Abstract
Inflows from unregulated tributaries change the physical, chemical, and biotic conditions in receiving regulated rivers, impacting microbial community structure and metabolic function. Understanding how tributary inflows affect bacterial carbon production (BCP) is integral to understanding energy transfer in riverine ecosystems. To investigate the role of tributary inflows on bacterial community composition and BCP, a ~90th percentile natural flow event was sampled over 5 days along the Lachlan River and its tributaries within the Murray-Darling Basin of eastern Australia. Increased tributary inflows after rainfall corresponded with a significantly different and more diverse bacterial community in the regulated mainstem. The major contributor to this difference was an increase in relative abundance of bacterial groups with a potential metabolic preference for humic substances (Burkholderiaceae Polynucleobacter, Alcaligenaceae GKS98 freshwater group, Saccharimonadia) and a significant decrease in Spirosomaceae Pseudarcicella, known to metabolise algal exudates. Increases in orthophosphate and river discharge explained 31% of community change, suggesting a combination of resource delivery and microbial community coalescence as major drivers. BCP initially decreased significantly with tributary inflows, but the total load of carbon assimilated by bacteria increased by up to 20 times with flow due to increased water volume. The significant drivers of BCP were dissolved organic carbon, water temperature, and conductivity. Notably, BCP was not correlated with bacterial diversity or community composition. Tributary inflows were shown to alter mainstem bacterial community structure and metabolic function to take advantage of fresh terrestrial dissolved organic material, resulting in substantial changes to riverine carbon assimilation over small times scales.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
River regulation interrupts the longitudinal flow of biota and resources from river headwaters and the surrounding catchment, which can subsequently impact riverine microbial community structure and function [1,2,3]. Dams and weirs create lentic environments along river channels where thermal stratification can create conditions that favour algal blooms or anoxia and nutrient release from sediments in the hypolimnion, which fuels primary production downstream and shifts rivers into net autotrophy [4,5,6]. River regulation also reduces connectivity to the catchment by reducing the frequency and magnitude of average annual flows and floods downstream of the dam, as well as changing the seasonal distribution of flows through the decoupling of rainfall and flows [7,8,9]. These altered flow regimes diminish the amount of dissolved organic carbon (DOC) exported from the floodplain, with models predicting reductions in annual DOC export of up to two-thirds of natural levels under regulated scenarios [10, 11].
Tributaries of major rivers are often unregulated or less regulated than the river mainstem, so their flow regimes and flow-related resource pulses can help to restore degraded river mainstems to a more natural state [12,13,14]. Tributary inflows alter the physical, chemical, and biological conditions in river mainstems through temperature shifts and the import of fresh dissolved organic matter (DOM), nutrients, sediment, and biota from tributary headwaters and the greater catchment [12, 14,15,16].
The changed physical and chemical environment in regulated rivers during tributary inflows causes a corresponding shift in planktonic microbial community composition and metabolic function towards one more representative of surrounding floodplain habitats [3]. Bacterioplankton communities shift almost immediately during tributary inflow events due to mobilisation of bacteria from catchment soil, the hyporheic zone, and upstream aquatic microhabitats [3, 17, 18]. These disparate microbial communities can come together to form a new community with emergent properties distinct from its components in a process known as microbial community coalescence [19]. The altered bacterial community structure and increased availability of complex terrestrial DOM can drive increased pelagic bacterial carbon production (BCP), which captures DOC and nutrients into bacterial biomass making them biologically available to higher trophic levels via the microbial loop [20,21,22,23]. Community mixing effects are typically short lived (~1–2 weeks, depending on catchment complexity), and over the following days and weeks, processes such as environmental filtering, competition, and predation become the dominant structuring forces of riverine bacterioplankton communities [24, 25]. However, frequent tributary inflows may lead to altered bacterial communities in mainstem rivers over time, and even transient resource pulses can continue to subsidise the food web for months after microbial assimilation [3, 26].
Linking tributary inflow characteristics with their impact on mainstem bacterioplankton communities and subsequent changes in BCP and microbial carbon assimilation is integral to modelling overall energy transfer in riverine ecosystems [27]. However, it is not well understood how effective these inflows are at supporting instream BCP during the crucial initial stages of a flow-based resource pulse. This is particularly so in Australia’s highly regulated Murray-Darling Basin, where nearly 70% of mean annual inflow is captured through dams and irrigation pumping [28]. For example, an 18-month seasonal study of BCP on the Murray River recorded the highest planktonic BCP rates (> 60 μg C L−1 h−1) directly below an unregulated tributary during an inflow event, but relationships with tributary inflow characteristics could not be identified due to infrequent sampling [29]. In addition, responses to resource pulses are often studied only during extreme hydrological events (i.e. large-scale flooding, drought) with comparably extreme and transient impacts on river ecology [30,31,32]. Moderate flow pulses (~90–95th percentile flows) are far more frequent in these systems and hence may be more important to the structure and function of microbial communities than larger, more infrequent events.
The aim of this study was to examine the immediate impact of moderate-sized inflows from tributaries on heterotrophic BCP and bacterioplankton community structure in a major regulated river and investigate the possible mechanisms behind any changes observed. Water column DOC, nutrients, physico-chemical parameters, BCP, and bacterial DNA were sampled daily on the Lachlan River and its tributaries in central New South Wales Australia before and during a four-day rainfall event of ~60 mm. It was hypothesised that tributary inflows would alter environmental conditions and bacterial community composition in the river mainstem, which would increase BCP per litre, thus increasing the total load of carbon assimilated by riverine bacteria.
Methods
Study Sites
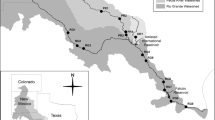
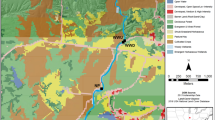
The Lachlan River is an inland floodplain river and major tributary of the Murray River in central New South Wales, Australia. This region experiences an average annual rainfall of 600 mm. Flows of the Lachlan River are regulated by Wyangala Dam (1217 GL, catchment size 8300 km2), with the river having a largely agricultural catchment, consisting mainly of grazing and dryland cropping [33]. Downstream of Wyangala Dam, the Lachlan River is fed by several unregulated (or only moderately regulated) tributaries. Of importance to this study are Boorowa River, entering the river 18 km downstream of the dam wall with a catchment size of 1500 km2, and Belubula River entering the river 102 km downstream of the dam wall with a catchment size of 1600 km2. The Belubula River is regulated by a small dam (Carcoar Dam 36 GL, catchment size 230 km2) ~130 km before its confluence with the Lachlan River but maintains an almost natural hydrograph at the confluence [34].
Samples were collected from six sites along the mid-reaches of the Lachlan River and two adjoining tributaries (Boorowa and Belubula Rivers) (Fig. 1). Four sites were spaced along the Lachlan River mainstem to capture instream conditions upstream and downstream of major tributary confluences. Two smaller tributaries, Crowther and Hovells Creek, were not sampled due to insufficient water depth at base flow (Crowther ~0.17 m3 s−1, Hovells ~0.06 m3 s−1).
Sample Collection and Analysis
Samples were collected as close to the same time of day as possible over 5 days, before and during a moderate natural flow event over the week of 23–27 October 2020. Sites were prioritised for sampling each morning, based on current hydrographs from local gauges. At each site, 10 L of water was collected from 20 to 30 cm below the surface. Because ecology at the microbial level is heterogenous at a small spatial scale [35], a composite sample was made up of six combined ~1.5 L subsamples and combined in an acid-rinsed (10% HCl) polyethylene bottle.
For DOC, nitrogen oxides (NOx), ammonia, and orthophosphate (RP) analyses, 200 mL of river water was filtered through pre-flushed 0.45 μm PTFE filters in triplicate, with filtered water then frozen until analysis. DOC was determined using a Shimadzu TOC-L analyser and NOx, ammonia, and RP were determined by flow injection analysis on a Lachat Quikchem 8500. Blanks, certified reference materials, spiked matrices, and analytical duplicates were used for quality control. Water column temperature, conductivity, dissolved oxygen, and pH were sampled with a Hydrolab Surveyor and MS5 Minisonde.
Bacterial Carbon Production
Bacterial carbon production (BCP) was determined using an amended version of the 3H-leucine incorporation method of Smith and Azam [36]. Optimum 3H-leucine concentration was determined by creating a saturation curve (15, 30, 60, 120 nM) with composite samples from the sample sites 2 weeks before the flow event as in Buesing and Gessner [37]. Sample water was filtered to 2.7 μm through a sample-flushed glass fibre filter to remove bacterial predators [38]. Triplicate 1 mL aliquots of sample water were then dark incubated in 2 mL sterile-screw cap centrifuge tubes with o-rings with 3H-leucine (120 nM 1:5 3H-leucine:unlabelled leucine). Incubations were conducted at the field sites in an insulated container at ~18–20 °C for 1 h before incubations were terminated with cold 5% w/v trichloroacetic acid (TCA). One killed control, pre-treated with cold TCA, was included for every 12 replicates. Samples were then centrifuged, the supernatant was siphoned off, and the pellets were rinsed with ice-cold 5% TCA twice before being combined with Ultima Gold II scintillation fluid and radioactivity measured using a PerkinElmer Tri-Carb 2810 TR scintillation analyser. BCP was calculated from bacterial protein production using a conversion factor of 0.86 [39] with an internal isotope dilution factor of one assumed due to the high concentration of 3H-leucine used [40]. BCP measurements are communicated as either volumetric BCP (BCPvol μg C L−1 h−1) or total load of BCP in the water column during the hour of sampling (BCPload g C). BCPload was calculated as BCPvol multiplied by hourly discharge at the time of sampling. To determine if BCP was nutrient limited, an amendment of Lindstrom’s [41] growth media L16 (containing nitrate, phosphate, and micro-elements) was made without carbon and added to replicates of 3H-leucine incubations to determine what the maximum BCP rate was when not nutrient limited [42].
16S rRNA Gene Sequencing
Selected days at sites M2, M3, M4, T1, and T2 were sequenced to capture important changes in the microbiome based on preliminary BCP results. Site M1 was not sequenced due to negligible change in discharge or water source. Sample water was filtered through 47 mm, 0.22 μm PVDF membrane filters using a peristaltic pump (100 rpm). Filters were snap frozen in liquid nitrogen and stored at -80 °C until DNA was extracted using Qiagen DNeasy PowerWater Kit, according to the manufacturer’s instructions. Negative controls were extracted from field and lab blank filters.
DNA was amplified using the 16S rRNA primers; Bakt_341F and Bakt_805R, which amplify the V3–V4 region [43], with the following cycling conditions: 95 °C for 3 min followed by 25 cycles of: 95 °C for 30 s, 55 °C for 30 s, 72 °C for 30 s, and then 72 °C for 5 min with a final hold at 4 °C [44]. Amplicons, including negative controls, were checked on agarose gel to ensure the integrity of the filter and DNA extraction. Then sample amplicons were sequenced on the Illumina Miseq platform (2 × 300 bp) following the manufacturer’s guidelines at the Australian Genome Research Facility, Melbourne (AGRF). Raw data files in FASTQ format were deposited in the NCBI Sequence Read Archive (SRA) under Bioproject number PRJNA943514.
Raw demultiplexed 16S rRNA gene data were processed using the Quantitative Insights into Microbial Ecology (QIIME 2 version 2020.6.0) pipeline [45]. Briefly, paired-ended 16S DNA sequences were imported, then trimmed and denoised using DADA2 version 2020.6.0, which also removes chimeras [46]. The classify-sklearn qiime feature classifier was used to assign taxonomy against the Silva v138 database [47] at the amplicon sequence variant (ASV) level. The dataset was further cleaned by removing ASVs with less than 50 reads (0.01% of the total reads) and those identified as chloroplasts, mitochondria, or as unassigned sequences. Cleaned data were then rarefied at 3530 reads per sample (Supplementary figure S1).
Statistical Analysis
All statistical analysis was carried out in R version 4.1.3 [48]. Hourly flow rates at each site were estimated by time-shifting data from the nearest discharge gauge [34] to accommodate estimated water travel time to/from the gauge. Water transit times were calculated based on the time difference of peaks in discharge at consecutive gauges and equated to river distance measured in QGIS 3.24.2 [49]. Water velocity was estimated at 0.49 m s−1. Estimates may vary in accuracy due to changes in water velocity with magnitude of discharge but provide a more accurate estimate than using unadjusted gauge readings.
Relationships were explored between hourly flow rate, basal resources, bacterial community diversity, bacterial taxa relative abundance, and BCPvol using Pearson’s correlation tests. For statistical comparison, hourly flow rate was categorised into three groups: base flow, small inflow, and event flow. These categories were based on hydrographs and field observations of rainfall and changes in river height. Briefly, small inflows were classed as when flows more than doubled from base flow, and event flow was classed as flows over 2.9 m3 s−1. Differences in BCP and basal resources between flow categories and sites were compared with ANOVA or t-tests. Where the assumption of equal variance was not satisfied, Kruskal-Wallis or Wilcoxon rank sum tests were used as non-parametric alternatives. Drivers of BCPvol were modelled using stepwise multiple regression in the MASS package, and best models were validated by running a simulated model fit and selecting the best RMSE and R2 [50, 51]. To understand how much additional carbon was captured by the system in total during a flow event, BCPload was calculated for the hour during which sampling occurred only, with the assumption that BCPvol rates remained relatively stable over that hour. Loads were not modelled over the entire study period due to uncertainty in the ecological significance of the correlation between discharge and BCPvol rates and the high variability in BCPvol rates over small time periods [35].
Bacteria community composition was SIMPROF clustered with no a priori assumptions using clustsig to determine whether spatial or temporal factors were more important in the similarity between communities [52]. Differences in community composition between flow categories were assessed by ANOSIM using Bray-Curtis dissimilarity, with multivariate dispersion first tested using betadisper in vegan version 2.6.4 [53]. ASVs contributing the most to these differences were then identified with SIMPER. A Bonferroni correction was applied for pairwise ANOSIM comparisons. Shannon diversity was also calculated using vegan and significance tested using Kruskal-Wallis and Dunn’s tests. Differences in the relative abundance of taxa were tested with ANOVA. Environmental drivers of bacterial community composition were modelled with nMDS and stepwise dbRDA using vegan with best model fit assessed by permutation. Correlations between the 50 most abundant ASVs and environmental variables were visualised using network analysis. Pearson’s correlations were first generated using MICtools and the network was visualised in Cytoscape version 3.9.1 [54].
Figures were generated using ggplot2, and maps created in QGIS [55].
Results
Hydrology and Resource Mobilisation
All rivers in the local catchment had been at base flow for ~4 months prior to sampling. During the study period, rainfall caused a local flow pulse down the Lachlan River mainstem and a substantial flow pulse down the T2 tributary (mean daily discharge 9.5 m3 s−1, 92nd percentile). A minor flow pulse also occurred on the T1 (2.2 m3 s−1, 82nd percentile), Hovells Creek (2.5 m3 s−1, 93rd percentile), and Crowther Creek (0.81 m3 s−1, 84th percentile) tributaries (Fig. 2). DOC concentrations ranged between 7 and 13 mg L−1 and were significantly higher during small inflows than base flows at all sites except M1 (all except M1 χ2 = 12.9, p = 0.002; M1 only t = 1.69, p = 0.14, Table 1). At M1, concentrations were significantly higher than other sites throughout the study period despite little change in discharge at this site (12.1 ± 1.2 mg L−1, F(5,44) = 5.2, p = 0.001, Table 1). RP concentrations ranged from 1 to 36 μg L−1 and were up to 7 times higher during event flow within tributaries and significantly elevated at sites directly downstream of tributary confluences during event flow (all except M1 χ2 = 11.6, p = 0.003, Table 1). NOx and ammonia significantly increased by up to 8-fold during event inflows at T1 and all other mainstem sites, but NOx did not increase with flow at T2 (NOx all except T2 χ2 = 22.8, p < 0.001; T2 only W = 8, p = 1, Table 1). Temperature dropped significantly in tributaries and at all sites below confluences over the course of the flow event from ~21 to 17 °C (all except M1 F(2,60) = 89.4, p < 0.0001).
Hydrographs for study sites from the Lachlan River catchment during 23–28 October 2020. Sample times are marked with (◆). Water NSW gauge number used to calculate discharge is listed for each site [34]
Bacterial Carbon Production
The rate of BCPvol (μg C L−1 h−1) significantly decreased during the first 2 days of increased flow, then began to recover significantly at most sites on the final day of sampling (F(4,70) = 24.3, p < 0.0001), except for M1 and T1 which continued to drop (Fig. 3). The initial rate of BCPvol at base flow was disparate between sites, with BCPvol being statistically highest at M1 and lowest at T2 (F(5,12) = 86.3, p < 0.0001), but rates became more similar across sites on the final day of sampling (BCPvol start range = 1.7–4.9, end range = 2.0– 2.6 μg C L−1 h−1, Fig. 3). There was a significant increase in BCPvol during base flow at T1, T2, and M3 on the second day of sampling that corresponded with observations of initial direct rainfall (not sufficient magnitude to register on discharge gauge) and increased river channel wetting (F(3,8) ≥ 939, p < 0.0001, Fig. 3). Stepwise multiple linear regression identified significant positive drivers of log10 BCPvol as DOC, temperature, and conductivity (adj R2 = 0.56, F(3,21) = 11.12, p = 0.0001). No association could be found between BCPvol and bacterial community parameters or relative abundance of bacterial taxa. There was no difference in BCP between L16 growth media amended incubations and controls (Supplementary table S2).
Despite BCPvol rates per litre dropping during the flow event, the total load of carbon assimilated by riverine bacteria during the hour of sampling (BCPload) increased significantly by 4–20 times during the flow event due to the increased volume of water in the river channel, except for at M1 due to little change in discharge at this site (all except M1 F(2,18) = 7.5, p = 0.004; M1 only t = −0.13, p = 0.91; Table 1).
Bacterial Community Relative Abundance and Composition
Shannon diversity was significantly higher during the flow event compared to base and small inflows at all sites (χ2 = 18.19, p = 0.0001, Dunn’s p < 0.008). nMDS ordination of bacterial ASVs showed a clear grouping of base flow and event flow communities (Fig. 4). Event flow communities were significantly different to those during base and small inflows (ANOSIM overall R = 0.49, p = 0.0001, event-base R = 0.32, p = 0.0001, event-small inflow R = 0.45, p = 0.0001), and across the study period the community at T2 was significantly different to that at T1 and all other upstream sites (ANOSIM overall R = 0.26, p = 0.0001, T2-upstream R > 0.42, p < 0.005). SIMPROF clustering indicated the closest level of bacterial community similarity was between each tributary and the site directly downstream of their confluence, usually on the same day (Supplementary figure S3). Orthophosphate concentration and hourly discharge explained 31% of the variation in bacterial community composition across all sites, with communities clearly grouped by flow category in a distance-based redundancy analysis ordination (F(2,11) = 2.43, p = 0.001, Fig. 5).
Distance-based redundancy analysis (dbRDA) using Bray-Curtis dissimilarity showing influence of environmental variables on bacterial community composition during a tributary-driven flow pulse at five sites in the Lachlan River catchment. Only significant variables orthophosphate (RP) and discharge (hourly discharge) (p = 0.001) are shown as vectors, and ellipses represent 95% CI of the mean of each flow category
The ASVs contributing most to the difference in bacterial communities between flow categories belonged to the genera Spirosomaceae (Pseudarcicella), Burkholderiaceae (Polynucleobacter), and Alcaligenaceae (GKS98 freshwater group) (Supplementary table S4). The top contributing ASVs in these genera were all less abundant during the flow event than base or small inflows, with the exception of one Polynucleobacter ASV and two Pseudarcicella ASVs, which were more abundant during the flow event (Supplementary table S4). ASVs within the same three bacterial genera contributed most to the difference between tributary communities, with all ASVs more abundant at T2 than T1 (Supplementary table S5).
At the genus level, mean relative abundance of Pseudarcicella decreased significantly from 12% ± 5 to 2% ± 0.8 during event flow (F(4,37) = 5.8, p = 0.001), where there was a significant increase in abundance of GKS98 and Polynucleobacter at the onset of event flow that was sustained in Polynucleobacter only (GKS98 F(4,37) = 5.7, p = 0.001, Polynucleobacter F(4,37) = 5.9, p = 0.001). At the class level, Saccharimonadia (phyla Patescibacteria) comprised only a small proportion of mean relative abundance at base flow (< 2%) but became one of the dominant classes during event flow at all sites (~25%), particularly at tributary sites (Fig. 6). Network analysis showed positive correlations between nearly all Polynucleobacter and GKS98 ASVs and hourly discharge but negative correlations with BCPvol (Fig. 7). Conversely, nearly all Pseudarcicella ASVs were negatively correlated with hourly discharge but positively correlated with temperature.
Mean relative abundance of bacterial classes throughout the study period at tributary sites T1, T2, and mainstem sites M2, M3, and M4 in the Lachlan River catchment NSW. Data from sites M2 and M3 has been pooled as 16S rRNA data was only available from site M2 on day 2 (base flow) and M3 on days 3–5 (event flow)
Network analysis of Pearson’s correlations between the 50 most abundant bacterial amplicon sequencing variants (ASVs) and environmental variables during a tributary-driven flow pulse in the Lachlan River catchment NSW. Dark grey edges represent positive correlations, and light grey edges represent negative correlations with width of the line representing the strength of the relationship (absolute r range = 0.12–0.76). Node fill shade denotes bacterial genus of the ASV, and node size represents relative abundance within the top 50 ASVs (range 1–4%). Only significant correlations are presented
Discussion
During this study, tributary inflows altered environmental conditions in the river mainstem, mainly via increased concentrations of DOC, phosphorus and nitrogen, and reduced temperatures. These environmental changes corresponded with a shift in bacterial community composition and an initial decrease in BCPvol. There was nevertheless an overall increase in the total amount of carbon assimilated by riverine bacteria at all sites below tributary confluences due to the increased volume of water in the river channel. These findings deliver critical new knowledge on the role that tributary inflows play in supporting BCP in a regulated river during moderate-sized flow-resource pulses.
Tributary Inflow Impact on Bacterial Carbon Production
During base flow, there was a gradient of high to low BCPvol from upstream to downstream, which is consistent with patterns observed in a longitudinal study of the Hudson River, USA [56]. This gradient could have been caused by a longitudinal reduction in the amount of labile DOM available. As water from the dam moves downstream, repeated biological and photochemical processing leaves mostly recalcitrant DOM available with few new terrestrial inputs due to a lack of lateral connectivity at low flow [57]. A previous study of the Lachlan River found no longitudinal pattern in biological oxygen demand (BOD5); however, this study was conducted in the lower reaches of the river where the system becomes distributary and flow is regularly interrupted by weirs [32]. Interestingly, measures of bacterial metabolism did not respond to nutrient amendments of nitrate and phosphate in the aforementioned or current study, suggesting BCP in the Lachlan River may not generally be limited by inorganic nutrients.
Tributary inflows did not initially appear to stimulate or suppress BCPvol rates at mainstem sites below confluences, even though BCPvol rates at T1 were much higher than those in the mainstem and those at T2 were lower. However, BCPvol rates eventually began to increase at all study sites downstream of tributary inflows on day 5. As diverse lotic bacterial communities can adapt quickly to changed conditions in the regulated mainstem [25], it is possible that the delayed response time in BCPvol stimulation by tributary inflows was influenced by shortened mixing distances during the flow event [58]. As the flow event set in, the increased water velocity and higher proportion of water sourced from tributaries vs dam releases may have facilitated more efficient mixing of tributary-sourced DOC, nutrients, and bacteria resulting in increased BCPvol rates at sites downstream of tributary confluences on day 5 [59]. This eventual increase in BCPvol was not observed at M1 and T1, where rates continued to drop on the final day of sampling. This may have been an artefact of the homogenisation in bacterial community composition and BCPvol rates seen across sites during the flow event, considering both sites began with higher rates of BCPvol than all other sites (Fig. 3). Convergence in bacterial community composition and metabolism has been seen in other landscape-scale studies of bacterial composition during rainfall events [3, 25].
Tributary Inflow Impacts on Bacterial Community Composition
Tributary inflows rapidly changed bacterial communities at all sites, increasing bacterial diversity and shifting the dominant taxa. Increases in discharge and orthophosphate were found to influence these changes. Bacterial communities at sites below tributary confluences more closely resembled those in the tributaries during the flow event compared to base flow, indicating that tributary inflows may also be importing bacteria to the regulated mainstem or altering conditions to support similar communities in both locations. These findings suggest that a combination of resource delivery and microbial community coalescence was responsible for the changes observed in bacterial communities during tributary inflows. The convergence in BCPvol across sites observed in this study may have been an emergent property of the new microbial community formed by the coalescence of the catchment, tributary, and mainstem microbial communities [60]. Tributaries can be sources of microbial diversity in mainstem regulated rivers, even at times of base flow [61]. Provision and support of a more diverse bacterial community by tributary inflows could enhance the functional stability of the regulated mainstem ecosystem, as greater species richness and evenness is associated with improved ecosystem resilience [62, 63].
ASVs identified as Spirosomaceae, Pseudarcicella were dominant at most study sites during base flow, but the relative abundance of members of this group decreased substantially during event flow. Pseudarcicella hirudinus is known to utilise a variety of substances found in algal exudates (carbohydrates, amino acids, pyruvate) as its sole carbon source [64], and the genus has been associated with algal blooms [65] and may prefer low DOC, nutrients, and turbidity [66,67,68]. This suggests the changed environment during the flow event could have created hostile conditions for Pseudarcicella, reducing its relative abundance through a combination of dilution, competition, and community coalescence [3, 60].
As Pseudarcicella populations decreased in relative abundance, both the Burkholderiaceae, Polynucleobacter, and Alcaligenaceae GKS98 increased significantly with tributary inflows. Polynucleobacter and GKS98 are small cosmopolitan bacterial genera that mainly rely on organic acids released by the photochemical degradation of complex DOM for growth [69,70,71,72]. Increases in Polynucleobacter abundance of up to 50-fold have been observed in microcosms within 4 days of terrestrial DOC addition [73]. This suggests that the increase in DOC inputs from tributaries may be beneficial for these taxa’s growth.
At the class level, relative abundances of Saccharimonadia (phyla Patescibacteria) increased greatly during event flow, particularly in tributaries and mainstem sites downstream of the confluences. Patescibacteria may be considerably important in the determination of microbial metabolism during tributary inflows due to their preferential mobilisation from soils and groundwater and ability to degrade both simple and complex sources of carbon [74, 75]. They have even been indicated as potential degraders of recalcitrant hydrocarbons in studies on the microbial degradation of PET and PVC [76, 77].
The changes observed in bacterial taxa during tributary inflows from Pseudarcicella dominated to Polynucleobacter and GKS98 dominated suggest a shift in bacterial community metabolic function across the catchment from one mostly reliant on algal exudates to one more adapted to the heterotrophic assimilation of complex terrestrial DOM. Similar responses to surface water inflows have been seen in hyporheic bacterial communities and demonstrate the metabolic adaptability of a diverse microbiome [18].
Environmental Influences on Bacterial Carbon Production
BCPvol rates in this study were positively correlated with DOC, temperature, and conductivity, similar to other global studies of riverine BCP [22, 31, 78]. Temperature positively affects BCPvol and the speed at which BCPvol responds to nutrient additions [22, 79]. The drop in temperature across all sites of ~4 °C during tributary inflows may have reduced BCPvol and contributed to delayed response times to the increased resource loads. The change in DOC with discharge was not uniform across the catchment, with concentrations at tributary sites and M2 increasing with flow but other mainstem sites remaining at a similar concentration to base flow. However, there may have been a change in DOC composition, rather than quantity, during the flow event. The bulk of DOC in river mainstems can become recalcitrant due to repeated biological and photochemical processing; hence, bacterial communities in river mainstems are often carbon or nutrient limited [80,81,82,83]. Rainfall-driven inflows can deliver a subsidy of fresh terrestrial DOM to rivers [84]. Along with the increased bacterial diversity provided by tributary inflows, this subsidy may have unified the Lachlan River catchment’s bacterial community metabolism, creating a carbon “sponge” to take advantage of this fleeting resource. There was no detectable relationship between bacterial community composition or diversity and BCP, which suggests that changes in BCP were occurring somewhat evenly throughout the community.
Dominant small taxa (cell size < 0.22 μm) that were detected in this study, including Polynucelobacter, GKS98, and Patescibacteria, play an important role in riverine DOM metabolism and are often mobilised in large quantities from catchment soils and groundwater during flow events [24, 75, 85]. The increased importance of these bacterial taxa during tributary inflows should imply an accompanying increase in BCP; however, the relative abundance of dominant GKS98 and Polynucleobacter ASVs were negatively correlated with BCPvol. This suggests that a related unsampled variable could be an important driver of BCP in the Lachlan River system. For example, a competitive, symbiotic, or predatory relationship between bacterial and protistan taxa can impact on bacterial production [86, 87]. Additionally, these genera’s weak utilisation of amino acids may have confounded the 3H-leucine incubations and erroneously caused this correlation. Both Polynucleobacter and GKS98 have a high metabolic specificity for a range of carboxylic acids and acetate but only low use of typical algal exudates, including amino acids [72]. For example, in a study of diverse freshwater bodies, Polynucleobacter only utilised 56% of leucine in incubations [88]. It is therefore possible that some of the fluctuations measured in BCP in this study were more related to changes in community utilisation of leucine than actual changes in BCP.
Despite the overall drop in BCPvol during the flow event compared to base flow, the total load of carbon assimilated by riverine bacteria (BCPload) increased up to 20 times during the flow event due to the larger volume of water in the river channel. This provides an important bioavailable carbon subsidy for higher trophic levels in the river food web, especially after a lengthy period of low flow with limited new resources entering the system [89, 90]. Bacterioplankton are predated upon by small zooplankton (rotifers, nauplii) and protists [21, 91]. This trophic transfer can occur extremely quickly, with heterotrophic nanoflagellates (HNFs) reaching bacterial clearance rates of > 100% day−1 during increased discharge [92, 93]. So, it is feasible that the increased energy captured by bacteria during a flow event could be available to higher-order consumers within 24 h [94, 95].
Implications
How important tributary inflows are for basal food web stimulation in comparison to managed environmental water releases from dams is yet to be determined. Managed dam releases can stimulate similar increases in DOC and nutrients to natural inflows and can be managed to provide additional benefits, such as increased longitudinal connectivity during important fish breeding times [10, 96]. Although DOM molecular composition may be altered due to lentic dynamics within the dam [5, 97], collection of terrestrial materials from dry river benches with increasing distance downstream could support the food web in similar ways to natural inflows [98]. However, there are other benefits to the protection of tributary inflows that environmental flows do not provide, such as superior fish production outcomes and maintenance of “hot spots” of biotic diversity and complexity below confluences [99, 100]. The combined benefit of these outcomes along with basal food web stimulation implicates protection of tributary flows as a valuable river management tool.
Conclusions
In this study, tributary inflows provided a significant subsidy of terrestrial carbon to the riverine food web in a regulated mainstem river. Inflows changed bacterial community composition and increased bacterial diversity in the mainstem through a combination of microbial community coalescence and environmental changes. Community changes did not appear to increase the capacity of the river microbiome to metabolise complex terrestrial DOM imported during the flow event on a per litre basis; however, the large increase in water volume in the river channel resulted in a parallel increase in riverine carbon assimilation. These findings highlight the key role tributary inflows play in maintaining microbial diversity and fuelling the food web in regulated rivers.
Data Availability
16S rRNA gene sequencing data from the current study are available in the NCBI Sequence Read Archive (SRA) under Bioproject number PRJNA943514. All other datasets generated during the current study are available from the corresponding author on reasonable request.
References
Ward JV, Stanford JA (1995) The serial discontinuity concept: extending the model to floodplain rivers. River Res Appl 10:159–168
Junk WJ, Bayley PB, Sparks RE (1989) The flood pulse concept in river-floodplain systems. Can Spec Publ Fish Aquat Sci 106:110–127
Doering M, Freimann R, Antenen N et al (2021) Microbial communities in floodplain ecosystems in relation to altered flow regimes and experimental flooding. Sci Total Environ 788. https://doi.org/10.1016/j.scitotenv.2021.147497
Growns I, Reinfelds I, Williams S, Coade G (2009) Longitudinal effects of a water supply reservoir (Tallowa Dam) on downstream water quality, substrate and riffle macroinvertebrate assemblages in the Shoalhaven River, Australia. Mar Freshw Res 60:594–606
Cortez DP, Growns IO, Mitrovic SM, Lim RP (2012) Effects of a gradient in river regulation on the longitudinal trends in water quality and benthic algal and macroinvertebrate assemblages in the Hunter River, Australia. Mar Freshw Res 63:494–504
Veraart AJ, Romaní AM, Tornes E, Sabater S (2008) Algal response to nutrient enrichment in a forested oligotrophic stream. J Phycol 44:564–572
Maheshwari B (1995) Effects of regulation on the flow regime of the river Murray, Australia. Regul Rivers: Res Manage 10:15–38
Wang Y, Rhoads BL, Wang D (2016) Assessment of the flow regime alterations in the middle reach of the Yangtze River associated with dam construction: potential ecological implications. Hydrol Process 30:3949–3966. https://doi.org/10.1002/hyp.10921
Wen L, Rogers K, Ling J, Saintilan N (2011) The impacts of river regulation and water diversion on the hydrological drought characteristics in the Lower Murrumbidgee River, Australia. J Hydrol (Amst) 405:382–391. https://doi.org/10.1016/j.jhydrol.2011.05.037
Westhorpe DP, Mitrovic SM (2012) Dissolved organic carbon mobilisation in relation to variable discharges and environmental flows in a highly regulated lowland river. Mar Freshw Res 63:1218–1230. https://doi.org/10.1071/MF12122
Baldwin DS, Colloff MJ, Mitrovic SM et al (2016) Restoring dissolved organic carbon subsidies from floodplains to lowland river food webs: a role for environmental flows? Mar Freshw Res 67:1387–1399. https://doi.org/10.1071/MF15382
Rohlfs AM, Mitrovic SM, Williams S, Coleman D (2016) Can tributary in-flows improve the recovery of the dissolved organic carbon regime in a snowmelt river regulated by a large reservoir? Mar Freshw Res 67:1338–1345. https://doi.org/10.1071/MF14230
Bruns DA, Minshall GW, Cushing CE et al (1984) Tributaries as modifiers of the river continuum concept: analysis by polar ordination and regression models. Archiv fur Hydrobiology 99:208–220
Sabo JL, Caron M, Doucett R et al (2018) Pulsed flows, tributary inputs and food-web structure in a highly regulated river. J Appl Ecol 55:1884–1895
McClain ME, Naiman RJ (2008) Andean influences on the biogeochemistry and ecology of the Amazon River. Bioscience 58:325–338. https://doi.org/10.1641/B580408
Ward JV, Stanford JA (1983) The serial discontinuity concept of lotic ecosystems. Dyn Lotic Ecosystems 10:29–42
Caillon F, Schelker J (2020) Dynamic transfer of soil bacteria and dissolved organic carbon into small streams during hydrological events. Aquat Sci 82. https://doi.org/10.1007/s00027-020-0714-4
Graham EB, Crump AR, Resch CT et al (2017) Deterministic influences exceed dispersal effects on hydrologically-connected microbiomes. Environ Microbiol 19:1552–1567. https://doi.org/10.1111/1462-2920.13720
Mansour I, Heppell CM, Ryo M, Rillig MC (2018) Application of the microbial community coalescence concept to riverine networks. Biol Rev 93:1832–1845. https://doi.org/10.1111/brv.12422
Degerman R, Lefébure R, Byström P et al (2018) Food web interactions determine energy transfer efficiency and top consumer responses to inputs of dissolved organic carbon. Hydrobiologia 805:131–146. https://doi.org/10.1007/s10750-017-3298-9
Meyer JL (1994) The microbial loop in flowing waters. Microb Ecol 28:195–199. https://doi.org/10.1007/BF00166808
Adams HE, Crump BC, Kling GW (2015) Isolating the effects of storm events on arctic aquatic bacteria: temperature, nutrients, and community composition as controls on bacterial productivity. Front Microbiol 6:1–13. https://doi.org/10.3389/fmicb.2015.00250
Guillemette F, Mccallister SL, del Giorgio PA (2016) Selective consumption and metabolic allocation of terrestrial and algal carbon determine allochthony in lake bacteria. ISME J 10:1373–1382. https://doi.org/10.1038/ismej.2015.215
Hermans SM, Buckley HL, Case BS, Lear G (2020) Connecting through space and time: catchment-scale distributions of bacteria in soil, stream water and sediment. Environ Microbiol 22:1000–1010. https://doi.org/10.1111/1462-2920.14792
Adams HE, Crump BC, Kling GW (2014) Metacommunity dynamics of bacteria in an arctic lake: the impact of species sorting and mass effects on bacterial production and biogeography. Front Microbiol 5:1–10. https://doi.org/10.3389/fmicb.2014.00082
Baldwin DS, Whitworth KL, Hockley CL (2014) Uptake of dissolved organic carbon by biofilms provides insights into the potential impact of loss of large woody debris on the functioning of lowland rivers. Freshw Biol 59:692–702
Rolls RJ, Baldwin DS, Bond NR et al (2017) A framework for evaluating food-web responses to hydrological manipulations in riverine systems. J Environ Manage 203:136–150. https://doi.org/10.1016/j.jenvman.2017.07.040
MDBA (Murray-Darling Basin Authority) (2011) Guide to the proposed Basin Plan: technical background Part 1. Canberra
Rees GN, Beattie G, Bowen PM, Hart BT (2005) Heterotrophic bacterial production in the lower Murray River, south-eastern Australia. Mar Freshw Res 56:835–841. https://doi.org/10.1071/MF04232
Burford MA, Cook AJ, Fellows CS et al (2008) Sources of carbon fuelling production in an arid floodplain river. Mar Freshw Res 59:224–234. https://doi.org/10.1071/MF07159
Carney RL, Mitrovic SM, Jeffries T et al (2015) River bacterioplankton community responses to a high inflow event. Aquat Microb Ecol 75:187–205. https://doi.org/10.3354/ame01758
Moran NP, Ganf GG, Wallace TA, Brookes JD (2014) Flow variability and longitudinal characteristics of organic carbon in the Lachlan River, Australia. Mar Freshw Res 65:50–58. https://doi.org/10.1071/MF12297
Green D, Petrovic J, Moss P, Burrell M (2011) Water resources and management overview: Lachlan catchment. NSW Office of Water, Sydney
Water NSW (2022) Real time data - rivers and streams. https://realtimedata.waternsw.com.au/. Accessed 18 Oct 2018
del Giorgio P a, Pace ML, Fischer D (2006) Relationship of bacterial growth efficiency to spatial variation in bacterial activity in the Hudson River. Aquat Microb Ecol 45:55–67. https://doi.org/10.3354/ame045055
Smith D, Azam F (1992) A simple, economical method for measuring bacterial protein synthesis rates in seawater using. Mar Microb Food Webs 6:107–114
Buesing N, Gessner MO (2003) Incorporation of radiolabeled leucine into protein to estimate bacterial production in plant litter, sediment, epiphytic biofilms, and water samples. Microb Ecol 45:291–301. https://doi.org/10.1007/s00248-002-2036-6
Guillemette F, del Giorgio PA (2011) Reconstructing the various facets of dissolved organic carbon bioavailability in freshwater ecosystems. Limnol Oceanogr 56:734–748. https://doi.org/10.4319/lo.2011.56.2.0734
Simon M, Azam F (1989) Protein content and protein synthesis rates of planktonic marine bacteria. Mar Ecol Prog Ser 51:201–213. https://doi.org/10.3354/meps051201
Giering SLC, Evans C (2022) Overestimation of prokaryotic production by leucine incorporation—and how to avoid it. Limnol Oceanogr 67:726–738
Lindstrom K (1991) Nutrient requirements of the Dinoflagellate Peridinium gatenense. J Phycol 27:207–219
Soares ARA, Bergström A, Sponseller RA et al (2017) New insights on resource stoichiometry: assessing availability of carbon , nitrogen , and phosphorus to bacterioplankton. Biogeosciences 14:1527–1539. https://doi.org/10.5194/bg-14-1527-2017
Herlemann DPR, Labrenz M, Jürgens K et al (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. https://doi.org/10.1038/ismej.2011.41
Illumina (2013) 16S metagenomic sequencing library preparation: preparing 16S ribosomal RNA gene amplicons for the illumina MiSeq system. Illumina
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
R Core Team (2021) R: A language and environment for statistical computing
QGIS.org (2022) QGIS Geographic Information System
Venables WN, Ripley BD (2002) Modern Applied Statistics with S. Fourth, Springer, New York
Kassambara A (2018) Stepwise regression. Machine learning essentials: practical guide in R. First, STHDA, pp 82–85
Whitaker D, Christman M (2014) clustsig: significant cluster analysis. R package version 1.1, <https://CRAN.R-project.org/package=clustsig>
Oksanen J, Blanchet FG, Friendly M, et al (2022) vegan: community ecology package. R package version 2.6.4, <https://CRAN.R-project.org/package=vegan>
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
Maranger RJ, Pace ML, del Giorgio PA et al (2005) Longitudinal spatial patterns of bacterial production and respiration in a large river-estuary: implications for ecosystem carbon consumption. Ecosystems 8:318–330. https://doi.org/10.1007/s10021-003-0071-x
Mulholland PJ (2003) Large-scale patterns in dissolved organic carbon concentration, flux, and sources. In: Sinsabaugh RL (ed) Findlay SEG. Aquatic Ecosystems, Interactivity of Dissolved Organic Matter. Elsevier, pp 139–159
Velimirov B, Milosevic N, Kavka GG et al (2011) Development of the bacterial compartment along the Danube River: a continuum despite local influences. Microb Ecol 61:955–967. https://doi.org/10.1007/s00248-010-9768-5
Turunen K, Räsänen T, Hämäläinen E et al (2020) Analysing contaminant mixing and dilution in river waters influenced by mine water discharges. Water Air Soil Pollut 231:1–15. https://doi.org/10.1007/s11270-020-04683-y
Rillig MC, Antonovics J, Caruso T et al (2015) Interchange of entire communities: microbial community coalescence. Trends Ecol Evol 30:470–476
Cruaud P, Vigneron A, Dorea CC et al (2020) Rapid changes in microbial community structures along a meandering river. Microorganisms 8:1–21. https://doi.org/10.3390/microorganisms8111631
Bell T, Newman JA, Silverman BW et al (2005) The contribution of species richness and composition to bacterial services. Nature 436:1157–1160
Wittebolle L, Marzorati M, Clement L et al (2009) Initial community evenness favours functionality under selective stress. Nature 458:623–626. https://doi.org/10.1038/nature07840
Kämpfer P, Busse HJ, Longaric I et al (2012) Pseudarcicella hirudinis gen. nov., sp. nov., isolated from the skin of the medical leech hirudo medicinalis. Int J Syst Evol Microbiol 62:2247–2251. https://doi.org/10.1099/ijs.0.037390-0
Shao Q, Lin Z, Zhou C et al (2020) Succession of bacterioplankton communities over complete Gymnodinium-diatom bloom cycles. Sci Total Environ 709. https://doi.org/10.1016/j.scitotenv.2019.135951
Guo D, Liang J, Chen W et al (2020) Bacterial community analysis of two neighboring freshwater lakes originating from one lake. Pol J Environ Stud 30:111–117. https://doi.org/10.15244/pjoes/119094
Yang Y, Li S, Gao Y et al (2019) Environment-driven geographical distribution of bacterial communities and identification of indicator taxa in Songhua River. Ecol Indic 101:62–70. https://doi.org/10.1016/j.ecolind.2018.12.047
Yu S, He R, Song A et al (2019) Spatial and temporal dynamics of bacterioplankton community composition in a subtropical dammed karst river of southwestern China. Microbiologyopen 8. https://doi.org/10.1002/mbo3.849
Watanabe K, Komatsu N, Ishii Y, Negishi M (2009) Effective isolation of bacterioplankton genus Polynucleobacter from freshwater environments grown on photochemically degraded dissolved organic matter. FEMS Microbiol Ecol 67:57–68. https://doi.org/10.1111/j.1574-6941.2008.00606.x
Hahn MW, Scheuerl T, Jezberová J et al (2012) The passive yet successful way of planktonic life: genomic and experimental analysis of the ecology of a free-living polynucleobacter population. PLoS One 7. https://doi.org/10.1371/journal.pone.0032772
Jezberová J, Jezbera J, Brandt U et al (2010) Ubiquity of Polynucleobacter necessarius ssp. asymbioticus in lentic freshwater habitats of a heterogenous 2000 km2 area. Environ Microbiol 12:658–669. https://doi.org/10.1111/j.1462-2920.2009.02106.x
Watanabe K, Komatsu N, Kitamura T et al (2012) Ecological niche separation in the Polynucleobacter subclusters linked to quality of dissolved organic matter: a demonstration using a high sensitivity cultivation-based approach. Environ Microbiol 14:2511–2525. https://doi.org/10.1111/j.1462-2920.2012.02815.x
Burkert U, Warnecke F, Babenzien D et al (2003) Members of a readily enriched β-proteobacterial clade are common in surface waters of a humic lake. Appl Environ Microbiol 69:6550–6559. https://doi.org/10.1128/AEM.69.11.6550-6559.2003
Vigneron A, Cruaud P, Langlois V et al (2020) Ultra-small and abundant: candidate phyla radiation bacteria are potential catalysts of carbon transformation in a thermokarst lake ecosystem. Limnol Oceanogr Lett 5:212–220. https://doi.org/10.1002/lol2.10132
Herrmann M, Wegner CE, Taubert M et al (2019) Predominance of Cand. Patescibacteria in groundwater is caused by their preferential mobilization from soils and flourishing under oligotrophic conditions. Front Microbiol 10:1–15. https://doi.org/10.3389/fmicb.2019.01407
Wu X, Pan J, Li M et al (2019) Selective enrichment of bacterial pathogens by microplastic biofilm. Water Res 165:114979
Zhelezova AD, Zverev AO, Zueva AI et al (2021) Prokaryotic community formation on polyethylene films incubated for six months in a tropical soil. Environ Pollut 269:116126. https://doi.org/10.1016/j.envpol.2020.116126
Becker JC, Rodibaugh KJ, Hahn D, Nowlin WH (2017) Bacterial community composition and carbon metabolism in a subtropical riverscape. Hydrobiologia 792:209–226. https://doi.org/10.1007/s10750-016-3058-2
Watanabe K, Ishii Y, Komatsu N et al (2017) Growth rates and tolerance to low water temperatures of freshwater bacterioplankton strains: ecological insights from shallow hypereutrophic lakes in Japan. Hydrobiologia 792:67–81. https://doi.org/10.1007/s10750-016-3045-7
Benner R, Opsahl S, Chin-Leo G et al (1995) Bacterial carbon metabolism in the Amazon River system. Limnol Oceanogr 40:1262–1270
Westhorpe DP, Mitrovic SM, Ryan D, Kobayashi T (2010) Limitation of lowland riverine bacterioplankton by dissolved organic carbon and inorganic nutrients. Hydrobiologia 652:101–117. https://doi.org/10.1007/s10750-010-0322-8
Soares ARA, Kritzberg ES, Custelcean I, Berggren M (2018) Bacterioplankton responses to increased organic carbon and nutrient loading in a boreal estuary—separate and interactive effects on growth and respiration. Microb Ecol 76:144–155. https://doi.org/10.1007/s00248-017-1115-7
Yeh T, Krennmayr K, Liao C et al (2020) Effects of terrigenous organic substrates and additional phosphorus on bacterioplankton metabolism and exoenzyme stoichiometry. Freshw Biol 65:1973–1988. https://doi.org/10.1111/fwb.13593
Robertson AI, Burns A, Hillman TJ (2016) Scale-dependent lateral exchanges of organic carbon in a dryland river during a high-flow experiment. Mar Freshw Res 67:1293–1301. https://doi.org/10.1071/MF15371
Fujii M, Kojima H, Iwata T et al (2012) Dissolved organic carbon as major environmental factor affecting bacterioplankton communities in mountain lakes of Eastern Japan. Microb Ecol 63:496–508. https://doi.org/10.1007/s00248-011-9983-8
Šimek K, Horňák K, Jezbera J et al (2008) Spatio-temporal patterns of bacterioplankton production and community composition related to phytoplankton composition and protistan bacterivory in a dam reservoir. Aquat Microb Ecol 51:249–262. https://doi.org/10.3354/ame01193
Carney RL, Seymour JR, Westhorpe D, Mitrovic SM (2016) Lotic bacterioplankton and phytoplankton community changes under dissolved organic-carbon amendment: evidence for competition for nutrients. Mar Freshw Res 67:1362–1373. https://doi.org/10.1071/MF15372
Jezbera J, Jezberová J, Koll U et al (2012) Contrasting trends in distribution of four major planktonic betaproteobacterial groups along a pH gradient of epilimnia of 72 freshwater habitats. FEMS Microbiol Ecol 81:467–479. https://doi.org/10.1111/j.1574-6941.2012.01372.x
Roach KA (2013) Environmental factors affecting incorporation of terrestrial material into large river food webs. Freshw Sci 32:283–298. https://doi.org/10.1899/12-063.1
Bino G, Brandis K, Kingsford RT, Porter J (2020) Waterbird synchrony across Australia’s highly variable dryland rivers – risks and opportunities for conservation. Biol Conserv 243:108497. https://doi.org/10.1016/j.biocon.2020.108497
Meira BR, Lansac-Toha FM, Segovia BT et al (2018) The importance of herbivory by protists in lakes of a tropical floodplain system. Aquat Ecol 1–18. https://doi.org/10.1007/s10452-018-9654-7
Weitere M, Arndt H (2002) Water discharge-regulated bacteria-heterotrophic nanoflagellate (HNF) interactions in the water column of the River Rhine. Microb Ecol 44:19–29. https://doi.org/10.1007/s00248-002-2010-3
Dolan JR, Gallegos CL (1991) Trophic coupling of rotifers, microflagellates, and bacteria during fall months in the Rhode River Estuary. Mar Ecol Prog Ser 77:147–156. https://doi.org/10.3354/meps077147
Kaminskas S, Humphries P (2009) Diet of murray cod (Maccullochella peelii peelii) (mitchell) larvae in an Australian lowland river in low flow and high flow years. Hydrobiologia 636:449–461. https://doi.org/10.1007/s10750-009-9973-8
Balcombe SR, Bunn SE, McKenzie-Smith FJ, Davies PM (2005) Variability of fish diets between dry and flood periods in an arid zone floodplain river. J Fish Biol 67:1552–1567. https://doi.org/10.1111/j.1095-8649.2005.00858.x
Stuart IG, Sharpe CP (2022) Ecohydraulic model for designing environmental flows supports recovery of imperilled Murray cod (Maccullochella peelii) in the Lower Darling-Baaka River following catastrophic fish kills. Mar Freshw Res 73:247–258. https://doi.org/10.1071/MF20377
Westhorpe DP, Mitrovic SM, Growns IO et al (2015) Disruption in water quality patterns along the river continuum by a large bottom release dam. Aust J Environ Manag 22:400–416
Rohlfs AM, Mitrovic SM, Williams S et al (2016) Dissolved organic carbon delivery from managed flow releases in a montane snowmelt river. Aquat Sci 78:793–807. https://doi.org/10.1007/s00027-016-0472-5
Beesley L, King AJ, Gawne B et al (2014) Optimising environmental watering of floodplain wetlands for fish. Freshw Biol 59:2024–2037. https://doi.org/10.1111/fwb.12404
Rice SP, Kiffney P, Greene C, Pess GR (2008) The ecological importance of tributaries and confluences. River Confluences, Tributaries Fluvial Netw:209–242
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and data collection were performed by Lauren O’Brien and Matthew Balzer. Sample analysis and statistical analysis were performed by Lauren O’Brien and Nachshon Siboni. The first draft of the manuscript was written by Lauren O’Brien, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
ESM 1
(DOCX 1480 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
O’Brien, L., Siboni, N., Seymour, J.R. et al. Tributary Inflows to a Regulated River Influence Bacterial Communities and Increase Bacterial Carbon Assimilation. Microb Ecol 86, 2642–2654 (2023). https://doi.org/10.1007/s00248-023-02271-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02271-1