Abstract
Sugar beet is the most important crop for sugar production in temperate zones. The plant microbiome is considered an important factor in crop productivity and health. Here, we investigated the bacterial diversity of seeds, roots, and rhizosphere of five sugar beet hybrids named Eduarda (ED), Koala (KO), Tibor (T), Tajfun (TF), and Cercospora-resistant (C). A culture-independent next-generation sequencing approach was used for the further investigation of seed-borne endophytes. Hybrid-associated bacteria were evaluated for their plant growth–promoting (PGP) characteristics, antagonistic activity towards Cercospora beticola and several Fusarium strains in dual culture assays, and drought and salinity tolerance. High-throughput sequencing revealed that the Proteobacteria phylum was most dominant in the seeds of all hybrids, followed by Cyanobacteria and Actinobacteriota. The predominant genus in all hybrids was Pantoea, followed by Pseudomonas, Acinetobacter, Chalicogloea, Corynebacterium, Enterobacter, Enterococcus, Glutamicibacter, Kosakonia, and Marinilactibacillus. Unique genera in the hybrids were Pleurocapsa and Arthrobacter (T), Klebsiella (TF), Apibacter (ED), and Alloscardovia (KO). The genera that were most represented in one hybrid were Weissella and Staphylococcus (TF); Streptococcus (T); Gardnerella, Prevotella, and Rothia (KO); and Gilliamella, Lactobacillus, and Snodgrassella (ED). Thirty-two bacteria out of 156 isolates from the rhizosphere, roots, and seeds were selected with respect to various plant growth–promoting activities in vitro, i.e., nitrogen fixation, phosphate solubilization, siderophore production, indole-3-acetic acid production, 1-aminocyclopropane-1-carboxylic acid deaminase activity, hydrogen cyanide production, exoenzymatic activity (amylase, protease, lipase, cellulase, xylanase, mannanases, gelatinase, and pectinase), mitigation of environmental stresses, and antifungal activity. Mixta theicola KO3-44, Providencia vermicola ED3-10, Curtobacterium pusillum ED2-6, and Bacillus subtilis KO3-18 had the highest potential to promote plant growth due to their multiple abilities (nitrogen fixation, phosphate solubilization, production of siderophores, and IAA). The best antagonistic activity towards phytopathogenic fungi was found for Bacillus velezensis C3-19, Paenibacillus polymyxa C3-36 and Bacillus halotolerans C3-16/2.1. Only four isolates B. velezensis T2-23, B. subtilis T3-4, B. velezensis ED2-2, and Bacillus halotolerans C3-16/2.1 all showed enzymatic activity, with the exception of xylanase production. B. halotolerans C3-16/2.1 exhibited the greatest tolerance to salinity, while two B. subtilis strains (C3-62 and TF2-1) grew successfully at the maximum concentration of PEG. The current study demonstrates that sugar beet–associated bacteria have a wide range of beneficial traits and are therefore highly promising for the formulation of biological control and PGP agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants live in a close symbiotic relationship with their microbiota and are thus regarded as holobionts where the cells of the plant host are typically outnumbered by the related microorganisms [1, 2]. Depending on the plant environment, plant-associated microbiota can be found on the exterior of plants, such as the rhizosphere or phyllosphere, or in the interior of plants, referred to as the endosphere [2]. The soil harbors a wide variety of microorganisms, with the root microbiota mainly acquired from the soil through horizontal transfer. Additionally, seeds serve as another crucial source of microorganisms, vertically transmitting their microbiota, and some bacteria can thrive in the roots of growing plants [3]. The presence of particular co-occurrence patterns and microbial networks highlights the fact that the colonization of plants by microbes is a targeted process [2]. Distinct plant species grew in symbiosis with remarkably different microbial communities in the rhizosphere and root compartments while occupying the same soil environment [3]. The structure of plant roots and their exudates, such as amino acids, phenolic compounds, organic acids, sugars, and other small secondary metabolites, plays an important role in shaping the rhizosphere microbiome. Variations in the plant genome affect the composition of exudates and root structure, resulting in different microbiome assemblies. It has been reported that even a minor genetic variation of the host plant can affect its microbiome composition [4]. The composition of plant secondary metabolites is also influenced by their associated microbiota which results in the development of new metabotypes as has been shown recently [2]. Some plants under the phosphate or iron deficiency tend to increase the secretion of citrate, malate, or oxalate to enrich the rhizosphere with organic carbon, which attracts beneficial microorganisms [5]. While previous research was focused more on the plant-pathogen relationship, discoveries from the past decade elucidate significance of the beneficial interaction between microbes and their hosts. Strong involvement of microbial diversity in the development of these interactions was recognized [6]. Plant growth–promoting bacteria (PGPB) are beneficial bacteria that can stimulate plant growth through various mechanisms such as nitrogen fixation, phosphate solubilization, production of siderophores, indole-3-acetic acid (IAA), and 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase [7]. Furthermore, PGPBs are recognized for the protection of host plants against phytopathogens via different mechanisms that involve the production of antimicrobial components, hydrolytic enzymes, competition for nutrients, and induction of resistance in the host plant [8]. Drought and high salinities are one of the limiting factors in the growth and productivity of agricultural crops, especially in arid and semi-arid regions. As a natural hazard, drought in Serbia in the summer of 2017 severely affected agricultural production, reducing yields of corn, sunflower, soybeans, and sugar beets between 30 and 60%, while the total loss was estimated at $1.5 billion [9]. In this regard, it has been reported that PGPBs enhanced tolerance to drought and salt stress of many different crops [10]. Moreover, recent research has revealed that the plant microbiome also contributes to cold acclimation, a key element restricting plant geographic range as well as crop development and yield in specific regions [2]. Thus, the plant microbiome is acknowledged as a significant driver of plant productivity and health.
The population of the world is anticipated to increase steadily and will total 9.8 billion in 2050 and 11.2 billion in 2100. Simultaneously, there will be less area accessible for crop production, indicating that agricultural management and technologies must be used to enable intensification of crop production [11]. A more sustainable approach to farming may result from manipulating the plant microbiome, which also has the potential to boost agricultural output, decrease chemical inputs, and lower greenhouse gas emissions. This goal is considered essential for supporting the world’s expanding population [12].
Sugar beet is the most important crop for sugar production in temperate zones, while it is second in the world, covering 30% of the world sugar production. Recently, sugar beet has also been recognized as a source for biofuel production [13, 14]. Unfortunately, sugar beet production is exposed to several phytosanitary problems which include various fungal pathogens resulting in significant losses in crop yield and quality [15]. Cercospora leaf spot is one of the most widespread and destructive foliar diseases of sugarbeet caused by Cercospora beticola [15]. Seeds, seedlings, roots, and leaves are affected by Fusarium species, among which F. oxysporum causes various diseases including Fusarium yellow, seedling wilt, and root rot [16]. Fusarium wilt was also recorded for been, tomato, and cowpea, whereas F. equiseti poses a serious threat for significant yield losses [17]. Fusarium solani is a causal agent of Fusarium root rot of sugar beets in the USA [18]. Recently, several Fusarium species were isolated from sugar beet roots that showed symptoms of root rot, including F. graminearum, F. nygamai, F. equiseti, and F. verticillioides [18]. To date, they have not been characterized as sugar beet pathogens, but in the pathogenicity test, all listed species caused root rot symptoms similar to natural infections [18]. The use of fungicides has been effective in controlling these diseases under field conditions [15]. However, the use of fungicides for crop protection is associated with several negative consequences, such as the emergence of resistant strains of pathogens, the reduction of the number of beneficial microorganisms in the microbiome, and the accumulation of toxic substances in the soil [19]. The use of microorganisms such as bacteria to control fungal diseases and stimulate crop growth is a strategy that is part of sustainable and eco-friendly production.
Exploring plant-microbe interactions and community building provides vital insights into plants as metaorganisms. This holistic understanding enhances our knowledge of mutual benefits between plants and microbes, leading to sustainable agriculture and improved crop resilience in challenging environments. With more information about the plant microbiota in relation to biotic and abiotic stresses, plant genotypes, and environmental conditions, it may be possible to identify more suitable candidates or approaches for inoculation in a given environment. The microbiome of the rhizosphere, roots, and seeds of various sugar beet cultivars has been reported [20,21,22]. We have closely examined the bacterial diversity in the rhizosphere, roots, and seeds of the most commonly used sugar beet hybrids (Eduarda, Koala, Tibor, Tajfun, and the Cercospora-resistant hybrids) in Serbia (Fig. 1). In addition, our study represents a pioneering investigation of the diversity of total seed-borne endophytic bacteria using the next-generation sequencing (NGS) approach. By culturing endophytic (roots and seed) and rhizospheric bacteria and screening them for plant growth–promoting properties, enzyme production, stress tolerance, and antifungal potential, we also intend to identify important microbial strains with beneficial properties. Our results should contribute to the development of targeted agricultural strategies by providing insights into specific microbial traits that can improve sugar beet health and overall crop performance.
Materials and Methods
Plant Material
Five hybrids of sugar beet, i.e., Eduarda, Koala, Tibor, Tajfun, and Cercospora-resistant, were used in this study, hereafter referred as ED, KO, T, TF, and C, respectively. Plant samples and their surrounding rhizosphere of sugar beet hybrids were collected in May 2021 at the experimental field of Sunoko company in Kovačica, Serbia (45°05′06.7″N 20°38′07.2″E) at V2.1 growth stage (two leaves are unrolled, and the third leaf is just visible) [23] and transported to the laboratory in bags.
DNA Extraction, Library Preparation, and NGS
Sugar beet seeds were obtained commercially by the Sunoko Research and Development Center. The seeds (10 seeds per hybrid) were sterilized with 70% ethanol, shaken for 1–2 min and then washed with water. The procedure was repeated with 3% sodium hypochlorite, and the samples were washed with water several times. After surface sterilization, the seed samples were crushed in a mortar. Extraction of total DNA was performed using a Quick-DNATM Faecal/Soil Microbe Miniprep Kit (Zymo Research, Irvine, CA, USA) according to the manufacturer’s instructions. The DNA was quantified using Qubit fluorometric quantitation (Qubit 4 fluorometer, Invitrogen™, Waltham, MA, USA). DNA samples were commercially sequenced by Novogene Co., Ltd. (Cambridge, UK) using a 2 × 250-bp paired-end run on a MiSeq Sequencer, according to the manufacturer’s instructions (Illumina, San Diego, CA, USA). The 16S rRNA gene-specific sequences to target the V3 and V4 regions (470 bp) were used in this study, with 341F (5′-CCTACGGGNGGCWGCAG-3′) and 805R (5′-GACTACHVGGGTATCTAATCC-3′) primers [24]
Sequence Data Processing and Taxonomy Annotation
Following the initial quality check, primers were eliminated using cutadapt 3.4 in paired-end mode with default settings. Only read pairs containing both primers were subjected to further processing. Subsequently, sequence inference was performed using the dada2 R package [25]. Quality trimming of reads was accomplished using default parameters in the filterAndTrim function, with the additional criteria of right trimming after 223 nt for forward and reverse reads, and discarding reads shorter than 150 nt. Sequences with more than three expected errors for the forward and reverse strands, calculated using the quality score (Q), were discarded (maxEE = c(3, 3), where Q is the quality score represented as sum(10^(−Q/10))). To join paired sequences, a minimum overlap of 12 nt (default) was used. Chimera removal was performed with default options in removeBimeraDenovo. Taxonomy assignment up to the genus level was accomplished using the SILVA 138.1 (silva_nr99_v138.1) strain set (https://zenodo.org/record/4587955) with IDTAXA [26] using default parameters and a confidence threshold set to 50. After annotating using Silva 138.1 as described above, the taxonomy was further supplemented by conducting a blast search against the NCBI 16S ribosomal RNA database (accessed at May 5, 2022). Blast best hits (BBH) with at least 98% identical matches and at least 99% query coverage were used to annotate missing taxonomies, but only if the entire taxonomy between the BBH and Silva annotation, up to the highest annotated taxonomic level, matched. Species-level annotation was performed by exact sequence matching (0 mismatches) against the combined Silva species (silva_species_assignment_v138.1 at https://zenodo.org/record/4587955) and NCBI 16S ribosomal RNA databases (accessed at May 5, 2022). In cases where the amplicon sequence variants (ASVs) matched multiple sequences, a concatenated string of all exactly matched species was employed as the species-level annotation.
Bioinformatic and Statistical Analyses
Alpha diversity, assessed at ASV, species, genus, family, and phylum levels using the phyloseq R package [27], was determined after rarefaction to a consistent depth (101,470 reads, corresponding to the sample with the lowest reads). Chao, ACE, OBS, Simpson, and Shannon indices were used for evaluation. Kruskal–Wallis one-way analysis of variance compared alpha diversity among the five hybrids, with Dunn’s test presenting significant differences denoted by distinct letters above the bars. Beta diversity, evaluated at ASV, species, and genus levels, utilized double principal coordinate analysis (DPCoA) [28]. Prior to ordination, singleton ASVs and those with a read sum below 30 across all samples were removed. Multivariate homogeneity of group dispersions was assessed via PERMDISP2 [29] on the DPCoA distance matrix. PERMANOVA (ADONIS) [30] tested compositional similarities among groups, and pairwise ADONIS analyses examined dissimilarities between groups. Differential abundance analysis, aggregated up to the species level following the same ordination as for DPCoA, was performed with the microbiomeMarker package [31] using DESeq2 [32], which employed default parameters. The Bonferroni method adjusted p-values, considering p < 0.01 as statistically significant. Differential abundance, aggregated up to the species level, excluded species present in only one sample or with a read sum below 30 across all samples before analysis.
Isolation of Endophytic and Rhizospheric Bacteria from Sugar Beet Hybrids
Samples of the lower part of the roots, approx. 100 mg (five roots of one hybrid plant) and seeds (five seeds of each hybrid), were washed, cleaned of soil particles with tap water, and sterilized as described above. After surface sterilization, the root samples were dried and chopped into a previously sterilized mortar with sterile tweezers and scissors. The roots and seeds were then crushed in the mortar and transferred to the 1.5 mL of Luria–Bertani (LB) medium supplemented with glycerol (final concentration 20%). The prepared stocks were stored at −80 °C until the further use. Eight different nitrogen-free media (NFb, LGI, JNFb, LGI-P, modified LGI with acids, JMV, M-medium, and M-medium without biotin) were used to isolate culturable endophytic bacteria from seeds and roots, as summarized in the work of Baldani et al. [33]. Rich culture media such as tryptic soy agar (TSA), nutrient agar (NA), and glucose-yeast calcium carbonate (GYC) were also used. Of the starting material obtained from the seed sample, 100 μL was spread onto each isolation medium, while the root material was diluted 1:10 before application to the media and then incubated at 30 °C for 10 days. After the incubation period, morphologically distinct colonies were restreaked onto NA and incubated under the same conditions to obtain axenic cultures. For selective isolation of Bacillus species from the rhizosphere, the thermal inactivation method was used. In short, the soil sample of each hybrid was transferred to test tubes with 3 mL of nutrient broth medium (NB) and incubated at 80 °C for 10 min. Furthermore, the tubes were incubated at 30 °C until growth appeared. Cultures were streaked on the Luria–Bertani agar plates (LA), and colonies of different morphologies were further selected. Pure cultures of the selected bacterial isolates from all origins were stored in LB glycerol stocks at −80 °C until further use.
Molecular Identification of Endophytic and Rhizospheric Bacteria from Sugar Beet Hybrids
Genomic DNA obtained from a bacterial colony was used for PCR amplification [34]. From pure cultures growing on NA plates, a single colony was picked with the top of the pipette tips and resuspended in 25 μL of sterile deionized water. The prepared samples were incubated at 98 °C for 10 min. Colony PCR amplifications were performed in a 25-μL reaction mixture containing 12.5 μL FastGene Taq ready mix with dye (NIPPON Genetics Europe, GmbH, Düren, Germany), 0.5 μL of each primer (10 μM), 5 μL of DNA template, and PCR water to final volume. Primers used for the amplification of 16S rRNA gene (1500 bp) were the following: fD1Funi-16SF (5′-AGAGTTTGATCCTGGCTCAG-3′) and Rp2Runi-16SR (5′-ACGGCTACCTTGTTAGGACTT-3′). The following cycling conditions were used: one step of initial denaturation at 95 °C for 5 min followed by 30 cycles of denaturation at 95 °C for 40 s, annealing at 54 °C for 40 s and extension at 72 °C for 90 s and final extension step at 72 °C for 7 min. All PCR amplicons were purified using a purification kit (Euroclone spinNAker Gel&PCR DNA Purification Kit, Italy) according to the manufacturer’s instructions, and later sequenced by Eurofins Genomics Europe Sequencing service (Wien, Austria) using the 907R-16S primer (5′-CCGTCAATTCMTTTRAGTTT-3′). The obtained sequences were searched for sequence homologs of the reference strains in the NCBI GenBank database using the BLASTn algorithm. Sequences of the most closely related strains were used to ensure taxonomic identification. Sequences were aligned using the CLUSTAL W algorithm implemented in the BioEdit 7.2.5 program, and nucleotide position was checked manually.
Evaluation of Endophytic and Rhizospheric Sugar Beet Bacteria for Their Plant Growth-Promoting (PGP) Properties In Vitro
Unless otherwise stated, 5 μL of the overnight culture was used in all tests. The ability of the isolates to fix nitrogen was determined by subculturing on nutrient media with and without NH4Cl as a source of nitrogen [35]. Screening of phosphate solubilization was performed on the NBRIP agar medium according to the method of Nautiyal [36]. The potential of sugar beet isolates to produce indole-3-acetic acid (IAA) was evaluated by the modification of the method described by Gordon and Weber [37]. A full loop (1 μL size) of bacterial culture was inoculated into the 1 mL of IAA detection medium and shaken for 24 h at 30 °C. Medium without bacteria was used as a negative control. After incubation, tubes were centrifuged at 10,000 rpm for 10 min. Subsequently, 1 mL of supernatant was transferred to the new tube and mixed with 1 mL of Salkowski reagent. Prepared samples were incubated for 2 h and then evaluated based on the color. Depending on the intensity of the color change (light orange, dark orange and red), IAA production was scored as follows: low (+), medium (++), and high (+++). Siderophore production was characterized by the modification of the O-CAS method described by Pérez-Miranda et al. [38]. Of bacterial overnight culture, 10 μL was poured into sterile wells previously placed on NA plates and incubated at 30 °C for 24 h. After incubation, 10 mL of CAS medium was poured on each plate. Following CAS medium solidification, the wells were removed and the plates were incubated at 30 °C for 24 h. The appearance of clear zones around bacterial colonies was a confirmation of siderophore production. Qualitative assessment of 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase activity was conducted following the methodology outlined by Gupta et al. [39]. Exopolysaccharide (EPS) production was determined by subculturing isolates on yeast mannitol agar (YMA) plates, using the streak-plate procedure [40]. The motility assay was performed according to the method of Kim and Surette [41]. The ability of the isolates to produce hydrogen cyanide (HCN) was determined using Cyantesmo indicator strips (Machery-Nagel GmbH&Co., Germany) as described by Knežević et al. [42].
Evaluation of Endophytic and Rhizospheric Sugar Beet Bacteria for Extracellular Enzyme Production In Vitro
The ability of the isolates to produce extracellular enzymes was assessed by inoculating 5 μL of the overnight bacterial culture onto the specific solid media. After 24 or 48 h of incubation at 30 °C, the enzymatic index (EI) was determined. The EI was calculated according to the following formula: EI = diameter of hydrolytic zone / colony diameter (mm), as described by Saroj and Narasimhulu [43]. For EI > 1, the activity was considered good; for EI = 1–2, the activity was very good; and for EI > 2, the activity was considered high. The amylase, protease, xylanase, and mannanase production of sugar beet isolates was performed as previously described [44]. Lipolytic activity was tested on plates containing TSA and 1% glycerol tributyrate, while production was confirmed by appearance of the illuminated zones around bacterial growth [45]. Furthermore, sugar beet isolates were evaluated for their ability to produce cellulases [46], gelatinases [47], and pectinases [48].
Stress Tolerance of Endophytic and Rhizospheric Sugar Beet Bacteria In Vitro
Salinity stress tolerance was evaluated by the modification of the method described by Gupta et al. [39]. NA plates, previously enriched with 1%, 3%, 5%, 8%, and 10% NaCl, were spot inoculated with 5 μL of the overnight bacterial culture and incubated for 24 h at 30 °C, after which the growth of isolates was evaluated. The results are presented using positive and negative symbols relative to the growth observed at 1% NaCl in the substrate. For drought tolerance, a method adapted from Ali et al [49] was used. Overnight cultures were grown in diluted TSB medium (10 g/L), and after measuring the optical density (OD600), all isolates were diluted to 0.4. The tolerance of strains to desiccation was studied in TSB medium (10 g/L) amended with 5, 10, 20, and 30% PEG in flasks containing 1% bacterial cultures (OD600 = 0.4). The flasks were incubated overnight at 30 °C (150 rpm). Evaluation of the percent growth of isolates in the various concentrations of PEG was performed in comparison to a control, with the optical density of the control (OD600) set at a value of 100%. These percentage results were then labelled with symbols corresponding to the different concentration intervals: + for the 0–30% range, ++ for the 30–60% range, and +++ for the 60–100% range.
Antifungal Potential of Endophytic and Rhizospheric Sugar Beet Bacteria Against Phytopathogenic Fungi In Vitro
The antifungal potential against the 22 phytopathogenic fungi was evaluated using the initial screening and co-culture plate test [50]. The fungal pathogens tested were Fusarium sp. TS1, Fusarium oxysporum TS2, Fusarium equiseti TS3, Fusarium subglutinans TS5, Fusarium nigamai TS6, Fusarium solani TS8, and Cercospora beticola TS4 from sugar beet as well as Fusarium semitectum TS7, F. graminearum GD1, F. graminearum S3-7, F. graminearum CIK, F. oxysporum S4-2, F. verticillioides K67 5.1, F. venatum IB1I-12, F. ipomoeae IB6I-10, F. foetens IP27, F. falciforme IP31, F. coffeatum IP32, and F. denticulatum IP39 from different origins. The origin of all phytopathogenic fungi is listed in Table S1. The existing collection is part of the Department of Algology and Mycology, Institute of Botany and Botanical Garden “Jevremovac,” University of Belgrade — Faculty of Biology. Effects on mycelial growth were evaluated by calculating the percent of growth inhibition, PGI % = 100(KR − R1)/KR, where KR represents the distance (measured in mm) from the point of inoculation to the colony margin on the control dishes, and R1 is the distance of fungal growth from the point of inoculation to the colony margin on the treated dishes in the direction of the antagonist [50]. The experiments were repeated twice independently, with three replications for each fungus. Following the Kolmogorov–Smirnov test for normality, standard variance analysis was used to analyze the obtained PGI% results (one-way ANOVA test). In vitro mycelial growth inhibition percentages were averaged out using the Tukey’s HSD test. Statistical significance applied in all tests was p < 0.05. The statistical analysis was performed according to the general procedures of IBM SPSS Statistics v.23 (SPSS, Inc., Armonk, NY, USA).
Results
The Analysis of Alpha and Beta Diversities of the Seed Endophytic Bacteria
The phylogenetic composition of bacterial communities associated with seeds of five sugar beet cultivars was analyzed by amplification and sequencing of the V3–V4 region of the 16S rRNA gene. The average number of amplicon sequence variants (ASVs) after removal of chimeras per hybrid was 75838.33 (C), 71714.33 (ED), 77315.67 (KO), 73495.00 (T), and 75020.67 (TF) (Table S6). Alpha diversity indexes were calculated for the number of observed features (OBS), Chao1, ACE, and Shannon and Simpson at family, genus, species, and ASV levels (Fig. 2). Higher alpha diversity was evident in ED, KO, and T hybrid across all taxonomic levels, in contrast to C and TF hybrid, with the observed richness showing correlation with estimated richness, except at the ASV level. ED hybrid displayed the highest richness based on Chao1 and ACE indices at the ASV level, while both ED and T hybrids exhibited the highest observed richness according to OBS. Conversely, TF hybrid consistently exhibited the lowest richness at all taxonomic levels based on both estimated and observed richness estimators. Shannon and Simpson indices indicated no statistically significant changes in alpha diversity at the family level between TF and C hybrid and between KO and T hybrid. However, at the species level, statistically significant differences were observed between all hybrids, with Shannon indices being the discriminant factor. On the other hand, at the ASV level, no statistically significant differences were found between ED, KO, and T hybrids, whereas C and TF hybrids showed significant differences from all other hybrids. The PCoA plot demonstrated that KO hybrid at the ASV level exhibited separation from the other hybrids along axis 1, explaining 67.7% of the variability, while ED, C, and T hybrids were grouped together (Fig. 3). TF hybrid showed distinct separation from the others along axis 2, accounting for 12% of the variance. Better separation of hybrids was observed at the genus level. The first axis (explaining 72.6% of variability) indicated that C and TF were distinct from the remaining hybrids based on genus composition, with KO hybrid being the most distant from all hybrids. Additionally, along the second axis (explaining 9.9% of variability), a clear separation of C hybrid from TF hybrid and the other hybrids, as well as its association with ED hybrid, was observed. At the genus and ASV levels, KO and TF appeared to be more distant from the remaining hybrids. PERMDISP2 results revealed a non-significant difference in hybrids dispersions at all taxonomic levels. However, the PERMANOVA test showed statistically significant variability between hybrids (p = 0.001) at all taxonomic levels.
The alpha diversity in the five hybrids Tajfun (TF), Eduarda (ED), Koala (KO), Tibor (T), and Cercospora-resistant (C) was compared at all tax-level using Kruskal–Wallis one-way analysis of variance, while a Dunn post hoc test was run for pairwise group comparison. The values marked with the same letter within the diagram columns do not indicate statistically significant differences
The Composition of the Seed Endophytic Bacteria Determined Using the NGS Approach
Of the 16 phyla detected, the phylum Proteobacteria was the most widespread in all five hybrids with relative abundance (RA) ranging from 38.63% in the KO hybrid up to 77.90% in the C hybrid (Fig. 4). The following major phyla in all hybrids were Cyanobacteria and Actinobacteriota, except in the TF hybrid. A highest representation of the phylum Firmicutes was detected in the TF hybrid with a RA of 24.65%, while Bacteroidota was the most abundant in the KO (4.20%). The most prevalent genus was Pantoea detected in the C, ED, and TF hybrids. Pseudomonas was the second most abundant genus in all hybrids, except for TF. In general, TF hybrid was characterized by Pantoea (30.23%), Enterobacter (13.63%), Kosakonia (10.35%), Acinetobacter (9.82%), Enterococcus (9.22%), Weissella (7.14%), Staphylococcus (6.47%), and Erwinia (2.31%). Genera Gardnerella (4.88%), Rubrobacter (3.71%), Glutamicibacter (2.28%), Sphingomonas (2.24%), Rothia (2.18%), and Prevotella (2.02%) were most abundant in KO hybrid. Most dominant genera in the T hybrid were Pantoea (13.87%), Pseudomonas (12.09%), and Acinetobacter (11.74%), followed by Actinomycetospora and Streptococcus (4.25% and 3.38%, respectively). The C hybrid had the lowest genera diversity, and it was dominated by Pantoea (36.09%), Kosakonia (19.15%), and Pseudomonas (15.10%). Most detected genera in ED were Pantoea (26.17%), Pseudomonas (16.76%), Gilliamella (4.37%), and Lactobacillus (4.11%) and two unidentified belonging to the families Chroococcidiopsaceae and Microcystaceae, with 7.81% and 3.48%, respectively. The genus Songrasella (2.72%) was most abundant in ED.
The highest average reads of Pantoea agglomerans/ananatis/conspicua/eucalypti/vagans at ASV level were recorded for TF, C, and T (Fig. 5). Kosakonia cowanii with the highest percentage of reads was detected in C and TF, while its abundance was significantly reduced in ED, KO, and T hybrids. Acinetobacter lwoffii was detected in T hybrid. Furthermore, ASV related to Acinetobacter baumannii/junii had a similar abundance in T and KO, while Acinetobacter bouvetii/haemolyticus/johnsonii/oryzae was characteristic for the TF. Weissella was highly detected in the TF hybrid with 7% of reads assigned as Weissella soli. Enterococcus saccharolyticus was highly presented in the TF hybrid. A higher percentage of ASV reads for Pseudomonas oryzihabitans and Pseudomonas fulva/oryzihabitans was detected in C and ED hybrid.
The Differential Abundance Analysis of the Seed Endophytic Bacteria Between Hybrids
The differential abundance estimation identified 23 taxa that were significantly more abundant in some hybrids compared to others (Fig. 6). Specifically, Actinobacteria and Alphaproteobacteria were found to be significantly more abundant in KO and T hybrids. On the other hand, families such as Pseudomonadaceae, Paenibacillaceae, Pseudonocardiaceae, Enterococcaceae, and Staphylococcaceae exhibited statistically different representation in the hybrids. Notably, TF and C hybrids exhibited a significantly higher presence of the Enterobacteriaceae family compared to the other hybrids. Family Pseudomonadaceae was detected in all samples but displayed a statistically higher abundance in ED, KO, T, and E hybrids compared to TF hybrid. Similarly, Paenibacillaceae showed greater abundance in T and C hybrids, while Pseudonocardiaceae was more prevalent in KO and T hybrids. Enterococcaceae and Staphylococcaceae were found to be the most abundant families in TF hybrid, in comparison to the other hybrids. The separation of hybrids was influenced by the orders Burkholderiales, Paenibacillales, Pseudonocardiales, and Staphylococcales. Burkholderiales showed statistical enrichment in ED, KO, and T hybrids, while Paenibacillales was the characteristic of all hybrids when compared to TF hybrid. Pseudonocardiales was more prevalent in KO and T hybrid, whereas Staphylococcales dominated in TF hybrid. Additionally, specific genera like Weissella, Staphylococcus, and Enterococcus were significantly present only in TF hybrid. Additionally, W. soli exhibited the same pattern with highest abundance in TF hybrid. The genus Kosakonia with species K. cowanii displayed statistical significance in abundance in TF and C hybrids compared to other hybrids, with its highest prevalence observed in C hybrid. Furthermore, the genus Pseudomonas showed higher prevalence in ED and KO compared to other hybrids, and this pattern was consistent for P. oryzihabitans. In summary, the differential abundance analysis revealed distinct taxonomic differences among the hybrids, indicating significant variations in the representation of specific taxa and genera, thus providing valuable insights into the microbial composition and diversity in the studied populations.
Differential abundance analysis of significantly more abundant taxa among in the seeds of five sugar beet hybrids Tajfun (TF), Eduarda (ED), Koala (KO), Tibor (T), and Cercospora-resistant (C). Bonferroni method was used for p-adjustment, and values of p < 0.01 were considered to be statistically significant
Diversity of Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria of Sugar Beet Hybrids
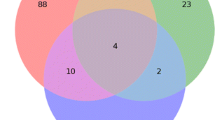
Each sugar beet hybrid’s rhizosphere contained species specific only to that particular hybrid, with one common species shared among all hybrids (Fig. 1). The isolates obtained from the roots of the hybrids did not exhibit any shared species. Notably, the root of the T hybrid displayed the highest species diversity, with 13 species unique to this hybrid. Comparing different hybrids, the roots of ED, KO, and C hybrids shared one or two species solely with the T hybrid. The ED hybrid showed the highest seed diversity, followed by C and KO hybrids with 12, 11, and 10 unique species, respectively. A total of 156 isolates were obtained from all hybrids, comprising 22, 31, and 63 unique species from the rhizosphere, roots, and seeds, respectively (Table S2).
The rhizosphere of sugar beet hybrids showed Lysinibacillus macroides to be a fundamental species, serving as the core member of this ecological niche. Among the Lysinibacillus genus, L. macroides, L. pakistanensis, and L. fusiformis were specifically isolated from the rhizosphere of the TF hybrid (Table 1; Table S3). The roots were particularly abundant in the genera Paenibacillus, Curtobacterium, Mycetocola, Knoellia, Neorhizobium, Microbacterium, Rhodococcus, Massilia, and Rathayibacter, with a total of 93 strains. The seeds displayed the highest diversity, especially in the case of ED (Table S3), KO (Table S3), and C (Table S3) hybrids. Bacillus subtilis was a unique species found in the seeds of all hybrids (Table 1). Additionally, prominent seed endophytes included species from the genera Bacillus, Lysinibacillus, Kocuria, Sanguibacter, Pantoea, Glutamicibacter, Pseudomonas, Erwinia, Providencia, and Pseudoclavibacter. Moreover, Curtobacterium pusillum was a unique species for C, ED, and KO seeds, while Bacillus sonorensis was isolated exclusively from the seeds of the T hybrid (Table 1; Table S3). Analyzing the diversity of culturable bacteria in the rhizosphere, roots, and seeds of each hybrid individually revealed minimal species overlap.
Evaluation of the Plant Growth–Promoting Properties In Vitro of Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria of Sugar Beet Hybrids
Examination of extracellular enzyme production and PGP characteristics (nitrogen fixation, phosphate solubilization, EPS production, swarming and swimming ability) was initially performed on 156 isolates (Table S4). The vast majority of isolates were able to grow on nitrogen-free media (86%), while 38% were phosphate-solubilizing bacteria. Swimming motility was recorded in a larger number of isolates (69%) in relation to the form of swarming (44%). Production of EPS was exhibited by 46 isolates (29%). Gelatinases were the most commonly secreted enzymes (44%), followed by amylases (30%), mannanases (17%), proteinases (17%), cellulases (16%), and pectinases (14%). Xylanase production was achieved by a significantly limited proportion of isolates, representing only 6% of the total. Isolates with a good and wider spectrum of PGP ability and secreted enzymes were selected for further testing on additional PGP characteristics (production of IAA, HCN, ACC deaminase, lipase and siderophores, drought and salinity tolerance).
Preliminary screening of sugar beet isolates for plant growth ability resulted in the selection of 32 bacterial strains with good PGP characteristics, while potential pathogens were classified as risk group 2 and excluded from the selection (Table 2).
All 32 isolates had the ability to grow on nitrogen-free media, whereas some isolates exhibited more intensive growth, especially Mixta theicola KO3-44, indicating better nitrogen fixation abilities. The most remarkable solubilization capacity observed in P. oryzihabitans KO3-19. Isolate B. subtilis KO3-18 had the highest PGP potential under in vitro conditions. Excellent producers of IAA were Providencia vermicola ED3-10 and M. theicola KO3-44. The best siderophore production was recorded for B. subtilis KO3-18, B. subtilis C3-59, B. subtilis C3-62, and Bacillus velezensis T2-23. ACC deaminase production was observed only for B. subtilis KO3-18, C. pusillum ED2-6, and Priestia aryabhattai T1-2. Significant production of EPS was recorded for M. theicola KO3-44 and Erwinia tasmaniensis ED3-79, while E. tasmaniensis ED3-79 and Glutamicibacter mishrai ED3-75 showed the greatest potential in HCN production.
Evaluation of the Exoenzymatic Activity In Vitro of Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria of Sugar Beet Hybrids
The most frequently secreted enzyme was gelatinase detected in 27 isolates with recorded EI values from 1.1 to 2.1, whereas the highest value was exhibited by Bacillus pacificus/paranthracis KO1-1. Twenty isolates showed amylolytic activity (1.1–2.5) with the highest activity observed in Paenibacillus taichungensis ED2-1. For cellulase production with EI range from EI 1.1 to 2.2, peak cellulase activity was exhibited by B. subtilis TF3-32. Among 14 isolates that were able to degrade the pectin (1.2–2.4), B. subtilis KO3-18 had the highest pectinolytic activity. Lipolytic activity was observed in the same number of isolates (1.1–1.8), and P. oryzihabitans KO3-19 stood out with the highest EI index value. Extracellular production of proteinases (1.1–2.0) and mannanases (1.1–1.8) were recorded for 13 isolates; and. G. mishrai ED3-75 and B. velezensis T2-23 exhibited the highest activity. The xylanolytic activity was recorded only for six isolates with EI values in the range 1.7–2.3, with the highest activity observed in B. subtilis TF3-6. Isolates B. velezensis T2-23, B. subtilis T3-4, B. velezensis ED2-2, and Bacillus halotolerans C3-16/2.1 were positive for all tested enzymatic activity, except for xylanase production. For Paenibacillus polymyxa C3-36, only proteolytic activity was missing. In contrast, isolates identified as L. macroides, M. theicola, P. vermicola, P. oryzihabitans, and E. tasmaniensis did not exhibit any exoenzymatic activity.
Assessment of Salt and Drought Tolerance In Vitro of Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria of Sugar Beet Hybrids
The majority of isolates exhibited successful growth up to an 8% NaCl concentration (Table S5). Out of 32 tested, 24 isolates could tolerate 8% NaCl, while half of them grew at 10% NaCl. B. halotolerans C3-16/2.1 exhibited the greatest tolerance to salinity, even at 10% NaCl. Almost all 32 isolates successfully tolerated drought in 5% and 10% PEG concentrations (Table S5). Eighteen isolates showed a tolerance range from moderate to complete resistance when exposed to a medium containing 20% PEG. Of the 32 isolates, two grew successfully at the maximum concentration of PEG with no visible reduction in growth compared to the control (B. subtilis C3-62 and B. subtilis TF2-1), while the growth of B. halotolerans C3-16/2.1, Bacillus amyloliquefaciens C3-19, Bacillus zhangzhouensis C3-50, B. velezensis ED2-2, B. velezensis T2-23, M. aloeverae T2-26, and B. sonorensis T3-5 was reduced to 50% of the control.
Evaluation of Antifungal Activity In Vitro of Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria of Sugar Beet Hybrids
According to the results obtained for PGP potential and exoenzymatic activity, 60 isolates were selected for initial screening against Fusarium sp. TS1, Fusarium equiseti TS2, Fusarium oxysporum TS3, and Cercospora beticola TS4 obtained from infected sugar beets. For further screening of antagonistic activity using the dual cultivation method, 32 isolates were selected from an original 60 that showed good inhibitory activity in the initial screening. Out of 32 isolates, 19 isolates showed moderate to exceptional antagonistic activity against all tested fungi (Table 3).
Among them, over 40% of growth inhibitions of all fungal strains were exhibited by B. amyloliquefaciens C3-19 (42–68%), P. polymyxa C3-36 (49–67%), B. velezensis ED2-2 (46–52%), B. subtilis ED3-89 (40–55%), B. velezensis T2-23 (49–56%), and B. subtilis TF3-6 (43–58%). The remaining isolates were active against two or three fungal pathogens, exhibiting moderate to weak activity. C. beticola TS4 was the most sensitive fungus, inhibited mostly by Bacillus species, recorded with an inhibition rate of over 50%. Among them, the most statistically significant antagonistic activity was achieved by B. amyloliquefaciens C3-19 and B. subtilis TF3-32 with 68% of inhibition of TS4. Apparently, F. equiseti TS2 was the most resistant pathogen to the tested isolates. Moderate to outstanding growth inhibition of this pathogen was detected for antagonistic isolates showing broad range antifungal activity with the highest inhibition rate exhibited by P. polymyxa C3-36 (67%). In addition, P. polymyxa C3-36 had the most statistically significant inhibition of F. equiseti TS2 and F. oxysporum TS3, while against Fusarium sp. TS1 that significance was achieved by B. velezensis T2-23.
In addition, the antagonistic activity of the selected 32 isolates tested against 15 different Fusarium strains in an initial screening test. Based on the results obtained, the 10 best strains were selected, and their antagonistic potential was subsequently quantified using the dual culture method (Table 4).
F. coffeatum IP32 was the most sensitive strain showing statistically significant susceptibility to all tested bacteria. Isolate B. velezensis T2-23 exhibited the highest statistically significant efficacy in suppressing the proliferation of a greater spectrum of Fusarium strains, followed by B. amyloliquefaciens C3-19, B. velezensis ED2-2, and B. subtilis TF2-1. Significantly, the strongest inhibition of all three isolates of F. graminearum was achieved equally with B. velezensis isolates (ED2-2 and T2-23). In general, the growth of even nine Fusarium spp. was successfully inhibited by all tested bacteria with an inhibition rate above 40%.
Discussion
Profiling Bacterial Diversity in Sugar Beet Hybrid Seeds by Next-Generation Sequencing Technology
In our study, a metabarcoding approach was used to characterize bacterial diversity in seeds of sugar beet hybrids. Notably, this study represents the pioneering investigation into the bacterial diversity of these hybrids using two distinct methodologies, with the exception of the rhizosphere of the Eduarda hybrid, which has been previously explored [51]. Bacterial community richness and alpha diversity analysis showed that the microbial communities of sugar beet hybrids were more diverse in ED, KO, and T seeds than in C and TF hybrids. PCoA analysis revealed that KO and TF are more distant from the remaining hybrids at the level of genus and ASV. The results of our study indicated that the genotype greatly influences the composition of the bacterial community inhabiting the sugar beet seeds as it was reported for Rhizoctonia susceptible and tolerant sugar beet cultivars [22]. The representation of phyla Actinobacteriota, Chloroflexi, Firmicutes, and Bacteroidetes in sugar beet seeds of hybrids tested in this study with Proteobacteria as the most represented phylum was also reported for sugar beet cultivar H7IM15 [21]. The similarity between cultivar H7IM15 and TF hybrid was reflected in the high presence of the phylum Firmicutes. In contrast to the H7IM15 cultivar, Acidobacteriota and Chloroflexi were detected in ED, KO, and T hybrids. In our study, genera Pantoea and Pseudomonas were detected in all five hybrids with high RA, with the exception of the low prevalence of Pseudomonas in TF. These genera have been highlighted as seed endophytes of different cultivars of sugar beet [21, 52]. Pantoea spp. include various life forms such as plant pathogens, plant growth stimulants, and strains used for commercial biocontrol of phytopathogens. Thus, Pantoea spp. is a good example of a group that has adapted to a specific niche [22]. Along with Pantoea spp. and Pseudomonas spp., each hybrid was represented by the dominance of one or two other genera, such as Kosakonia (C), Acinetobacter (T), and unknown genus from family Chroococcidiopsaceae (ED). Wolfgang et al. [22] reported that the abundance of Kosakonia, a plant growth–promoting genus, correlates with Rhizoctonia tolerance in sugar beet seeds. The observed correlation was in accordance with our result obtained on Cercospora-resistant hybrid suggesting that the genus Kosakonia plays an important role in sugar beet resistance to the major fungal pathogens of sugar beet. The role of Acinetobacter spp. in promoting plant growth, bioremediation and biodegradation was highlighted [53,54,55]. The TF hybrid was highly dominated by species of the genera Enterobacter, Kosakonia along with Pantoea. These taxa all belong to the Enterobacterales order, highlighting a taxonomic similarity among the dominant members of the bacterial community. The codominance of these genera suggests potential ecological interactions and niche specialization within this bacterial order. There are many PGP characteristics that Enterobacter spp. are known to possess, including the capability to fix nitrogen; solubilize phosphorus in soil; produce antibiotics; secrete siderophore products, chitinase, ACC deaminase, and hydrolytic enzymes in addition to exopolysaccharides; and increase soil porosity [56]. These properties are expressed by a variety of Enterobacter strains and contribute to plant growth and control of soil-borne plant diseases [57]. Other genera apart of Acinetobacter, Enterobacter, Enterococcus, and Kosakonia, detected in all hybrids, were Glutamicibacter and Marinilactibacillus, although their abundance depended on the hybrids. Representatives of these genera are associated with PGP traits [22, 56, 58,59,60,61], with most species classified as psychrotolerant/halotolerant bacteria [58,59,60,61]. In previous studies, Weissella spp. were reported as an endophyte of sugar beet seeds [21], and its high abundance in TF hybrids was the main characteristic that separated it from other analyzed hybrids. Hybrid-specific genera such as Apibacter (ED), Alloscardovia (KO), and Pleurocapsa (T) are mentioned as components of the marine, human, and animal microbiome [62,63,64,65], while their occurrence in the bacteriome of plants and in PGP capabilities is not yet known. The hybrid-specific genera Arthrobacter (T) and Klebsiella (TF) are known for their potential to promote plant growth [66, 67].
Diversity of Culturable Bacteria from Rhizosphere, Roots, and Seeds of Sugar Beet Hybrids
Within the field of biology, the concept of phylosymbiosis suggests that closely related species tend to display greater similarity in their microbiomes in comparison to distantly related species [68]. This aligns with research showing that plant genotypes, cultivars, developmental cycles, and other factors strongly influence endophytic microbial diversity and community structure [21]. Studying the microbiota of hybrid plants can reveal insights into the phylosymbiotic patterns within the plant family. The bacterial diversity present in the rhizosphere, roots, and seeds of five sugar beet hybrids, known as Eduarda, Koala, Tibor, Tajfun, and Cercospora-resistant, was thoroughly examined. Analysis of culturable bacteria showed that rhizospheres of all hybrids shared the common species L. macroides, while there were also species similarities between individual hybrids. The presence of Lysinibacillus strains in the rhizosphere has several benefits for the host plant, such as preventing cadmium uptake in plants [69] and attenuating the virulence of the pathogenic species Pectobacterium carotovorum by degrading the AHL signal and interrupting the pathogen's quorum sensing [70]. The rhizosphere is a region of rich, mostly soil derived, microbial diversity, influenced by plant roots through rhizodeposition of exudates, mucilage, and sloughed cells [12]. Considering that the composition of root exudates is influenced by the plant genome [4], as has been shown for the rhizosphere microbiome of Arabidopsis thaliana [12], the different genotypes of these five hybrids could be a determinant of bacterial diversity, as all hybrids had at least one hybrid-specific species in their rhizospheres. In the study by Zachow et al. [20] investigating differences in the rhizosphere microbiome of wild sugar beet (Beta vulgaris ssp. maritima) and modern varieties, it was found that greater bacterial diversity was associated with wild beet. Moreover, the same study revealed that they shared similarities in Pseudomonas and Stenotrophomonas species. The rhizosphere of the sugar beet was earlier showed to be colonized with B. subtilis, B. halotolerans, B. amyloliquefaciens, and Bacillus safensis [13], while the first three species were associated with the roots and seeds of investigated hybrids. Although Bacillus is ubiquitous in various environments, its abundance in cultivation can be easily overestimated due to copiotrophy and endospore formation. To the best of our knowledge, this is the first study to investigate the diversity of culturable bacteria in sugar beet seeds. Previous research also indicated a high diversity of bacterial taxa within the roots of sugar beet [71, 72]. Besides the genus Bacillus, species from genera Arthrobacter, Micrococcus, Microbacterium, Curtobacterium, Rhodococcus, and Staphylococcus were previously reported as root endophytes [71], which was in accordance with our study.
Sugar Beet–Associated Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria with Plant Growth–Promoting Properties
Plants harbor diverse bacterial communities whose interactions determine plant health and productivity. Determining whether bacteria promote or inhibit plant growth is an important step in the study to develop a PGPB-based inoculant. Moreover, indigenous bacterial strains are more suitable as growth promoters than new strains introduced from another location because plant-microbe interaction depends on different environmental parameters and indigenous PGPB are highly adapted to habitat [73]. Today, to meet the nutritional needs of plants and protect them from phytopathogens, large amounts of chemical fertilizers and pesticides are increasingly used [8]. However, it is well known that their excessive use leads to groundwater and soil pollution and a growing problem of resistance to pathogens. Since the use of chemicals in agriculture can never be completely eliminated, there is an increasing reliance on beneficial bacteria to alleviate this problem [74]. To investigate PGP abilities of culturable collection of bacteria isolated from five sugar beet hybrids, a panel of tests was performed. All 32 selected sugar beet isolates exhibited the ability to fix nitrogen to varying degrees. This is very important from the applicative standpoint, because the lack of available nitrogen in the soil on which the plant grows is one of the main factors that limit its development [75]. The majority of the soluble inorganic phosphorus used as chemical fertilizer is also immobilized quickly after application, rendering it unavailable to plants and resulting in its ineffectiveness [7]. As a result, one of the key characteristics of PGPB is the solubilization and mineralization of phosphorus by phosphate-solubilizing bacteria. In sugar beet isolates, the vast majority possessed this ability. P. vermicola ED3-10 and M. theicola KO3-44 exhibited excellent IAA production which is in accordance with the previous reports [76, 77]. IAA synthesized by bacteria increases root surface area and length, giving the plant better access to soil-derived nutrients. M. theicola has been reported with the significant improvement of root elongation in Zea mays related to IAA, in addition to many other stimulatory performances to this host [76]. In addition, bacterial IAA loosens plant cell walls, facilitating increased root exudation that provides additional nutrients to support the growth of rhizosphere bacteria [7]. Both P. vermicola and M. theicola showed to be great candidates for stimulation of plant growth as they both shared other PGP characteristics such as nitrogen fixation, phosphate solubilization, siderophore production, motility, and salinity tolerance up to 8% of NaCl. Also, M. theicola proved to be an excellent producer of EPS. Still, the production of hydrolytic enzymes was not recorded for these two isolates. For P. vermicola, a wide range of promoting characteristics for plant growth has been shown, of which siderophore production, phosphate solubilization, and nitrogen fixation [77,78,79] were also confirmed in our study. Furthermore, its biocontrol potential towards phytopathogenic soil-borne fungi and nematodes was reported [77]. Previous studies have demonstrated the efficacy of PGP bacteria in stimulating overall plant growth of sugar beet, including enzyme activity, sugar content, fiber and storage root development, and leaf yield, as well as leaf chlorophyll and nutrient content under drought stress [80]. Considering that iron deficiency leads to disruption of plant nutrition [73], the excellent siderophore production observed in this study, especially for the isolates of B. subtilis and B. velezensis, is a beneficial trait that could increase iron uptake by the host plant. These PGPB can also act as biocontrol agents through competition for iron as micronutrient, since their siderophores have a much greater affinity for iron than the fungal pathogens, making them unavailable [7].The ability of PGP’s bacteria to produce HCN, which is crucial for protecting crop plants from disease, is another important feature [48]. The known effects of HCN include disruption of the energy supply, inhibition of electron transport, and cell death [81]. The antifungal activity of B. halotolerans C3-16/2.1, B. zhangzhouensis C3-50, B. subtilis C3-72, and B. subtilis T3-4, apart from the likely existence of antifungal secondary metabolites, might be related to HCN synthesis, as well since these strains exhibited strong HCN production and suppression of fungal pathogens in this study. Suppression of Fusarium oxysporum f. sp. lycopersici spores has been previously reported for HCN produced by Pseudomonas fluorescens [82].
Sugar Beet–Associated Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria with Exoenzymatic Activity
In addition, PGPBs are known to protect host plants from phytopathogens via other mechanisms, including the production of hydrolytic enzymes [8]. Sugar beet isolates exhibited various enzymatic activities involving amylase, protease, lipase, xylanase, mannanase, cellulase, gelatinase, and pectinase production. Gelatinases which were often associated with biocontrol of plant pathogens such as insects and nematodes [83], were the most commonly produced enzymes in this study. Furthermore, the importance of gelatinases in disease protection of rice plants against spore-dispersing pathogenic fungi has been highlighted [83]. In another view regarding benefits to the plant, microorganisms with ability of producing cellulases, pectinases, xylanases, and amylases play an important role in the decomposition of organic matter and nutrient mineralization. In this study, B. halotolerans C3-16/2.1 and strains of B. velezensis (T2-23, ED2-2), B. subtilis T3-4, and P. polymyxa C3-36 showed the best exoenzymatic activity, while in some cases, the production of lipases, xylanases, and proteinases was absent. The enzymes such as cellulose, pectinase, xylanase, and protease are responsible for hydrolytic processes that allow endophytes to colonize plant tissues and establish a symbiotic relationship between endophytes and host plants [84]. The production of these enzymes was found in almost all Bacillus and Paenibacillus isolates from sugar beet, which was expected due to the fact that most were endophytes. In addition, the cellulolytic, lipolytic, and proteolytic activities of endophytes are known to protect host plants from pathogenic microorganisms by cell wall lysis of pathogens [7].
Sugar Beet–Associated Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria Tolerate Abiotic Stress
Salinity, flooding, drought, cold, and heavy metals are a few examples of the various abiotic stressors to which plants are exposed [85]. Crops exposed to these stressors can suffer significant production losses. Soil salinization is believed to have negative impacts on over 20% of all agricultural land worldwide. High soil salinity, primarily caused by the presence of NaCl, negatively impacts the growth and yield of most crops due to their limited salt tolerance. This is because the elevated salt levels interfere with the plants’ hormonal and nutritional processes, leading to reduced productivity and hindered growth [85]. Tolerance of sugar beet isolates to salt stress was evident as they grew successfully at elevated salt concentrations in the medium. B. halotolerans C3-16/2.1 proved to be an excellent candidate for stimulating plant growth in saline environments, showing the greatest tolerance to salinity, even at 10% NaCl. B. halotolerans was reported to protect plant growth and alleviate salt stress in wheat by modulating phytohormone synthesis and regulating osmotic balance, ion homeostasis, and gene expression [85]. Another important abiotic stress which leads to negative impact on crop production is agricultural drought as a period of soil moisture deficit resulting from a combination of rainfall deficiency and excessive evapotranspiration in a particular region [9]. Twenty-one isolates from sugar beets could be classified as drought-tolerant bacteria based on their ability to grow under high moisture deficit stress conditions. B. subtilis C3-62 and TF2-1 grew efficiently at the maximum concentration of PEG with no visible reduction in growth compared to the control and therefore could be used as stress-tolerant PGPR in soils facing drought. Bacteria can survive under stress conditions due to the production of EPS, which protects microorganisms from water stress by enhancing water retention and regulating the diffusion of organic carbon sources [49]. Since both B. subtilis strains were EPS-producing bacteria, this could be a reason for their high drought tolerance. EPS also helps microorganisms to irreversibly attach to and colonize the root, as it forms a network of fibrillar material that permanently connects the bacteria to the root surface [49], which is another important factor that favors these two strains from an application point of view. Regulation of plant hormone levels, especially auxin and ethylene, is perhaps the most important mechanism PGPB uses to promote plant development, especially in the presence of environmental conditions such as drought [86]. The amount of ACC in stressed plants is reduced by ACC deaminase–containing PGPR, which in turn reduces the amount of stress-induced ethylene production and subsequently plant damage. Since plants are frequently exposed to stress factors that trigger the synthesis of ethylene [49], the ACC deaminase–producing bacteria P. aryabhattai T1-2, B. subtilis KO3-18, and C. pusillum ED2-6 could be good candidates for the production of bioinoculants to control abiotic stress in plants.
Sugar Beet–Associated Cultured Endophytic (Seed and Roots) and Rhizospheric Bacteria as Potential Biocontrol Agents Based on Antifungal Potential In Vitro
The antagonistic activity of selected isolates was evaluated against several fungal pathogens known to cause Fusarium wilt [16], root rot [18], and Cercospora leaf spot disease [15] in sugar beet plants, as well as against other potential pathogens of Fusarium spp. [57, 87, 88]. Growth of fungal pathogens isolated from sugar beet was highly suppressed by P. polymyxa, B. subtilis, B. halotolerans, and B. velezensis strains. The great antagonistic potential of different Bacillus strains from the sugar beet rhizosphere against the most devastating pathogen C. beticola has been reported with the greatest activity achieved with Bacillus spp. strains (60-71% depending on the strain) [89]. Among 12 strains of B. subtilis tested, the best antifungal candidate against C. beticola was TF2-1. B. halotolerans, B. velezensis, B. subtilis, and P. polymyxa were the most successful in inhibiting growth of F. equiseti, where P. polymyxa C3-36 had the most statistically significant inhibition. Also, P. polymyxa had the statistically greatest activity against F. oxysporum. Inhibition of mycelial growth of Cercospora spp. and F. oxysporum [90, 91] has been reported for B. velezensis. The antifungal activity of B. velezensis is related to the production of secondary metabolites and hydrolytic enzymes [91], with bacillomycin D being recognized as the main compound potentially responsible for the broad antifungal activity [92]. In addition, isolate P. polymyxa C3-36 had excellent antifungal activity against all Fusarium strains tested, which already confirmed for various strains of P. polymyxa against F. oxysporum f. sp. lycopersici [93]. Both strains of B. halotolerans in our study were able to suppress the growth of each pathogen tested. Strains of B. halotolerans were already linked to the antifungal activity against various plant pathogens, including species of Fusarium, Rhizoctonia, and Botrytis genera [94, 95]. All tested strains of F. graminearum, which is known as a causative agent of head blight disease of cereal crops and sugar beet [16, 96], were highly sensitive to our B. subtilis, B. halotolerans, B. velezensis, and P. polymyxa strains. In general, the isolates of B. velezensis and P. polymyxa showed excellent antagonistic potential against all fungal pathogens tested.
Conclusion
The present study provides an interesting insight into the composition of seed-borne endophytes in five different sugar beet hybrids. The differences in the occurrence and abundance of certain genera in hybrid seeds as well as the presence of unique species clearly indicate that plant genotypes contribute to the structure of the seed-associated bacterial community of sugar beet. The prevalence of Pantoea and Pseudomonas species in the seeds of sugar beet hybrids was strikingly high, with the exception of the TF hybrid, which had a distinct profile characterized by a considerable presence of Weissella sp. Furthermore, the abundance of the Kosakonia genus in the Cercospora-resistant hybrid potentially contributes significantly to improved sugar beet resistance, which is consistent with its occurrence in other fungus-resistant sugar beet hybrids. This is also underlined by the fact that isolates with robust antifungal properties were predominantly derived from this particular hybrid. Furthermore, the hybrid genetic influence on the composition of the cultured rhizosphere and root endophyte community is visible through the diversity of isolates obtained isolates from different hybrids. The present study shows that the seed and root endophytes, as well as rhizospheric bacterial strains exhibit a broad spectrum of activities and thus offer a high potential for the development of plant probiotics and biological control agents. Based on their investigated activities, the obtained endophytes can be described as drought and halotolerant, phosphate solubilizing bacteria as well as bioproducers of siderophores, exopolysaccharides, indole-3-acetic acid, gelatinases, and amylases. Further studies will focus on testing the compatibility of the best candidates, forming viable “bottom-up” consortia, and monitoring their effects on sugar beet development and growth and pathogen resistance under controlled conditions and in the field.
Data Availability
The data presented in this study are openly available as BioProject ID: PRJNA954324 in the NCBI repository (https://www.ncbi.nlm.nih.gov/bioproject/954324).
References
Vandenkoornhuyse P, Quaiser A, Duhamel M et al (2015) The importance of the microbiome of the plant holobiont. New Phytol 206:1196–1206. https://doi.org/10.1111/NPH.13312
Berg G, Rybakova D, Grube M, Köberl M (2016) The plant microbiome explored: implications for experimental botany. J Exp Bot 67:995–1002. https://doi.org/10.1093/JXB/ERV466
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: ecology, functions, and emerging trends in microbial application. J Adv Res 19:29–37. https://doi.org/10.1016/j.jare.2019.03.004
Gupta R, Anand G, Gaur R, Yadav D (2021) Plant–microbiome interactions for sustainable agriculture: a review. Physiol Mol Biol Plants 27:165–179. https://doi.org/10.1007/S12298-021-00927-1/FIGURES/4
Santoyo G (2022) How plants recruit their microbiome? New insights into beneficial interactions. J Adv Res 40:45–58. https://doi.org/10.1016/J.JARE.2021.11.020
Berg G, Erlacher A, Grube M (2015) The edible plant microbiome: importance and health issues. Princ Plant-Microbe Interact Microbes Sustain Agric 419–426. https://doi.org/10.1007/978-3-319-08575-3_44/COVER
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica (Cairo) 2012:1–15. https://doi.org/10.6064/2012/963401
Dimkić I, Janakiev T, Petrović M et al (2022) Plant-associated Bacillus and Pseudomonas antimicrobial activities in plant disease suppression via biological control mechanisms - a review. Physiol Mol Plant Pathol 117:101754. https://doi.org/10.1016/J.PMPP.2021.101754
Mimić G, Živaljević B, Blagojević D et al (2022) Quantifying the effects of drought using the crop moisture stress as an indicator of maize and sunflower yield reduction in Serbia. Atmos 13(13):1880. https://doi.org/10.3390/ATMOS13111880
Yang J, Kloepper JW, Ryu CM (2009) Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 14:1–4. https://doi.org/10.1016/J.TPLANTS.2008.10.004
Sessitsch A, Brader G, Pfaffenbichler N et al (2018) The contribution of plant microbiota to economy growth. Microb Biotechnol 11:801. https://doi.org/10.1111/1751-7915.13290
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14:1–10. https://doi.org/10.1186/GB-2013-14-6-209/FIGURES/1
Farhaoui A, Adadi A, Tahiri A et al (2022) Biocontrol potential of plant growth-promoting rhizobacteria (PGPR) against Sclerotiorum rolfsii diseases on sugar beet (Beta vulgaris L.). Physiol Mol. Plant Pathol 119:101829. https://doi.org/10.1016/j.pmpp.2022.101829
Tsurumaru H, Okubo T, Okazaki K et al (2015) Metagenomic analysis of the bacterial community associated with the taproot of sugar beet. Microbes Environ 30:63–69. https://doi.org/10.1264/JSME2.ME14109
Srivastava SN (2004) Management of sugar beet diseases. Fruit Veg Dis 307–355. https://doi.org/10.1007/0-306-48575-3_9
Hanson LE, Hill AL (2004) Fusarium species causing Fusarium yellows of sugarbeet. J Sugar Beet Res 41:163–178
Haddoudi I, Cabrefiga J, Mora I et al (2021) Biological control of Fusarium wilt caused by Fusarium equiseti in Vicia faba with broad spectrum antifungal plant-associated Bacillus spp. Biol Control 160:104671. https://doi.org/10.1016/J.BIOCONTROL.2021.104671
Cao S, Yang N, Zhao C et al (2018) Diversity of Fusarium species associated with root rot of sugar beet in China. J Gen Plant Pathol 84:321–329. https://doi.org/10.1007/S10327-018-0792-5/TABLES/2
Smirnova I, Sadanov A (2019) Application of agriculturally important microorganisms for biocontrol of root rot infection of sugar beet. Arch Phytopathol Plant Prot 52:698–713
Zachow C, Müller H, Tilcher R, Berg G (2014) Differences between the rhizosphere microbiome of Beta vulgaris ssp. maritima-ancestor of all beet crops-and modern sugar beets. Front Microbiol 5:415. https://doi.org/10.3389/FMICB.2014.00415/ABSTRACT
Li M, Yang F, Wu X et al (2020) Effects of continuous cropping of sugar beet (Beta vulgaris L.) on its endophytic and soil bacterial community by high-throughput sequencing. Ann Microbiol 70:1–12. https://doi.org/10.1186/S13213-020-01583-8/FIGURES/6
Wolfgang A, Zachow C, Müller H et al (2020) Understanding the impact of cultivar, seed origin, and substrate on bacterial diversity of the sugar beet rhizosphere and suppression of soil-borne pathogens. Front Plant Sci 11:1450. https://doi.org/10.3389/FPLS.2020.560869/BIBTEX
Neamatollahi E, Bannayan M, Jahansuz MR et al (2012) Agro-ecological zoning for wheat (Triticum aestivum), sugar beet (Beta vulgaris) and corn (Zea mays) on the Mashhad plain, Khorasan Razavi province. Egypt J Remote Sens Sp Sci 15:99–112. https://doi.org/10.1016/J.EJRS.2012.05.002
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1):e1–e1. https://doi.org/10.1093/nar/gks808
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 137(13):581–583. https://doi.org/10.1038/nmeth.3869
Murali A, Bhargava A, Wright ES (2018) IDTAXA: A novel approach for accurate taxonomic classification of microbiome sequences. Microbiome 6:1–14. https://doi.org/10.1186/S40168-018-0521-5/FIGURES/6
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/JOURNAL.PONE.0061217
Pavoine S, Dufour AB, Chessel D (2004) From dissimilarities among species to dissimilarities among communities: a double principal coordinate analysis. J Theor Biol 228:523–537. https://doi.org/10.1016/J.JTBI.2004.02.014
Anderson MJ (2006) Distance-based tests for homogeneity of multivariate dispersions. Biometrics 62:245–253. https://doi.org/10.1111/J.1541-0420.2005.00440.X
McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: A comment on distance-based redundancy analysis. Ecology 82:290. https://doi.org/10.2307/2680104
Yang (2020) yiluheihei/microbiomeMarker: microbiomeMarker 0.0.1. https://doi.org/10.5281/ZENODO.3749415
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21. https://doi.org/10.1186/S13059-014-0550-8/FIGURES/9
Baldani JI, Reis VM, Videira SS et al (2014) The art of isolating nitrogen-fixing bacteria from non-leguminous plants using N-free semi-solid media: a practical guide for microbiologists. Plant Soil 384:413–431. https://doi.org/10.1007/S11104-014-2186-6/FIGURES/4
Woodman ME (2008) Direct PCR of intact bacteria (colony PCR). Curr Protoc Microbiol 9:A.3D.1-A.3D.6. https://doi.org/10.1002/9780471729259.MCA03DS9
Penrose DM, Glick BR (2003) Methods for isolating and characterizing ACC deaminase-containing plant growth-promoting rhizobacteria. Physiol Plant 118:10–15. https://doi.org/10.1034/J.1399-3054.2003.00086.X
Nautiyal CS (1999) An efficient microbiological growth medium for screening phosphate solubilizing microorganisms. FEMS Microbiol Lett 170:265–270. https://doi.org/10.1111/J.1574-6968.1999.TB13383.X
Gordon SA, Weber RP (1951) Colorimetric estimation of indoleacetic acid. Plant Physiol 26:192. https://doi.org/10.1104/PP.26.1.192
Pérez-Miranda S, Cabirol N, George-Téllez R et al (2007) O-CAS, a fast and universal method for siderophore detection. J Microbiol Methods 70:127–131. https://doi.org/10.1016/J.MIMET.2007.03.023
Gupta S, Pandey S (2019) ACC deaminase producing bacteria with multifarious plant growth promoting traits alleviates salinity stress in French bean (Phaseolus vulgaris) plants. Front Microbiol 10:1506. https://doi.org/10.3389/FMICB.2019.01506/BIBTEX
Zlosnik JEA, Hird TJ, Fraenkel MC et al (2008) Differential mucoid exopolysaccharide production by members of the Burkholderia cepacia complex. J Clin Microbiol 46:1470–1473. https://doi.org/10.1128/JCM.02273-07
Kim W, Surette MG (2003) Swarming populations of Salmonella represent a unique physiological state coupled to multiple mechanisms of antibiotic resistance. Biol Proced Online 5:189–196. https://doi.org/10.1251/BPO61/METRICS
Knežević MM, Stajković-Srbinović OS, Assel M et al (2021) The ability of a new strain of Bacillus pseudomycoides to improve the germination of alfalfa seeds in the presence of fungal infection or chromium. Rhizosphere 18:100353. https://doi.org/10.1016/J.RHISPH.2021.100353
Saroj P, Manasa P, Narasimhulu K (2018) Characterization of thermophilic fungi producing extracellular lignocellulolytic enzymes for lignocellulosic hydrolysis under solid-state fermentation. Bioresour Bioprocess 5:1–14. https://doi.org/10.1186/S40643-018-0216-6/FIGURES/7
Berić T, Urdaci MC, Stanković S, Knežević-Vukčević J (2009) RAPD analysis of genetic diversity and qualitative assessment of hydrolytic activities in a collection of Bacillus sp. isolate. Arch Biol Sci 61:645–652. https://doi.org/10.2298/ABS0904645B
Huber B, Riedel K, Hentzer M et al (2001) The cep quorum-sensing system of Burkholderia cepacia H111 controls biofilm formation and swarming motility. Microbiology 147:2517–2528. https://doi.org/10.1099/00221287-147-9-2517/CITE/REFWORKS
Ben SH, Cherif-Silini H, Bouket AC et al (2019a) Screening for Fusarium antagonistic bacteria from contrasting niches designated the endophyte Bacillus halotolerans as plant warden against Fusarium. Front Microbiol 10:3236. https://doi.org/10.3389/FMICB.2018.03236/BIBTEX
Medina P, Baresi L (2007) Rapid identification of gelatin and casein hydrolysis using TCA. J Microbiol Methods 69:391–393. https://doi.org/10.1016/J.MIMET.2007.01.005
Slama H, Ben TMA, Bouket AC et al (2019b) Screening of the high-rhizosphere competent Limoniastrum monopetalum’ culturable endophyte microbiota allows the recovery of multifaceted and versatile biocontrol agents. Microorg 7(7):249. https://doi.org/10.3390/MICROORGANISMS7080249
Ali SZ, Sandhya V, Rao LV (2014) Isolation and characterization of drought-tolerant ACC deaminase and exopolysaccharide-producing fluorescent Pseudomonas sp. Ann Microbiol 64:493–502. https://doi.org/10.1007/S13213-013-0680-3/FIGURES/68/FIGURES/9
Dimkić I, Berić T, Stević T et al (2015) Additive and synergistic effects of Bacillus spp. isolates and essential oils on the control of phytopathogenic and saprophytic fungi from medicinal plants and marigold seeds. Biol Control 87:6–13. https://doi.org/10.1016/J.BIOCONTROL.2015.04.011
Krstić Tomić T, Atanasković I, Nikolić I et al (2023) Culture-dependent and metabarcoding characterization of the sugar beet (Beta vulgaris L.) microbiome for high-yield isolation of bacteria with plant growth-promoting traits. Microorganisms 11:1538. https://doi.org/10.3390/MICROORGANISMS11061538/S1
Bertoldo G, Della Lucia MC, Squartini A et al (2021) Endophytic microbiome responses to sulfur availability in Beta vulgaris (L.). Int J Mol Sci 22(22):7184. https://doi.org/10.3390/IJMS22137184
Rojas-Tapias DF, Bonilla R, Dussán J (2014) Effect of inoculation and co-inoculation of Acinetobacter sp. RG30 and Pseudomonas putida GN04 on growth, fitness, and copper accumulation of maize (Zea mays). Water Air Soil Pollut 225:1–13. https://doi.org/10.1007/S11270-014-2232-2/FIGURES/4
Silambarasan S, Vangnai AS (2016) Biodegradation of 4-nitroaniline by plant-growth promoting Acinetobacter sp. AVLB2 and toxicological analysis of its biodegradation metabolites. J Hazard Mater 302:426–436. https://doi.org/10.1016/J.JHAZMAT.2015.10.010
Abbas S, Javed MT, Shahid M et al (2020) Acinetobacter sp. SG-5 inoculation alleviates cadmium toxicity in differentially Cd tolerant maize cultivars as deciphered by improved physio-biochemical attributes, antioxidants and nutrient physiology. Plant Physiol Biochem 155:815–827. https://doi.org/10.1016/j.plaphy.2020.08.024
Jha CK, Aeron A, Patel BV et al (2011) Enterobacter: Role in plant growth promotion. Bact Agrobiol Plant Growth Responses 159–182. https://doi.org/10.1007/978-3-642-20332-9_8
Silva SGA, Costa MM, Cardoso AMS et al (2023) Fusarium falciforme and Fusarium suttonianum cause root rot of melon in Brazil. Plant Pathol. https://doi.org/10.1111/PPA.13701
Panwar M, Tewari R, Nayyar H (2016) Native halo-tolerant plant growth promoting rhizobacteria Enterococcus and Pantoea sp. improve seed yield of mungbean (Vigna radiata L.) under soil salinity by reducing sodium uptake and stress injury. Physiol Mol Biol Plants 22:445–459. https://doi.org/10.1007/S12298-016-0376-9/FIGURES/2
Borker SS, Thakur A, Kumar S et al (2021) Comparative genomics and physiological investigation supported safety, cold adaptation, efficient hydrolytic and plant growth-promoting potential of psychrotrophic Glutamicibacter arilaitensis LJH19, isolated from night-soil compost. BMC Genomics 22:1–17. https://doi.org/10.1186/S12864-021-07632-Z/FIGURES/5
Huang XX, Xu L, Shang J, Sun JQ (2022) Marinilactibacillus kalidii sp. nov., an indole acetic acid-producing endophyte isolated from a shoot of halophyte Kalidium cuspidatum. Curr Microbiol 79:1–7. https://doi.org/10.1007/S00284-022-02894-6/TABLES/2
Patel P, Gajjar H, Joshi B et al (2022) Inoculation of salt-tolerant Acinetobacter sp (RSC9) improves the sugarcane (Saccharum sp. hybrids) growth under salinity stress condition. Sugar Tech 24:494–501. https://doi.org/10.1007/S12355-021-01043-W/TABLES/4
Waterbury JB, Stanier RY (1978) Patterns of growth and development in pleurocapsalean cyanobacteria. Microbiol Rev 42:2. https://doi.org/10.1128/MR.42.1.2-44.1978
Killer J, Ročková Š, Vlková E et al (2013) Alloscardovia macacae sp. nov., isolated from the milk of a macaque (Macaca mulatta), emended description of the genus Alloscardovia and proposal of Alloscardovia criceti comb. nov. Int J Syst Evol Microbiol 63:4439–4446. https://doi.org/10.1099/IJS.0.051326-0/CITE/REFWORKS
Kwong WK, Steele MI, Moran NA (2018) Genome sequences of Apibacter spp., gut symbionts of Asian honey bees. Genome Biol Evol 10:1174–1179. https://doi.org/10.1093/GBE/EVY076
Mattarelli P, Biavati B (2015) Alloscardovia. Bergey’s Man Syst Archaea Bact 1–3. https://doi.org/10.1002/9781118960608.GBM00018
Xu X, Xu M, Zhao Q et al (2018) Complete genome sequence of Cd(II)-resistant Arthrobacter sp. PGP41, a plant growth-promoting bacterium with potential in microbe-assisted phytoremediation. Curr Microbiol 75:1231–1239. https://doi.org/10.1007/S00284-018-1515-Z/TABLES/4
Sharma S, Gang S, Schumacher J et al (2021) Genomic appraisal of Klebsiella PGPB isolated from soil to enhance the growth of barley. Genes Genomics 43:869–883. https://doi.org/10.1007/S13258-021-01099-8/FIGURES/7
Lim SJ, Bordenstein SR (2020) An introduction to phylosymbiosis. Proc R Soc B 287(1922):20192900. https://doi.org/10.1098/rspb.2019.2900
Zhu L, Guo J, Sun Y et al (2021) Acetic acid-producing endophyte Lysinibacillus fusiformis orchestrates jasmonic acid signaling and contributes to repression of cadmium uptake in tomato plants. Front Plant Sci 12:1041. https://doi.org/10.3389/FPLS.2021.670216/BIBTEX
Stephens K, Bentley WE (2020) Synthetic biology for manipulating quorum sensing in microbial consortia. Trends Microbiol 28:633–643. https://doi.org/10.1016/J.TIM.2020.03.009
Lilley AK, Fry JC, Bailey MJ, Day MJ (1996) Comparison of aerobic heterotrophic taxa isolated from four root domains of mature sugar beet (Beta vulgaris). FEMS Microbiol Ecol 21:231–242. https://doi.org/10.1111/J.1574-6941.1996.TB00350.X
Okazaki K, Iino T, Kuroda Y et al (2014) An assessment of the diversity of culturable bacteria from main root of sugar beet. Microbes Environ 29:220–223. https://doi.org/10.1264/JSME2.ME13182
Natsagdorj O (2019) Biochemical characteristics and plant tissue localization of plant growth-promoting bacteria isolated from sugar beet (Beta vulgaris L.) Dissertation,. Iwate University
Kumar S (2012) Biopesticides: a need for food and environmental safety. J Biofertil Biopestic 3:1–3. https://doi.org/10.4172/2155-6202.1000e107
Galloway JN, Schlesinger WH, Levy H et al (1995) Nitrogen fixation: anthropogenic enhancement-environmental response. Global Biogeochem Cycles 9:235–252. https://doi.org/10.1029/95GB00158
Hagaggi NSA, Mohamed AAA (2020) Enhancement of Zea mays (L.) growth performance using indole acetic acid producing endophyte Mixta theicola isolated from Solenostemma argel (Hayne). South African J Bot 134:64–71. https://doi.org/10.1016/J.SAJB.2020.02.034
Panpatte D, Deepak P, Harsh S et al (2020) Providencia vermicola AAU PR1-a new bioinoculant for agriculture with multiple utility. Ind J Pure App Biosci 8:185–194. https://doi.org/10.18782/2582-2845.8160
Hussain K, Hameed S, Shahid M et al (2015) First report of Providencia vermicola strains characterized for enhanced rapeseed growth attributing parameters. Int J Agric Biol 17:1110–1116. https://doi.org/10.17957/IJAB/15.0033
Cavalcante Da Silva MJ, Ferreira S, Junior P et al (2020) IAA production of indigenous isolate of plant growth promoting rhizobacteria in the presence of tryptophan. AJCS 14:1835–2707. https://doi.org/10.21475/ajcs.20.14.03.p2239
Karagöz H, Çakmakçi R, Hosseinpour A, Kodaz S (2018) Alleviation of water stress and promotion of the growth of sugar beet (Beta vulgaris L.) plants by multi-traits rhizobacteria. Appl Ecol. Environ Res 16:6801–6813. https://doi.org/10.15666/AEER/1605_68016813
Ramyabharathi SA, Raguchander T (2014) Mode of action of Bacillus subtilis EPCO16 against tomato fusarium wilt. Biochem Cell Arch 14:47–50
Mwangi FM, Rüdiger H, Sikora RA, Mutitu E (2002) Does HCN from Pseudomonas fluorescens isolate T58 contribute in biocontrol of Fusarium oxysporum f. sp lycopersici?
Shimoi S, Inoue K, Kitagawa H et al (2010) Biological control for rice blast disease by employing detachment action with gelatinolytic bacteria. Biol Control 55:85–91. https://doi.org/10.1016/J.BIOCONTROL.2010.07.008
Hassan SED (2017) Plant growth-promoting activities for bacterial and fungal endophytes isolated from medicinal plant of Teucrium polium L. J Adv Res 8:687–695. https://doi.org/10.1016/J.JARE.2017.09.001
Wu X, Fan Y, Wang R et al (2022) Bacillus halotolerans KKD1 induces physiological, metabolic and molecular reprogramming in wheat under saline condition. Front Plant Sci 13. https://doi.org/10.3389/FPLS.2022.978066/FULL
Kim YC, Glick BR, Bashan Y, Ryu CM (2012) Enhancement of plant drought tolerance by microbes. Plant Responses to Drought Stress From Morphol to Mol Featur 9783642326530:383–413. https://doi.org/10.1007/978-3-642-32653-0_15/FIGURES/1
Schroers HJ, Baayen RP, Meffert JP et al (2004) Fusarium foetens, a new species pathogenic to begonia elatior hybrids (Begonia x hiemalis) and the sister taxon of the Fusarium oxysporum species complex. Mycologia 96:393–406. https://doi.org/10.1080/15572536.2005.11832984
Xu M, Zhang X, Yu J et al (2021) First report of Fusarium ipomoeae causing peanut leaf spot in China. Plant Dis 105:3754. https://doi.org/10.1094/PDIS-01-21-0226-PDN
Arzanlou M, Mousavi S, Bakhshi M et al (2016) Inhibitory effects of antagonistic bacteria inhabiting the rhizosphere of the sugarbeet plants, on Cercospora beticola Sacc., the causal agent of Cercospora leaf spot disease on sugarbeet. J Plant Prot Res 56:6–14. https://doi.org/10.1515/JPPR-2016-0002
Lee SY, Weon HY, Kim JJ, Han JH (2016) Biocontrol of leaf mustard powdery mildew caused by Erysiphe cruciferarum using Bacillus velezensis YP2. Korean J Pestic Sci 20:369–374. https://doi.org/10.7585/KJPS.2016.20.4.369
Torres M, Llamas I, Torres B et al (2020) Growth promotion on horticultural crops and antifungal activity of Bacillus velezensis XT1. Appl Soil Ecol 150:103453. https://doi.org/10.1016/J.APSOIL.2019.103453
Borriss R, Wu H, Gao X (2019) Secondary metabolites of the plant growth promoting model rhizobacterium Bacillus velezensis FZB42 are involved in direct suppression of plant pathogens and in stimulation of plant-induced systemic resistance. Second Metab Plant Growth Promot Rhizomicroorganisms Discov Appl 147–168. https://doi.org/10.1007/978-981-13-5862-3_8/FIGURES/2
Son SH, Khan Z, Kim SG, Kim YH (2009) Plant growth-promoting rhizobacteria, Paenibacillus polymyxa and Paenibacillus lentimorbus suppress disease complex caused by root-knot nematode and fusarium wilt fungus. J Appl Microbiol 107:524–532. https://doi.org/10.1111/J.1365-2672.2009.04238.X
Sagredo-Beltrán J, De La Cruz-Rodríguez Y, Alvarado-Rodríguez M et al (2018) Genome sequence of Bacillus halotolerans strain MS50-18A with antifungal activity against phytopathogens, isolated from Saline soil in San Luís Potosí, Mexico. Genome Announc 6. https://doi.org/10.1128/GENOMEA.00135-18
Tsalgatidou PC, Thomloudi EE, Baira E et al (2022) Integrated genomic and metabolomic analysis illuminates key secreted metabolites produced by the novel endophyte Bacillus halotolerans Cal.l.30 involved in diverse biological control activities. Microorg 399(10):399. https://doi.org/10.3390/MICROORGANISMS10020399
Hanson LE (2007) Fusarium yellowing of sugar beet caused by Fusarium graminearum from Minnesota and Wyoming. :686. https://doi.org/10.1094/PD-90-0686A
Funding
This work was supported by the Ministry of Education, Science and Technological developments of the Republic of Serbia, Collaborative Research Programme (CRP) — ICGEB Research Grants Programme, and by the FERTICO D.O.O. enterprise within the project “Development and application of new agricultural products based on bacterial endophytic and epiphytic inoculants, biofertilizers and biocontrol agents in sustainable agriculture” (Contracts Nos.: 451-03-47/2023-01/200178, CRP/SRB19-02 and 749/1-23.06.2021).
Author information
Authors and Affiliations
Contributions
MP, TJ, NU, TS, and SV — samples collection; SV and ID — project development; MP, TJ, MLG, NU, and TS — methodology and formal analysis; MP and ID — writing the original draft; MP, TJ, MLG, NU, and ID — review and editing; all authors have read and agreed to the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
A study was not required ethics approval.
Competing Interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Supplementary Information
ESM 1
(DOCX 107 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Petrović, M., Janakiev, T., Grbić, M.L. et al. Insights into Endophytic and Rhizospheric Bacteria of Five Sugar Beet Hybrids in Terms of Their Diversity, Plant-Growth Promoting, and Biocontrol Properties. Microb Ecol 87, 19 (2024). https://doi.org/10.1007/s00248-023-02329-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-023-02329-0