Abstract
Many insects are associated with endosymbionts that influence the feeding, reproduction, and distribution of their hosts. Although the small green mirid, Nesidiocoris tenuis (Reuter) (Hemiptera: Miridae), a zoophytophagous predator that feeds on plants as well as arthropods, is a globally important biological control agent, its microbiome has not been sufficiently studied. In the present study, we assessed the microbiome variation in 96 N. tenuis individuals from 14 locations throughout Japan, based on amplicon sequencing of the 16S ribosomal RNA gene. Nine major bacteria associated with N. tenuis were identified: Rickettsia, two strains of Wolbachia, Spiroplasma, Providencia, Serratia, Pseudochrobactrum, Lactococcus, and Stenotrophomonas. Additionally, a diagnostic PCR analysis for three typical insect reproductive manipulators, Rickettsia, Wolbachia, and Spiroplasma, was performed on a larger sample size (n = 360) of N. tenuis individuals; the most prevalent symbiont was Rickettsia (69.7%), followed by Wolbachia (39.2%) and Spiroplasma (6.1%). Although some symbionts were co-infected, their prevalence did not exhibit any specific tendency, such as a high frequency in specific infection combinations. The infection frequency of Rickettsia was significantly correlated with latitude and temperature, while that of Wolbachia and Spiroplasma was significantly correlated with host plants. The predominance of these bacteria and the absence of obligate symbionts suggested that the N. tenuis microbiome is typical for predatory arthropods rather than sap-feeding insects. Rickettsia and Wolbachia were vertically transmitted rather than horizontally transmitted from the prey. The functional validation of each symbiont would be warranted to develop N. tenuis as a biological control agent.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many insect species are closely associated with multiple endosymbionts that can affect the feeding, reproduction, and distribution of their hosts. Some of these associations have borne dependencies between the host and symbiont. For example, some herbivorous insects rely exclusively on nitrogen-poor substrates and require their symbionts for nutritional compensation [1], such as aphids with Buchnera [2], whiteflies with Portiera [3], and psyllids with Carsonella [4]. Furthermore, some blood-feeding insects have similar associations, such as tsetse flies with Wigglesworthia [5] and bedbugs with Wolbachia [6]. Such nutritional endosymbionts contribute significantly to the diversification of insect diets. Some endosymbionts have non-trophic effects on their hosts. Wolbachia, Rickettsia, Spiroplasma, and Cardinium induce reproductive phenotypes, such as cytoplasmic incompatibility (CI), male killing (MK), parthenogenesis induction (PI), and feminization (Fem), which are considered to be “selfish strategies” for the endosymbionts [7,8,9,10,11,12]. Some other endosymbionts are known to contribute to improving host fitness by increasing reproduction and development [13], conferring tolerance to thermal stress [14], and conferring resistance to pathogens [15].
Symbiotic bacteria often co-infect an individual in the same host population and show considerable variation in their infection patterns [16, 17]. The main factors that shape such patterns and symbiont community structure include host species [18], host plant [19], and geography [17]. For example, Macrolophus (Hemiptera: Miridae) are known to harbor one strain of Wolbachia and two species of Rickettsia (relatives of Rickettsia bellii and Rickettsia limoniae). Macrolophus pygmaeus harbors all these symbionts, whereas Macrolophus melanotoma (syn. Macrolophus caliginosus) harbors only Wolbachia and R. limoniae [18, 20, 21]. Although M. pygmaeus populations are geographically separated, their microbiomes are homogeneous, whereas the microbiomes of M. melanotoma are diverse [18, 21]. To fully understand the ecology and evolution of such species, it is important to understand how this variation in the symbiotic microbiota is involved in host adaptation. This is especially important given the potential role that many predatory insects play as biological control agents.
The small green mirid, Nesidiocoris tenuis (Hemiptera: Miridae), is a cosmopolitan species commonly used in the control of agricultural pests [22, 23]. They are zoophytophagous, which allows them to survive by feeding not only on arthropods but also on plants, which can augment their biological control activities but can also cause damage to crops [23,24,25]. N. tenuis are often found on Sesamum indicum (sesame) and Cleome hassleriana (cleome) in warm regions of Japan [23, 25]. In N. tenuis, two genera of symbionts, Wolbachia and Rickettsia, have been detected in Israeli populations and commercially available strains [26, 27]. Of these, the infection frequency of Rickettsia was found to be high (93–100%) in Israeli populations [26], whereas the infection frequency of Wolbachia remains unknown. Caspi-Fluger et al. [26] suggested that Rickettsia plays a nutritional role in zoophytophagous N. tenuis due to its high prevalence and abundance in adults and localization in the gut.
The aim of the present study was to elucidate the population structure of N. tenuis in Japan in terms of microbiome composition. We revealed the diversity of the microbiome in N. tenuis by 16S rRNA amplicon sequencing and diagnostic PCR assay. We also investigated whether the infection frequencies of Wolbachia, Rickettsia, and Spiroplasma were correlated with geography, climate, host plant, and host sex. These results reveal the complex relationships between N. tenuis and its symbionts, which may potentially contribute to improve the use of this species as a biological control agent.
Materials and Methods
Insect Collection
In total, 360 wild-caught adults of N. tenuis were collected from Sesamum indicum (sesame) or Cleome hassleriana (cleome) from 15 farms in Japan between 2017 and 2021 (Table S1). All individuals were stored in 99.5% ethanol at − 80 °C until DNA extraction was performed.
DNA Extraction
The 360 DNA samples were extracted from the whole insect bodies using the Wizard® Genomic DNA Purification Kit (Promega Corporation, Madison, WI, USA) according to the manufacturer’s protocol. DNA was dissolved in 100 μL of Tris-EDTA (pH 8.0) and stored at − 30 °C until use.
Amplicon Sequencing
For the selected 96 samples (Table S1), hypervariable V3/V4 regions of the 16S rRNA gene were amplified using the KAPA HiFi HotStart ReadyMix (Kapa Biosystems Inc., Wilmington, MA, USA) with V3V4_F primer (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′) and V3V4_R primer (5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′). The reactions were initiated by denaturation at 95 °C for 3 min, followed by 25 cycles of 30 s at 95 °C, 30 s at 55 °C, 30 s at 72 °C, and a final extension step of 5 min at 72 °C. After purification of the PCR products using AMPure XP beads (Beckman Coulter Inc., Brea, CA, USA), eight cycles of a second PCR were performed to add barcode sequences to each product using the TG Nextera XT Index Kit v2 Set A (Illumina Inc., San Diego, CA, USA). All barcoded amplicons were pooled in equal concentrations and sequenced on the Illumina MiSeq platform using the MiSeq Reagent Nano Kit v3 (600 cycles) according to the manufacturer’s recommended protocol (https://icom.illumina.com/) to produce 300-bp paired-end reads.
Raw Sequencing Data Analysis
The Illumina sequence data were processed using QIIME2 ver. qiime2-2020.11 [28]. The Illumina reads were demultiplexed based on the barcode sequences using “qiime demux emp-paired.” Denoizing and clustering were performed to obtain representative sequences and the feature table using “qiime dada2 denoise-paired” command. Taxonomic assignment to the representative sequences was then performed using “qiime feature-classifier classify-blast.” Sequences not identified as bacteria and all features with an abundance of < 0.01% were filtered out for further analysis. Data visualization was performed using the “qiime metadata tabulate” command, and the “qiime taxa barplot” command was used to generate a taxonomic bar plot. Alpha- and beta-diversity analyses were performed using the “qiime diversity alpha-rarefaction” and “qiime diversity core-metrics-phylogenetic” commands.
Diagnostic PCR for Rickettsia, Wolbachia, and Spiroplasma
Diagnostic PCR for the insect reproductive manipulators Rickettsia, Wolbachia, and Spiroplasma was performed on 360 N. tenuis individuals from 15 farms in Japan (Table S1). PCR was performed using the Go Taq Green Master Mix (Promega) with 528-F (5′-ACTAATCTAGAGTGTAGTAGGGGATGATGG-3′) and 1044-R (5′-GTTTTCTTATAGTTCCTGGCATTACCC-3′) for Rickettsia [29], wsp81F (5′- TGGTCCAATAAGTGATGAAGAAAC-3′) and wsp691R (5′- AAAAATTAAACGCTACTCCA-3′) for Wolbachia [30], and Spiro_Nt_124F (5′-GACGGTACCTTACCAGAAAG-3′) and Spiro_Nt_409R (5′-TTCGTGCCTAAACGTCAGTG-3′) for Spiroplasma in N. tenuis. Reactions were initiated by denaturation at 95 °C for 3 min, followed by 35 cycles of 30 s at 95 °C, 30 s at 60 °C (for 528-F/1044-R) or 55 °C (for wsp81F/wsp691R) or 56 °C (for Spiro_Nt_124F/Spiro_Nt_409R), 60 s at 72 °C, and a final extension step of 10 min at 72 °C. DNA was detected by electrophoresis on a 2% agarose gel prestained with Midori Green Xtra (Nippon Genetics Co., Ltd., Tokyo, Japan) in Tris-acetate-EDTA buffer.
Molecular Phylogenetic Analysis
Partial 16S rRNA sequences of Wolbachia, Rickettsia, and Spiroplasma isolated through amplicon sequencing were used for phylogenetic analyses. The datasets were registered in DDBJ (accession numbers: LC769520–LC769523). Phylogenetic trees based on the nucleotide sequences were constructed using the maximum likelihood method in MEGA 7.0 [31]. Kimura’s two-parameter model, evaluated with the best-fit method, was applied for the calculation [32].
Statistical Analysis
The PCR-based presence or absence of each bacterium within a mirid individual was analyzed based on a generalized linear model (GLM) with a binomial distribution (with a logit link function). Latitude, longitude, annual mean temperature, host plant, and sex were analyzed as explanatory variables. Data with unidentified sex were excluded from the analysis. Based on the GLM, an analysis of variance (ANOVA) was performed to evaluate the effects of each explanatory variable. Geographical and climatic factors that showed significant correlations with the GLM analysis were plotted to explicitly evaluate differential infection frequency. In addition, graphical visualization and Fisher’s exact test for the presence/absence of each bacterium were performed for each host plant. Geographical data were obtained from the Geospatial Information Authority of Japan (https://maps.gsi.go.jp), and climatic data were obtained from the Automated Meteorological Data Acquisition System administered by the Japan Meteorological Agency (https://www.jma.go.jp). Co-infection of Rickettsia, Wolbachia, and Spiroplasma was analyzed using an association screening approach as previously described [33]. The envelope function from the boot package in R software was used to estimate the 95% confidence envelope for the distribution profile of the combination counts, simultaneously including all infection patterns. A global test based on the 95% confidence envelope was then performed. All of the above analyses were performed using R version 4.2.2 [34].
Vertical Transmission Analysis
An isofemale line, K11, co-infected with Wolbachia and Rickettsia, was established from a female collected from population no. 1 in 2022 (Table S1). In the laboratory, K11 was reared using Ephestia kuehniella eggs (purchased in a frozen state from Agrisect Inc., Ibaraki, Japan) as the food source and Crassula ovata leaves as the oviposition substrate. The founder female and one of her G2 generation offspring were subjected to DNA extraction and amplicon sequencing as described above. All breeding was performed at 25 ± 1 °C with a light:dark regime of 14:10 h.
Results
Microbiomes of N. tenuis Inferred from 16S rRNA Gene Amplicon Sequencing
For the 96 N. tenuis individuals, the microbiomes were analyzed by amplicon sequencing of the hypervariable V3/V4 region of 16S rRNA, and a total of 4,625,099 reads were clustered into 77 operational taxonomic units (OTUs). The nine major OTUs (> 25,000 total reads and > 2000 reads per observed sample) were Rickettsia sp., two strains of Wolbachia sp., Providencia sp., Serratia marcescens, Pseudochrobactrum sp., Lactococcus lactis, Stenotrophomonas sp., and Spiroplasma sp., in order of frequency (Fig. 1, Table S2). Assuming that a mirid individual has the bacterium when it represented more than 1% of the tags analyzed, 69 out of 96 individuals had Rickettsia (71.9%), 30 individuals had Wolbachia sp. A (wNtenA, 31.3%), 4 individuals had Wolbachia sp. B (wNtenB, 4.2%), 25 individuals had Providencia (26.0%), 26 individuals had Serratia marcescens (27.1%), 18 individuals had Pseudochrobactrum (18.8%), 10 individuals had Lactococcus lactis (10.4%), 9 individuals had Stenotrophomonas (9.4%), and 5 individuals had Spiroplasma (5.2%).
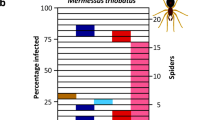
Proportion of bacterial sequences in 96 N. tenuis individuals collected from 14 regions in Japan. Sequences were obtained by amplicon sequencing of the hypervariable V3/V4 region of 16S rRNA. Assigned bacterial taxa are color coded as shown in the box on the right. Sequences with less than 25,000 total reads or 2000 reads per observed sample are categorized as “others.” The numbers at the bottom represent the geographic populations shown in Table S1
Infection Status of Rickettsia, Wolbachia, and Spiroplasma Inferred from Diagnostic PCR
We further investigated the prevalence of Rickettsia, Wolbachia, and Spiroplasma in 360 individuals from 15 populations of N. tenuis. We found that 293 of the 360 individuals (81.4%) were infected with at least one bacterium. Rickettsia was the most prevalent being detected in 251/360 individuals (69.7%), followed by Wolbachia in 142/360 individuals (39.4%) and Spiroplasma in 22/360 individuals (6.1%), and the frequency of infection varied between populations (Fig. 2a, 2b; Table S3). Some N. tenuis were co-infected with multiple bacteria; 104 individuals were doubly infected with Rickettsia and Wolbachia, 9 individuals were doubly infected with Rickettsia and Spiroplasma, 1 individual was doubly infected with Wolbachia and Spiroplasma, and 4 individuals were triply infected (Fig. 2b).
Infection frequencies of Rickettsia, Wolbachia, and Spiroplasma in each population of N. tenuis based on the diagnostic PCR assay. a Infection frequencies of Rickettsia (left panel), Wolbachia (center panel), and Spiroplasma (right panel). The frequencies of positive (black) and negative (white) individuals are shown with bar graphs. b Venn diagrams illustrating the co-infection status of Rickettsia, Wolbachia, and Spiroplasma. Each inner circle indicates the number of N. tenuis individuals infected with Rickettsia (red), Wolbachia (blue), and Spiroplasma (green). Overlapping circles indicate multiple infections. The outer circles represent the total number of N. tenuis individuals that were examined. Population numbers correspond to those in Table S1
Correlation of Rickettsia, Wolbachia, and Spiroplasma with Latitude, Temperature, and Host Plants
GLMs showed that the infection frequency of Rickettsia was significantly correlated with latitude and annual mean temperature (Table 1; Fig. 2a). Regression analyses showed a higher frequency of Rickettsia at lower latitude and higher temperature (Fig. 3a). Furthermore, GLMs indicated that the host plant significantly affected the infection frequency of Wolbachia and Spiroplasma (Table 1). Wolbachia infection frequency was significantly higher on C. hassleriana (cleome) than on S. indicum (sesame), while Spiroplasma was not found on C. hassleriana (Fig. 3b). The association screening approach showed that no significant association was detected between the co-infection status of Rickettsia, Wolbachia, or Spiroplasma from 360 individuals of N. tenuis, and this was also the case when the analysis was run by area (Table 2).
Relationship between infection frequencies of each symbiont (Rickettsia, Wolbachia, or Spiroplasma) in N. tenuis and each variable (latitude, temperature, or host plant). a A generalized linear model (GLM) with binomial error and logit-link function was plotted to estimate the effects of the correlation between Rickettsia and latitude or annual mean temperature for those significant differences detected (Table 1). The difference in deviance between the null hypothesis and the estimated model explained by each GLM is shown as ΔD, and the 95% confidence intervals are shaded in gray. b Infection frequencies of each endosymbiont in the host plants Sesamum indicum (sesame) and Cleome hassleriana (cleome). Error bars indicate 95% bootstrap percentiles (10,000 replicates). Asterisks indicate significant differences based on Fisher’s exact test (*P < 0.05; ***P < 0.0005)
Molecular Phylogenetic Analysis of Rickettsia, Wolbachia, and Spiroplasma
To infer the phylogenetic position of Rickettsia, Wolbachia, and Spiroplasma, nucleotide sequences (360, 360, and 384 bp, respectively) obtained through amplicon sequencing were subjected to maximum-likelihood tree reconstruction. In the Rickettsia phylogeny, the Rickettsia in the N. tenuis from Japanese populations was identical to that from the Israeli population [26], which was closely related to Rickettsia bellii (Fig. 4a). Of the two Wolbachia isolates in N. tenuis, one is the major isolate (wNtenA), which was detected in 30 out of 96 individuals, and the other is the minor isolate (wNtenB), which was detected in 4 out of 96 individuals. In the Wolbachia phylogeny, wNtenA and wNtenB both belonged to the Wolbachia supergroup B (Fig. 4b). wNtenA was closely related to the Wolbachia from the whitefly Bemisia tabaci, and wNtenB was closely related to those from Macrolophus pygmaeus, Cadra cautella, and Culex pipiens. In the Spiroplasma phylogeny, the Spiroplasma in N. tenuis fell into the Citri-Poulsonii clade, a large group consisting of S. citri, S. melliferum, S. kunkelli, S. penaei, S. insolitum, S. leucomae, S. phoeniceum, and S. poulsonii (Fig. 4c).
Phylogenetic trees based on the 16S rRNA gene sequences of Rickettsia, Wolbachia, and Spiroplasma. These trees were generated using the maximum likelihood method based on the Kimura 2-parameter model [32] with 1000 bootstrap replicates. Bootstrap values < 50% are not shown. The symbionts from N. tenuis are shown in red. The host organisms are given in parentheses, whereas the accession number is provided after each OTU. The scale bar indicates 0.02 substitutions per site. a Phylogenetic tree of Rickettsia based on 360 nucleotide sites. MK and PI represent Rickettsia isolates that cause male killing and parthenogenesis induction, respectively. The OTUs from insect symbionts are shaded gray. The outgroups are Wolbachia pipientis and Orientia tsutsugamushi. b Phylogenetic tree of Wolbachia based on 360 nucleotide sites. Wolbachia supergroups are depicted on the right side. The outgroup is Ehrlichia ruminantium. c Phylogenetic tree of Spiroplasma based on 384 nucleotide sites. MK and CI represent Spiroplasma isolates that cause male killing and cytoplasmic incompatibility, respectively. The Citri-Poulsonii clade is shaded green. The outgroup is Erysipelothrix larvae
Vertical Transmission of Rickettsia and Wolbachia
A total of 35,102 reads were obtained from the amplicon sequence analysis of the founder female of strain K11. Two major OTUs were classified as Rickettsia (17,396 reads) and Wolbachia (17,642 reads), respectively (Fig. S1). In G2, a total of 35,397 reads were clustered into Rickettsia (11,158 reads) and Wolbachia (24,097 reads) (Fig. S1). These nucleotide sequences of Rickettsia and Wolbachia were identical to those of Rickettsia and wNtenA obtained from N. tenuis in Fig. 4, respectively.
Discussion
This study demonstrated the high prevalence of Rickettsia and Wolbachia in Japanese N. tenuis populations (Fig. 1; Table S2), which is consistent with the results of a previous study on Israeli N. tenuis [26]. These symbionts induce reproductive phenotypes in insect hosts, and some of them can improve host fitness [9, 12, 13, 35]. Similarly, Spiroplasma manipulates host reproduction in some insects and can confer resistance to various parasites [10, 36, 37]. To the best of our knowledge, our study is the first to detect Spiroplasma in N. tenuis. In addition, Providencia, Serratia marcescens, Pseudochrobactrum, Stenotrophomonas, and Lactococcus lactis were found to be relatively abundant bacterial taxa in the N. tenuis population in Japan (Fig. 1; Table S2). S. marcescens and Providencia are commonly present in the environment [38, 39], and S. marcescens was also isolated from N. tenuis in a previous study [27]. Our study showed widespread infection of S. marcescens and Providencia among individuals but with low sequence reads per individual (Table S2), which may suggest opportunistic pathogenic properties of these bacteria [39, 40]. In the mosquito species Aedes aegypti, S. marcescens is present as a gut commensal bacterium that influences viral vector competence [41]. Stenotrophomonas, Pseudochrobactrum, and Lactococcus lactis have also been reported to be latent in the environment [42,43,44] and insect gut [45]. These results reveal a diversity of endosymbiotic microbes in natural populations of N. tenuis. The fact that none of the bacterial species found in this study were fixed in zoophytophagous N. tenuis suggests the absence of obligate symbionts in N. tenuis, a trait more typical for predatory arthropods rather than sap-feeding insects.
Rickettsia, Wolbachia, and Spiroplasma manipulate host reproduction in various insects [7,8,9,10, 12]. We found all possible combinations of these genera in N. tenuis individuals. Given that there was no correlation between the frequency of Rickettsia, Wolbachia, or Spiroplasma and the host sex (Table 1), it is unlikely that these symbionts induce MK, PI, or Fem in N. tenuis, which would otherwise result in a female-biased sex ratio and preferential presence of the symbiont in females. Coexisting symbionts may engage in interactions that are either negative or positive [46, 47]. Although no significant association with infection frequency was found (Table 2), further analysis of reproductive phenotypes or life history traits in various symbiont combinations is needed to understand the complex symbiotic system of N. tenuis populations and to propose the optimal biological control agent. Rickettsia, Wolbachia, or Spiroplasma have been detected in other carnivorous arthropods, such as mirids [18, 21, 26], coccinellids [48], and lacewings [7]. Feeding on other arthropods may have increased the chance of acquiring the symbionts common to prey species for N. tenuis [49].
The Rickettsia found in the present study is identical based on the partial sequence of the 16S rRNA gene to the Rickettsia sequence previously reported in N. tenuis [26], which is closely related to the R. bellii group. Previously, Rickettsia was detected in the gut lumen along the digestive tract of N. tenuis, while Wolbachia was detected in the surrounding epithelial cells [26]. A similar distribution of Rickettsia was elucidated in Macrolophus; both R. bellii and R. limoniae were found in the gut of M. pygmaeus and M. melanotoma [20, 21]. Although no correlation was found between the infection frequency of Rickettsia and the host plant, future studies should investigate the possible involvement of Rickettsia in the nutrient metabolism, including the zoophytophagous trait, of this species. It should be noted that no significant effects of Rickettsia and Wolbachia on the fitness traits of nymphal development and fecundity were detected in M. pygmaeus [18]. Although the Israeli populations harbored Rickettsia at a consistently high frequency (93–100%) [26], Japanese populations harbored Rickettsia at variable and relatively low frequencies (20.8–95.8%) (Fig. 2; Table S3). The fact that the high infection frequency of Rickettsia was associated with lower latitude and higher annual mean temperature (Fig. 3; Table 1) suggests the possibility that Rickettsia may provide positive effects to the host, such as heat tolerance, under high temperature [50] or negative effects under low temperature. Rickettsia infection is known to upregulate the expression of stress response genes in B. tabaci, which may underlie the mechanism of heat tolerance [50]. Furthermore, the supercooling point of M. pygmaeus exhibited a decrease upon the removal of its symbionts (two Rickettsia species and Wolbachia); however, it remains uncertain which bacterium influenced to the freezing susceptibility [51]. These possible effects of Rickettsia on hosts may explain the variable frequency of Rickettsia in Japanese populations of N. tenuis. Alternatively, Rickettsia may have no effect on host temperature sensitivity and our observation simply reflects the temperature sensitivity of Rickettsia itself [52].
Of the two supergroup B Wolbachia strains identified in this study, the major strain wNtenA was identical in terms of the partial 16S rRNA gene sequence to the Wolbachia strain found in B. tabaci. Despite the existence of a predator–prey relationship between N. tenuis and B. tabaci [23, 24], it is unlikely that the detected Wolbachia bacteria are exclusively derived from undigested B. tabaci remaining in the gut. This is because a large number of Wolbachia sequence reads were obtained using amplicon sequencing. Furthermore, vertical transmission was confirmed by breeding individuals under controlled laboratory conditions where they were not exposed to B. tabaci (Fig. S1). In B. tabaci, Wolbachia can be transmitted horizontally through plants and subsequently transmitted vertically to offspring [53]. The possibility that N. tenuis acquired Wolbachia from plants might be supported by the observed correlation between the frequency of Wolbachia and host plants.
The other strain, wNtenB, was identical with respect to the partial sequence of the 16S rRNA gene to the Wolbachia strain found in M. pygmaeus, which is known to induce strong CI [54]. Although strong CI is generally considered to cause widespread infection of the symbiont within the host population [55], wNtenB was rare (4 out of 96) in the N. tenuis populations in Japan. Furthermore, we did not observe co-infection of wNtenA and wNtenB, so whether they are in conflict or not remains unclear.
In the present study, we detected Spiroplsma from N. tenuis for the first time. Spiroplasma has also been detected in other hemipteran species, such as planthoppers, leafhoppers, and Orius predatory bugs [56,57,58]. In leafhoppers, Spiroplasma is transmitted horizontally between plants and insects [56]. Interestingly, we found that the infection frequency of Spiroplasma differed depending on the host plant (Fig. 3; Table 1), and the partial sequence of the 16S rRNA gene of Spiroplasma from N. tenuis was related to that of another mirid bug from Taiwan, Trigonotylus ruficornis (Fig. 4c). Future studies should aim to directly test whether Spiroplasma can be horizontally transmitted via plants.
The presence of symbionts may have important implications for the practical use of the predators as biological control agents. In particular, the high infection frequency of Rickettsia and Wolbachia may indicate their ability to manipulate host reproduction or their positive effects on host fitness. Significant differences in infection rates among host plants and geographic regions may affect the effectiveness of its use as a biological control agent, including its choice of insectary plants and its ability to propagate in the regions where it is used, both of which remain unexplored. Our findings highlight the potential importance of these symbionts, which may strongly affect the intrinsic rate of increase and confound the population dynamics of N. tenuis. We encourage future studies to determine the impact of each symbiont on this important biological control agent.
Data Availability
The datasets presented in this study can be found in online repositories. Repository names and accession numbers can be found at https://www.ddbj.nig.ac.jp/, DRR480549–DRR480644, LC769520–LC769523.
References
Skidmore IH, Hansen AK (2017) The evolutionary development of plant-feeding insects and their nutritional endosymbionts. Insect Sci 24:910–928. https://doi.org/10.1111/1744-7917.12463
Hansen AK, Moran NA (2014) The impact of microbial symbionts on host plant utilization by herbivorous insects. Mol Ecol 23:1473–1496. https://doi.org/10.1111/mec.12421
Sloan DB, Moran NA (2012) Endosymbiotic bacteria as a source of carotenoids in whiteflies. Biol Lett 8:986–989. https://doi.org/10.1098/rsbl.2012.0664
Spaulding AW, von Dohlen CD (2001) Psyllid endosymbionts exhibit patterns of co-speciation with hosts and destabilizing substitutions in ribosomal RNA. Insect Mol Biol 10:57–67. https://doi.org/10.1046/j.1365-2583.2001.00231.x
Michalkova V, Benoit JB, Weiss BL et al (2014) Vitamin B6 generated by obligate symbionts is critical for maintaining proline homeostasis and fecundity in tsetse flies. Appl Environ Microbiol 80:5844–5853. https://doi.org/10.1128/AEM.01150-14
Hosokawa T, Koga R, Kikuchi Y et al (2010) Wolbachia as a bacteriocyte-associated nutritional mutualist. Proc Natl Acad Sci 107:769–774. https://doi.org/10.1073/pnas.0911476107
Hayashi M, Watanabe M, Yukuhiro F et al (2016) A nightmare for males? A maternally transmitted male-killing bacterium and strong female bias in a green lacewing population. PLoS One 11:e0155794. https://doi.org/10.1371/journal.pone.0155794
Narita S, Kageyama D, Nomura M, Fukatsu T (2007) Unexpected mechanism of symbiont-induced reversal of insect sex: feminizing Wolbachia continuously acts on the butterfly Eurema hecabe during larval development. Appl Environ Microbiol 73:4332–4341. https://doi.org/10.1128/AEM.00145-07
Perlman SJ, Hunter MS, Zchori-Fein E (2006) The emerging diversity of Rickettsia. Proc R Soc B Biol Sci 273:2097–2106. https://doi.org/10.1098/rspb.2006.3541
Pollmann M, Moore LD, Krimmer E et al (2022) Highly transmissible cytoplasmic incompatibility by the extracellular insect symbiont Spiroplasma. iScience 25:104335. https://doi.org/10.1016/j.isci.2022.104335
Zchori-Fein E, Perlman SJ (2004) Distribution of the bacterial symbiont Cardinium in arthropods. Mol Ecol 13:2009–2016. https://doi.org/10.1111/j.1365-294X.2004.02203.x
Kaur R, Shropshire JD, Cross KL et al (2021) Living in the endosymbiotic world of Wolbachia: a centennial review. Cell Host Microbe 29:879–893. https://doi.org/10.1016/j.chom.2021.03.006
Himler AG, Adachi-Hagimori T, Bergen JE et al (2011) Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science 332:254–256. https://doi.org/10.1126/science.1199410
Dunbar HE, Wilson ACC, Ferguson NR, Moran NA (2007) Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol 5:e96. https://doi.org/10.1371/journal.pbio.0050096
Jaenike J, Unckless R, Cockburn SN et al (2010) Adaptation via symbiosis: recent spread of a Drosophila defensive symbiont. Science 329:212–215. https://doi.org/10.1126/science.1188235
Nishide Y, Sugimoto TN, Watanabe K et al (2022) Genetic variations and microbiome of the poultry red mite Dermanyssus gallinae. Front Microbiol 13:1031535. https://doi.org/10.3389/fmicb.2022.1031535
Toju H, Fukatsu T (2011) Diversity and infection prevalence of endosymbionts in natural populations of the chestnut weevil: relevance of local climate and host plants. Mol Ecol 20:853–868. https://doi.org/10.1111/j.1365-294X.2010.04980.x
Machtelinckx T, Van Leeuwen T, Van De Wiele T et al (2012) Microbial community of predatory bugs of the genus Macrolophus (Hemiptera: Miridae). BMC Microbiol 12:S9. https://doi.org/10.1186/1471-2180-12-S1-S9
Xu S, Jiang L, Qiao G, Chen J (2020) The bacterial flora associated with the polyphagous aphid Aphis gossypii Glover (Hemiptera: Aphididae) is strongly affected by host plants. Microb Ecol 79:971–984. https://doi.org/10.1007/s00248-019-01435-2
Dally M, Lalzar M, Belausov E et al (2020) Cellular localization of two Rickettsia symbionts in the digestive system and within the ovaries of the mirid bug, Macrolophous pygmaeus. Insects 11:530. https://doi.org/10.3390/insects11080530
Dally M, Izraeli Y, Belausov E et al (2023) Rickettsia association with two Macrolophus (Heteroptera: Miridae) species: a comparative study of phylogenies and within-host localization patterns. Front Microbiol 13:1107153. https://doi.org/10.3389/fmicb.2022.1107153
Calvo FJ, Lorente MJ, Stansly PA, Belda JE (2012) Preplant release of Nesidiocoris tenuis and supplementary tactics for control of Tuta absoluta and Bemisa tabaci in greenhouse tomato. Entomol Exp Appl 143:111–119. https://doi.org/10.1111/j.1570-7458.2012.01238.x
Yano E (2022) Biological control using zoophytophagous bugs in Japan. J Pest Sci 95:1473–1484. https://doi.org/10.1007/s10340-022-01561-w
Calvo J, Bolckmans K, Stansly PA, Urbaneja A (2009) Predation by Nesidiocoris tenuis on Bemisia tabaci and injury to tomato. BioControl 54:237–246. https://doi.org/10.1007/s10526-008-9164-y
Nakano R, Morita T, Okamoto Y et al (2021) Cleome hassleriana plants fully support the development and reproduction of Nesidiocoris tenuis. BioControl 66:407–418. https://doi.org/10.1007/s10526-021-10079-6
Caspi-Fluger A, Inbar M, Steinberg S et al (2014) Characterization of the symbiont Rickettsia in the mirid bug Nesidiocoris tenuis (Reuter) (Heteroptera: Miridae). Bull Entomol Res 104:681–688. https://doi.org/10.1017/S0007485314000492
Ferguson KB, Visser S, Dalíková M et al (2021) Jekyll or Hyde? The genome (and more) of Nesidiocoris tenuis, a zoophytophagous predatory bug that is both a biological control agent and a pest. Insect Mol Biol 30:188–209. https://doi.org/10.1111/imb.12688
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Chiel E, Zchori-Fein E, Inbar M et al (2009) Almost there: transmission routes of bacterial symbionts between trophic levels. PLoS One 4:e4767. https://doi.org/10.1371/journal.pone.0004767
Braig HR, Zhou W, Dobson SL, O’Neill SL (1998) Cloning and characterization of a gene encoding the major surface protein of the bacterial endosymbiont Wolbachia pipientis. J Bacteriol 180:2373–2378. https://doi.org/10.1128/JB.180.9.2373-2378.1998
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Vaumourin E, Vourc’h G, Telfer S et al (2014) To be or not to be associated: power study of four statistical modeling approaches to identify parasite associations in cross-sectional studies. Front Cell Infect Microbiol 4:62. https://doi.org/10.3389/fcimb.2014.00062
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria https://www.r-project.org/
Hagimori T, Abe Y, Date S, Miura K (2006) The first finding of a Rickettsia bacterium associated with parthenogenesis induction among insects. Curr Microbiol 52:97–101. https://doi.org/10.1007/s00284-005-0092-0
Arai H, Takamatsu T, Lin S-R et al (2023) Diverse molecular mechanisms underlying microbe-inducing male killing in the moth Homona magnanima. Appl Environ Microbiol 0:e02095–e02022. https://doi.org/10.1128/aem.02095-22
Ballinger MJ, Perlman SJ (2019) The defensive Spiroplasma. Curr Opin Insect Sci 32:36–41. https://doi.org/10.1016/j.cois.2018.10.004
Cristina ML, Sartini M, Spagnolo AM (2019) Serratia marcescens infections in neonatal intensive care units (NICUs). Int J Environ Res Public Health 16:610. https://doi.org/10.3390/ijerph16040610
O’Hara CM, Brenner FW, Miller JM (2000) Classification, identification, and clinical significance of Proteus, Providencia, and Morganella. Clin Microbiol Rev 13:534–546. https://doi.org/10.1128/cmr.13.4.534
Raymann K, Coon KL, Shaffer Z et al (2018) Pathogenicity of Serratia marcescens strains in honey bees. mBio 9:e01649–e01618. https://doi.org/10.1128/mBio.01649-18
Wu P, Sun P, Nie K et al (2019) A gut commensal bacterium promotes mosquito permissiveness to arboviruses. Cell Host Microbe 25:101–112.e5. https://doi.org/10.1016/j.chom.2018.11.004
Fakri M, Lani M, Chuah T-S et al (2018) In vitro antifungal potential of Lactococcus lactis isolated from agricultural soils in terengganu against anthracnose pathogen, Colletotrichum capsici. Malays Appl Biol 47:169–182
Ryan RP, Monchy S, Cardinale M et al (2009) The versatility and adaptation of bacteria from the genus Stenotrophomonas. Nat Rev Microbiol 7:514–525. https://doi.org/10.1038/nrmicro2163
Siddique K, Shahid M, Shahzad T et al (2021) Comparative efficacy of biogenic zinc oxide nanoparticles synthesized by Pseudochrobactrum sp. C5 and chemically synthesized zinc oxide nanoparticles for catalytic degradation of dyes and wastewater treatment. Environ Sci Pollut Res 28:28307–28318. https://doi.org/10.1007/s11356-021-12575-9
Ali HRK, Hemeda NF, Abdelaliem YF (2019) Symbiotic cellulolytic bacteria from the gut of the subterranean termite Psammotermes hypostoma Desneux and their role in cellulose digestion. AMB Express 9:111. https://doi.org/10.1186/s13568-019-0830-5
Vautrin E, Vavre F (2009) Interactions between vertically transmitted symbionts: cooperation or conflict? Trends Microbiol 17:95–99. https://doi.org/10.1016/j.tim.2008.12.002
White JA, Kelly SE, Perlman SJ, Hunter MS (2009) Cytoplasmic incompatibility in the parasitic wasp Encarsia inaron: disentangling the roles of Cardinium and Wolbachia symbionts. Heredity 102:483–489. https://doi.org/10.1038/hdy.2009.5
Werren JH, Hurst GD, Zhang W et al (1994) Rickettsial relative associated with male killing in the ladybird beetle (Adalia bipunctata). J Bacteriol 176:388–394. https://doi.org/10.1128/jb.176.2.388-394.1994
Le Clec’h W, Chevalier FD, Genty L et al (2013) Cannibalism and predation as paths for horizontal passage of Wolbachia between terrestrial isopods. PLoS One 8:e60232. https://doi.org/10.3389/fmicb.2022.1031535
Brumin M, Kontsedalov S, Ghanim M (2011) Rickettsia influences thermotolerance in the whitefly Bemisia tabaci B biotype. Insect Sci 18:57–66. https://doi.org/10.1111/j.1744-7917.2010.01396.x
Maes S, Machtelinckx T, Moens M et al (2012) The influence of acclimation, endosymbionts and diet on the supercooling capacity of the predatory bug Macrolophus pygmaeus. BioControl 57:643–651. https://doi.org/10.1007/s10526-012-9446-2
Galletti MFBM, Fujita A, Rosa RD et al (2016) Virulence genes of Rickettsia rickettsii are differentially modulated by either temperature upshift or blood-feeding in tick midgut and salivary glands. Parasit Vectors 9:331. https://doi.org/10.1186/s13071-016-1581-7
Li S-J, Ahmed MZ, Lv N et al (2017) Plant mediated horizontal transmission of Wolbachia between whiteflies. ISME J 11:1019–1028. https://doi.org/10.1038/ismej.2016.164
Machtelinckx T, Van Leeuwen T, Vanholme B et al (2009) Wolbachia induces strong cytoplasmic incompatibility in the predatory bug Macrolophus pygmaeus. Insect Mol Biol 18:373–381. https://doi.org/10.1111/j.1365-2583.2009.00877.x
Shropshire JD, Leigh B, Bordenstein SR (2020) Symbiont-mediated cytoplasmic incompatibility: what have we learned in 50 years? eLife 9:e61989. https://doi.org/10.7554/eLife.61989
Mello AFS, Wayadande AC, Yokomi RK, Fletcher J (2009) Transmission of different isolates of Spiroplasma citri to carrot and citrus by Circulifer tenellus (Hemiptera: Cicadellidae). J Econ Entomol 102:1417–1422. https://doi.org/10.1603/029.102.0403
Sanada-Morimura S, Matsumura M, Noda H (2013) Male killing caused by a Spiroplasma symbiont in the small brown planthopper, Laodelphax striatellus. J Hered 104:821–829. https://doi.org/10.1093/jhered/est052
Watanabe M, Yukuhiro F, Maeda T et al (2014) Novel strain of Spiroplasma found in flower bugs of the genus Orius (Hemiptera: Anthocoridae): transovarial transmission, coexistence with Wolbachia and varied population density. Microb Ecol 67:219–228. https://doi.org/10.1007/s00248-013-0335-8
Acknowledgements
We thank the researchers at the Research Center of Genetic Resources in NARO, Prefectural Agricultural Experiment Stations in Fukuoka and Okinawa, and local farmers for their help with collecting N. tenuis samples.
Funding
This work was supported by a Grant-in-Aid for Scientific Research (C) from the Japan Society for the Promotion of Science (19K06076) and Cabinet Office, Government of Japan, Moonshot Research and Development Program for Agriculture, Forestry, and Fisheries (funding agency: Bio-oriented Technology Research Advancement Institution) (no. JPJ009237).
Author information
Authors and Affiliations
Contributions
Tetsuya Adachi-Hagimori, Toma Minami, and Daisuke Kageyama designed the research. Tetsuya Adachi-Hagimori, Toma Minami, Yuta Owashi, and Ryohei Nakano collected the materials and Toma Minami conducted the PCR assay. Taisei Kikuchi and Akemi Yoshida contributed the NGS assay. Yuta Owashi, Toma Minami, Taisei Kikuchi, Daisuke Kageyama, and Tetsuya Adachi-Hagimori analyzed the data. Yuta Owashi and Daisuke Kageyama wrote the first draft of the manuscript. All authors critically reviewed the manuscript and approved the final submission.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Owashi, Y., Minami, T., Kikuchi, T. et al. Microbiome of Zoophytophagous Biological Control Agent Nesidiocoris tenuis. Microb Ecol 86, 2923–2933 (2023). https://doi.org/10.1007/s00248-023-02290-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02290-y