Abstract
Denitrification in sediments is a key microbial process that removes excess fixed nitrogen, while dissimilatory nitrate reduction to ammonium (DNRA) converts nitrate to ammonium. Although microorganisms are responsible for essential nitrogen (N) cycling, it is not yet fully understood how these microbially mediated processes respond to toxic hydrophobic organic compounds (HOCs) and metals. In this study, we sampled long-term polluted sediment from the outer harbor of Oskarshamn (Baltic Sea), measured denitrification and DNRA rates, and analyzed taxonomic structure and N-cycling genes of microbial communities using metagenomics. Results showed that denitrification and DNRA rates were within the range of a national reference site and other unpolluted sites in the Baltic Sea, indicating that long-term pollution did not significantly affect these processes. Furthermore, our results indicate an adaptation to metal pollution by the N-cycling microbial community. These findings suggest that denitrification and DNRA rates are affected more by eutrophication and organic enrichment than by historic pollution of metals and organic contaminants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrogen loading has increased globally in many coastal seascapes, mainly due to the increase in agricultural food production and use of chemical fertilizers [1]. This has implications for coastal ecosystems resulting in enhanced primary productivity and eutrophication with episodic hypoxia in bottom water and sediments [1, 2]. Particularly the Baltic Sea ecosystem has a history of exposure to eutrophication with bottom water and sediment hypoxia and anoxia [3]. Microbial denitrification is the main route of nitrate (NO3−) and nitrite (NO2−) removal in coastal zones [1]. This anaerobic process reduces NO3− and NO2− into gaseous nitrogen compounds and typically occurs directly below the oxic sediment surface in coastal environments [4]. Bottom water NO3− concentrations and nitrification in the oxic sediment surface regulate the availability of NO3− for denitrification in the underlying sediment [4]. Through several redox reduction steps, NO3− is ultimately reduced to nitrogen gas (N2) [5]. Each step is carried out by enzymes encoded by specific genes and this information has been used in various studies to investigate and compare denitrification gene diversity, abundance, or gene expression. For example, the genes nirS and nirK encode for nitrite reductases (NO2− to NO), and nosZ encodes for nitrous oxide reductase responsible for the final step in the denitrification pathway (N2O to N2) [e.g., 6, 7, 8]. While denitrification provides a route for removal of bioavailable nitrogen (N), dissimilatory nitrate reduction to ammonium (DNRA) instead recycles NO3− to NH4+ [5, 6]. This is an important process that internally recycles fixed N via mineralization/release of NH4+ followed by subsequent nitrification to NO3−. Compared to denitrification, DNRA thus prevents NO3− being lost as N2-gas, potentially supporting eutrophication [7]. Studies investigating DNRA target the gene nrfA that encodes for a nitrite reductase enzyme used in DNRA [8]. In addition, the gene nirB encodes for a nitrate reductase used in DNRA but might also be used in assimilatory nitrate reduction [9]. The key roles of sediment microorganisms in these processes of cycling essential nitrogen [10] between sediment and water [11], together with the prevalent industrial pollution of urbanized coastal areas, call for a better understanding of how and whether the microbial community’s functioning is impacted in industrially polluted coastal sediments.
Both pollution by toxic compounds (e.g., metals and organic contaminants) and nutrient loading of coastal ecosystems have dramatically increased during the last century as a result of industrial processes, land use changes, increased agricultural practices, and the excessive use of fertilizers [2, 12, 13]. Of particular environmental and societal concern is the pollution by toxic metals and persistent hydrophobic organic compounds (HOCs), such as polychlorinated dibenzodioxins (PCDD) and dibenzofurans (PCDF), because of their high toxicity and long-term effect on biota [14, 15]. Industrially emitted metals often reach concentrations in the sediment that have been shown to be toxic to macrofauna but may also select for species with a tolerance to metals [14, 16]. Previous research indicates that at least some microscopic animals (e.g., specific nematodes) can develop tolerance and survive in sediment with persistent organic contaminants and/or toxic metals [17,18,19]. Tolerance to metals and hydrophobic organic compounds has, however, been shown to be considerably more common among bacteria due to highly diverse bacterial metabolic capacities and rapid genetic and physiological adaptation [20, 21].
There is currently a lack of studies investigating how high concentrations of metals in sediments affect DNRA rates; however, some previous studies have shown that high concentrations of, e.g., Cd, lead (Pb), copper (Cu), and zinc (Zn) in sediment can inhibit microbial denitrification [22, 23]. Broman et al. [24], on the other hand, found increased denitrification rates in sediment exposed to small amendments of Cd (1 mg Cd kg−1 ww sediment) and suggested that this was a result of fluctuating oxygen conditions and cadmium sulfide formation causing reduced metal bioavailability. The presence of toxic metals has been found to change the diversity and reduce the abundance of denitrifying bacteria [25], which might explain some of the different effects on denitrification rates observed in previous studies. Compared to metals, less is known about how persistent organic contaminants may influence denitrification rates. In a 60-day laboratory study, Lindgren et al. [26] observed lower denitrification rates over time, indicating that petroleum hydrocarbons might decrease denitrification rates. However, it is unknown if and how long-term exposures of benthic microbial communities in the field to historical emissions of both metals and persistent organic pollutants (such as PCDD/Fs) may affect the metabolic capacity of benthic microbial communities for nitrogen transformation processes denitrification and DNRA in the sediment.
In this study, we sampled coastal polluted Baltic Sea sediments with the aim of investigating to what extent the nitrogen transformation by sediment microbial communities is affected by historic industrial emissions of toxic metals and dioxins (PCDD/Fs). To address this aim, we analyzed sediment from the contaminated outer harbor of Oskarshamn in south-west Sweden at two stations with different levels of pollution. For reference sediment, we used one of the Swedish marine field reference sites that has little or no industrial pollution, the Uttervik Bay near the Askö Marine Laboratory in the western Baltic Sea. From the collected sediment, we (i) measured denitrification and DNRA rates and (ii) investigated the microbial community composition, diversity, and relative abundance of genes for nitrogen cycling, metal resistance, and degradation of organic pollutants using a metagenomics approach. We hypothesized that the polluted harbor sediment would have (1) lower denitrification and DNRA production rates than reported for unpolluted sites in the Baltic Sea; (2) a low relative abundances of denitrification and DNRA genes, but high genetic capacity for resistance to metals and to degrade organic pollutants; and (3) different bacterial and archaeal taxonomic diversity when compared with sediment communities from less polluted sediments.
Materials and Methods
Field Sampling
Polluted sediment was collected on June 27 in 2020, with the research vessel R/V Electra, from the outer harbor of the town Oskarshamn, western Baltic Sea, Sweden. This harbor has a 100 years of history of pollution of metals and persistent organic contaminants due to an adjacent copper plant (no longer active), a Cd-nickel (Ni) battery factory, and shipyards [27]. While the sediment in the inner harbor (not sampled in this study) has been suction-dredged to reduce contamination levels of metals and some of the highest PCDD/PCDFs concentrations so far noted in Sweden [27,28,29,30,31], the outer harbor sediment, which we sampled in this study, has not been remediated [for a map of the harbor area, see [32]. The outer harbor sediments have been reported by environmental consultants to have high concentrations of metals (e.g., Cu, Zn, and Pb with values > 5 times above national reference standard values) and organic pollutants such as dioxins [33]. Sediment was collected from the outer harbor for denitrification and DNRA rate measurements (isotope pairing technique (IPT) incubations), DNA extractions, total organic carbon (TOC) and nitrogen (TN), and metals and PCDD/PCDFs analyses. Samples were collected from two locations ca 130 m apart in the outer harbor: station OSK 1 (16 m water depth; WGS84 coordinates: Lat 57.264483, Long 16.481950; Fig. S1) and station OSK 2 (15.5 m water depth; WGS84 coordinates: Lat 57.265733, Long 16.482300). The sediment was collected using a modified box corer [34] with insert transparent acrylic liners (surface area: 756.25 cm2; length and width: 27.5 × 27.5 cm). To collect undisturbed bottom water and sediment, the box cores were subsampled on deck immediately after collection using small transparent acrylic sediment tube cores (cylinders; inner diameter 4.6 cm, length 30 cm). From each station, two box cores were collected. Nine tube cores were subsampled from the first box and 8 tube cores were sampled from the second box. For the station OSK 1, four cores from box 1 and three cores from box 2 were sliced directly on the boat (top 0–1 cm sediment surface) into 50-mL centrifuge tubes (Sarstedt), while for station OSK 2 four cores that were sampled from box 1 were sliced. The sediment samples were stored at − 20 °C and were later used for DNA extraction (OSK 1 n = 7, OSK 2 n = 4). The remaining sampled sediment cores that had not been sliced (n = 10 per station) were closed with rubber stoppers, kept at 8 °C overnight and later used for the 15NO3− IPT incubation to measure denitrification and DNRA rates. Bottom water (~ 25 L per station) was collected before the sediment collection using 10-L Niskin bottles connected to a Rosette sampler. This water was later used to submerse the sediment cores for the IPT incubation in the laboratory. In parallel, a CTD connected to the Rosette was used to measure in situ temperature, oxygen, and salinity profiles of the water column (SEABird SBE 911 plus). All samples for DNA extraction were transported back to the university on dry ice, and intact sediment cores were transported in a cooling box filled with ice packs. Surficial sediment samples (0–4 cm) for PCDD/Fs and metal analyses (As, Ba, Cd, Co, Cr, Cu, Hg, Ni, Pb, V, Zn) were collected on the same day from additional boxes containing bottom water and sediment (one box was collected from each station using the box corer as described above). The boxes were part of another experiment and were kept aerated until November 18 at 8–16 °C before being subsampled. A total of four randomly selected locations inside each box were subsampled using a syringe (5 mL sediment from each spot), pooled and stored at − 20 °C until further analysis.
An identical sampling approach using the same research vessel, sampling equipment, and type of material was conducted at the Swedish national marine monitoring reference site stations, near the Askö Laboratory station [35, 36], in the Uttervik Bay: station ASK (17 m water depth, WGS84 coordinates: Lat 58.843577, Long 17.544748; Fig. S1), on August 10 during 2020. There is no industry in this area and ASK has no known history of being affected by local metal pollution sources and previous Cd concentration measurements have shown Cd concentrations of ~ 19 µg kg−1 dry weight (dw) at the 0–1 cm sediment surface [24], which is well below the national threshold of 0.5 mg kg−1 dw set by the Swedish Environmental Protection Agency [37]. Similarly to OSK 1, two box cores were collected and 9 tube cores were subsampled from the first box while 8 tube cores were subsampled from the second box. Four of the cores from box 1 and three cores from box 2 were sliced directly on the boat to be used for DNA extraction (n = 7), while the remaining 10 cores were used for IPT incubation. Water column properties were determined and bottom water was sampled as described above. Finally, one additional box was collected, kept aerated, and subsampled for metals and PCDD/Fs as described above.
Denitrification and DNRA Rate Measurements
Following sampling, the ten collected cores that were not used for slicing were stored, kept in darkness at 10 °C, and submersed in a bath consisting of collected in situ bottom water that was aerated with air stones for 6 days. Externally driven magnetic bars inside the cores were used to continuously mix the water inside the cores to ensure aeration. To increase microbial activity during the IPT incubation, the temperature was changed to 13 °C until the IPT incubation started 5 days later.
The IPT incubation followed the experimental design described by Nielsen [38] and closely followed the protocol described in Broman et al. [24]. In brief, the cores were spiked with 15NO3 and pre-incubated for 1.5 h to allow 15N to homogenize with the in situ 14N inside the cores [39]. The cores were then closed using a rubber stopper and incubated for 6 h, followed by collection of water samples from each core for N2 analysis (n = 9 per station) and dissolved 15NH4+ (n = 9 per station). Full details of the methods used for the IPT incubation are available in the supplementary information.
Sediment Characteristics and Water Analyses
Sediment porosity was measured on a portion of sampled sediment collected in the field (0–1 cm) and was calculated by dividing the water content (sediment dried at 60 °C for 3 days) by the known bulk volume (5 mL). Organic carbon and nitrogen content were measured by acidifying dry sediment (20 mg) with HCl (1 M) over night at 60 °C to remove inorganic carbon and analyzed using a C/N elemental analyzer (Thermo Scientific Flash 2000). Samples for metal analyses were sent to ALS Scandinavia, Sweden and analyzed according to the standards ISO 11464:2006 [40] and SS-EN ISO 17294–2:2016 [41]. In brief, the sediment was dried, sieved (< 2 mm), heated and digested in 7 M HNO3, and analyzed by inductively coupled plasma (ICP) (see Table 1 for a list of metals that were measured). Because of the large difference in contaminants between the sites (i.e., much less at ASK), the ASK station required more sediment for PCDD/F analysis. Therefore, in addition to the samples collected with syringes, the ASK box was also subsampled on January 25 in five random locations with acrylic cores (inner diameter: 4.6 cm, length: 30 cm length) and the top 0–3 cm sliced (ca 50 mL sediment per slice) and pooled. Samples for PCDD/Fs were sent to OEKOMETRIC GmbH, Germany and analyzed according to the SS-EN 16,190:2019 standard [42]. In brief, samples were dried, extracted with toluene using pressurized liquid extraction (ASE 180 °C, 140 bar), multicolumn clean-up (multi-silica, ALOX), and analyzed using a gas chromatography high-resolution mass spectrometry (GC-HRMS) (see Table 2 for a list of dioxins and furans measured). A total of 17 congeners were analyzed known to be of toxic relevance and bio-accumulative by organisms in marine sediments [43].
Samples from the water phase of the cores were analyzed for NH4+ and NO2− + NO3− that were measured with standard colorimetric methods indophenol-blue and naphthalene-based azo dye, respectively [44], using a spectrophotometer.
DNA Extraction and Sequencing
DNA was extracted from 2 g of sediment using the DNeasy PowerMax Soil kit (Qiagen), and the DNA was stored at − 20 °C in 2 mL elution buffer (solution C6). DNA was precipitated and concentrated × 10 with NaCl, EtOH, and Tris as described in the DNeasy PowerMax Soil kit. The final DNA quality and quantity were determined with a NanoDrop 1000 and Qubit 1 (ThermoFisher), respectively. The extracted DNA was prepared with the TruSeq PCR-free kit (Illumina) and sequenced at Science for Life Laboratory at Stockholm on one NovaSeq6000 S4 lane using a 2 × 150 paired-end setup. Sequencing yielded on average 104.9 million reads (min: 80.9, max: 133.8). See Data S1 for a list of sample names and obtained number of sequences per sample. The raw sequence data has been uploaded to the NCBI BioProject PRJNA721298.
Bioinformatic Analysis
The sequence data was quality trimmed as described previously using SeqPrep and trimmomatic [7]. The functional gene annotation followed the SAMSA2 pipeline [45]. The taxonomic annotation was done on 16S rRNA reads extracted from the metagenomes using the software combo Kraken + Bracken2 [46, 47] as described in Broman et al. [48]. Full details of the bioinformatic analyses are available in the supplementary information.
Statistics
Shannon’s H alpha diversity index was calculated from the 16S rRNA gene data using the software Explicet by subsampling to the lowest sample size (16S rRNA gene data: 30,819 counts) and with bootstrap × 100 (with the mean reported). Beta diversity was tested using the Bray–Curtis dissimilarity index and Sørensen-Dice coefficient and pairwise PERMANOVA tests (9999 permutations) with Bonferroni corrected P values using the software past 4.05 [49]. For the beta diversity analyses, the input data was based on either relative abundance (bacteria and archaea, Bray–Curtis) or normalized read counts for the KEGG KO identifiers. Shapiro–Wilk test was used to check if data had a normal distribution using R 3.6.2 [50] with the shapiro.test function (R package: car 3.0–11 [51]) and the homogeneity of variance was analyzed with Levene’s test using the leveneTest function (R package: stats 3.0–4). One-way ANOVA and post hoc Tukey tests were used to test significant differences between the stations using R with the res.aov and TukeyHSD functions. Boxplots with jitter alongside one-way ANOVA and t-test were produced with the R package ggpubr 0.4.0. Comparisons of As, Cd, Cu, and Zn metal-related genes between stations were based on listed genes in Yan et al. [52]. The R package Hmisc 4.7.0 [53] was used to conduct Pearson correlations between the normalized read counts for nitrogen cycling genes, and the sum of the metal-related genes, and the sum of dioxin degradation genes. In addition, the edgeR package in R [54] was used to normalize counts (as CPM) and test for statistical differences (false discovery rate, FDR < 0.05) between KEGG KOs by using the raw count data as input. All data are reported as mean ± standard error.
Results
Field Conditions, Metal and PCDD/F Concentrations in the Sediments
The sampled polluted area in Oskarshamn (stations OSK 1 and OSK 2) and the national reference station (ASK) both had oxic bottom waters, salinity 6.27–7.21, and a temperature of 16.05–17.17 °C (Table 1). Dissolved NO2− + NO3− was higher in the bottom water at Oskarshamn compared to the reference station ASK (n = 3; one-way ANOVA, df = 5, F = 223.9, P < 0.001), while dissolved NH4+ concentrations in the bottom water were similar among stations (n = 3; one-way ANOVA, df = 5, F = 0.204, P = 0.675). The sediment porosity (Φ) was similar for each station, while sediment organic carbon (Corg) and nitrogen content (N) were different among all stations (n = 1 per station; Table 1).
Harbor sediments at OSK 1 and 2 were found to contain twice as high concentrations of As, Cu, Hg, Pb, and Zn, respectively, and seven times higher concentrations of Cd when compared with the reference site ASK (based on the average for OSK 1 and OSK 2, Zn was only higher at OSK 1; Table 1). The highest concentrations of Cd (3.0), Cu (86.5), Hg (1.4), Pb (10.40), and Zn (227.0) were found at OSK 1 (values shown as mg kg−1 dw). Compared with the two OSK stations, the sediment at ASK contained higher concentrations of Ba, Co, Cr, Ni, and V (Table 1). However, most of these metals at ASK were below what is classified as a high significant deviation from the national threshold by the Swedish Environmental Protection Agency, except for Zn and Cu at all stations. Notably, Cd, Hg, and Pb were categorized as “high” at the OSK stations; however, Cr was categorized as “high” at the ASK station (Table 1). For dioxin concentrations, taken together, there were no large differences between the two OSK stations as compared with ASK (on average 0.6–1.3 times difference per compound); however, concentrations were higher at OSK 1 than at OSK 2 (Table 2). In contrast, furan concentrations were on average 7 and 3 times higher at OSK 1 and OSK 2, respectively, compared with ASK (Table 2). The highest concentrations of furans were octachlorodibenzofuran (OCDF) and total HeptaCDF with OCDF at OSK 1 (693.0), OSK 2 (208.0), ASK (23.0), and total HeptaCDF at OSK 1 (276.0), OSK 2 (90.0), and ASK (26.0), respectively (all values shown as ng kg−1 dw).
Denitrification and DNRA Rates
The 15NO3− isotope labeling incubations showed that there was no significant difference in denitrification rates among the stations (OSK 1 189.1 ± 30.4, OSK 2 131.9 ± 23.4, and ASK 210.6 ± 33.0 µmol N m−2 d−1; one-way ANOVA, df = 25, F = 1.59, P = 0.23, n = 9 per station except OSK 1 n = 8; Fig. 1A). This represents 92.3, 92.6, and 93.7% denitrification of the total nitrate reduction, respectively (i.e., % denitrification = denitrification / (denitrification + DNRA)). Similarly, no significant difference in DNRA rates was found among the stations (OSK 1 15.0 ± 2.2, OSK 2 11.0 ± 2.4, and ASK 14.2 ± 2.0 µmol N m−2 d−1; one-way ANOVA, df = 26, F = 0.75, P = 0.48; n = 9 for each station; Fig. 1B). This represents 7.7, 7.4, and 6.3% DNRA of the total nitrate reduction, respectively (i.e., % DNRA = DNRA / (denitrification + DNRA)). See Data S2 for a full list of all data for each replicate. Even though no significant differences were found for denitrification and DNRA rates, the calculated denitrification derived from nitrification (D14N) was higher at the reference station ASK (96.9 ± 0.3%) compared with the polluted harbor stations OSK 1 (92.1 ± 0.7%) and OSK 2 (90.9 ± 0.8%) (one-way ANOVA tests, df = 16 and 17, F = 11.9 and 12.6, P < 0.01 for both OSK 1 and OSK 2; Fig. 1C).
Boxplots showing N2 production rates from 15NO3.− labeled incubations conducted on sediment sampled from the polluted harbor of Oskarshamn (stations OSK 1 and OSK 2) and a reference station Askö (station ASK). The subpanels show A denitrification rates; B DNRA rates; and C denitrification derived from nitrification (D14N). The middle line inside each box denotes the median, the top and bottom of each box represent the 25th and 75th quartile, while the whiskers represents the maximum and minimum values. Number of replicates n = 9 per station except for A OSK 1 denitrification n = 8 and C OSK 1 D14N n = 8
Nitrogen Cycling Pathways and Genetic Signatures
Metagenome data analysis resulted, on average, in 19.8 million reads (range: 14.7–25.6) per sample that could be classified to KEGG KO identifiers (assigned reads in MEGAN). After normalization (subsampling to lowest sample size), on average 173,000 reads could be assigned to KEGG KO identifiers belonging to the nitrogen metabolism pathway. A full list of KEGG nitrogen metabolism classifications with normalized read counts is available in Data S3. Full pathways for DNRA, assimilatory nitrate reduction, denitrification, N2 fixation, nitrification, and anammox were identified at all three stations (Fig. S2). Analysis of the Bray–Curtis dissimilarity index of all KEGG KO identifiers in the nitrogen metabolism map showed that the stations were different from each other (PERMANOVA, df = 17, pseudo-F = 22.17, P < 0.001; with all stations being different from each other when tested with pairwise comparison, Bonferroni corrected P values < 0.05).
The majority of nitrogen cycling genes that could be classified to taxa were attributed to different Proteobacteria classes (Alpha, Beta, Delta, Gamma), Bacteroidetes, and archaeal Thaumarchaeota at all stations (Data S4). Looking closer at abundances of genes for reductive N-cycling processes, differences were observed both for denitrification and DNRA. The statistical tests, edgeR, and one-way ANOVA with t-tests showed similar results and the one-way ANOVA tests are reported below (see methods for more details and Data S5 for edgeR results). Polluted harbor sediments OSK 1 and OSK 2 showed a lower relative abundance of narG, nirS, and nirK genes, as well as nosZ for OSK 1, when compared to the reference station ASK (Student’s t-test, P < 0.05; Fig. 2A–E). Analyses of differences in the denitrification pathways within the polluted stations showed that there was no difference in nitrate reduction genes between OSK 1 and OSK 2 (for example, the narG gene; Fig. 2A); nitrite reduction was significantly different only for nirS gene abundance but not nirK (with more nirS genes in OSK 2 compared to OSK 1; one-way ANOVA Student’s pairwise t-test, P < 0.05, Fig. 2B–C), while nitric oxide and nitrous oxide reduction (norB and nosZ, respectively) showed no difference between the two OSK sites (Fig. 2D–E). However, the nirS and nirK community composition was similar between OSK 1 and OSK 2 with most of the classified genes attributed to Gammaproteobacteria, Thaumarchaeota, and Chloroflexi (Fig. S3 and Data S4). In addition, more nirK genes were attributed to the archaeal phylum Thaumarchaeota at ASK when compared with both OSK 1 and OSK 2 (one-way ANOVA tests, F = 177 and 91, respectively, P < 0.01; Data S4). General differences in taxonomic profile attributed to these genes included a higher relative abundance of Nitrospirae carrying the narG gene at ASK, whereas at OSK 1 the narG gene was predominantly attributed to Deltaproteobacteria (Data S4). Most of the nosZ genes could not be classified (ca 60%) but the majority of the sequences that could be classified belonged to Bacteroidetes, with no difference among stations (Data S4).
Boxplots with jitter showing selected nitrogen cycling KEGG KO identifiers and their associated gene names, each sample is indicated as a dot (i.e., jitter). A shows nitrate reduction (narG gene), two genes for nitrite reduction B nirS and C nirK, D nitric oxide reduction (norB), E nitrous oxide reduction (nosZ), two genes for DNRA F nrfA and G nirB, and H nitrogen fixation (nifH). The data is based on normalized read counts, and the one-way ANOVA results show the results for the whole model (including all three stations), while the P value between stations is based on pairwise t-tests. P values < 0.05 have been marked with bold text. The middle line inside each box denotes the median, the top and bottom of each box represent the 25th and 75th quartile, while the whiskers represents the maximum and minimum values. Outliers are shown outside the whiskers if the distance is longer than 1.5 × box height. Number of replicates: ASK n = 7, OSK 1 n = 7, OSK 2 n = 4
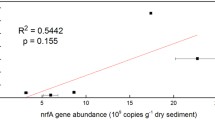
For sequence reads that could be matched to DNRA, there were more identified genes coding for nitrate reductase cytochrome C (nrfA) at OSK 1 compared with both ASK and OSK 2 (Student’s t-test, P < 0.05: Fig. 2F), while reads for nitrite reductase NADH (nirB), another enzyme in DNRA, were overrepresented at OSK 2 compared with ASK and OSK 1 (Student’s t-test, P < 0.05: Fig. 2G). Within taxa attributed to nrfA, the analyses showed that Bacteroidetes had a higher relative abundance at OSK 1 when compared with ASK and OSK 2 (one-way ANOVA tests, F = 248 and 11, respectively, P < 0.01; Fig. S3 and Data S4). Furthermore, taxa attributed to nirB were found to be dominated by Gammaproteobacteria at OSK 1 and OSK 2 but could not be further classified to family level (Data S4). Finally, we also observed a difference between stations in the nitrogen fixation gene nifH that was highest at OSK 1 (Fig. 2H), ammonia oxidation genes amoAB that were highest at ASK, and anammox genes hzs and hdh that had low counts (< 30) at all stations (Fig. S4). See supplementary information for full details of these additional results.
Metal and PCDD/F Resistance and Genetic Signatures
Both polluted harbor stations OSK 1 and OSK 2 were found to have higher numbers of reads attributed to several metal-related KEGG KOs compared with the reference station ASK, e.g., genes encoding for arsenate reductase (glutaredoxin/thioredoxin), P-type Cu2+ transporter, two-component system OmpR family heavy metal sensor histidine kinase CusS, and Zn2+/Cd2+-exporting ATPase (edgeR, FDR < 0.5; Fig. 3A). Furthermore, the relative abundance of reads attributed to some of these genes was higher at OSK 1 compared with OSK 2, including the genes coding for copper homeostasis protein, copper/silver efflux system protein, and Zn2+/Cd2+-exporting ATPase, respectively (Fig. 3A). Taxa at OSK 1 and OSK 2 that could be affiliated to the metal-related genes were most closely related to members of the order Desulfuromonadales (that was also attributed to N2-fixation, nifH) and were classified to, e.g., P-type Cu2+ transporter, two-component system OmpR family heavy metal sensor histidine kinase CusS, Zn2+/Cd2+-exporting ATPase, and copper/silver efflux system protein (Data S4). Several phyla and classes that were attributed to denitrification and DNRA genes were also affiliated with the metal-related genes included, e.g., Bacteroidetes, Chloroflexi, Alpha-, Delta-, and Gammaproteobacteria (Data S4). Examples of taxa that both denitrification (nirS or nirK) or DNRA (nrfA or nirB) and metal-related genes could be affiliated with included Pseudomonas, Thiotrichaceae, Gemmatimonadaceae, Rhodobacteraceae, Desulfobacterales, and Cytophagales (Data S4).
Heatmap of A KEGG KOs attributed to metal-related genes for As, Cd, Cu, and Zn for each station and replicate sample (denoted with a number on the top x-axis) and B KEGG KOs included in the dioxin degradation KEGG pathway, respectively. The list of selected genes for the metals is based on Yan et al. [52]. The data shown is based on normalized read counts. The edgeR results show which KEGG KOs were significantly higher between stations (FDR < 0.05). Note that the color gradients are based on the lowest and highest values per row
The dioxin degradation pathway included dibenzofuran dioxygenase enzymes involved in both PCDD and PCDF degradation with genes dxnA1, dxnA2, dbfA1, and dbfA2 (KEGG KOs: K22715, K22716, K14599, K14600, respectively). None of these KEGG KOs was, however, statistically significantly different among stations (Fig. 3B). Nor was there any clear pattern that showed that the harbor stations had a higher gene abundance for later steps in these pathways (Fig. 3B). Overall, reads attributed to PCDD and PCDF degradation were significantly fewer than those attributed to metal-related genes (~ 5000 reads per sample compared to ~ 65,000 attributed to the toxic metals). Taxa attributed to dibenzofuran dioxygenase enzymes included mainly Actinobacteria and Gammaproteobacteria with no further classification to class level (Data S4).
Pearson correlations based on the normalized read counts for the metal resistance, dioxin degradation, and the nitrogen cycling genes showed that narG, nirS, nirK, and nosZ were negatively correlated with the sum of all metal-related genes (r = − 0.50, − 0.70, − 0.91, − 0.83, all P < 0.05, respectively). In contrast, the genes norB, nrfA, and nirB were positively correlated with the metal-related genes (r = 0.85, 0.84, 0.67, all P < 0.05, respectively). The sum of the dioxin degradation genes did not correlate significantly with any of the nitrogen cycling genes.
Microbial Diversity and Community Composition
Considering the difference in pollution between the harbor stations and the national reference station, we also investigated whether there were differences in taxonomic diversity among the bacterial and archaeal communities. Shannon’s H diversity index was significantly lower at the reference ASK station (6.16 ± 0.05 Shannon’s H, n = 7) when compared with both OSK 1 (6.82 ± 0.04, n = 7) and OSK 2 (6.87 ± 0.04, n = 4) (tested on the 16S rRNA gene data; one-way ANOVA, df = 17, F = 82.4, P < 0.001; and with post hoc TukeyHSD multiple comparison test: P < 0.001 for both harbor stations when compared with ASK). There was, however, no difference in Shannon’s H diversity index between the two harbor stations OSK 1 and OSK 2 (one-way ANOVA with post hoc Tukey test, P = 0.81). The Bray–Curtis dissimilarity index was tested between all stations and showed that there was a significant difference in community composition (PERMANOVA, df = 17, pseudo-F = 20.98, P < 0.001). Pairwise comparison showed that the ASK station was different from both OSK 1 and OSK 2 (Bonferroni corrected P values < 0.05), and that OSK 1 and OSK 2 were different (Bonferroni corrected P values < 0.05).
Overall, the bacterial and archaeal communities were dominated by Proteobacteria (classes Gamma, Delta, and Alpha in order of highest relative abundance) at all three stations. Additional top dominant phyla included Bacteroidetes, Chloroflexi, Firmicutes, and Thaumarchaeota (Fig. 4). Closer analyses of some examples of the dominant genera that were different among stations showed that Pseudomonas had the highest relative abundance at ASK (on average 8.7%), followed by OSK 2 (6.0%) and OSK 1 (4.1%) (one-way ANOVA, df = 17, F = 37.96, P < 0.05; with post hoc TukeyHSD test, P < 0.05 for all three stations; Fig. 5). An uncultured genus belonging to the family Anaerolineaceae had a higher relative abundance at station ASK (5.0%) when compared with OSK 2 (3.8%) (F = 5.089, P < 0.05, TukeyHSD, P < 0.05) but there was no major difference between OSK 1 (4.4%) and OSK 2. The MND1 genus belonging to the family Nitrosomonadaceae was significantly different among all stations, with the lowest relative abundance in the polluted sediment at OSK 1 (0.7%) and OSK 2 (1.7%) compared with the reference station ASK (8.1%) (F = 811.3, P < 0.001; TukeyHSD, P < 0.05; Fig. 5). An uncultured genus belonging to the family Thiotrichaceae was also significantly different among all stations, most abundant at OSK 2 (4.2%) followed by OSK 1 (2.7%), and lowest at ASK (0.3%) (F = 73.61, P < 0.001; TukeyHSD, P < 0.05; Fig. 5).
Stacked bars showing the bacterial and archaeal community composition as relative abundance (%) detected in the collected sediment samples (top 1 cm sediment slice). The data is based on metagenome extracted 16S rRNA gene sequences classified against the SILVA database (using the Kraken2 + Bracken2 software combo). OSK 1 = Oskarshamn station 1, OSK 2 = Oskarshamn station 2, ASK = Askö station. The numbers in parenthesis denote the replicate number (individual sediment cores)
Heatmap showing the dominant bacterial and archaeal taxa in the sediment based on the metagenome 16S rRNA gene data. The heatmap shows data based on lowest classified taxonomic level (i.e., genera). The data was delimited to genera > 1% (average of all samples). OSK 1 = Oskarshamn station 1, OSK 2 = Oskarshamn station 2, ASK = Askö station. The numbers in parenthesis denote the replicate number (individual sediment cores). The labels inside the heatmap denote the relative abundances (%), with dark cells having white labels to enhance reading clarity
Moreover, the class Dehalococcoidia that consists of known bacterial members capable of carrying out dechlorination had a higher relative abundance at the OSK 1 and OSK 2 stations when compared with ASK (one-way ANOVA tests, df = 13 and 10, F = 138.5 and 50.0, P < 0.01 for both, respectively). The relative abundance of Dehalococcoidia was, however, low at all stations with ASK 0.01 ± 0.02%, OSK 1 0.27 ± 0.05%, and OSK 2 0.36 ± 0.15%. A full list of all taxonomy classifications with read counts can be found in Data S6.
Discussion
This study shows that the benthic microbial community from the historically polluted sediment in Oskarshamn’s outer harbor has adapted to the metal pollution. These results reflect the harbor’s history of pollution over the last century [55], with historic emissions of contaminants derived from various industrial processes, e.g., a copper plant operated between 1918 and 1969, and a battery factory (since 1917, still in operation) [27]. While the inner harbor has recently been remediated through suction-dredging to decrease the level of pollutants [29,30,31], the outer harbor remains unremedied and, as this study shows, is still polluted with high levels of metals and PCDD/Fs. Concentrations of Cd, Cu, Hg, and Pb in the harbor sediments were significantly higher compared with the national reference station near Askö (ASK), and significantly above the national thresholds [37]. Furans (PCDF) concentrations were 3–7 times higher at the harbor stations (OSK) compared with ASK. Calculated PCDD/F toxic equivalencies (ng kg−1 TEQ dw) showed that OSK 1 (15 ng kg−1) was three times higher than at both OSK 2 (5 ng kg−1) and ASK (5 ng kg−1) [TEF values to calculate TEQ taken from 56]. Given the large differences in pollution between polluted OSK stations and the national reference station, and considering that mixed metal pollution, as well as petroleum exposure experiments, have been shown to decrease the rate of denitrification [22, 23], we expected lower denitrification and DNRA rates in the polluted harbor sediments. However, we detected no statistically significant differences in denitrification or DNRA rates among the stations.
Observed denitrification and DNRA rates (Fig. 1) were similar to those previously shown at unpolluted sites in the southern and central Baltic Sea [57, 58], which indicates that the polluted stations had denitrification and DNRA rates within the range generally found in the Baltic Sea. In more detail, for the two OSK stations, the average denitrification rates calculated per hour (5.5–7.9 µmol N m−2 h−1) and DNRA rates (0.5–0.6 µmol N m−2 h−1) are at similar levels to what has previously been reported for the Himmerfjärden area nearby the reference station Askö [58]. These values are also comparable to denitrification rates measured in the central Baltic Sea and in coastal sediments in the more saline southern Baltic Sea [57]. There was a time difference (of ca 6 weeks between June 27 and August 10), due to logistical reasons, between sampling the OSK and ASK stations; however, our results do not indicate a significant inhibition effect on denitrification and DNRA at the polluted Oskarshamn stations. Moreover, previous research has shown that denitrification rates at Askö station were significantly lower in June than in August [58]; thus, denitrification and DNRA rates would have been expected to be ~ 3 times lower at Askö if the station had been sampled 6 weeks earlier (i.e., in June), further supporting that inhibition was only minor at the OSK stations. In this study, we investigated two stations within one polluted harbor in the Baltic Sea, and potentially other polluted sites with different environmental conditions might yield different results when compared to the reference station. Nevertheless, the high concentrations of contaminations at our polluted sites indicate that our results are likely relevant for other polluted coastal areas. Possibly anaerobic oxidation of ammonium (anammox) rates, which were not analyzed in this study, could contribute to the total nitrate reduction but since several previous studies have shown that anammox activity is typically of minor importance in shallow estuarine and coastal sediments [59], as shown at, for example, Askö [58], the process is not expected to be significant at the Oskarshamn stations. Indeed, the metagenome data indicated a very low capacity for anammox in the studied sediments, with < 30 read counts at ASK and < 15 counts at respective OSK station. Anammox rate measurements to estimate its contribution to total nitrate reduction would be relevant; however, in studies of polluted sediments at deeper water depths, this process has been shown to be of less importance below 100 m water depth [59]. To summarize, our findings suggest that the long-term metal and organochloride pesticide pollution did not have any significant inhibitory effect on reductive N cycling processes at the time of sampling.
The genetic adaptation of the microbial communities at OSK 1 and 2 to the high concentrations of pollutants was revealed by the metagenome results. This was evident by the gene data with an overrepresentation of metal resistance genetic prerequisites at OSK 1 and 2, and the presence of taxa that could be matched to both denitrification/DNRA genes and the metal-related genes, such as Thiotrichaceae and Pseudomonas. The family Thiotrichaceae, which was significantly more abundant in the polluted sediments, has previously been described to include members with genetic prerequisites for both metal tolerance and DNRA production [9, 60,61,62]. The genus Pseudomonas, which was present at all stations, is known to include species with both metal tolerance and denitrification capacity [63,64,65]. However, with the data obtained in this study, it was not possible to differentiate among species of this genus. Results of metal resistance capacity complement previous findings, which showed that Baltic Sea deep water sediments were overrepresented in Cd-Zn-Hg resistance pathways in comparison with several other deep waters and sediments globally [66, 67], and suggest a broader genetic adaptation to metal pollution in the Baltic Sea. Even though we detected differences in denitrification gene abundances (such as nirK and nirS) among stations, these genes have previously been described to also be present in, e.g., anaerobic ammonium-oxidizing bacteria and methane-oxidizing bacteria [68], which we identified at all stations, possibly partly explaining less correlation with denitrification rates. Here we used the KEGG KO annotations from software combo DIAMOND + MEGAN [69]. We have previously used these tools in conjunction with metagenome assemblies, RNA-seq, and measured environmental variables to validate our findings [70,71,72]. Differences observed in N-cycling genes in our dataset (e.g., norB being high at OSK while nirS being high at ASK) are therefore potentially a result of the environmental conditions at each station selecting for specific microbes carrying these genes. Our metagenomics results furthermore confirm that the abundance of genes encoding proteins of enzymes involved in nitrate reduction does not directly relate to the biogeochemical rates, as previously shown for several biogeochemical processes [73].
In addition to microbial community traits, environmental conditions at sampled sites may explain denitrification rates. For example, differences in organic matter content, grain size, and redox potential are more likely than metals to influence denitrification rates [74, 75]. The sampled stations were all oxic at the sampled depth, but presented natural differences in, e.g., temperature, salinity, and marked differences in quantity/quality of organic matter (see Corg and N data). Especially this latter factor may explain a part of the variation in our data similarly to what has been previously observed in Baltic Sea sediments [76]. Since there was more Corg and N (as well as a higher Corg/N ratio) at the polluted sites than at the reference site, we argue that the stimulatory effect of more C and N on denitrification rates might have been negatively offset by the high levels of contaminants. Even though NO2− + NO3− in the bottom water were low at all stations, it was slightly higher at the polluted sites (on average 1.5 µM) compared with the reference site (0.6 µM), which might explain small differences in nitrogen cycling processes between the sites. While O2 and NH4+ concentrations at the bottom water revealed oxic conditions, O2, H2S, and NH4+ concentrations in the sediment surface were not determined and differences in the sediment redox profile among stations might also have influenced denitrification and DNRA rates. However, we have a good grasp of how much these sediments consume in terms of O2 consumption over time since extensive microprofiling in all these sediment has been conducted in our previous works [58, 76,77,78,79]. When pre-incubating the sediments for 1.5 h followed by 6 h at 13 °C, these sediments are estimated to consume on average 27% of the initial oxygen available in the cores, meaning that no cores went hypoxic. Also, due to the very low oxygen penetration depth of only a mm [see, for example, 78], N2 production rates were likely linear over time. Furthermore, we observed higher denitrification derived from nitrification (i.e., D14N) at ASK compared with OSK 1 and OSK 2, which was also in accordance with a higher relative abundance of amoABC gene counts, and Nitrosomonadaceae bacteria at ASK. Even though the cores were “flooded” with N during the IPT incubation (an increase from ~ 0.6 µM to ~ 20 µM NO3−), this could indicate that nitrification limitation is regulating denitrification rates at ASK even under ambient NO3− concentrations [80].
Even though there was an obvious difference in metal resistance genes between the polluted stations and the national reference station, the metagenome data showed that the capacity to degrade PCDD/Fs in the sediment was limited at all three studied stations. The latter result was less expected as there are known specific bacterial populations, such as Pseudomonas species, that have been shown to tolerate PCDD addition [81], and some bacteria, such as Dehalococcoides, degrade PCDD/Fs (so-called dehalogenation) using hydrogen as an electron donor [82,83,84]. That there was no observed difference in denitrification rates among the studied sites suggests that the PCDD/Fs not noteworthy influenced the nitrate-reducing populations. Explanations for a limited capacity to degrade PCDD/Fs in our dataset include, e.g., that (1) these substances have a low solubility and therefore sorb to particles [85] resulting in less available substances for the microbial community and (2) bacterial transformation of PCDD/F depends on H2 as an electron donor [82, 83], which at the high concentration of sulfate and abundance of sulfate-reducing bacteria (that consume H2) possibly made degradation of PCDD/Fs unfavorable [83]. This was also indicated by a low relative abundance attributed to Dehalococcoides (< 0.1% per station, Data S6). Potentially, results would be different in freshwater systems where H2 might be more readily available at the sediment surface. Our results therefore indicate that microbial transformation of PCDD/Fs was likely not a major metabolic process at any of the studied coastal sites.
Conclusions
Our findings indicate that benthic denitrification and DNRA do not seem to be affected by historic contaminant emissions of metals and dioxins in Baltic Sea sediment collected during the summer. We found that the polluted sediment at the outer harbor of Oskarshamn holds a N-cycling microbial community that is adapted to pollution. This was indicated by an overrepresentation of metal resistance genes, some of which could be matched to putative denitrifiers, which might partly explain the denitrification rate results. However, the same pattern was not observed for PCDD/Fs degradation genes, which might be due to the fact that the dioxins are adsorbed to sediment particles and have a relatively low bioavailability, or due to sulfate-reducing bacteria competing for H2 as an electron donor (that is also used in bacterial transformation of PCDD/Fs). Even though the polluted sediment revealed less sequences attributed to denitrification genes, such as nirK, nirS, and nosZ, this was not reflected in the N-cycling rates when compared to a national reference station and other sites in the Baltic Sea. Our results of denitrification/DNRA rates and metal genes indicate an important adaptation of nitrate-reducing bacteria to metals resulting in a more metal-tolerant microbial community with persistent nitrate reduction in the polluted harbor sediment. Consequently, the microbial N-cycling might be more impacted by the access to organic carbon and nutrients than by pollution of metals and dioxins.
Data Availability
All data supporting this study is available in the published article or supplementary information. The raw sequencing data has been uploaded to the NCBI BioProject PRJNA721298.
References
Galloway JN, Dentener FJ, Capone DG, Boyer EW, Howarth RW, Seitzinger SP, Asner GP, Cleveland CC, Green PA, Holland EA, Karl DM, Michaels AF, Porter JH, Townsend AR, Vöosmarty CJ (2004) Nitrogen cycles: past, present, and future. Biogeochemistry 70:153–226. https://doi.org/10.1007/s10533-004-0370-0
Lotze HK, Lenihan HS, Bourque BJ, Bradbury RH, Cooke RG, Kay MC, Kidwell SM, Kirby MX, Peterson CH, Jackson JBC (2006) Depletion, degradation, and recovery potential of estuaries and coastal seas. Science 312:1806–1809. https://doi.org/10.1126/science.1128035
Carstensen J, Andersen JH, Gustafsson BG, Conley DJ (2014) Deoxygenation of the Baltic Sea during the last century. Proc Natl Acad Sci USA 111:5628–5633. https://doi.org/10.1073/pnas.1323156111
Cornwell JC, Kemp WM, Kana TM (1999) Denitrification in coastal ecosystems: methods, environmental controls, and ecosystem level controls, a review. Aquat Ecol 33:41–54. https://doi.org/10.1023/A:1009921414151
Payne WJ (1973) Reduction of nitrogenous oxides by microorganisms. Bacteriol Rev 37:409–452. https://doi.org/10.1128/br.37.4.409-452.1973
Hietanen S, Jäntti H, Buizert C, Jürgens K, Labrenz M, Voss M, Kuparinen J (2012) Hypoxia and nitrogen processing in the Baltic Sea water column. Limnol Oceanogr 57:325–337
Broman E, Zilius M, Samuiloviene A, Vybernaite-Lubiene I, Politi T, Klawonn I, Voss M, Nascimento FJA, Bonaglia S (2021) Active DNRA and denitrification in oxic hypereutrophic waters. Water Research 194:116954. https://doi.org/10.1016/j.watres.2021.116954
Mohan SB, Schmid M, Jetten M, Cole J (2004) Detection and widespread distribution of the nrfA gene encoding nitrite reduction to ammonia, a short circuit in the biological nitrogen cycle that competes with denitrification. FEMS Microbiol Ecol 49:433–443. https://doi.org/10.1016/j.femsec.2004.04.012
Pandey CB, Kumar U, Kaviraj M, Minick KJ, Mishra AK, Singh JS (2020) DNRA: a short-circuit in biological N-cycling to conserve nitrogen in terrestrial ecosystems. Science of The Total Environment 738:139710. https://doi.org/10.1016/j.scitotenv.2020.139710
Burdige DJ (2006) Geochemistry of marine sediments. PRINCETON University Press
Griffiths JR, Kadin M, Nascimento FJA, Tamelander T, Törnroos A, Bonaglia S, Bonsdorff E, Brüchert V, Gårdmark A, Järnström M, Kotta J, Lindegren M, Nordström MC, Norkko A, Olsson J, Weigel B, Žydelis R, Blenckner T, Niiranen S, Winder M (2017) The importance of benthic–pelagic coupling for marine ecosystem functioning in a changing world. Glob Change Biol 23:2179–2196. https://doi.org/10.1111/gcb.13642
Breitburg D, Levin LA, Oschlies A, Grégoire M, Chavez FP, Conley DJ, Garçon V, Gilbert D, Gutiérrez D, Isensee K, Jacinto GS, Limburg KE, Montes I, Naqvi SWA, Pitcher GC, Rabalais NN, Roman MR, Rose KA, Seibel BA, Telszewski M, Yasuhara M, Zhang J (2018) Declining oxygen in the global ocean and coastal waters. Science 359:eaam7240. https://doi.org/10.1126/science.aam7240
Howarth RW, Marino R (2006) Nitrogen as the limiting nutrient for eutrophication in coastal marine ecosystems: evolving views over three decades. Limnol Oceanogr 51:364–376. https://doi.org/10.4319/lo.2006.51.1_part_2.0364
Qian Y, Zhang W, Yu L, Feng H (2015) Metal pollution in coastal sediments. Current Pollution Reports 1:203–219. https://doi.org/10.1007/s40726-015-0018-9
Czuczwa JM, Hites RA (1984) Environmental fate of combustion-generated polychlorinated dioxins and furans. Environ Sci Technol 18:444–450
Ryu J, Khim JS, Kang S-G, Kang D, Lee C-h, Koh C-h (2011) The impact of heavy metal pollution gradients in sediments on benthic macrofauna at population and community levels. Environ Pollut 159:2622–2629. https://doi.org/10.1016/j.envpol.2011.05.034
Kang T, Oh JH, Hong J-S, Kim D (2018) Response of intertidal meiofaunal communities to heavy metal contamination in laboratory microcosm experiments. J Coast Res 85:361–365
Somerfield P, Gee J, Warwick R (1994) Soft sediment meiofaunal community structure in relation to a long-term heavy metal gradient in the Fal estuary system. Mar Ecol Prog Ser 105:79–88
Iburg S, Nybom I, Bonaglia S, Karlson AM, Sobek A, Nascimento FJ (2020) Organic contaminant mixture significantly changes microbenthic community structure and increases the expression of PAH degradation genes. Front Environ Sci 8:128
Nies DH (1999) Microbial heavy-metal resistance. Appl Microbiol Biotechnol 51:730–750. https://doi.org/10.1007/s002530051457
Ma J, Ullah S, Niu A, Liao Z, Qin Q, Xu S, Lin C (2021) Heavy metal pollution increases CH4 and decreases CO2 emissions due to soil microbial changes in a mangrove wetland: microcosm experiment and field examination. Chemosphere 269:128735. https://doi.org/10.1016/j.chemosphere.2020.128735
Sakadevan K, Zheng H, Bavor HJ (1999) Impact of heavy metals on denitrification in surface wetland sediments receiving wastewater. Water Sci Technol 40:349–355. https://doi.org/10.1016/S0273-1223(99)00471-0
Liu Y, Liu Y, Zhou H, Li L, Zheng J, Zhang X, Zheng J, Pan G (2016) Abundance, composition and activity of denitrifier communities in metal polluted paddy soils. Sci Rep 6:19086. https://doi.org/10.1038/srep19086 (https://www.nature.com/articles/srep19086#supplementary-information)
Broman E, Motwani NH, Bonaglia S, Landberg T, Nascimento FJA, Sjöling S (2019) Denitrification responses to increasing cadmium exposure in Baltic Sea sediments. Aquat Toxicol 217:105328. https://doi.org/10.1016/j.aquatox.2019.105328
Guo Q, Li N, Bing Y, Chen S, Zhang Z, Chang S, Chen Y, Xie S (2018) Denitrifier communities impacted by heavy metal contamination in freshwater sediment. Environ Pollut 242:426–432. https://doi.org/10.1016/j.envpol.2018.07.020
Lindgren JF, Hassellöv I-M, Dahllöf I (2014) PAH effects on meio- and microbial benthic communities strongly depend on bioavailability. Aquat Toxicol 146:230–238. https://doi.org/10.1016/j.aquatox.2013.11.013
Tobiasson S, Andersson S (2013) Avgångshastighet för metaller och organiska miljögifter i förorenade sediment från Oskarshamns hamn, 2011–2013. Rapport 2013:9. Linnéuniversitetet, Institutionen för biologi och miljö
Bank A (2005) Preliminär miljö- och hälsoriskbedömning av föroreningar i sediment inom hamnbassängen. Rapport nr Oskarshamns hamn 2004:14. Oskarshamns kommun, 1–30
Fathollahzadeh H, Kaczala F, Bhatnagar A, Hogland W (2015) Significance of environmental dredging on metal mobility from contaminated sediments in the Oskarshamn Harbor, Sweden. Chemosphere 119:445–451. https://doi.org/10.1016/j.chemosphere.2014.07.008
Oskarshamns Kommun (2016) Saneringen av Oskarshamns Hamnbassäng: Projekt- och erfarenhetsrapport 1996–2015. In: Schultzén, J (ed.), Oskarshamn, 1–96
Jönsson BL (2020) Information om saneringen av Oskarshamns Hamnbassäng: Nyhetsbrev 19 - April 2020. Oskarshamn Kommun
Rämö R, Bonaglia S, Nybom I, Kreutzer A, Witt G, Sobek A, Gunnarsson JS (2022) Sediment remediation using activated carbon: effects of sorbent particle size and resuspension on sequestration of metals and organic contaminants. Environ Toxicol Chem 41:1096–1110. https://doi.org/10.1002/etc.5292
Björinger P (2011) Sedimentprovtagning delområde E (Djuphåla). O-hamn 2011:13 NIRAS Environment
Blomqvist S, Ekeroth N, Elmgren R, Hall PO (2015) Long overdue improvement of box corer sampling. Mar Ecol Prog Ser 538:13–21
Blomqvist M, Leonardsson K (2016) A probability based index for assessment of benthic invertebrates in the Baltic Sea. WATERS report.
Leonardsson K, Blomqvist M, Rosenberg R (2009) Theoretical and practical aspects on benthic quality assessment according to the EU-Water Framework Directive–examples from Swedish waters. Mar Pollut Bull 58:1286–1296
Swedish EPA (2000) Environmental quality criteria: coasts and seas. Swedish Environ Protection Agency Report 5052:138
Nielsen LP (1992) Denitrification in sediment determined from nitrogen isotope pairing. FEMS Microbiol Lett 86:357–362. https://doi.org/10.1016/0378-1097(92)90800-4
De Brabandere L, Bonaglia S, Kononets MY, Viktorsson L, Stigebrandt A, Thamdrup B, Hall POJ (2015) Oxygenation of an anoxic fjord basin strongly stimulates benthic denitrification and DNRA. Biogeochemistry 126:131–152. https://doi.org/10.1007/s10533-015-0148-6
International Organization for Standardization (2006) Soil quality — pretreatment of samples for physico-chemical analysis. ISO 11464:2006.
Swedish Institute for Standards (2016) Water quality - application of inductively coupled plasma mass spectrometry (ICP-MS) - part 2: determination of selected elements including uranium isotopes. vol. SS-EN ISO 17294–2:2016. Swedish Standard
Swedish Institute for Standards (2019) Soil, treated biowaste and sludge - determination of dioxins and furans and dioxin-like polychlorinated biphenyls by gas chromatography with high resolution mass selective detection (HR GC-MS). vol. SS-EN 16190:2019. Swedish Standard, 56
Schaanning MT, Beylich B, Gunnarsson JS, Eek E (2021) Long-term effects of thin layer capping in the Grenland fjords, Norway: reduced uptake of dioxins in passive samplers and sediment-dwelling organisms. Chemosphere 264:128544. https://doi.org/10.1016/j.chemosphere.2020.128544
Valderrama JC (1995) Methods of nutrient analysis. In: Hallegraeff GM, ADaCA (ed.) Manual on harmful marine microalgae. Intergovernmental Oceanographic Commision of UNESCO, Paris, 251–268
Westreich ST, Treiber ML, Mills DA, Korf I, Lemay DG (2018) SAMSA2: a standalone metatranscriptome analysis pipeline. BMC Bioinformatics 19:175–175. https://doi.org/10.1186/s12859-018-2189-z
Wood DE, Salzberg SL (2014) Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol 15:1–12
Lu J, Breitwieser FP, Thielen P, Salzberg SL (2017) Bracken: estimating species abundance in metagenomics data. PeerJ Computer Science 3:e104
Broman E, Bonaglia S, Norkko A, Creer S, Nascimento FJ (2020) High throughput shotgun sequencing of eRNA reveals taxonomic and derived functional shifts across a benthic productivity gradient. Mol Ecol 30:3023–3039
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.
Fox J, Weisberg S, Adler D, Bates D, Baud-Bovy G, Ellison S, Firth D, Friendly M, Gorjanc G, Graves S (2012) Package ‘car’. Vienna: R Foundation for Statistical Computing: 16
Yan C, Wang F, Geng H, Liu H, Pu S, Tian Z, Chen H, Zhou B, Yuan R, Yao J (2020) Integrating high-throughput sequencing and metagenome analysis to reveal the characteristic and resistance mechanism of microbial community in metal contaminated sediments. Sci Total Environ 707:136116. https://doi.org/10.1016/j.scitotenv.2019.136116
Harrell Jr FE, Dupont C (2008) Hmisc: Harrell miscellaneous. R package version 3
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Oskarshamns Kommun (2005) Huvudstudierapport: Sanering av hamnbassängen i Oskarshamn. In: Bank, A, Carlsson, B (eds.), Oskarshamn, 1–77
Van den Berg M, Birnbaum LS, Denison M, De Vito M, Farland W, Feeley M, Fiedler H, Hakansson H, Hanberg A, Haws L (2006) The 2005 World Health Organization reevaluation of human and mammalian toxic equivalency factors for dioxins and dioxin-like compounds. Toxicol Sci 93:223–241
Deutsch B, Forster S, Wilhelm M, Dippner JW, Voss M (2010) Denitrification in sediments as a major nitrogen sink in the Baltic Sea: an extrapolation using sediment characteristics. Biogeosciences 7:3259–3271. https://doi.org/10.5194/bg-7-3259-2010
Bonaglia S, Deutsch B, Bartoli M, Marchant HK, Brüchert V (2014) Seasonal oxygen, nitrogen and phosphorus benthic cycling along an impacted Baltic Sea estuary: regulation and spatial patterns. Biogeochemistry 119:139–160. https://doi.org/10.1007/s10533-014-9953-6
Thamdrup B (2012) New pathways and processes in the global nitrogen cycle. Annu Rev Ecol Evol Syst 43:407–428. https://doi.org/10.1146/annurev-ecolsys-102710-145048
Zárate A, Dorador C, Valdés J, Molina V, Icaza G, Pacheco AS, Castillo A (2021) Benthic microbial diversity trends in response to heavy metals in an oxygen-deficient eutrophic bay of the Humboldt current system offshore the Atacama Desert. Environ Pollut 286:117281. https://doi.org/10.1016/j.envpol.2021.117281
Tsai Y-P, You S-J, Pai T-Y, Chen K-W (2005) Effect of cadmium on composition and diversity of bacterial communities in activated sludges. Int Biodeterior Biodegradation 55:285–291. https://doi.org/10.1016/j.ibiod.2005.03.005
Prokopenko MG, Hirst MB, De Brabandere L, Lawrence DJP, Berelson WM, Granger J, Chang BX, Dawson S, Crane Iii EJ, Chong L, Thamdrup B, Townsend-Small A, Sigman DM (2013) Nitrogen losses in anoxic marine sediments driven by Thioploca–anammox bacterial consortia. Nature 500:194–198. https://doi.org/10.1038/nature12365
Zhang N, Chen H, Lyu Y, Wang Y (2019) Nitrogen removal by a metal-resistant bacterium, Pseudomonas putida ZN1, capable of heterotrophic nitrification–aerobic denitrification. J Chem Technol Biotechnol 94:1165–1175. https://doi.org/10.1002/jctb.5863
Raja CE, Anbazhagan K, Selvam GS (2006) Isolation and characterization of a metal-resistant Pseudomonas Aeruginosa strain. World J Microbiol Biotechnol 22:577–585. https://doi.org/10.1007/s11274-005-9074-4
Carlson CA, Ingraham JL (1983) Comparison of denitrification by Pseudomonas stutzeri, Pseudomonas aeruginosa, and Paracoccus denitrificans. Appl Environ Microbiol 45:1247–1253. https://doi.org/10.1128/aem.45.4.1247-1253.1983
Thureborn P, Franzetti A, Lundin D, Sjöling S (2016) Reconstructing ecosystem functions of the active microbial community of the Baltic Sea oxygen depleted sediments. PeerJ 4:e1593
Thureborn P, Lundin D, Plathan J, Poole AM, Sjoberg BM, Sjoling S (2013) A metagenomics transect into the deepest point of the Baltic Sea reveals clear stratification of microbial functional capacities. PLOS One 8:e74983. https://doi.org/10.1371/journal.pone.0074983
Kuypers MMM, Marchant HK, Kartal B (2018) The microbial nitrogen-cycling network. Nat Rev Microbiol 16:263–276. https://doi.org/10.1038/nrmicro.2018.9
Bağcı C, Patz S, Huson DH (2021) DIAMOND+MEGAN: fast and easy taxonomic and functional analysis of short and long microbiome sequences. Current Protocols 1:e59. https://doi.org/10.1002/cpz1.59
Broman E, Sun X, Stranne C, Salgado MG, Bonaglia S, Geibel M, ..., Nascimento FJ (2020) Low abundance of methanotrophs in sediments of shallow boreal coastal zones with high water methane concentrations. Front Microbiol 11:1536
Broman E, Bonaglia S, Holovachov O, Marzocchi U, Hall PO, Nascimento FJ (2020) Uncovering diversity and metabolic spectrum of animals in dead zone sediments. Communications Biology 3:106. https://doi.org/10.1038/s42003-020-0822-7
Broman E, Izabel-Shen D, Rodríguez-Gijón A, Bonaglia S, Garcia SL, Nascimento FJA (2022) Microbial functional genes are driven by gradients in sediment stoichiometry, oxygen, and salinity across the Baltic benthic ecosystem. Microbiome 10:126. https://doi.org/10.1186/s40168-022-01321-z
Rocca JD, Hall EK, Lennon JT, Evans SE, Waldrop MP, Cotner JB, Nemergut DR, Graham EB, Wallenstein MD (2015) Relationships between protein-encoding gene abundance and corresponding process are commonly assumed yet rarely observed. ISME J 9:1693–1699. https://doi.org/10.1038/ismej.2014.252
Magalhães C, Costa J, Teixeira C, Bordalo AA (2007) Impact of trace metals on denitrification in estuarine sediments of the Douro River estuary, Portugal. Mar Chem 107:332–341. https://doi.org/10.1016/j.marchem.2007.02.005
Wittorf L, Roger F, Alsterberg C, Gamfeldt L, Hulth S, Sundbäck K, Jones CM, Hallin S (2020) Habitat diversity and type govern potential nitrogen loss by denitrification in coastal sediments and differences in ecosystem-level diversities of disparate N2O reducing communities. FEMS Microbiol Ecol 96:fiaa091
Albert S, Bonaglia S, Stjärnkvist N, Winder M, Thamdrup B, Nascimento FJA (2021) Influence of settling organic matter quantity and quality on benthic nitrogen cycling. Limnol Oceanogr 66:1882–1895. https://doi.org/10.1002/lno.11730
Bonaglia S, Nascimento FJA, Bartoli M, Klawonn I, Brüchert V (2014) Meiofauna increases bacterial denitrification in marine sediments. Nat Commun 5:5133. https://doi.org/10.1038/ncomms6133
Bonaglia S, Rämö R, Marzocchi U, Le Bouille L, Leermakers M, Nascimento FJA, Gunnarsson JS (2019) Capping with activated carbon reduces nutrient fluxes, denitrification and meiofauna in contaminated sediments. Water Res 148:515–525. https://doi.org/10.1016/j.watres.2018.10.083
Bonaglia S, Bartoli M, Gunnarsson JS, Rahm L, Raymond C, Svensson O, Yekta SS, Brüchert V (2013) Effect of reoxygenation and Marenzelleria spp. bioturbation on Baltic Sea sediment metabolism. Mar Ecol Prog Ser 482:43–55
Jenkins MC, Kemp WM (1984) The coupling of nitrification and denitrification in two estuarine sediments 1, 2. Limnol Oceanogr 29:609–619
Shiang FuQ, Barkovskii AL, Adriaens P (2005) Microbial dechlorination of dioxins in estuarine enrichment cultures: effects of respiratory conditions and priming compound on community structure and dechlorination patterns. Mar Environ Res 59:177–195. https://doi.org/10.1016/j.marenvres.2004.04.003
Holliger C, Wohlfarth G, Diekert G (1998) Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol Rev 22:383–398. https://doi.org/10.1111/j.1574-6976.1998.tb00377.x
Zanaroli G, Negroni A, Häggblom MM, Fava F (2015) Microbial dehalogenation of organohalides in marine and estuarine environments. Curr Opin Biotechnol 33:287–295. https://doi.org/10.1016/j.copbio.2015.03.013
Bunge M, Lechner U (2009) Anaerobic reductive dehalogenation of polychlorinated dioxins. Appl Microbiol Biotechnol 84:429–444. https://doi.org/10.1007/s00253-009-2084-7
Gruden CL, Fu QS, Barkovskii AL, Albrecht ID, Lynam MM, Adriaens P (2003) Dechlorination of sediment dioxins: catalysts, mechanisms, and implications for remedial strategies and dioxin cycling. In: Häggblom MM, Bossert ID (eds) Dehalogenation: microbial processes and environmental applications. Springer, US, Boston, MA, pp 347–372
Braker G, Zhou J, Wu L, Devol AH, Tiedje JM (2000) Nitrite reductase genes (nirK and nirS) as functional markers to investigate diversity of denitrifying bacteria in pacific northwest marine sediment communities. Appl Environ Microbiol 66:2096–2104
Throbäck IN, Enwall K, Jarvis Å, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417. https://doi.org/10.1016/j.femsec.2004.04.011
Acknowledgements
The authors acknowledge support from the National Genomics Infrastructure in Stockholm funded by Science for Life Laboratory, the Knut and Alice Wallenberg Foundation and the Swedish Research Council, and SNIC/Uppsala Multidisciplinary Center for Advanced Computational Science for assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. The bioinformatic analyses were enabled by resources in project SNIC 2021/22-137 provided by the Swedish National Infrastructure for Computing (SNIC) at UPPMAX, partially funded by the Swedish Research Council through grant agreement no. 2018-05973.
Funding
Open access funding provided by Stockholm University. The study was supported by the Foundation for Baltic and East European Studies Project 3150–3.1.1–2017 to SS, FN, and SB. The study was also co-funded by two grants to JG: the Swedish Environmental Protection Agency, PROMISE grant nr 2020–0002; and the Swedish Geotechnical Institute, TUFFO program, grant nr 1.1–1602-0106.
Author information
Authors and Affiliations
Contributions
Elias Broman: conceptualization, methodology, investigation, formal analysis, visualization, writing—original draft, writing—review and editing. Mohanad Abdelgadir: methodology, investigation. Stefano Bonaglia: conceptualization, methodology, investigation, formal analysis, funding acquisition. Sara C. Forsberg: investigation. Johan Wikström: investigation. Jonas S. Gunnarsson: conceptualization, methodology, investigation, review and editing, funding acquisition, supervision. Francisco J.A. Nascimento: conceptualization, funding acquisition, supervision. Sara Sjöling: conceptualization, funding acquisition, supervision, writing—review and editing. All authors improved and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Broman, E., Abdelgadir, M., Bonaglia, S. et al. Long-Term Pollution Does Not Inhibit Denitrification and DNRA by Adapted Benthic Microbial Communities. Microb Ecol 86, 2357–2372 (2023). https://doi.org/10.1007/s00248-023-02241-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02241-7