, BioProject, and BioSample. In total, 151 A. hydrophila strains from 18 countries between 1980 and 2022 were analyzed in this study. Average nucleotide identity (ANI) in all A. hydrophila genomes and six other Aeromonas species (Aeromonas caviae, Aeromonas salmonicida, Aeromonas sobria, Aeromonas veronii, Aeromonas jandaei, Aeromonas dhakensis) were calculated using FastANI (version 1.34) [40]. A heatmap and dendrogram were created using the “gplots” package in R (version 4.2.3), based on ANI results among strains [41]. The dendrogram analysis employed the “Euclidean” method for distance calculation and clustered using the “average” method. Based on the isolation source, all strains were categorized into either aquatic or terrestrial environments. Among the 151 strains, 108 were classified as originating from aquatic environments, while the remaining 43 were categorized as terrestrial strains (Table S1). Flagellin homologs in A. hydrophila were examined, and three subunits of flagellin genes, annotated as flagellin A (flaA), flagellin B (flaB), and flagellin (fla), were identified using the NCBI Prokaryotic Genome Annotation Pipeline [42]. The number and type of flagella genes varied among strains [43].
Flagellin Homologs
The number of flaA, flaB, and fla gene copies from each strain was used for the dendrogram analysis using “ape” and “phytools” packages in R (version 4.2.3) [44,45,46]. Five distinct groups were identified based on the composition and number of flagellin homologs. This study named each strain named into fla(+ 1) to fla(+ 4) groups based on the number of flagellin copies. These groups encompassed strains with two to four copies of fla (abbreviated as fla(+ 2), fla(+ 3), or fla(+ 4)), strains with a combination of a single copy of flaA and flaB (abbreviated as flaA(+ 1)/flaB(+ 1)), and strains with a single copy each of the three flagellin subunits (abbreviated as fla(+ 1)/flaA(+ 1)/flaB(+ 1)). The relative proportion of the aquatic strains was determined using the following formula:
Chi-square analysis was performed using the relative proportion of strains from the aquatic environment (AE) and relative proportion of strains from terrestrial environment (TE) using the “gmodels” package in R [47]. p-value less than 0.05 was considered a statistical significance.
Phylogenetic Analysis of Flagellin Homologs
The sequences of all flagellin homologs were obtained from genomic coding sequences and GFF/GTF files. Approximately, 400 flagellin nucleotide sequences were utilized for phylogenetic analysis using Nextstrain [48]. Briefly, all flagellin sequences are grouped by type of flagellin composition, isolation year, and isolation site. Also, “sequences-per-group,” which specifies the maximum number of sequences per group, was assigned to 50. All sequences were aligned by the option of “–file-gap” without reference sequence for constructing the phylogenetic tree. Time-resolved tree was generated through the option of “data-confidence”, “coalescent = opt”, and “data-inference = marginal”. The trait, ancestral sequence inference, and mutation sites were annotated and profiled using the “traits”, “ancestral”, and “translate” commands. The phylogenetic results were then visualized in auspice using the “nextstrain view” command line.
Flagellin Expression Level and Global Transcriptome
The investigation into the transcriptomic characteristics of various flagellin genes involved the selection of five strains (23-C-23, ATCC7966, NJ-35, SCV-1, and WCX23). These strains were chosen given their composition of different flagellin genes in their genome and availability in open databases. At least one of selected five strains has been confirmed to harbor the fla, flaA, and flaB genes. The flagellin gene and global bacterial transcriptomic expressions under ordinary cultural environments currently available in SRA (Sequence Read Archive) were profiled from five A. hydrophila strains using raw transcriptomic files (FastQ) [22,23,24]. Three to four biologically replicated FastQ files derived from the transcriptome of each A. hydrophila strain cultured under ordinary culturable conditions (e.g., tryptone soy or Luria–Bertani medium at 28 °C for ~ 1 day) were downloaded from SRA [20,21,22,23,24]. The strain WCX23 and 23-C-23 belong to flaA(+ 1)/flaB(+ 1) flagellin group, and ATCC7966 harbored single flaA, flaB, and fla gene in the genome (flaA(+ 1), flaB(+ 1), and fla(+ 1) group). On the other hand, NJ-35 and SCV-1 strain have three fla genes belonging to a flagellin fla(+ 3) group (Table 1).
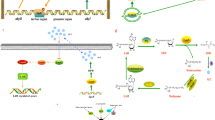
In this study, two distinct bioinformatics pipelines were utilized to assess the diverse levels of flagellin expressions across each strain, as well as to conduct a comparative analysis of the global transcriptome among the strains. To quantify the expression of various subunits of flagellin present in individual strains, transcriptomic reads were aligned to the genome of each strain (Fig. 1). In detail, FastQ files after trimming bad quality sequence reads below Q20 scores using Trimmomatic (Galaxy version 0.38.1) [49] were mapped onto each reference genome (WCX23; NZ_CP038463, 23-C-23; NZ_CP038465, ATCC7966; NZ_JAGDEM010000001, NJ-35; NZ_CP006870, and SCV-1; DANIKG010000001) using Bowtie2 with very sensitive end-to-end option (–very-sensitive) (Galaxy version 2.5.0 + galaxy0) [50, 51]. The transcriptome assembly, merge, and quantification were carried out using cufflinks (Galaxy version 2.2.1.3), cuffmerge (Galaxy version 0.0.5), and cuffdiff (Galaxy version 2.2.1.6) [52]. The FPKM (fragments per kilobase per million mapped fragments) value was validated to TPM (transcripts per million) value using below formula. TPM values of flagellin homologs from each sample were utilized for the level of flagellin gene expression as sample traits in this study. Welch’s t-test and Duncan’s multiple range test (analysis of variance; ANOVA) were carried out for the transcriptomic levels of each type of flagellin using an “aricolae” package in R (version 4.2.3) [53].
Schematic for profiling the expression of each flagellin gene and global gene expressions among the strains used in this study. The figure was drawn using BioRender (https://biorender.com/)
On the other hand, as for the global gene expression for the comparison among strains, all transcription reads were mapped onto one specific reference genome (FDAARGOS_916; NZ_CP065651). Except for the reference genome, the same pipeline was used as described above for obtaining TPM values (Fig. 1).
Bioinformatics
Differentially Expressed Genes (DEGs)
Based on global transcriptome results, the differentially expressed genes between flaA(+ 1)/flaB(+ 1) (WCX23 and 23-C-23 strains) and fla(+ 3) (NJ-35 and SCV-1 strains) groups were observed. The p-value of each gene between flagellin groups was calculated using edgeR in R. The gene lower than p-value 1.14 × × 10 −5 from the Bonferroni adjustment (0.05/total number of genes observed in this study) was regarded as DEGs in this study. The DEGs were annotated and mapped onto Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways [54].
Weighted Correlation Network Analysis (WGCNA)
The transcriptomic results from the strains that harbored flaA and flaB genes (WCX23, 23-C-23, and ATCC7966) were utilized for WGCNA. FlaA and flaB TPM results were used as sample traits. The signed network with the option of verbose = 3, merge cut height = 0.25, minimum module size = 30, and soft threshold power = 20 was generated to identify the gene cluster shared similar pattern of expression. Each module eigengene was used for correlation analysis with the expression of flaA and flaB to identify modules significantly correlated with their pattern of expression (p-value < 0.05). All genes belonging to significant modules were annotated using KEGG. Similarly, WGCNA was conducted under the same conditions as described above, utilizing NJ-35 and SCV-1 strains, which possess three fla genes (fla-2, fla-3, and fla-4) belonging to the fla(+ 3) group. All analyses and data visualization were carried out using “matrixStats”, “impute”, and “WGCNA” packages in R (version 4.2.3) [55].
Correlation Analysis
Correlation coefficients between sample traits (flaA and flaB) and all gene expressions were used for the clustering analysis. The heatmap with dendrogram was visualized using “RColorBrewer”, “gplot”, “pheatmap”, and “d3heatmap” in R (version 4.2.3). Based on the pattern of correlation coefficients between all genes and flaA and flaB, eight gene clusters were divided. Based on the pattern of overall correlation coefficients with flaA and flaB in a cluster, positive coefficients were denoted with “ +” , while negative coefficients were denoted with “ −” . Gene clusters harbored both positive correlation coefficients in flaA and flaB (flaA + /flaB +), opposite direction of correlation coefficients between homologs (flaA + /flaB- or flaA-/flaB +), or both negative correlation coefficients (flaA-/flaB-) were annotated through KEGG pathways.
KEGG Pathway Enrichment
Genes in significant WGCNA modules and gene clusters were mapped onto KEGG pathways. Regarding the DEG results, genes showing significantly higher expression in the flaA(+ 1)/flaB(+ 1) groups compared to the fla(+ 3) groups were denoted as upregulated, while the reverse scenario was labeled as downregulated. The significance of up- and downregulated DEG numbers in each pathway was evaluated using the Chi-square analysis, and the pathway with a p-value less than 0.05 was visualized. Likewise, top five pathways enriched by genes from significant modules through WGCNA and gene clusters with the different patterns of flaA and flaB correlation coefficients were drawn by “EnrichmentMap” in Cytoscape 3.10.0 [56, 57]. The KEGG pathways relevant to human diseases were not used for the analyses in this study. As for the WGCNA and gene cluster analyses, the KEGG two-component system, which includes many genes and has relatively broad biological meaning and implications, was not included in data visualization.
Results
Average Nucleotide Identity (ANI)
In this study, ANI results of 151 genomes were in a range of 95.3555–99.9997%, with one exception in B11 strain. The ANI between the label A. hydrophila strain B11 and other A. hydrophila strains was between 93.2555 and 94.1305%. However, it exhibits a higher ANI with A. dhakensis (97.1336%) (Figure S1; Table S2). The ANI among five strains (WCX23, 23-C-23, ATCC7966, NJ-35, and SCV-1) utilized for transcriptomic analysis in this study ranged from 96.6255 to 99.9980% (Table S2). Among them, a high ANI sharing of over 99.9% was observed between the 23-C-23 and WCX23 strains, as well as between the NJ-35 and SCV-1 strains. Interestingly, among these five strains, those sharing high ANI values correspondingly belong to the same flagellin group (flaA(+ 1)/flaB(+ 1) and fla(+ 3)).
The patterns of Flagellin Homolog Compositions and Phylogenetics in A. hydrophila
The number and type of flagellin genes in each strain were examined, revealing that 37 strains belonged to the fla(+ 3) group, 8 strains to the fla(+ 4) group, 56 strains to the flaA(+ 1)/flaB(+ 1) group, 12 strains to the fla(+ 1)/flaA(+ 1)/flaB(+ 1) group, 32 strains to the fla(+ 2) group, and the rest 6 strains that did not belong the majority. The strains harboring flaA and flaB were found to originate predominantly from terrestrial environments, whereas the fla(+ 3) and fla(+ 4) flagellin groups predominantly consisted of strains from aquatic environments. Specifically, the relative proportion of aquatic strains within the fla(+ 3) group was significantly higher compared to strains from terrestrial environments (Fig. 2). Based on phylogenetic analysis, the flagellin genes were classified into six subunits: four fla groups (named as fla-1, fla-2, fla-3, and fla-4, which were annotated same term but phylogenetically far distance flagellin), flaA, and flaB (Fig. 3). Interestingly, a notable observation was that the majority of strains did not possess a single type of flagellin gene, but instead exhibited a combination of different flagellin subunits. For instance, in the fla(+ 2) and fla(+ 3) flagellin groups, it was found that most strains harbored fla-1, fla-2, fla-3, and/or fla-4 genes, rather than multiple copies of the same flagellin type (Fig. 3A). The majority of strains belonging to the fla(+ 2), fla(+ 3), and fla(+ 4) groups possess at least one fla-4 gene. However, there were slight phylogenetic differences observed between fla-4 genes from the fla(+ 2) and fla(+ 3) groups (Fig. 3A, B). Additionally, it was noted that fla(+ 2) strains tended to be isolated more recently compared to fla(+ 3) strains (Fig. 3B). In terms of gene expression levels, flaB and flaA exhibited the highest and second-highest expression levels among the seven flagellin homologs, respectively. In contrast, fla-2 to fla-4 displayed relatively similar lower levels of expression. Conversely, the fla-1 gene showed very low or no expression (Fig. 3C). The phylogeographic results revealed that a significant number of strains positive for flaA and flaB originated from terrestrial sources, particularly strains from China and the USA (Fig. 4). On the other hand, most fla-2 to fla-4 genes were derived from aquatic strains, and notably, no terrestrial strain was observed in fla-2. In contrast, fla-1, which displayed the most distant phylogenetic relationship with other flagellin genes, exhibited a higher proportion of terrestrial strains compared to fla-2 to fla-4 (Fig. 4).
Dendrogram based on the type and number of flagellin genes by Aeromonas hydrophila strains. The blue and red circles indicate the isolated site of each strain (either aquatic or terrestrial environment). Bar color and length denote the type and number of flagellin genes. The relative proportion of strains originating from aquatic environment was described by the group sharing the same number of type of flagellin genes (fla(+3), fla(+4), fla(+1)/flaB(+1), fla(+1)/flaA(+1)/flaB(+1), and fla(+2)). Asterisks (***) indicates the significantly different number of aquatic strains compared to terrestrial strains in same flagellin group using Chi-square analysis (p-value < 0.001)
Phylogenetic tree of all flagellin genes in Aeromonas hydrophila observed in this study (A). Zoom in on the fla-4 clade with the label of isolation years (B). The transcription level of flagellin paralogs (fla-1, flaB, flaA, fla-2, fla-3, and fla-4) in 23-C-23, ATCC7966, NJ-35, SCV-1, and WCX23 strain based on transcripts per kilobase million (TPM) value (C). There was no statistically different expression between or among the strain within a same type of flagellin based on Welch’s t-test or one-way analysis of variance test using Duncan’s multiple comparison test
DEGs and Featured Pathways in the Flagellin Group
In total, 486 DEGs were identified between flaA(+ 1)/flaB(+ 1) and fla(+ 3) group, and 459 DEGs showed higher expression in flaA(+ 1)/flaB(+ 1) group. Five featured KEGG pathways, namely two-component system (ko02020), ABC transporters (ko02010), amino sugar and nucleotide sugar metabolism (ko00520), galactose metabolism (ko00052), and phosphotransferase system (PPTS) (ko02060), were identified as the most significantly different numbers of up- and downregulated DEGs across different flagellin groups. Notably, the flaA(+ 1)/flaB(+ 1) group exhibited a high number of upregulated DEGs in all five identified pathways (Fig. 5A). In particular, the two-component system pathway exhibited the highest number of DEGs among the identified pathways. Within this pathway, the flagellar motor switch adaptation system played a significant role, involving several DEGs such as methyl-accepting chemotaxis (mcp); aerotaxis receptor (aer); purine-binding chemotaxis protein (cheW); two-component system, chemotaxis family, sensor kinase (cheA); and two-component system, chemotaxis family, chemotaxis protein (cheY). In the A. hydrophila ASM162068 genome, five copies of mcp and two copies of aer were identified. The average gene expression levels of these genes were significantly higher in the flaA(+ 1)/flaB(+ 1) flagellin group. Similarly, the flaA(+ 1)/flaB(+ 1) group exhibited significantly higher TPM values in the genes cheW, cheA, and cheY compared to the fla(+ 3) group (Fig. 5B).
The KEGG enrichment analysis revealed a significantly different number of DEGs (differentially expressed genes) based on Chi-square analysis (p-value < 0.05) between flaA(+1)/flaB(+1) and fla(+3) strains (A). The two-component system relevant to flagellar motor switch adaptation based on the KEGG pathway for the prokaryote (B). The TPM level of relevant genes (mcp, aer, cheY, cheA, and cheW) was shown in raincloud plots. The expression of five homologous or paralogous mcp (Gene id: ①I6G73_RS10070, ②I6G73_RS19490, ③I6G73_RS04110, ④ I6G73_RS08330, and ⑤ I6G73_RS16810) and two aer (Gene id: ①I6G73_RS19930 and ②I6G73_RS05190) was shown as dot plots in accordance with different strains and groups. Asterisks (*p < 0.05, **p < 0.01, and ***p < 0.001) showed statistical difference between flaA(+1)/flaB(+1) and fla(+3) groups based on Welch two sample t-test
Module Identification Correlated with Flagellin Expressions Through WGCNA
The dendrogram showed overall transcriptomic expression of ATCC7966, WCX23, and 23-C-23, along with the relative expression levels of flaA and flaB utilized in the WGCNA (Fig. 6A). In total, eight modules were identified, and the eigengene of yellow and turquoise modules was positively and negatively correlated with the expression of flaA (Figure S2; Fig. 6B). On the other hand, flaB showed only a negatively significant correlation with brown eigengene (Fig. 6B). In total, 2125, 366, and 188 genes were labeled and identified as the turquoise, brown, and yellow modules, respectively. Also, since two-component system was the pathway composed of the highest numbers belonging to each module, regardless of module’s significance, it was not visualized in this study. ABC transporters and quorum sensing are the pathways with high number of genes in turquoise modules. But, featured pathways in brown module were mainly linked to biomolecule degradation (e.g., fatty acid, valine, leucine, and isoleucine) and metabolism (e.g., pyrimidine and nitrogen metabolism). Although the yellow module consisted of a relatively small number of genes compared to others, these genes were found to be closely associated with flagellar assembly, biofilm formation, and bacterial chemotaxis. On the other hand, fla-2, fla-3, and fla-4 expression patterns were almost identical among samples, and their correlation coefficient values with six detected modules were also similar (Figure S3).
Clustering dendrogram with the flaA and flaB expression in each strain (A). The thicker red color denotes the higher flaA or flaB expression. The correlation between module eigengene and flaA or flaB expression (B). The highest enriched KEGG pathways in a turquoise (C-a), brown (C-b), and yellow module (C-c)
Correlation Analysis Between All Gene and flaA/flaB Expressions
The correlation coefficients between all gene expressions and flagellin genes (flaA and flaB) are shown in Fig. 7. Eight clusters (cluster 1–cluster 8) were identified based on correlation coefficient values between flaA and flaB. The top five pathways, each consisting of more than three genes from each cluster, were selected. Cluster 8, with both high expressions in flaA and flaB (flaA + /flaB +), comprised the genes that positively correlated with the expression of both flaA and flaB and were relevant to the propanoate and butanoate metabolism, valine, and leucine and isoleucine degradation. On the other hand, all genes that have both flaA and flaB negative correlation coefficients belonging to cluster 3, with both downregulations in flaA and flaB, (flaA-/flaB-), were mainly mapped onto the pyrimidine and purine metabolism, bacterial secretion system, etc. Clusters 1 and 2 showed a positive flaA but negative flaB correlation coefficient (flaA + /flaB-), and ribosome was the only pathway shared between clusters. Although many KEGG pathways share the same number of genes, relatively, many pathways were visualized in cluster 2, O-antigen nucleotide sugar biosynthesis, nitrogen metabolism, and ABC transporters were the highest pathways (Fig. 7). On the other hand, quorum sensing and ABC transporters were one of the featured pathways in cluster 6, composed of genes with negative flaA but positive flaB correlation coefficients (Fig. 7).
Discussion
In this study, we maximized the use of genomic and transcriptomic data to investigate the characteristics of different subunits of flagellin in A. hydrophila. This study incorporated comparative genomics analyses and conducted comprehensive profiling of transcriptomic responses across five representative strains of A. hydrophila. These strains harbored various copies of flagellin genes with characteristics that have yet to be clearly elucidated in their genomes. To assess the expression levels of individual flagellin genes, we mapped the transcriptomic data to their respective reference genomes. On the other hand, in examining the overall and global transcriptomic responses across strains, we analyzed the transcriptomic data by aligning it to a single reference genome. This approach facilitates a thorough investigation of the global transcriptomic expression across strains, enabling the preservation of diverse flagellin expression profiles within each genome. Additionally, it helps to the identification of correlated genes associated with flagellin expression.
Most of the A. hydrophila genomes exhibited a high ANI of over 95%; however, the B11 strain showed ANI values in the range of 93 to 94%, which are lower than the ANI comparison with A. dhakensis (~ 97%). The mislabeling issue of Aeromonas species in the GenBank database has been raised previously. Beaz-Hidalgo et al. [58] found that mislabeled A. hydrophila genomes in the GenBank, which had ANI values ranging from 93.70 to 93.84% with the A. hydrophila ML09-119 strain, actually belonged to A. dhakensis. Taken together, although the B11 strain has been labeled and utilized as A. hydrophila, it actually belongs to A. dhakensis. Considering that A. dhakensis has been documented as a highly virulent species capable of causing mortality in both humans and fish [59,60,61], further studies on the classification and understanding of A. dhakensis are necessary. However, the B11 strain did not harbor any distinctiveness in the pattern of flagellin subunit composition compared to other A. hydrophila strains. Among the five strains used for the transcriptomics study, the 23-C-23 and WCX23 strains, as well as the NJ-35 and SCV-1 strains, shared a high ANI of over 99.9%. Given that former studies [62, 63] reported that the isolation of clonal related A. hydrophila strains sharing high ANI (> 99%) in the fish, the strains over 99% ANI in this study may be genotypically clonal-related strains.
Flagella play a crucial role not only in the motility and virulence of bacteria but also in their evolutionary adaptation to diverse environmental conditions [64, 65]. Six flagellin clusters were phylogenetically identified among the flagellin sequences obtained from 151 A. hydrophila genomes, including flaA, flaB, and four additional flagellin. Notably, this study revealed significant phylogenetic variations among four fla genes annotated with the same term. Consequently, these flagellins were arbitrarily designated as fla-1 to fla-4. In general, the different subunits of flagellin composition can affect the swimming behaviors of bacteria [66]. Kühn et al. [67] investigated the phenotypic characteristics of the different assemblies of flaA and flaB in Shewanella putrefaciens, and flaA mutant strain (ΔflaA) and flaB mutant strains (ΔflaB) showed completely different patterns of swimming and colony structure. An impaired swimming capacity with significantly reduced filament length was observed in the ΔflaB strain, while ΔflaA strain did not exhibit substantial differences compared to the wild-type strains. Conversely, in the case of Helicobacter pylori, the ΔflaA strains showed a loss of motility, whereas the ΔflaB mutants retained their motility [68]. This indicates that the significance of flaA and flaB concerning bacterial motility may vary depending on the species and could be directly associated with bacterial motility. Interestingly, this study revealed that the majority of fla(+ 3) strains were derived from aquatic sources rather than terrestrial environments. In contrast, A. hydrophila strains in the flaA(+ 1)/flaB(+ 1) and fla(+ 1)/flaA(+ 1)/flaB(+ 1) groups tend to be more frequently isolated from terrestrial environments and animals. In Shewanella putrefaciens, diverse fractions of cells demonstrating screw motion in a high-viscosity environment were found to be influenced by the specific flagellin type [67]. Similarly, in the case of Aeromonas hydrophila, the observed variations in flagellin subunits harbored by different strains are likely to be a consequence of adaptive responses to distinct environmental challenges. Likewise, Nakamura and Minamino [36] suggested that flaA can be the principal component in soil bacteria such as Sinorhizobium meliloti and Rhizobium lupine. These results highlight the potential impact of flagellin diversity on the motility behavior of microorganisms and the adaptive significance of such variations in coping with varying environmental conditions.
In Aeromonas species, the sequence and genetic diversity in flagellin genes have greatly influenced the activity of biofilm formation, pathogenicity, and swimming motility [69]. Miyagi et al. [69] investigated genetic diversity in flaA in A. hydrophila, and the phylogenetic clusters were clearly divided in accordance with the type of origins, either clinical or environmental basis. The different level of motility and biofilm activity observed by flaA clusters indicated the genetic changes in the flagellin gene could drive different phenotypic characteristics. Given that flagella-relevant proteins, which directly exposed to outer environments, they can exhibit high variability due to external conditions. Based on the massive sequence information in Vibrio harveyi, the group incubated in seawater at 30 °C revealed a possibility that changes the genetic mutation in the flagellar relevant gene (putative flagellar motor switch protein) [29]. This study also verified some flagellin genes, for example fla-4, were clustered with the time of isolation (isolation year). This implied that the status of flagellin genes in A. hydrophila is not static but highly mutable and changeable, which can lead to environmental adaptation. However, the composition or nucleotide sequence of flagellin subunits did not show a significantly different pattern by country, not like subunits of environments such as aquatic and terrestrial traits.
The expression level of flagellin genes in ordinary cultural environments differed significantly depending on the flagellin type. FlaA and flaB showed relatively high expression, while fla-1 to fla-4 was not highly expressed like flaA and flaB. Interestingly, the strains belonging to the flaA(+ 1)/flaB(+ 1) group comprised many upregulated DEGs relevant to two-component system. Among many subordinated two-component systems, bacterial chemotaxis relevant to the flagellar motor switch adaptation pathway is featured and enriched subordinated systems in flaA(+ 1)/flaB(+ 1) group. In general, bacteria utilize chemotaxis to control their movement toward favorable environments by detecting the levels of chemoeffectors (e.g., attractants and repellents) through chemoreceptors [70]. The level of chemoeffectors can be detected by mcp and aer, and these signals were delivered through the cheW and cheA [71]. The level of autophosphorylated cheA is greatly influenced by signal transduction, which affects the stimulation of cheY by phosphorylation. Ultimately, phosphorylated cheY interacts with the flagellar motor and switches the direction of rotation in the flagellar [70, 72]. The high expression of flaA and flaB, along with the elevated expression of chemoreceptors and response regulator proteins (cheW, cheA, and cheY), might indicate that flaA(+ 1)/flaB(+ 1) group may have more robust flagella activity and/or movement, compared to the fla(+ 3) group. In addition, given that more than half number of strains in flaA(+ 1)/flaB(+ 1) group originated from terrestrial habitats where high variations of temperature and humidity were observed [73], it has been speculated that A. hydrophila can evolve and develop the type of flagellin fitting into the facing environment. Also, bacterial flagellins are known to the important epitope that causes the strong immuno-stimulatory response through toll-like receptor in aquatic animals [64, 74, 75]. Gao et al. [64] investigated that the level of immuno-stimulation effects varies depending on the subunits of flagellin (flaA, flaB, flaC, flaD, and flaE). This implies that the composition of flagellin genes in A. hydrophila could also have a significant impact on infection and prognosis.
In general, the construction of relationship networks based on the pattern of co-expression through pairwise measurement could be a robust approach for investigating biologically meaningful pathways. WGCNA, as the more advanced method, has received great attention to clarify the relationship between multidimensional results and featured sample traits [55, 76]. In this study, we applied the WGCNA method to identify the gene clusters that are highly correlated with flaA and flaB expression. FlaA and flaB were not typically co-expressed in A. hydrophila. For example, the yellow module, which positively correlated with the expression pattern of flaA but showed no significance with flaB, consisted of genes related to flagellar assembly, bacterial chemotaxis, and biofilm formation. On the other hand, a brown module, which showed a negative correlation with flaB expression but lacked significance with flaA, harbored the same flagellar assembly pathway. This does not necessarily imply that flaA and flaB have opposing functions in flagella assembly, but rather indicates that they do not share identical functions and characteristics. While both flaA and flaB may be associated with the flagellar assembly pathway, they may act in potentially different capacities or manners. Mohari et al. [77] investigated that Agrobacterium tumefaciens have several subunits of flagellin from flaA to flaD. FlaA in A. tumefaciens showed the highest importance that affected flagellin motility and functions, but others (flaB–flaD) had accessory functions [77]. Likewise, transcriptional expression of flaA and flaB in H. pylori is controlled by different sigma factors (ς28 and ς54), which suggests that both flagellin were regulated in a different timely manner [68, 78, 79]. Considering gene clusters or gene correlation analysis, it is a high possibility that flaA is probably associated with biofilm formation, bacterial chemotaxis, and bacterial secretion systems. Besides, flaB might be more associated with quorum sensing activities. But further studies are necessary to confirm this hypothesis.
Conclusion
This study conducted the first comprehensive investigation of hundreds of A. hydrophila genomes to profile flagellin subunits and clusters. We identified flaA, flaB, and four other phylogenetically different units of flagellin (fla-1 to fla-4). The composition of flagellin genes in A. hydrophila genomes exhibited significant diversity among strains, with a predominant presence of 3 to 4 fla subunits (fla(+ 3) and fla(+ 4) groups) originating from aquatic environments. In contrast, many terrestrial strains tended to harbor different flagellin subunits (flaA and flaB) in their genomes. Through multiple transcriptomic analyses conducted on strains with different flagellin subunits compositions under ordinary culture conditions, this study demonstrated significantly higher expression of flaA and flaB compared to the other four flagellins. Furthermore, genes associated with flagellin motor switch adaptation, controlling bacteria chemotaxis movement, showed significantly higher expression in the flaA(+ 1)/flaB( +) group compared to fla(+ 3). While further studies are necessary to elucidate the specific functions of each flagellin subunit, the pattern of flagellin composition and expression level found in this study provide valuable insights into the understanding of the role of flagellin and function and the adaptive mechanisms of microorganisms in diverse environments.
References
Chauret C, Volk C, Creason R, Jarosh J, Robinson J, Warnes C (2001) Detection of Aeromonas hydrophila in a drinking-water distribution system: a field and pilot study. Can J Microbiol 47:782–786. https://doi.org/10.1139/w01-070
Zhang Y, Ong C, Leung K (2000) Molecular analysis of genetic differences between virulent and avirulent strains of Aeromonas hydrophila isolated from diseased fish. Microbiology 146:999–1009. https://doi.org/10.1099/00221287-146-4-999
Fadl AA, Galindo CL, Sha J, Zhang F, Garner HR, Wang H-Q, Chopra AK (2007) Global transcriptional responses of wild-type Aeromonas hydrophila and its virulence-deficient mutant in a murine model of infection. Microb Pathog 42:193–203. https://doi.org/10.1016/j.micpath.2007.02.002
Ghatak S, Blom J, Das S, Sanjukta R, Puro K, Mawlong M, Shakuntala I, Sen A, Goesmann A, Kumar A (2016) Pan-genome analysis of Aeromonas hydrophila, Aeromonas veronii and Aeromonas caviae indicates phylogenomic diversity and greater pathogenic potential for Aeromonas hydrophila. Antonie Van Leeuwenhoek 109:945–956. https://doi.org/10.1007/s10482-016-0693-6
Zhao Xl WuG, Chen H, Li L, Kong Xh (2020) Analysis of virulence and immunogenic factors in Aeromonas hydrophila: towards the development of live vaccines. J Fish Dis 43:747–755. https://doi.org/10.1111/jfd.13174
Dong J, Zhang L, Liu Y, Zhou S, Yang Y, Xu N, Yang Q, Ai X (2021) Resveratrol influences the pathogenesis of Aeromonas hydrophila by inhibiting production of aerolysin and biofilm. Food Control 126:108083. https://doi.org/10.1016/j.foodcont.2021.108083
Chandrarathna H, Nikapitiya C, Dananjaya S, Wijerathne C, Wimalasena S, Kwun HJ, Heo G-J, Lee J, De Zoysa M (2018) Outcome of co-infection with opportunistic and multidrug resistant Aeromonas hydrophila and A. veronii in zebrafish: identification, characterization, pathogenicity and immune responses. Fish Shellfish Immunol 80:573–581. https://doi.org/10.1016/j.fsi.2018.06.049
Semwal A, Kumar A, Kumar N (2023) A review on pathogenicity of Aeromonas hydrophila and their mitigation through medicinal herbs in aquaculture. Heliyon:9. https://doi.org/10.1016/j.heliyon.2023.e14088
Lee S, Kim S, Oh Y, Lee Y (2000) Characterization of Aeromonas hydrophila isolated from rainbow trouts in Korea. J Microbiol 38:1–7
Dharmaratnam A, Swaminathan TR, Kumar R, Basheer V (2018) Aeromonas hydrophila associated with mass mortality of adult goldfish, Carassius auratus (Linnaeus 1758) in ornamental farms in India. Indian J Fish 65:116–126. https://doi.org/10.21077/ijf.2018.65.4.72719-14
Harikrishnan R, Balasundaram C (2005) Modern trends in Aeromonas hydrophila disease management with fish. Rev Fish Sci 13:281–320. https://doi.org/10.1080/10641260500320845
Neyts K, Huys G, Uyttendaele M, Swings J, Debevere J (2000) Incidence and identification of mesophilic Aeromonas spp. from retail foods. Lett Appl Microbiol 31:359–363. https://doi.org/10.1046/j.1472-765x.2000.00828.x
Stratev D, Odeyemi OA (2016) Antimicrobial resistance of Aeromonas hydrophila isolated from different food sources: a mini-review. J Infect Public Health 9:535–544. https://doi.org/10.1016/j.jiph.2015.10.006
Abulhamd AT (2009) Characterization of Aeromonas hydrophila isolated from aquatic environments using phenotypic and genotyping methods. Res J Agric Biol Sci 5:923–931
Awan F, Dong Y, Wang N, Liu J, Ma K, Liu Y (2018) The fight for invincibility: environmental stress response mechanisms and Aeromonas hydrophila. Microb Pathog 116:135–145. https://doi.org/10.1016/j.micpath.2018.01.023
Rasmussen-Ivey CR, Figueras MJ, McGarey D, Liles MR (2016) Virulence factors of Aeromonas hydrophila: in the wake of reclassification. Front Microbiol:1337. https://doi.org/10.3389/fmicb.2016.01337
Roh H, Kim BS, Lee MK, Park CI, Kim DH (2020) Genome-wide comparison of Carnobacterium maltaromaticum derived from diseased fish harbouring important virulence-related genes. J Fish Dis 43:1029–1037. https://doi.org/10.1111/jfd.13208
Roh H, Kim D-H (2021) Genotypic and phenotypic characterization of highly alkaline-resistant Carnobacterium maltaromaticum V-type ATPase from the dairy product based on comparative genomics. Microorganisms 9:1233. https://doi.org/10.3390/microorganisms9061233
Wardell SJ, Gauthier J, Martin LW, Potvin M, Brockway B, Levesque RC, Lamont IL (2021) Genome evolution drives transcriptomic and phenotypic adaptation in Pseudomonas aeruginosa during 20 years of infection. Microb Genom:7. https://doi.org/10.1099/mgen.0.000681
Liu J, Xiao G, Zhou W, Yang J, Wang Y, Wu Y, Cheng X, Sun Z (2021) Various novel colistin resistance mechanisms interact to facilitate adaptation of Aeromonas hydrophila to complex colistin environments. Antimicrob Agents Chemother:65. https://doi.org/10.1128/aac.00071-21
Zhu F, Yang Z, Zhang Y, Hu K, Fang W (2017) Transcriptome differences between enrofloxacin-resistant and enrofloxacin-susceptible strains of Aeromonas hydrophila. PLoS ONE 12:e0179549. https://doi.org/10.1371/journal.pone.0179549
Teng T, Xi B, Chen K, Pan L, Xie J, Xu P (2018) Comparative transcriptomic and proteomic analyses reveal upregulated expression of virulence and iron transport factors of Aeromonas hydrophila under iron limitation. BMC Microbiol 18:1–17. https://doi.org/10.1186/s12866-018-1178-8
Dong Y, Li S, Zhao D, Liu J, Ma S, Geng J, Lu C, Liu Y (2020) IolR, a negative regulator of the myo-inositol metabolic pathway, inhibits cell autoaggregation and biofilm formation by downregulating RpmA in Aeromonas hydrophila. npj Biofilms and Microbiomes 6:22. https://doi.org/10.1038/s41522-020-0132-3
Dong Y, Liu J, Nie M, Zhao D, Huang H, Geng J, Wan X, Lu C, Liu Y (2022) Comparative transcriptome combined with morphophysiological analyses revealed the molecular mechanism underlying Tetrahymena thermophila predation-induced antiphage defense in Aeromonas hydrophila. Virulence 13:1650–1665. https://doi.org/10.1080/21505594.2022.2127186
Montánchez I, Ogayar E, Plágaro AH, Esteve-Codina A, Gómez-Garrido J, Orruño M, Arana I, Kaberdin VR (2019) Analysis of Vibrio harveyi adaptation in sea water microcosms at elevated temperature provides insights into the putative mechanisms of its persistence and spread in the time of global warming. Sci Rep 9:289. https://doi.org/10.1038/s41598-018-36483-0
Yoon JB, Hwang S, Baek S-W, Lee S, Bang WY, Moon KH (2020) In vitro Edwardsiella piscicida CK108 transcriptome profiles with subinhibitory concentrations of phenol and formalin reveal new insights into bacterial pathogenesis mechanisms. Microorganisms 8:1068. https://doi.org/10.3390/microorganisms8071068
Lee Y, Kim N, Roh H, Kim A, Han H-J, Cho M, Kim D-H (2021) Transcriptome analysis unveils survival strategies of Streptococcus parauberis against fish serum. PLoS ONE 16:e0252200. https://doi.org/10.1371/journal.pone.0252200
Ruiz-Rodríguez M, Scheifler M, Sanchez-Brosseau S, Magnanou E, West N, Suzuki M, Duperron S, Desdevises Y (2020) Host species and body site explain the variation in the microbiota associated to wild sympatric Mediterranean teleost fishes. Microb Ecol 80:212–222. https://doi.org/10.1007/s00248-020-01484-y
Roh H (2022) A genome-wide association study of the occurrence of genetic variations in Edwardsiella piscicida, Vibrio harveyi, and Streptococcus parauberis under stressed environments. J Fish Dis 45:1373–1388. https://doi.org/10.1111/jfd.13668
Aigle A, Prosser JI, Gubry-Rangin C (2019) The application of high-throughput sequencing technology to analysis of amoA phylogeny and environmental niche specialisation of terrestrial bacterial ammonia-oxidisers. Environmental Microbiome 14:1–10. https://doi.org/10.1186/s40793-019-0342-6
Schiml VC, Delogu F, Kumar P, Kunath B, Batut B, Mehta S, Johnson JE, Grüning B, Pope PB, Jagtap PD (2023) Integrative meta-omics in Galaxy and beyond. Environmental Microbiome 18:56. https://doi.org/10.1186/s40793-023-00514-9
Beaz-Hidalgo R, Figueras M (2013) Aeromonas spp. whole genomes and virulence factors implicated in fish disease. J Fish Dis 36:371–388. https://doi.org/10.1111/jfd.12025
Canals R, Vilches S, Wilhelms M, Shaw JG, Merino S, Tomas JM (2007) Non-structural flagella genes affecting both polar and lateral flagella-mediated motility in Aeromonas hydrophila. Microbiology 153:1165–1175. https://doi.org/10.1099/mic.0.2006/000687-0
Wilhelms M, Vilches S, Molero R, Shaw JG, Tomás JM, Merino S (2009) Two redundant sodium-driven stator motor proteins are involved in Aeromonas hydrophila polar flagellum rotation. J Bacteriol 191:2206–2217. https://doi.org/10.1128/jb.01526-08
Sun L, Dong Y, Shi M, Jin M, Zhou Q, Luo Z-Q, Gao H (2014) Two residues predominantly dictate functional difference in motility between Shewanella oneidensis flagellins FlaA and FlaB. J Biol Chem 289:14547–14559. https://doi.org/10.1074/jbc.M114.552000
Nakamura S, Minamino T (2019) Flagella-driven motility of bacteria Biomolecules 9:279. https://doi.org/10.3390/biom9070279
Hu D, Reeves PR (2020) The remarkable dual-level diversity of prokaryotic flagellins. Msystems 5:e00705-00719. https://doi.org/10.1128/msystems.00705-19
Ro C, Altarriba M, Vilches S, Horsburgh G, Shaw JG, Tomás JM, Merino S (2006) Analysis of the lateral flagellar gene system of Aeromonas hydrophila AH-3. J Bacteriol 188:852–862. https://doi.org/10.1128/jb.188.3.852-862.2006
Ro C, Ramirez S, Vilches S, Horsburgh G, Shaw JG, Tomás JM, Merino S (2006) Polar flagellum biogenesis in Aeromonas hydrophila. J Bacteriol 188:542–555. https://doi.org/10.1128/jb.188.2.542-555.2006
Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S (2018) High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun 9:5114. https://doi.org/10.1038/s41467-018-07641-9
Warnes MGR, Bolker B, Bonebakker L, Gentleman R, Huber W, Liaw A (2016) Package ‘gplots’. Various R programming tools for plotting data:112–119. https://citeseerx.ist.psu.edu/document?repid=rep1&type=pdf&doi=61fde6caaed22f7be7032086f6130e65b505f9f4
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2012) GenBank. Nucleic Acids Res 41:D36–D42. https://doi.org/10.1093/nar/gks1195
R Core Team (2023) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna https://www.R-project.org/
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
Revell LJ (2012) phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 217–223. https://doi.org/10.1111/j.2041-210X.2011.00169.x
Warnes GR, Bolker B, Lumley T, Johnson RC (2024) gmodels: various R programming tools for model fitting. https://CRAN.R-project.org/package=gmodels
Hadfield J, Megill C, Bell SM, Huddleston J, Potter B, Callender C, Sagulenko P, Bedford T, Neher RA (2018) Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34:4121–4123. https://doi.org/10.1093/bioinformatics/bty407
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:1–10. https://doi.org/10.1186/gb-2009-10-3-r25
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, Van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515. https://doi.org/10.1038/nbt.1621
Mendiburu FD (2023) Agricolae: statistical procedures for agricultural research. https://cran.rproject.org/web/packages/agricolae/index.html
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:W182–W185. https://doi.org/10.1093/nar/gkm321
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:1–13. https://doi.org/10.1186/1471-2105-9-559
Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N, Workman C, Christmas R, Avila-Campilo I, Creech M, Gross B (2007) Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2:2366–2382. https://doi.org/10.1038/nprot.2007.324
Merico D, Isserlin R, Stueker O, Emili A, Bader GD (2010) Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE 5:e13984. https://doi.org/10.1371/journal.pone.0013984
Beaz-Hidalgo R, Hossain MJ, Liles MR, Figueras M-J (2015) Strategies to avoid wrongly labelled genomes using as example the detected wrong taxonomic affiliation for Aeromonas genomes in the GenBank database. PLoS ONE 10:e0115813. https://doi.org/10.1371/journal.pone.0115813
Huang M, Chen H, Li C, Liu Y, Gan C, El-Sayed Ahmed MAE-G, Liu R, Shen C, Zhong R, Tian G-B (2020) Rapid fulminant progression and mortality secondary to Aeromonas dhakensis septicemia with hepatitis B virus infection following the ingestion of snakehead fish in mainland China: a case report. Foodborne Pathog Dis 17:743–749. https://doi.org/10.1089/fpd.2019.2780
Bartie KL, Ngô TP, Bekaert M, Hoang Oanh DT, Hoare R, Adams A, Desbois AP (2023) Aeromonas hydrophila ST251 and Aeromonas dhakensis are major emerging pathogens of striped catfish in Vietnam. Front Microbiol 13:1067235. https://doi.org/10.3389/fmicb.2022.1067235
Soto-Rodriguez S, Lozano-Olvera R, Garcia-Gasca M, Abad-Rosales S, Gomez-Gil B, Ayala-Arellano J (2018) Virulence of the fish pathogen Aeromonas dhakensis: genes involved, characterization and histopathology of experimentally infected hybrid tilapia. Dis Aquat Org 129:107–116. https://doi.org/10.3354/dao03247
Hossain MJ, Sun D, McGarey DJ, Wrenn S, Alexander LM, Martino ME, Xing Y, Terhune JS, Liles MR (2014) An Asian origin of virulent Aeromonas hydrophila responsible for disease epidemics in United States-farmed catfish. MBio 5 https://doi.org/10.1128/mbio.00848-14
Rasmussen-Ivey CR, Hossain MJ, Odom SE, Terhune JS, Hemstreet WG, Shoemaker CA, Zhang D, Xu D-H, Griffin MJ, Liu Y-J (2016) Classification of a hypervirulent Aeromonas hydrophila pathotype responsible for epidemic outbreaks in warm-water fishes. Front Microbiol 7:216817. https://doi.org/10.3389/fmicb.2016.01615
Gao Q, Yi S, Li Y, Luo J, Xing Q, Yang X, Zhao M, Min M, Wang Q, Wang Y (2021) The role of flagellin B in Vibrio anguillarum-induced intestinal immunity and functional domain identification. Front Immunol 12:774233. https://doi.org/10.3389/fimmu.2021.774233
Chaban B, Hughes HV, Beeby M (2015) The flagellum in bacterial pathogens: for motility and a whole lot more. Semin Cell Dev Biol, vol. 46. Elsevier, pp. 91–103. https://doi.org/10.1016/j.semcdb.2015.10.032
Grognot M, Taute KM (2021) More than propellers: how flagella shape bacterial motility behaviors. Curr Opin Microbiol 61:73–81. https://doi.org/10.1016/j.mib.2021.02.005
Kühn MJ, Schmidt FK, Farthing NE, Rossmann FM, Helm B, Wilson LG, Eckhardt B, Thormann KM (2018) Spatial arrangement of several flagellins within bacterial flagella improves motility in different environments. Nat Commun 9:5369. https://doi.org/10.1038/s41467-018-07802-w
Suerbaum S, Josenhans C, Labigne A (1993) Cloning and genetic characterization of the Helicobacter pylori and Helicobacter mustelae flaB flagellin genes and construction of H. pylori flaA-and flaB-negative mutants by electroporation-mediated allelic exchange. J Bacteriol 175:3278–3288. https://doi.org/10.1128/jb.175.11.3278-3288.1993
Miyagi K, Shimoji N, Oshiro H, Hirai I (2023) Differences in flaA gene sequences, swimming motility, and biofilm forming ability between clinical and environmental isolates of Aeromonas species. Environ Sci Pollut Res 30:11740–11754. https://doi.org/10.1007/s11356-022-22871-7
Porter SL, Wadhams GH, Armitage JP (2011) Signal processing in complex chemotaxis pathways. Nat Rev Microbiol 9:153–165. https://doi.org/10.1038/nrmicro2505
Borkovich KA, Simon MI (1990) The dynamics of protein phosphorylation in bacterial chemotaxis. Cell 63:1339–1348. https://doi.org/10.1016/0092-8674(90)90429-I
Sourjik V, Berg HC (2002) Receptor sensitivity in bacterial chemotaxis. Proc Natl Acad Sci 99:123–127. https://doi.org/10.1073/pnas.011589998
Or D, Smets BF, Wraith J, Dechesne A, Friedman S (2007) Physical constraints affecting bacterial habitats and activity in unsaturated porous media–a review. AdWR 30:1505–1527. https://doi.org/10.1016/j.advwatres.2006.05.025
Li Y, Yao H, Liu S, Song D, Wu C, Zhang S, Gao Q, Zhang L (2023) The role of flagellin F in Vibrio parahaemolyticus-induced intestinal immunity and functional domain identification. Int J Biol Macromol 244:125404. https://doi.org/10.1016/j.ijbiomac.2023.125404
Li Y, Liu C, Sun Y, Wang R, Wu C, Zhao H, Zhang L, Song D, Gao Q (2024) Construction of the flagellin F mutant of Vibrio parahaemolyticus and its toxic effects on silver pomfret (Pampus argenteus) cells. Int J Biol Macromol 259:129395. https://doi.org/10.1016/j.ijbiomac.2024.129395
Pei G, Chen L, Zhang W (2017) WGCNA application to proteomic and metabolomic data analysisMethods in enzymology. Elsevier, pp 135–158. https://doi.org/10.1016/bs.mie.2016.09.016
Mohari B, Thompson MA, Trinidad JC, Setayeshgar S, Fuqua C (2018) Multiple flagellin proteins have distinct and synergistic roles in Agrobacterium tumefaciens motility. J Bacteriol:200. https://doi.org/10.1128/jb.00327-18
Leying H, Suerbaum S, Geis G, Haas R (1992) Cloning and genetic characterization of a Hellcobacter pylori flagellin gene. Mol Microbiol 6:2863–2874. https://doi.org/10.1111/j.1365-2958.1992.tb01466.x
Allan E, Dorrell N, Foynes S, Anyim M, Wren BW (2000) Mutational analysis of genes encoding the early flagellar components of Helicobacter pylori: evidence for transcriptional regulation of flagellin A biosynthesis. J Bacteriol 182:5274–5277. https://doi.org/10.1128/jb.182.18.5274-5277.2000
Acknowledgements
The authors thank Institute of Marine Research (Havforskningsinstituttet) for supporting the open access of this article.
Funding
Open access funding provided by Institute Of Marine Research
Author information
Authors and Affiliations
Contributions
HR and DK conceptualized the study. HR carried out data curation, method development, data visualization, results analysis, and writing the original draft. DK edited and revised the manuscript. All authors read and approved the final version of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of Interest
The authors declare that they have no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Roh, H., Kannimuthu, D. Genomic and Transcriptomic Diversification of Flagellin Genes Provides Insight into Environmental Adaptation and Phylogeographic Characteristics in Aeromonas hydrophila. Microb Ecol 87, 65 (2024). https://doi.org/10.1007/s00248-024-02373-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02373-4