Abstract
Desertification leads to the extreme fragility of ecosystems and seriously threatens ecosystem functioning in desert areas. The planting of xerophytes, especially leguminous shrubs, is an effective and common means to reverse desertification. Soil microorganisms play a crucial role in nutrient cycling and energy flow in ecosystems. However, the effects of introducing leguminous shrubs on soil microbial diversity and the relevant mechanisms are not clear. Here, we employed the high-throughput absolute quantification 16S rRNA sequencing method to analyze the diversity of soil bacteria in sand-fixing areas of mixed shrublands with three combinations of shrubs, i.e., C. korshinskii × Corethrodendron scoparium (CaKCoS), C. korshinskii × Calligonum mongolicum (CaKCaM), and C. scoparium × C. mongolicum (CoSCaM), in the south of the Mu Us Sandy Land, China. This area suffered from moving dunes 20 years ago, but after introducing these shrubs to fix the dunes, the ecosystem was restored. Additionally, the effects of soil physicochemical properties on soil bacterial composition and diversity were analyzed with redundancy analysis (RDA) and structural equation modeling (SEM). It was found that the Shannon index of soil bacteria in CaKCoS was significantly higher than that in CaKCaM and CoSCaM, and the abundance of the dominant phyla, including Actinobacteria, Proteobacteria, Acidobacteria, Chloroflexi, Planctomycetes, Thaumarchaeota, Armatimonadetes, candidate_division_WPS-1, and Nitrospirae, increased significantly in CaKCoS and CaKCaM compared to that in CoSCaM. RDA showed that the majority of soil properties, such as total nitrogen (TN), available potassium (AK), N:P ratio, soil moisture (SM), and available phosphorus (AP), were important soil environmental factors affecting the abundance of the dominant phyla, and RDA1 and RDA2 accounted for 56.66% and 2.35% of the total variation, respectively. SEM showed that the soil bacterial α-diversity was positively affected by the soil organic carbon (SOC), N:P ratio, and total phosphorus (TP). Moreover, CaKCoS had higher SM, total carbon (TC), total potassium (TK), and AP than CaKCaM and CoSCaM. Collectively, these results highlight a conceptual framework in which the combination of leguminous shrubs can effectively drive soil bacterial diversity by improving soil physicochemical properties and maintaining ecosystem functioning during desertification reversal.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Desertification is a worldwide environmental problem and poses a great threat to ecosystems in dryland regions [91]. The increasing expansion of desertification not only limits economic development and reduces people’s quality of life but also causes vegetation degradation and ecological imbalance [10]. Desertification results in serious soil erosion, low vegetation coverage and sparse ground surface soil, and a continuous reduction in soil nutrients and moisture, resulting in barren sandy land and frequent sandstorms [46, 91]. These changes have strongly affected the ecological processes and functions of the sandy land ecosystem [77, 105, 111]. Thus, desertification control has captured great attention from all over the world. Different means of reversing desertification have been used in different regions of the world, for example, in the Mediterranean region, the establishment of shrubs is a crucial step in the process of desertification reversal [7, 47]. In Central Asian deserts, Haloxylon is widely used in contributing to desertification reversal due to its excellent sand-fixing properties [54]. In the Chihuahua deserts of South America, shrubs are often identified as pioneer “nurse” plants, forming species-rich patches [70]. In the sandy soil of the northern Sahara in Algeria, the richness of plant species increases with increasing vegetation coverage, and the protective effect of vegetation plays an important role in the restoration of sandy soil [48]. Following the conference on desertification control sponsored by the United Nations in 1977, the Chinese government has taken positive actions to prevent and control desertification, including ecological restoration projects such as the Three-North Shelter Forest Program, the “Grain for Green” Program, and the Natural Forest Protection Program, aimed at saving these fragile desertified areas [45]. For example, in northern China, the legume C. korshinskii was used to control large areas of sandy land through sand fixation restoration [83, 85]. After several stages of afforestation restoration practices, desertification in northern China has been effectively controlled and has achieved remarkable feedback [46].
Many studies have shown that afforestation projects, such as the introduction of drought-tolerant plants and the sand-fixing method, have effectively stopped the process of desertification and have gradually showed a trend of desertification reversal [47, 59, 100, 111, 114, 115]. Among them, straw checkerboard barrier measures have been widely applied to control the spread of moving dunes, allowing for sand fixation all over the world [58, 82]. One of the effective applications is the planting of xerophytic shrubs in straw checkerboards [36, 74, 104]. These sand-fixing patterns of combining xerophytic shrubs with straw checkerboards can effectively promote the physical and chemical properties of the soil [36, 74], and the nursing effect of the shrub canopy can enhance the development of herbaceous plant species [39], thus forming fascinating and resource-rich harbors known as “islands of fertility” [1, 6]. In extremely arid and semiarid areas, shrubs, especially nitrogen-fixing leguminous shrubs [40, 65], promote the formation of “islands of fertility” and improve soil nutrient accumulation, which is an extensive method of land degradation restoration [66, 87]. The leguminous shrub Retama sphaerocarpa is a commonly used species [55]. Nitrogen (N) fixation by leguminous shrubs, such as Caragana korshinskii [93, 95, 103], has been shown to improve soil N content and accumulate rich carbon (C) sources from leaf litter and root exudates. Plants of Caragana and Corethrodendron are xerophytic leguminous shrubs and are the dominant plants with the largest afforestation area in northern China; they are the most common shrubs used in sand fixation [5, 85]. Long-term afforestation practices and research have indicated that the leguminous shrub C. korshinskii has a strong ability for regrowth [17] and water acquisition [88], as well as excellent performance regarding sand fixation [18, 19]. Therefore, the sand-fixation mode of planting leguminous shrubs within straw checkerboards is of great significance for understanding the underlying mechanism of soil microecology in sandy areas with harsh environments.
Soils harbor diverse microbial communities, among which bacteria and fungi play an essential role in regulating C and N cycling processes for ecosystem functioning [28]. The diverse soil microbiota is frequently accompanied by different vegetation types in sandy land because aboveground vegetation can not only effectively improve soil texture, physicochemical properties, and the microenvironment but also input C stores from litter and root exudates [76, 95, 103]. Bacteria are an important part of the ecosystem that participate in nutrient cycling, fight against pathogenic microorganisms, improve the health of plants, and enhance ecological functions [22, 79]. In desert ecosystems, planting xerophytic plants can effectively enrich the soil bacterial community and diversity. Cao et al. [5] suggested that xerophytic plants, including C. microphylla, Artemisia halodendron, Hedysarum fruticosum, Pinus sylvestris var. mongolica, Populus simonii, and Salix gordejevii, have been artificially planted for sand fixation for 32 years, and the soil bacterial community composition was similar to that of the native community [110]. Zhou et al. [116] found that the rhizocompartments of two leguminous shrubs (Hedysarum scoparium and H. mongolicum) strongly influenced the soil bacterial communities and had filtration and enrichment effects on beneficial soil microbiota. Mediterranean wild leguminous shrubs, with their N-fixing rhizobia and mycorrhizal properties, play the most important role in contributing to land restoration and desertification reversal [7]. Most of the shrubs involved in sand-fixing afforestation are mainly legumes that can fix atmospheric N and may be the key factor affecting the formation of beneficial soil bacteria in the process of desertification reversal [60]. Evidence has been found that afforestation with the legumes C. korshinskii and Robinia pseudoacacia effectively increased the soil bacterial abundance and diversity compared with abandoned land on the Loess Plateau of China, and the Proteobacteria phylum gradually evolved into the dominant phylum [63], mainly due to the change in the soil N:P ratio [61, 64, 107, 108]. Therefore, the formation of soil nutrient pools may be mediated by legumes to a large extent and exert a large influence on the evolution of soil bacterial communities.
However, our knowledge about the effects of the combination of leguminous and nonleguminous shrubs on soil bacterial diversity during desertification reversal is still poor. Given the contribution of leguminous shrubs to soil properties and microhabitats in sandy land, as well as the excellent performance of the xeric leguminous shrubs in afforestation in sand-fixing areas, we hypothesized that the combination of leguminous shrubs in sand-fixing afforestation is more effective in ameliorating soil physical and chemical properties and enriching soil bacterial diversity than the combination of one leguminous and nonleguminous shrub after the long-term restoration of sandy land. The objective of this study is to test the following hypothesis: the combination of two leguminous shrubs effectively improves soil bacterial diversity by ameliorating soil environmental factors such as soil physical properties and nutrient contents during the process of desertification reversal. Our study contributes to addressing the issues of integrating strategies of shrub afforestation in nutrient-deficient sandy lands all over the world.
Materials and Methods
Study Area and Sampling Design
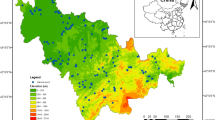
The study area was located in the Baijitan National Nature Reserve in Ningxia and belongs to the core area for sand fixation via straw checkerboards combined with shrubs in the south of the Mu Us Sandy Land, China (37°58′26″N, 106°24′03″E) (Fig. 1). The Mu Us Sandy Land is one of the largest sandy lands in China and is characterized by a fragile ecological environment and drought conditions due to desertification [91]. This area has a typical continental monsoon climate, and the annual average precipitation is approximately 230 mm and is mostly concentrated from July to September. The average annual accumulated temperature is approximately 3334.8 °C, with an average annual temperature of 6.7–8.8 ℃ and a frost-free period of 157 days. The soil type in this area is mainly aeolian sandy soil, and the vegetation is sparse. In 2001, moving dunes were covered with straw checkerboards (half-buried in sand) with each checkerboard covering an area of 1.0 m × 1.0 m. In the checkerboard, xeric shrubs dominated by C. korshinskii, C. scoparium, C. mongolicum, and C. fruticosum var. mongolicum were planted in 2002 with a row spacing of 2 m × 2 m. After years of combating desertification, the sand-fixing model of straw checkerboard-coupled vegetation was successfully implemented, and the moving dune was effectively fixed. Currently, it is considered a typical sand-fixing area worldwide. For example, the sand-fixing model of planting two xerophytic shrubs on a checkerboard is widespread in northern China [20]; in particular, leguminous shrubs are widely used in sand fixation due to its N fixation capacity [23], and the most typical representative shrubs are C. korshinskii [17], C. scoparium [5], and Corethrodendron mongolicum [116].
It was observed that these sand-fixing measures effectively prevented the shift of dunes and the occurrence of sandstorms [102, 114]. Thus, we selected 3 typical combinations of shrubs for sand-fixing afforestation conducted in 2002 as our experimental treatments on the southwestern margin of the Mu Us Sandy Land of China, that is, C. korshinskii and Corethrodendron scoparium mixed shrubland (2 m × 2 m, row pitch), defined as two leguminous shrubs containing C. korshinskii (CaKCoS),C. korshinskii and Calligonum mongolicum mixed shrubland (2 m × 2 m, row pitch), defined as one leguminous shrub containing C. korshinskii (CaKCaM); and C. scoparium and C. mongolicum mixed shrubland (2 m × 2 m, row pitch), defined as one leguminous shrub excluding C. korshinskii (CoSCaM). Three plots were established for each combination of shrubs (treatment), and approximately 1.2 hm2 of each plot was selected as the observation area, with a distance of 1 km from each plot. Each plot included 5 sampling quadrats of 10 m × 10 m. None of the areas had been subjected to anthropogenic disturbances since their establishment (Fig. 1).
Soil Sampling
In August 2019, from each treatment, we collected the 0–10 cm topsoil at five points (arranged in an “X” shape) in each quadrat using a soil auger (diameter, 4 cm) after removing the litter layer on the ground surface and completely mixed the five samples from the same quadrat into one uniform soil sample (Fig. 1). We removed stones and roots from all soil samples collected (i.e., 3 combinations of sand-fixing shrubs × 3 plots/combination × 5 quadrats/plot). The collected soil samples were divided into two parts, one of which was packed in a sterile bag for the analysis of the physical and chemical properties of the soil, and the soil moisture was determined immediately. The other part was passed through a sieve with an aperture of 2 mm, packed into a 15 ml sterile centrifuge tube, quickly put into a dry ice bucket, brought back to the laboratory and stored at − 80 °C for DNA extraction [62].
Analysis of Soil Physicochemical Properties
Soil moisture was determined by oven drying to a constant mass at 105 ℃. The air-dried soil samples were divided into two parts and passed through soil sieves with apertures of 1 mm and 0.149 mm. The soil pH and electronic conductivity (EC) were determined by a pH meter (PHS-3D, Shanghai Sanxin Instrument, China) and a conductivity meter (DDS-307A, Shanghai Youke Instrument Co., Ltd., China), respectively. The soil total carbon (TC), soil organic carbon (SOC), total nitrogen (TN), available nitrogen (AN), total phosphorus (TP), available phosphorus (AP), total potassium (TK), and available potassium (AK) were measured as described by Bao [2]. Soil C, N, and P and their stoichiometric ratios (SOC/TN, C:N; SOC/TP, C:P; TN/TP, and N:P) were also calculated.
Soil DNA Extraction, Amplification, and Absolute Quantification Sequencing
The DNA extraction of soil bacteria was performed using a Power Soil™ DNA Isolation Kit (MOBIO Laboratories, Carlsbad, CA, USA). Agarose-gel electrophoresis and Nanodrop 2000 (Thermo Fisher Scientific, USA) were used to detect the integrity and purity of the genomic DNA from the extracted samples, and high-fidelity PCR amplification was performed for the V4-V5 region of the samples in triplicate. It has been reported that the amplified bacterial sequences in the V4-V5 region show the least intragenomic heterogeneity, thus maintaining good specificity and complete database information and minimizing overestimation, making it the best sequencing region for bacteria [8, 75]. Therefore, targeting the V4-V5 regions of 16S rRNA for PCR amplification was conducted using primers 515F (5′-GTGCCAGCMGCCGCGG-3′) and 907R (5′-CCGTCAATTCMTTTRAGTTT-3′) [84]. Sequencing was carried out using the Illumina MiSeq 2 × 250 bp, technically supported by Genesky Biotechnologies Inc., Shanghai, China (201,315) [33]. Over the last few decades, high-throughput sequencing technology has accelerated our understanding of soil microbial diversity and its underlying mechanisms [113]. In this study, a high-throughput absolute quantification sequencing method was employed to obtain an accurate and reliable absolute abundance of soil bacteria because this method can better solve the problem of the relative quantification of microbes in environmental samples while obtaining the exact species composition information, especially with respect to the soil bacterial community and dynamics [99]. The absolute quantification sequencing method eliminates the difficulty in finding specific primers for qPCR validation after relative quantification [73]. In addition, absolute abundance can be used to calculate relative abundance, but the reverse is not true [99].
Illumina Read Data Processing
To obtain high-quality sequencing data and improve the accuracy of subsequent bioinformatic analysis, the raw sequencing data were quality controlled and filtered. The Trim Galore software was used to remove sequences with a base mass less than 20 at the end of the V4-V5 tag sequences (which contain adapters) and to remove short sequences less than 100 bp in length. FLASH2 software was used to splice paired sequences and remove low-quality sequences (over 90% of base mass below 20). Mothur and UCHIME software were used to find and remove primers, as well as chimeric sequences. Finally, sequences with a similarity of more than 97% were classified as the same OTU, and the resulting representative OTU sequences were used for subsequent species annotation. The proportion of spike-in sequences was marked and counted, and these sequences were removed during subsequent analysis. The standard curve equation was made according to the spike-in sequence of each sample, and the absolute copy number of each OTU in each sample was calculated. Then, the absolute copy number of species in a unit sample was calculated according to the amount of DNA in the sequencing template, the amount of DNA extracted from the sample, and the number of samples used for DNA extraction [84]. There are multiple copies of the 16S gene in a species to ensure survival. In high-throughput sequencing, post-PCR sequencing will amplify this cardinality effect due to the copy number of different bases, resulting in a deviation in the read number for each species. This analysis was based on the rrnDB database (Version V5.6) (https://rrndb.umms.med.umich.edu/). The gene copy number of each OTU was estimated according to the closest relative species, and then, the absolute copy number of OTUs divided by the estimated gene copy number was corrected [72, 92].
Data Analysis
The changes in soil physicochemical properties among three combinations of shrubs (CaKCoS, CaKCaM, and CoSCaM) were determined based on a one-way analysis of variance (ANOVA) with Duncan’s test by using SPSS 18.0 (IBM Corp., USA). Rarefaction curves, Circos diagrams for the composition of species, Venn diagrams, heatmaps, and comparative diagrams of significantly different phyla were created using the R packages “ggplot2” and “Vegan” (R 4.0.5; https://www.r-project.org/). The alpha diversity (Shannon index) of bacteria was calculated using QIIME2 [3], and a significant difference diagram was drawn using the R package “Vegan.” In the 3 types of leguminous shrub treatments, the top 700 OTUs with significant differences (P < 0.05) in phylum-level abundance were filtered out by using SPSS 18.0, and then, the ternary diagram was created using the R package “VCD.” Redundancy analysis (RDA) was performed to elucidate the relationship between bacterial abundance at the phylum level and soil environmental factors using Canonco 5. To demonstrate the relationship between key soil components (C, N, P, and their stoichiometric ratios) and soil bacterial diversity, we constructed a structural equation model (SEM) for all soil samples using the R packages “lavaan,” “haven,” “Hmisc,” and “semPlot” [16]. All of the images described above were processed by Adobe Illustrator CS6, and the significant differences are expressed as P < 0.05, P < 0.01, and P < 0.001.
Results
Effects of Leguminous Shrubs on Soil Bacterial Diversity in Sandy Land
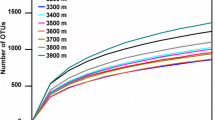
After quality screening, read removal, and statistical analysis, we obtained 10,137,510 reads from 45 soil samples by 16S rRNA gene sequencing. The number of clean reads per sample ranged from 126,903 to 762,429, and an average of 225,278 clean reads was isolated from each soil sample (Table S1). The Venn diagram showed that 10,023 OTUs were shared by the three combinations of shrubs from the absolute copy number of genes per gram of soil, and the CaKCoS treatment had 1124 unique OTUs, which was the highest among treatments (Fig. 2A). The rarefaction curve indicated that the sequences of all samples covered the diversity of bacterial populations across the three combinations of shrubs (Fig. 2B). The dominant phyla identified for absolute quantitative 16S rRNA sequencing from the three combinations of shrubs included Actinobacteria (20.57–25.4%), Acidobacteria (16.4–16.89%), Proteobacteria (13.92–16.77%), Chloroflexi (9.03–10.93%), Bacteroidetes (6.45–7.47%), Cyanobacteria/Chloroplast (2.07–6.53%), Planctomycetes (4.55–4.86%), Thaumarchaeota (3.27–3.82%), Gemmatimonadetes (1.39–2.29%), Armatimonadetes (1.96–2.31%), and candidate_division_WPS-1 (1.65–1.8%) (Fig. 2C). The Shannon index in CaKCoS was significantly higher than that in CaKCaM and CoSCaM (P < 0.01, Fig. 3A). The ternary diagram showed a more diverse OTU enrichment in CaKCoS than in CaKCaM and CoSCaM (Fig. 3B). These results suggested that the diversity of soil bacteria was strongly associated with the sand-fixing combination of the two leguminous shrubs (C. korshinskii × C. scoparium) rather than that with one leguminous shrub.
Effects of three combinations of shrubs on soil bacterial communities (A Venn diagram; B Rarefaction curve; and C Circos plots showing the abundance at the phylum level. All phyla are abbreviated by the first four letters of their names. Acti, Acid, Prot, Chlo, Bact, Cyan, Plan, Thau, Arma, Gemm, Cand, Firm, Verr, Nitr, and Eury are abbreviated forms of Actinobacteria, Acidobacteria, Proteobacteria, Chloroflexi, Bacteroidetes, Cyanobacteria/Chloroplast, Planctomycetes, Thaumarchaeota, Armatimonadetes, Gemmatimonadetes, candidate_division_WPS-1, Firmicutes, Verrucomicrobia, Nitrospirae, and Euryarchaeota, respectively). CaKCoS, CaKCaM, and CoSCaM represent the mixture of C. korshinskii and C. scoparium, the mixture of C. korshinskii and C. mongolicum, and the mixture of C. scoparium and C. mongolicum, respectively
Comparison of soil bacterial diversity in different combinations of sand-fixing shrubs (A Shannon index; B Ternary diagram. Solid dots represent significantly different OTUs (P < 0.05), the size of the dots indicates the abundance of the OTUs, and gray dots represent OTUs that are not significantly different). CaKCoS, CaKCaM, and CoSCaM represent the mixture of C. korshinskii and C. scoparium, the mixture of C. korshinskii and C. mongolicum, and the mixture of C. scoparium and C. mongolicum, respectively
A heatmap was constructed to show the phylum-level abundance of bacteria among the three combinations of shrubs (Fig. 4). This result indicated that phyla clustered into a clade according to their abundance. Crucially, the abundance of the most dominant phyla was higher in CaKCoS and CaKCaM than in CoSCaM, while the abundance of Cyanobacteria was higher in CaKCaM and CoSCaM than in CaKCoS. Among these dominant phyla, Actinobacteria, Acidobacteria, Proteobacteria, Chloroflexi, Planctomycetes, Thaumarchaeota, Armatimonadetes, candidate_division_WPS-1, Verrucomicrobia, BRC1, and Nitrospirae were significantly more abundant in CaKCoS and CaKCaM than in CoSCaM (P ≤ 0.05) (Fig. S1).
Heatmap and hierarchical cluster diagram based on Euclidean distance of the abundant phyla of the soil bacteria in different combinations of sand-fixing shrubs. The transition from blue to red reflects the gradual change in abundance from low to high. CaKCoS, CaKCaM, and CoSCaM represent the mixture of C. korshinskii and C. scoparium, the mixture of C. korshinskii and C. mongolicum, and the mixture of C. scoparium and C. mongolicum, respectively
Relationships Between Bacterial Communities and Soil Physicochemical Properties
RDA indicated that the majority of the soil factors were significantly positively correlated with the abundance of the dominant phyla, including Actinobacteria, Acidobacteria, Proteobacteria, Chloroflexi, Planctomycetes, Thaumarchaeota, and Armatimonadetes. Canonical RDA1 and RDA2 accounted for 56.66% and 2.35% of the total variation, respectively (Fig. 5A). TN, N:P, SM, and AP significantly affected the abundance of bacterial phyla in the soil. Considering the relationship with the sample plots, the promotion mainly occurred in CaKCoS- and CaKCaM-containing C. korshinskii.
Relationship between soil bacterial diversity and soil physicochemical properties (A Redundancy analysis indicating the relationship between the bacterial abundance of dominant phyla (blue arrows), edaphic factors (red arrows), and sample plot (solid dot with different colors). All species at the phylum level are abbreviated as the first four letters of their names; B Path diagram of the structural equation modeling (SEM) for all samples. Positive correlations are indicated by green arrows, and negative correlations are indicated by red arrows in SEM)
The SEM results showed that the model could well explain the relationship between soil bacterial diversity and soil C, N, and P nutrients (Fig. 5B). There were significant SEM paths based on all samples (P(χ2) = 0.689, chisq = 2.252, df = 4, gfi = 0.986, rmsea = 0.000, srmr = 0.083). SOC (0.92), N:P (0.52), and TP (0.18) were positively correlated with the bacterial Shannon index, which plays a crucial role in shaping bacterial diversity. However, TN (− 0.79), C:P (− 0.51), and C:N (− 0.47) were negatively correlated with soil bacterial diversity. Collectively, our study emphasizes that the combination of sand-fixing shrubs shapes bacterial diversity by changing the soil physical and chemical properties. Interestingly, this diversity occurred in the combination containing two leguminous shrubs but was not found in the combination containing one leguminous shrub.
Comparison of Soil Physical and Chemical Properties Among Different Combinations of Sand-Fixing Shrubs
The soil physical and chemical properties at a depth of 0–10 cm significantly differed among the three combinations of shrubs, as shown in Table 1 (P < 0.05). The CaKCoS and CaKCaM treatments had significantly higher SOC, TN, AK, EC, N:P, and C:P, while the contents of SM, TC, TK, and AP were all higher in CaKCoS than in CaKCaM and CoSCaM. Briefly, these results reinforced that the soil fertility was effectively enhanced in the sand-fixing combination containing two leguminous shrubs than in that containing one leguminous shrub, indicating that the combination of legumes is more beneficial for restoring sandy land ecosystems.
Discussion
The Restoration of Sandy Land with Leguminous Shrubs Could Effectively Drive Soil Bacterial Diversity
The novelty of this study was that we applied absolute quantification sequencing of 16S rRNA genes to accurately determine the changes in soil bacterial abundance in sand-fixing shrub combinations involving leguminous shrubs. In this study, we found that the dominant phyla were sequenced from the three combinations of shrub plantations (Fig. 2). After fixing the dunes with straw checkerboards, the combination of planting two leguminous shrubs (CaKCoS) significantly increased the diversity of soil bacteria and enriched the significantly different OTUs (Fig. 3). This result confirmed our hypothesis and provided a conceptual framework that the combination of leguminous shrubs can drive soil bacterial diversity during the process of desertification reversal. A large number of studies have shown that vegetation restoration involving legumes can effectively promote the α-diversity of soil bacteria, such as drought-tolerant legume shrub afforestation by using C. korshinskii [108], R. pseudoacacia, or C. korshinskii leguminous forestland [63, 64, 97], and 32-year-old C. microphylla plantations [5, 110], which supported the results of this study. This result might be because legumes enrich soil nutrients through N fixation and litter input and provide resource stores for bacterial communities to multiply in desert areas [100, 116]. C. korshinskii, R. pseudoacacia, and C. microphylla are typical leguminous shrubs used in forestation, and the dominant bacterial phyla found in these shrublands were almost consistent with the sequencing results in our study. The key factor driving soil bacterial diversity lay in the combination of species, especially legumes.
Another explanation could be that soil microbial diversity is closely related to soil biological crusts in dryland ecosystems, and litter that was produced by planting xerophytic shrubs to stabilize the moving dunes contributes to creating a suitable habitat for the occurrence of biological crusts [43]. Litter is retained on the soil surface for a long time and gradually forms the litter crust, which is the main contributor of C resources and plays an important role in maintaining the function of the soil ecosystem. The soil bacterial community and nutrient enrichment documented in Liu et al. [42] were closely related to the increase in SOC and TC, which were associated with litter crusts, and the diversity of bacteria in the litter crust was also significantly higher than that in bare soil. The key factor is that litter crusts are involved in nutrient regulation in the plant‒soil-microbe continuum and promote the accumulation of C and N on the soil surface [32, 42]. Studies have shown that litter crust depth is closely related to soil microbial diversity [41]. Li et al. [37] reported that the litter quality and soil enzyme activity of a C. korshinskii plantation were higher than those of a Platycladus orientalis plantation, indicating that legumes were the most effective means of afforestation in drylands. Therefore, the improvement of soil nutrients caused by litter crusts was likely to be a pathway for the enrichment of soil bacterial diversity.
Differences in plant functional composition contribute to differences in microbial community composition, which is a common indication of ecosystem functioning [11, 26]. In our study, the sand-fixing combination of two leguminous shrubs effectively increased the abundance of dominant soil phyla, including Actinobacteria, Acidobacteria, Proteobacteria, Chloroflexi, Planctomycetes, Thaumarchaeota, and Armatimonadetes (Fig. 4); among these phyla, the most obvious changes in order were found for Actinobacteria, Gemmatimonadetes, Proteobacteria, Chloroflexi, candidate_division_WPS-1, and BRC1 (Fig. S1). The cycling and balance of soil C and N are closely related to these bacterial phyla [21]. Studies have shown that Actinobacteria, as a copiotrophic phylum, had a high content in the soil of vegetation restoration areas, while Acidobacteria, as an oligotrophic phylum, was commonly abundant in cropland [28]. In general, copiotrophic and oligotrophic bacteria were taken as indicators for assessing the soil environment [38]. Actinobacteria is a typical phylum of gram-positive bacteria that helps to decompose dead organisms in the soil, promotes material circulation, produces antibiotics, and makes a great contribution to human health [67]. Acidobacteria are widely distributed bacteria with high variation and high abundance in low organic C environments [34]. Proteobacteria is a phylum of gram-negative bacteria, and some α- or β-Proteobacteria can also grow in low nutrient environments, inducing N fixation and forming symbionts with plants [56, 60]. Proteobacteria was one of the most important phyla in the leguminous shrub plantation in the present research. Ren et al. [63] found that soil bacterial diversity significantly increased when afforestation with C. korshinskii was compared with abandoned land and that afforestation effectively increased the abundance and dominance of Proteobacteria. These results indicate that the dominance of the N-fixing groups in the soil bacteria was closely related to the improvement of soil N with the introduction of legumes, which was consistent with the results of our study. Many studies have confirmed that the mineralization rate of soil C and the cycling of N could be predicted by the abundance of Proteobacteria, Acidobacteria, Bacteroidetes, and Actinobacteria; in particular, almost half of the Rhizobiales in Proteobacteria were directly associated with N fixation [21, 80]. In a study of the rhizobia of the leguminous plant Dalbergia, among 68 isolated bacteria, 65 belonged to Proteobacteria, especially Burkholderia, which can significantly induce the formation of nodules in the roots of leguminous plants [60]. Past studies also showed that the dominant phyla found in the present study, including Proteobacteria, Actinobacteria, and Bacteroidetes, were the main group in the rhizosphere of Hedysarum during afforestation with two leguminous shrubs [116]. Proteobacteria have been proven to be closely related to soil C availability [21],the input of C sources affected the content of soil nutrients, changed the C:P and N:P ratios, and ultimately influenced the change in microbial composition. Additionally, with the increase in forestation years of C. korshinskii, Proteobacteria in the soil became increasingly abundant, while Actinobacteria decreased correspondingly [108]. In the leguminous C. korshinskii and R. pseudoacacia plantations, the abundance of Actinobacteria was higher than that in grassland [95], which was similar to the results of this study. A large number of N-fixing microorganisms enriched the soil of the C. korshinskii plantation [98], which effectively increased the soil N. Therefore, the significant increase in the dominant bacteria in leguminous shrub combinations might also be related to the increase in N:P and C content in the present study.
Cyanobacteria are photosynthetic bacteria that play an important role when subjected to environmental stress. Biological soil crusts with Cyanobacteria can be isolated in many desert systems [78, 90]. After years of sand fixation with straw checkerboards combined with different shrubs, it was found that the abundance of Cyanobacteria was higher in the combination containing one leguminous and one nonleguminous shrub (CaKCaM) (Fig. 4). Strong habitat stress on the soil surface, especially water stress, can induce Cyanobacteria colonization, but Cyanobacteria must be equipped with mechanisms to protect themselves and avoiding high light intensities. More importantly, moderate fluctuation in environmental water creates conditions for Cyanobacteria to thrive, while continuous water conditions may lead to some cyanobacterial deaths [68, 90]. Cyanobacteria do not have the ability to mobilize water from deep soil layers and can only rely on the amount or frequency of precipitation to perform ecological functions such as N fixation [57], which might be related to the moderate SM content (Table 1) in the leguminous and nonleguminous shrub combination in this study. A previous study confirmed that Cyanobacteria in the bare substrate biocrusts of mountain vegetation types in the northern Urals were specific to different plant communities [52], which suggests that our future research needs to pay more attention to the effects of different sand-fixing plant communities on Cyanobacteria. Another possibility is that the sand-fixing combination of two leguminous shrubs inputs functional litter to the soil [71], leading to a significantly higher leaf C, N, P, and K content and a higher N:P ratio than those of nonleguminous plants [25], which further leads to a higher soil bacterial diversity and increases the abundance of bacterial phyla, such as Actinobacteria, Proteobacteria, Chloroflexi, Acidobacteria, and Bacteroidetes, in the soil instead of Cyanobacteria. This result indicates that leguminous plants had a strong competitiveness in leaf functional traits and changed the quality and properties of the litter.
The Improvement in Soil Properties Contributed Greatly to the Bacterial Diversity in Different Combinations of Sand-Fixing Shrubs
Our results showed that higher soil bacterial diversity was caused by improvements in soil physicochemical properties, which was attributed to the combination of leguminous shrubs used for sand fixation and afforestation. This may be due to the legume‒legume combinations triggering different N-fixing capacities and leading to the functional differentiation of soil nutrients [15, 69]. This result indicated that bacterial diversity was promoted by SOC, N:P, and TP but inhibited by TN, C:N, and C:P (Fig. 5B), and the abundance of dominant phyla was promoted by TN, AK, N:P, SM, and AP (Fig. 5A). This result might be because the combination of two leguminous shrubs produced litter, which initiated a considerable C pool, and the bacterial community, which was dominated by Proteobacteria, Actinobacteria, and Acidobacteria, depended crucially on the litter characteristics of specific plants [44], especially the high yield and quality of leguminous litter [29, 37], and its leaf N fixation traits [12]. Therefore, it has been documented that the properties of litter produced by legumes are key to the contribution of soil bacterial diversity. Similar studies have been reported, for example, in restoration patterns of legumes (R. pseudoacacia, C. korshinskii), litter characteristics significantly increased the Chao1 index [95] and community composition [94] of soil bacteria. The litter layer was the resource store of the ecosystem, containing nutrients such as C, N, and phosphorus (P), as well as other matter, which was driven by the microbial community [9], and litter quality was the crucial factor in promoting bacterial diversity in the topsoil layer [35].
The accumulation of SOC from leguminous shrub litter increased the metabolic function of Actinobacteria, while Acidobacteria was abundant in lower soil resources [38, 94]. Xu et al. [96] found that leguminous shrubs could significantly improve the Shannon and Chao1 indices of soil microorganisms, which were also significantly affected by SOC [100]. These findings were similar to our results. Thus, we suggest that C accumulation induced by leguminous litter might be a more favorable sand-fixing strategy that results in higher bacterial diversity. Recent studies supported that litter crusts positively promote the input of SOC and TN, which was strongly related to the bacterial community [42]. Viruel et al. [81] also found that SOC was one of the factors that determined the α-diversity of bacteria, which is consistent with our results, but the promoting effect of TN was contrary to that in the present study. The soil TN content effectively increased the abundance of bacteria (Fig. 5A), but TN and the C:N and C:P ratios decreased the diversity (Fig. 5B), which was different from some previous studies [62, 94]. Liu et al. [38] reported that R. pseudoacacia, a typical legume, significantly improved soil bacterial abundance when used alone, and this result was similar to those of our study. In summary, we proposed that the bacterial diversity might ultimately be due to the increased rate of C mineralization rather than N mineralization [76]. In contrast, in the desert oligotrophic environment, soil N was likely to stimulate the abundance of specific phyla but not the conditions that triggered increased diversity.
In this study, the N:P ratio could promote soil bacterial diversity under the combination of two leguminous shrubs. For example, in the Loess Plateau of China, the longer that C. korshinskii was planted, the more correlated the soil microbial diversity and dominant phyla abundance were with the N:P and C:P ratios, especially the N:P ratio [108]. The results suggested that the correlation between the N:P ratio and microbial diversity had a more useful effect on the changes in the soil microbial community than the C:N and C:P ratios in the leguminous plantations of C. korshinskii and R. pseudoacacia [61, 64], which was consistent with the results of this study. The negative correlation between the C:P ratio and bacterial diversity might be because N-fixing microbes increased continuously after long-term afforestation with leguminous shrubs in sandy land, resulting in imbalances in the soil C, N, and P. The soil bacterial diversity was closely related to P accumulation [14, 94]. The evidence shows that bacterial diversity peaks under the conditions of high P availability, soil organic matter reduction, and low C:P and C:N [14], and these soil conditions tend to accelerate nutrient cycling [13] and increase bacterial diversity by reducing fungal diversity [80]. Thus, these results supported our argument (Fig. 5).
The Restoration of Sandy Land with Leguminous Shrubs Could Effectively Improve the Soil Physical and Chemical Properties
In the present study, the combination of leguminous shrubs in sand fixation could effectively improve the soil properties (Table 1). Before 2001, our sampling plot suffered from severe disturbances due to moving dunes. After the long-term sand fixing practice was implemented, the combination of xerophytic shrubs gradually planted in the checkerboards stopped dune movement, and the physical and chemical properties of the soil gradually improved. Interestingly, we found that the combination of two leguminous shrubs (CaKCoS) significantly increased the soil physicochemical properties, such as SM, TC, TK, and AP (Table 1). Previous studies have confirmed that the long-term afforestation of C. korshinskii significantly promoted the accumulation of soil TC, SOC, TN, AK, and AN contents, which strongly supported our study results [37]. This was most likely related to the litter quality and soil enzymatic activity of legumes [30], suggesting that the use of specific species is the key to the improvement of soil properties. This might be driven by the improvement of the root exudates and the decomposition of the litter residues [53], and it might also be due to the dual N-fixation by the combination of legumes [49, 89]. The key factor is that legumes are special functional groups with higher productivity that feed back to the soil and promote soil fertility [23]. Most of the contributions to the soil surface layer come from N fixation and litter properties [100, 116]. The N-fixing function of leguminous leaves provides the soil with rich litter C and N resources [12, 15, 50], thus driving bacterial diversity by promoting soil fertility, which supported the hypothesis of this study. The functional traits and symbiotic systems of legumes explained the excellent performance of leguminous shrubs in soil improvement [94]; for example, the litter production and quality of arboreal legumes (Gliricidia sepium) (Herrera et al., 2020) and decomposition rete [31, 71] promoted soil nutrient cycling. The asymbiotic leaf N-fixation traits also contribute significantly to soil N and C accumulation by mediating litter properties [50]. We also found that the highest SOC and AN contents were in CaKCaM, which is consistent with Zhou et al. [114] and might be due to the abundance of Cyanobacteria in this combination during sand fixation.
In general, the long-term afforestation of xerophytic shrubs could release abundant nutrients to the soil, especially the combination of leguminous shrubs, which benefits from the unique functional traits of legume coupling. In the C. microphylla nebkha, which is a leguminous shrub, the enrichment rates of soil organic matter (SOM), AN, and AP were higher than those in nonlegumes (Atraphaxis manshurica and Salix gordejevii) [4]. Our previous studies have confirmed that soil C, N, and K are the key factors in vegetation restoration in desertification reversal areas [59], this result was well verified in our sample plots in this study. A meta-analysis by Gao and Huang [24] showed that the Three-North Shelter Forest construction project could significantly improve the soil AK content (72.93%) in the 0–20 cm soil layer of the Loess Plateau, especially in leguminous forests of R. pseudoacacia and C. korshinskii, which was similar to the result of higher potassium in this study. Crucially, our results suggested that the contents of SM, TC, TK, and AP were significantly higher in the combination of two leguminous shrubs than in the other treatments (Table 1). SM is the key driving factor of afforestation success in desert areas. Caragana shows strong water utilization adaptability in long-term afforestation practices [112], for instance, the water use capacity of Hedysarun fruticosum gradually yielded to that of C. microphylla [27]. A study confirmed that C. korshinskii had a high dependence on water content in the deep soil layer (80–100 cm) during the drought period [112], and the increase in water content in the surface layer (0–10 cm) in this study might be closely related to the nursing effect [39] of the canopy formed by the intermixing of C. korshinskii and C. scoparium. Under this shrub canopy, a higher content of fine particle fractions with strong water-holding capacity was formed [106], and the combination of these two shrubs (CaKCoS) gradually contributed to the formation of C, N, and P stores, called “resource islands” [1, 101]. Zhang et al. [109] found that C. korshinskii had a stronger effect on enhancing the preservation of soil fine particle composition than natural grassland restoration, and the SM content in the 0–100 cm soil layer was significantly positively correlated with the fine particle fraction (silt and clay). A large number of studies have confirmed that the content of soil nutrients is the highest in fine particles of silt or clay [51, 83, 86, 109].
In conclusion, our study demonstrated a conceptual framework in which the combination of leguminous shrubs enhanced soil bacterial diversity by improving soil physicochemical properties, which are key factors in the accumulation of soil C, N, and P. This study provides a theoretical basis for effectively promoting ecosystem functioning in the struggle to fix sandy dunes and provides a scientific basis for the practice of vegetation recovery in arid and semiarid regions.
Conclusion
In this study, we took three combinations of sand-fixing shrubs as research objects. The combination of two leguminous shrubs (CaKCoS) and combinations of one leguminous and one nonleguminous shrub (CaKCaM and CoSCaM) were analyzed to explore the soil physicochemical properties, bacterial diversity, and community composition, as well as their driving factors, based on high-throughput absolute quantification 16S rRNA sequencing.
Our results suggested that the combination of two leguminous shrubs (CaKCoS) significantly improved soil physicochemical properties and soil bacterial diversity (Fig. 6). The abundance of the dominant phyla in the bacterial community composition of CaKCoS was significantly higher than that of CaKCaM and CoSCaM, the dominant bacterial phyla of which were mainly Actinobacteria, Acidobacteria, Proteobacteria, Chloroflexi, Planctomycetes, and Thaumarchaeota. The RDA indicated that the majority of soil properties (TN, AK, N:P, SM, and AP) were important soil environmental factors affecting the abundance of the dominant phyla, and canonical RDA1 and RDA2 accounted for 56.66% and 2.35% of the total variation, respectively. SEM revealed that soil bacterial diversity was positively affected by SOC, N:P, and TP but negatively affected by C:P and C:N. Among these factors, SOC and N:P were the crucial factors influencing soil bacterial diversity, and soil P was probably the main limiting factor. This study elucidates that a combination of leguminous shrubs is the crucial driving factor for soil bacterial diversity and is most likely attributable to the litter properties of legumes.
The guiding principle diagram indicates the general conclusions of this study (Sig. represents the significant influence on the factor the arrow is pointing to; CaKCoS, CaKCaM, and CoSCaM represent a mixture of C. korshinskii and C. scoparium, mixture of C. korshinskii and C. mongolicum, and mixture of C. scoparium and C. mongolicum, respectively)
Overall, our results highlight a conceptual framework for sand-fixing afforestation and vegetation recovery in sandy land, that is, the combination of leguminous shrubs, which effectively improved soil properties, drove soil bacterial diversity, and maintained ecosystem functioning and balance during desertification reversal.
References
Bachar A, Soares MI, Gillor O (2012) The effect of resource islands on abundance and diversity of bacteria in arid soils. Microb Ecol 63(3):694–700. https://doi.org/10.1007/s00248-011-9957-x
Bao S (2000) Agricultural chemistry analysis of soil properties, 3rd edn. China Agriculture Press, Beijing
Bolyen E, Rideout J, Dillon M, Bokulich N, Abnet C, Al-Ghalith G, Alexander H, Alm E, Arumugam M, Asnicar F, Bai Y, Bisanz J, Bittinger K, Brejnrod A, Brislawn C, Brown C, Callahan B, Caraballo-Rodríguez A, Chase J, Cope E, Da Silva R, Diener C, Dorrestein P, Douglas G, Durall D, Duvallet C, Edwardson C, Ernst M, Estaki M, Fouquier J, Gauglitz J, Gibbons S, Gibson D, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley G, Janssen S, Jarmusch A, Jiang L, Kaehler B, Kang K, Keefe C, Keim P, Kelley S, Knights D, Koester I, Kosciolek T, Kreps J, Langille M, Lee J, Ley R, Liu Y, Loftfield E, Lozupone C, Maher M, Marotz C, Martin B, McDonald D, McIver L, Melnik A, Metcalf J, Morgan S, Morton J, Naimey A, Navas-Molina J, Nothias L, Orchanian S, Pearson T, Peoples S, Petras D, Preuss M, Pruesse E, Rasmussen L, Rivers A, Robeson M, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song S, Spear J, Swafford A, Thompson L, Torres P, Trinh P, Tripathi A, Turnbaugh P, Ul-Hasan S, van der Hooft J, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber K, Williamson C, Willis A, Xu Z, Zaneveld J, Zhang Y, Zhu Q, Knight R, Caporaso J (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Cao C, Abulajiang Y, Zhang Y, Feng S, Wang T, Ren Q, Li H (2016) Assessment of the effects of phytogenic nebkhas on soil nutrient accumulation and soil microbiological property improvement in semi-arid sandy land. Ecol Eng 91:582–589. https://doi.org/10.1016/j.ecoleng.2016.03.042
Cao C, Zhang Y, Cui Z, Feng S, Wang T, Ren Q (2017) Soil bacterial community responses to revegetation of moving sand dune in semi-arid grassland. Appl Microbiol Biotechnol 101(15):6217–6228. https://doi.org/10.1007/s00253-017-8336-z
Cao Y, Li Y, Li C, Huang G, Lü G (2016) Relationship between presence of the desert shrub Haloxylon ammodendron and microbial communities in two soils with contrasting textures. Appl Soil Ecol 103:93–100. https://doi.org/10.1016/j.apsoil.2016.03.011
Cardinale M, Brusetti L, Lanza A, Orlando S, Daffonchio D, Puglia AM, Quatrini P (2010) Rehabilitation of Mediterranean anthropogenic soils using symbiotic wild legume shrubs: Plant establishment and impact on the soil bacterial community structure. Appl Soil Ecol 46(1):1–8. https://doi.org/10.1016/j.apsoil.2010.05.007
Claesson MJ, Wang Q, O’Sullivan O, Greene-Diniz R, Cole JR, Ross RP, O’Toole PW (2010) Comparison of two next-generation sequencing technologies for resolving highly complex microbiota composition using tandem variable 16S rRNA gene regions. Nucleic Acids Res 38(22):e200. https://doi.org/10.1093/nar/gkq873
Crumsey JM, Capowiez Y, Goodsitt MM, Larson S, Moine JML, Bird JA, Kling GW, Nadelhoffer KJ (2015) Exotic earthworm community composition interacts with soil texture to affect redistribution and retention of litter-derived C and N in northern temperate forest soils. Biogeochemistry 126:379–395. https://doi.org/10.1007/s10533-015-0164-6
D’Odorico P, Bhattachan A, Davis KF, Ravi S, Runyan CW (2013) Global desertification: Drivers and feedbacks. Adv Water Resour 51:326–344. https://doi.org/10.1016/j.advwatres.2012.01.013
de Vries FT, Manning P, Tallowin JRB, Mortimer SR, Pilgrim ES, Harrison KA, Hobbs PJ, Quirk H, Shipley B, Cornelissen JHC, Kattge J, Bardgett RD (2012) Abiotic drivers and plant traits explain landscape-scale patterns in soil microbial communities. Ecol Lett 15(11):1230–1239. https://doi.org/10.1111/j.1461-0248.2012.01844.x
Delgado-Baquerizo M, Fry EL, Eldridge DJ, de Vries FT, Manning P, Hamonts K, Kattge J, Boenisch G, Singh BK, Bardgett RD (2018) Plant attributes explain the distribution of soil microbial communities in two contrasting regions of the globe. New Phytol 219(2):574–587. https://doi.org/10.1111/nph.15161
Delgado-Baquerizo M, Maestre FT, Gallardo A, Bowker MA, Wallenstein MD, Quero JL, Ochoa V, Gozalo B, Garcia-Gomez M, Soliveres S, Garcia-Palacios P, Berdugo M, Valencia E, Escolar C, Arredondo T, Barraza-Zepeda C, Bran D, Carreira JA, Chaieb M, Conceicao AA, Derak M, Eldridge DJ, Escudero A, Espinosa CI, Gaitan J, Gatica MG, Gomez-Gonzalez S, Guzman E, Gutierrez JR, Florentino A, Hepper E, Hernandez RM, Huber-Sannwald E, Jankju M, Liu J, Mau RL, Miriti M, Monerris J, Naseri K, Noumi Z, Polo V, Prina A, Pucheta E, Ramirez E, Ramirez-Collantes DA, Romao R, Tighe M, Torres D, Torres-Diaz C, Ungar ED, Val J, Wamiti W, Wang D, Zaady E (2013) Decoupling of soil nutrient cycles as a function of aridity in global drylands. Nature 502(7473):672–676. https://doi.org/10.1038/nature12670
Delgado-Baquerizo M, Reich PB, Khachane AN, Campbell CD, Thomas N, Freitag TE, Abu Al-Soud W, Sorensen S, Bardgett RD, Singh BK (2017) It is elemental: soil nutrient stoichiometry drives bacterial diversity. Environ Microbiol 19(3):1176–1188. https://doi.org/10.1111/1462-2920.13642
Dovrat G, Bakhshian H, Masci T, Sheffer E (2020) The nitrogen economic spectrum of legume stoichiometry and fixation strategy. New Phytol 227(2):365–375. https://doi.org/10.1111/nph.16543
Epskamp S (2015) semPlot: Unified visualizations of structural equation models. Struct Equ Modeling 22(3):474–483. https://doi.org/10.1080/10705511.2014.937847
Fang X, Li J, Xiong Y, Xu D, Fan X, Li F (2008) Responses of Caragana korshinskii Kom to shoot removal: mechanisms underlying regrowth. Ecol Res 23(5):863–871. https://doi.org/10.1007/s11284-007-0449-x
Fang XW, Turner NC, Li FM, Li WJ, Guo XS (2011) Caragana korshinskii seedlings maintain positive photosynthesis during short-term, severe drought stress. Photosynthetica 49(4):603–609. https://doi.org/10.1007/s11099-011-0067-2
Fang XW, Turner NC, Palta JA, Yu MX, Gao TP, Li FM (2013) The distribution of four Caragana species is related to their differential responses to drought stress. Plant Ecol 215(1):133–142. https://doi.org/10.1007/s11258-013-0285-8
Feng Q, Tian Y, Yu T, Yin Z, Cao S (2019) Combating desertification through economic development in northwestern China. Land Degrad Dev 30(8):910–917. https://doi.org/10.1002/ldr.3277
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364. https://doi.org/10.1890/05-1839|
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. P Natl Acad Sci USA 103:626–631. https://doi.org/10.1073/pnas.0507535103
Furey GN, Tilman D (2021) Plant biodiversity and the regeneration of soil fertility. Proc Natl Acad Sci U S A 118(49):e2111321118. https://doi.org/10.1073/pnas.2111321118
Gao H, Huang Y (2020) Impacts of the Three-North shelter forest program on the main soil nutrients in Northern Shaanxi China: a meta-analysis. For Ecol Manag 458:117808. https://doi.org/10.1016/j.foreco.2019.117808
Gomes AL, Revermann R, Meller P, Goncalves FMP, Aidar MPM, Lages F, Finckh M (2021) Functional traits and symbiotic associations of geoxyles and trees explain the dominance of detarioid legumes in miombo ecosystems. New Phytol 230(2):510–520. https://doi.org/10.1111/nph.17168
Grigulis K, Lavorel S, Krainer U, Legay N, Baxendale C, Dumont M, Kastl E, Arnoldi C, Bardgett RD, Poly F, Pommier T, Schloter M, Tappeiner U, Bahn M, Clément J, Hutchings M (2013) Relative contributions of plant traits and soil microbial properties to mountain grassland ecosystem services. J Ecol 101(1):47–57. https://doi.org/10.1111/1365-2745.12014
Guo J, Gong X, Fang L, Jiang D, Ala M, Bucci SJ, Scholz FG, Goldstein G, Hao G (2020) Switching of dominant positions between two sand-fixing shrub species during the dune revegetation process is underlain by their contrasting xylem hydraulics and water-use strategies. Land Degrad Dev 31(10):1195–1205. https://doi.org/10.1002/ldr.3493
Guo Y, Chen X, Wu Y, Zhang L, Cheng J, Wei G, Lin Y (2018) Natural revegetation of a semiarid habitat alters taxonomic and functional diversity of soil microbial communities. Sci Total Environ 635:598–606. https://doi.org/10.1016/j.scitotenv.2018.04.171
Herrera AM, de Mello ACL, Apolinário VXDO, DubeuxJúnior JCB, da Silva VJ, dos Santos MVF, da Cunha MV (2020) Decomposition of senescent leaves of signalgrass (Urochloa decumbens Stapf R Webster) and arboreal legumes in silvopastoral systems. Agroforestry Syst 94(6):2213–2224
Hong S, Piao S, Chen A, Liu Y, Liu L, Peng S, Sardans J, Sun Y, Penuelas J, Zeng H (2018) Afforestation neutralizes soil pH. Nat Commun 9(1):520. https://doi.org/10.1038/s41467-018-02970-1
Jaramillo DM, Dubeux JCB, Sollenberger L, Mackowiak C, Vendramini JMB, DiLorenzo N, Queiroz LMD, Santos ERS, Garcia L, Ruiz-Moreno M, Santen E (2021) Litter mass, deposition rate, and decomposition in nitrogen-fertilized or grass–legume grazing systems. Crop Sci 61(3):2176–2189. https://doi.org/10.1002/csc2.20475
Jia C, Huang Z, Miao H, Lu R, Li J, Liu Y, Shen W, He H, Wu G (2018) Litter crusts promote herb species formation by improving surface microhabitats in a desert ecosystem. CATENA 171:245–250. https://doi.org/10.1016/j.catena.2018.07.024
Jiang S, Yu Y, Gao R, Wang H, Zhang J, Li R, Long X, Shen Q, Chen W, Cai F (2019) High-throughput absolute quantification sequencing reveals the effect of different fertilizer applications on bacterial community in a tomato cultivated coastal saline soil. Sci Total Environ 687:601–609. https://doi.org/10.1016/j.scitotenv.2019.06.105
Kielak AM, Barreto CC, Kowalchuk GA, van Veen JA, Kuramae EE (2016) The ecology of acidobacteria: moving beyond genes and genomes. Front Microbiol 7:744. https://doi.org/10.3389/fmicb.2016.00744
Kooijman AM, Weiler HA, Cusell C, Anders N, Meng X, Seijmonsbergen AC, Cammeraat LH (2019) Litter quality and microtopography as key drivers to topsoil properties and understorey plant diversity in ancient broadleaved forests on decalcified marl. Sci Total Environ 684:113–125. https://doi.org/10.1016/j.scitotenv.2019.05.285
Li Y, Cui J, Zhang T, Okuro T, Drake S (2009) Effectiveness of sand-fixing measures on desert land restoration in Kerqin Sandy Land, northern China. Ecol Eng 35(1):118–127. https://doi.org/10.1016/j.ecoleng.2008.09.013
Li Y, Han C, Sun S, Zhao C (2021) Effects of tree species and soil enzyme activities on soil nutrients in dryland plantations. Forests 12(9):1153. https://doi.org/10.3390/f12091153
Liu J, Yang Z, Dang P, Zhu H, Gao Y, Ha VN, Zhao Z (2018) Response of soil microbial community dynamics to Robinia pseudoacacia L. afforestation in the loess plateau: a chronosequence approach. Plant Soil 423(1–2):327–338. https://doi.org/10.1007/s11104-017-3516-2
Liu L, Bai Y, She W, Qiao Y, Qin S, Zhang Y (2020) A nurse shrub species helps associated herbaceous plants by preventing shade-induced evaporation in a desert ecosystem. Land Degrad Dev 32(4):1796–1808. https://doi.org/10.1002/ldr.3831
Liu N, Guo Q (2012) Resource-use efficiencies of three indigenous tree species planted in resource islands created by shrubs: implications for reforestation of subtropical degraded shrublands. Plant Ecol 213(7):1177–1185. https://doi.org/10.1007/s11258-012-0075-8
Liu X, Huang Z, Havrilla CA, Liu Y, Wu G (2021) Plant litter crust role in nutrients cycling potentials by bacterial communities in a sandy land ecosystem. Land Degrad Dev 32(11):3194–3203. https://doi.org/10.1002/ldr.3973
Liu Y, Havrilla CA, Jia C, Liu X, Wu G (2021) Litter crusts enhance soil nutrients through bacteria rather than fungi in sandy ecosystems. CATENA 204:105413. https://doi.org/10.1016/j.catena.2021.105413
Liu Y, Li X, Xing Z, Zhao X, Pan Y (2013) Responses of soil microbial biomass and community composition to biological soil crusts in the revegetated areas of the Tengger Desert. Appl Soil Ecol 65:52–59. https://doi.org/10.1016/j.apsoil.2013.01.005
Liu YL, Zhu GY, Hai XY, Li JW, Shangguan ZP, Peng CH, Deng L (2020) Long-term forest succession improves plant diversity and soil quality but not significantly increase soil microbial diversity: Evidence from the Loess Plateau. Ecol Eng 142:105631. https://doi.org/10.1016/j.ecoleng.2019.105631
Lyu Y, Yang Y, Guo L, Liu L, Shi P, Zhang G, Qu Z, Hu X, Wang J, Xiong Y, Wen H, Lei J, Liang B, Dai J (2016) Desertification and blown sand disaster in China. J Agr Sci Tech-IRAN 6:363–371. https://doi.org/10.17265/2161-6256/2016.06.001
Lyu YL, Shi PJ, Han GY, Liu LY, Guo LL, Hu X, Zhang GM (2020) Desertification control practices in China. Sustainability 12(8):3258. https://doi.org/10.3390/su12083258
Maestre FT, Bowker MA, Puche MD, Hinojosa MB, Martinez I, Garcia-Palacios P, Castillo AP, Soliveres S, Luzuriaga AL, Sanchez AM, Carreira JA, Gallardo A, Escudero A (2009) Shrub encroachment can reverse desertification in semi-arid Mediterranean grasslands. Ecol Lett 12(9):930–941. https://doi.org/10.1111/j.1461-0248.2009.01352.x
Marouane A, Haroun C, El Mohammed HB, Lyès B, M’hammed B (2021) Diversity of psammophyte communities on sand dunes and sandy soils of the northern Sahara desert. J King Saud Univ - Sci 33(8):101656. https://doi.org/10.1016/J.JKSUS.2021.101656
Massawe PI, Mtei KM, Munishi LK, Ndakidemi PA (2017) Effects of Rhizobium inoculation and cropping systems on macronutrients uptake and partitioning in two legumes (Common bean and Lablab). Indian J Agric Res 51(3):206–213. https://doi.org/10.18805/ijare.v51i03.7908
Moreira JCF, Brum M, de Almeida LC, Barrera-Berdugo S, de Souza AA, de Camargo PB, Oliveira RS, Alves LF, Rosado BHP, Lambais MR (2021) Asymbiotic nitrogen fixation in the phyllosphere of the Amazon forest: changing nitrogen cycle paradigms. Sci Total Environ 773:145066. https://doi.org/10.1016/j.scitotenv.2021.145066
Mukherjee A, Lal R, Zimmerman AR (2014) Effects of biochar and other amendments on the physical properties and greenhouse gas emissions of an artificially degraded soil. Sci Total Environ 487:26–36. https://doi.org/10.1016/j.scitotenv.2014.03.141
Novakovskaya IV, Patova EN, Dubrovskiy YA, Novakovskiy AB, Kulyugina EE (2022) Distribution of algae and cyanobacteria of biological soil crusts along the elevation gradient in mountain plant communities at the Northern Urals (Russian European Northeast). J Mt Sci 19(3):637–646. https://doi.org/10.1007/s11629-021-6952-7
Oleghe E, Naveed M, Baggs EM, Hallett PD (2019) Residues with varying decomposability interact differently with seed or root exudate compounds to affect the biophysical behaviour of soil. Geoderma 343:50–59. https://doi.org/10.1016/j.geoderma.2019.02.023
Orlovsky N, Birnbaum E (2002) The role of Haloxylon species for combating desertification in Central Asia. Plant Biosyst - An Int J Deal all Aspects Plant Biol 136(2):233–240. https://doi.org/10.1080/11263500212331351139
Padilla FM, Pugnaire FI (2009) Species identity and water availability determine establishment success under the canopy of Retama sphaerocarpa shrubs in a dry environment. Restor Ecol 17(6):900–907. https://doi.org/10.1111/j.1526-100X.2008.00460.x
Paulitsch F, Dos Reis FB Jr, Hungria M (2021) Twenty years of paradigm-breaking studies of taxonomy and symbiotic nitrogen fixation by beta-rhizobia, and indication of Brazil as a hotspot of Paraburkholderia diversity. Arch Microbiol 203(8):1–19. https://doi.org/10.1007/s00203-021-02466-5
Qi, J., Li, X., Jia, R., Yang, H., Shi, Y., Sun, J., Fu, T.: Response of biocrust nitrogenase activity to the variation of rainfall regime in the Tengger Desert, northern China. Catena 212. https://doi.org/10.1016/j.catena.2022.106093
Qiu G, Lee I, Shimizu H, Gao Y, Ding G (2004) Principles of sand dune fixation with straw checkerboard technology and its effects on the environment. J Arid Environ 56(3):449–464. https://doi.org/10.1016/s0140-1963(03)00066-1
Qiu KY, Xie YZ, Xu DM, Pott R (2018) Ecosystem functions including soil organic carbon, total nitrogen and available potassium are crucial for vegetation recovery. Sci Rep 8:7607. https://doi.org/10.1038/s41598-018-25875-x
Rasolomampianina R, Bailly X, Fetiarison R, Rabevohitra R, Bena G, Ramaroson L, Raherimandimby M, Moulin L, De Lajudie P, Dreyfus P, Avarre JC (2005) Nitrogen-fixing nodules from rose wood legume trees (Dalbergia spp) endemic to Madagascar host seven different genera belonging to alpha- and beta-Proteobacteria. Mol Ecol 14(13):4135–4146
Ren C, Chen J, Deng J, Zhao F, Han X, Yang G, Tong X, Feng Y, Shelton S, Ren G (2017) Response of microbial diversity to C:N: P stoichiometry in fine root and microbial biomass following afforestation. Biol Fertil Soils 53:457–468. https://doi.org/10.1007/s00374-017-1197-x
Ren C, Liu W, Zhao F, Zhong Z, Deng J, Han X, Yang G, Feng Y, Ren G (2019) Soil bacterial and fungal diversity and compositions respond differently to forest development. CATENA 181:104071. https://doi.org/10.1016/j.catena.2019.104071
Ren C, Sun P, Kang D, Zhao F, Feng Y, Ren G, Han X, Yang G (2016) Responsiveness of soil nitrogen fractions and bacterial communities to afforestation in the Loess Hilly Region (LHR) of China. Sci Rep 6:28469. https://doi.org/10.1038/srep28469
Ren C, Zhao F, Kang D, Yang G, Han X, Tong X, Feng Y, Ren G (2016) Linkages of C:N: P stoichiometry and bacterial community in soil following afforestation of former farmland. For Ecol Manag 376:59–66. https://doi.org/10.1016/j.foreco.2016.06.004
Sandoval Pérez AL, Camargo-Ricalde SL, Montaño NM, García-Oliva F, Alarcón A, Montaño-Arias SA, Esperón-Rodríguez M (2016) Biocrusts, inside and outside resource islands of Mimosa luisana (Leguminosae), improve soil carbon and nitrogen dynamics in a tropical semiarid ecosystem. Eur J Soil Biol 74:93–103. https://doi.org/10.1016/j.ejsobi.2016.03.006
Segoli M, Ungar ED, Shachak M (2012) Fine-scale spatial heterogeneity of resource modulation in semi-arid “islands of fertility.” Arid Land Res Manag 26(4):344–354. https://doi.org/10.1080/15324982.2012.694397
Servin JA, Herbold CW, Skophammer RG, Lake JA (2008) Evidence excluding the root of the tree of life from the Actinobacteria. Mol Biol Evol 25(1):1–4. https://doi.org/10.1093/molbev/msm249
Shaw E, Hill DR, Brittain N, Wright DJ, Tauber U, Marand H, Helm RF, Potts M (2003) Unusual water flux in the extracellular polysaccharide of the cyanobacterium Nostoc commune. Appl Environ Microbiol 69(9):5679–5684. https://doi.org/10.1128/AEM.69.9.5679-5684.2003
Shin C, Sharif M, Lee H (2018) Evaluating the effect of bacterial inoculation and fertilization on the soil nutrient status of coal mine soil by growing soybean (Glycine max) and shrub Lespedeza (Lespedeza bicolor). Sustainability 10(12):4793. https://doi.org/10.3390/su10124793
Silvertown J, Wilson JB (1994) Community structure in a desert perennial community. Ecology 75(2):409–417. https://doi.org/10.2307/1939544
Song X, Cai J, Meng H, Ding S, Wang L, Liu B, Chang Q, Zhao X, Li Z, Wang D (2020) Defoliation and neighbouring legume plants accelerate leaf and root litter decomposition of Leymus chinensis dominating grasslands. Agr Ecosyst Environ 302:107074. https://doi.org/10.1016/j.agee.2020.107074
Stoddard SF, Smith BJ, Hein R, Roller Benjamin RK, Schmidt TM (2015) rrnDB: improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res 43(D1):D593–D598. https://doi.org/10.1093/nar/gku1201
Stokell JR, Hamp TJ, Steck TR (2016) Examining changes in bacterial abundance in complex communities using next-generation sequencing is enhanced with quantitative PCR. Antonie Van Leeuwenhoek 109(8):1161–1166. https://doi.org/10.1007/s10482-016-0707-4
Su YZ, Zhao WZ, Su PX, Zhang ZH, Wang T, Ram R (2007) Ecological effects of desertification control and desertified land reclamation in an oasis–desert ecotone in an arid region: A case study in Hexi Corridor, northwest China. Ecol Eng 29(2):117–124. https://doi.org/10.1016/j.ecoleng.2005.10.015
Sun DL, Jiang X, Wu QL, Zhou NY (2013) Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity. Appl Environ Microbiol 79(19):5962–5969. https://doi.org/10.1128/AEM.01282-13
Sun Y, Liu Z, Zhang Y, Lai Z, She W, Bai Y, Feng W, Qin S (2020) Microbial communities and their genetic repertoire mediate the decomposition of soil organic carbon pools in revegetation shrublands in a desert in northern China. Eur J Soil Sci 71(1):93–105. https://doi.org/10.1111/ejss.12824
Torres L, Abraham EM, Rubio C, Barbero-Sierra C, Ruiz-Pérez M (2015) Desertification research in Argentina. Land Degrad Dev 26(5):433–440. https://doi.org/10.1002/ldr.2392
Tsujimura S, Nakahara H, Kosaki T, Ishida N, Karbozova E (1998) Distribution of soil algae in salinized irrigation land in the arid region of Central Asia. Soil Science and Plant Nutrition 44(1):53–65. https://doi.org/10.1080/00380768.1998.10414426
Tyc O, Song C, Dickschat JS, Vos M, Garbeva P (2017) The ecological role of volatile and soluble secondary metabolites produced by soil bacteria. Trends Microbiol 25(4):280–292. https://doi.org/10.1016/j.tim.2016.12.002
Van Der Heijden MG, Bardgett RD, Van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11(3):296–310. https://doi.org/10.1111/j.1461-0248.2007.01139.x
Viruel E, Fontana CA, Puglisi E, Nasca JA, Banegas NR, Cocconcelli PS (2022) Land-use change affects the diversity and functionality of soil bacterial communities in semi-arid Chaco region. Argentina Appl Soil Ecol 172:104362. https://doi.org/10.1016/j.apsoil.2021.104362
Wang J, Wang R (2019) The physical and chemical properties of soil crust in straw checkerboards with different ages in the Mu Us sandland. Northern China Sustain 11(17):4755. https://doi.org/10.3390/su11174755
Wang J, Zhao W, Wang G, Yang S, Pereira P (2021) Effects of long-term afforestation and natural grassland recovery on soil properties and quality in Loess Plateau (China). Sci Total Environ 770:144833. https://doi.org/10.1016/j.scitotenv.2020.144833
Wang WH, Wang N, Dang KK, Dai W, Guan L, Wang BR, Gao JS, Cui ZL, Dong YH, Wang H (2020) Long-term nitrogen application decreases the abundance and copy number of predatory myxobacteria and alters the myxobacterial community structure in the soil. Sci Total Environ 708:135114. https://doi.org/10.1016/j.scitotenv.2019.135114
Wang XP, Li XR, Xiao HL, Pan YX (2006) Evolutionary characteristics of the artificially revegetated shrub ecosystem in the Tengger Desert. Northern China Ecological Research 21(3):415–424. https://doi.org/10.1007/s11284-005-0135-9
Wang YS, Li YH, Li YR (2020) Land engineering consolidates degraded sandy land for agricultural development in the largest sandy land of China. Land 9(6):199. https://doi.org/10.3390/land9060199
Ward D, Trinogga J, Wiegand K, du Toit J, Okubamichael D, Reinsch S, Schleicher J (2018) Large shrubs increase soil nutrients in a semi-arid savanna. Geoderma 310:153–162. https://doi.org/10.1016/j.geoderma.2017.09.023
Waseem M, Nie ZF, Yao GQ, Hasan M, Xiang Y, Fang XW (2021) Dew absorption by leaf trichomes in Caragana korshinskii: an alternative water acquisition strategy for withstanding drought in arid environments. Physiol Plant 172(2):528–539. https://doi.org/10.1111/ppl.13334
Wendlandt CE, Gano-Cohen KA, Stokes PJN, Jonnala BNR, Zomorrodian AJ, Al-Moussawi K, Sachs JL (2022) Wild legumes maintain beneficial soil rhizobia populations despite decades of nitrogen deposition. Oecologia. https://doi.org/10.1007/s00442-022-05116-9
Whitton BA (2012) Ecology of Cyanobacteria II: their diversity in space and time. Springer Press, London
Wu G, Jia C, Huang Z, López-Vicente M, Liu Y (2021) Plant litter crust appear as a promising measure to combat desertification in sandy land ecosystem. CATENA 206:105573. https://doi.org/10.1016/j.catena.2021.105573
Wu L, Yang Y, Chen S, Jason SZ, Zhao M, Zhu Z, Yang S, Qu Y, Ma Q, He Z, Zhou J, He Q (2017) Microbial functional trait of rRNA operon copy numbers increases with organic levels in anaerobic digesters. ISME J 11:2874–2878. https://doi.org/10.1038/ismej.2017.135
Xiang Y, Cheng M, Huang Y, An S, Darboux F (2017) Changes in soil microbial community and its effect on carbon sequestration following afforestation on the Loess Plateau, China. Int J Environ Res Public Health 14(8):948. https://doi.org/10.3390/ijerph14080948
Xu MP, Lu XQ, Xu YD, Zhong ZK, Zhang W, Ren CJ, Han X, Yang GH, Feng YZ (2020) Dynamics of bacterial community in litter and soil along a chronosequence of Robinia pseudoacacia plantations. Sci Total Environ 703:135613. https://doi.org/10.1016/j.scitotenv.2019.135613
Xu MP, Wang JY, Zhu YF, Han XH, Ren CJ, Yang GH (2021) Plant biomass and soil nutrients mainly explain the variation of soil microbial communities during secondary succession on the Loess Plateau. Microb Ecol. https://doi.org/10.1007/s00248-021-01740-9
Xu Y, Zhang W, Zhong Z, Guo S, Han X, Yang G, Ren C, Chen Z, Dai Y, Qiao W (2019) Vegetation restoration alters the diversity and community composition of soil nitrogen-fixing microorganisms in the Loess Hilly Region of China. Soil Sci Soc Am J 83(5):1378–1386. https://doi.org/10.2136/sssaj2019.03.0066
Xu Y, Zhong Z, Zhang W, Han X, Yang G, Ren C, Feng Y, Ren G, Wang X (2019) Responses of soil nosZ-type denitrifying microbial communities to the various land-use types of the Loess Plateau. China Soil Till Res 195:104378. https://doi.org/10.1016/j.still.2019.104378
Yan H, Huang YM, Jiang YL, Zhao T (2014) Soil mineralization under two kinds of shrub lands in mountainous area of sourthern Ningxia, Northwest China. Acta Sci Cirumstantiae 34:2111–2120. https://doi.org/10.13671/j.hjkxxb.2014.0538
Yang L, Lou J, Wang HZ, Wu LS, Xu JM (2018) Use of an improved high-throughput absolute abundance quantification method to characterize soil bacterial community and dynamics. Sci Total Environ 633:360–371. https://doi.org/10.1016/j.scitotenv.2018.03.201
Yu J, Liu F, Tripathi BM, Steinberger Y (2020) Changes in the composition of soil bacterial and fungal communities after revegetation with Caragana microphylla in a desertified semiarid grassland. J Arid Environ 182:104262. https://doi.org/10.1016/j.jaridenv.2020.104262
Yu K, Wang G (2018) Long-term impacts of shrub plantations in a desert-oasis ecotone: Accumulation of soil nutrients, salinity, and development of herbaceour layer. Land Degrad Dev 29(8):2681–2693. https://doi.org/10.1002/ldr.3009
Zhang C, Li Q, Zhou N, Zhang J, Kang L, Shen Y, Jia W (2016) Field observations of wind profiles and sand fluxes above the windward slope of a sand dune before and after the establishment of semi-buried straw checkerboard barriers. Aeolian Res 20:59–70. https://doi.org/10.1016/j.aeolia.2015.11.003
Zhang C, Liu GB, Xue S, Song ZL (2011) Rhizosphere soil microbial activity under different vegetation types on the Loess Plateau. China Geoderma 161(3–4):115–125. https://doi.org/10.1016/j.geoderma.2010.12.003
Zhang J, Zhang C, Zhou N, Ma X (2011) Spatial pattern of grain-size distribution in surface sediments as a result of variations in the aeolian environment in China’s Shapotou railway protective system. Aeolian Res 3(3):295–302. https://doi.org/10.1016/j.aeolia.2011.07.005
Zhang JY, Gu PF, Li LY, Zong LY, Zhao WJ (2016) Changes of soil particle size fraction along a chronosequence in sandy desertified land: a fundamental process for ecosystem succession and ecological restoration. J Soils Sediments 16(12):2651–2656. https://doi.org/10.1007/s11368-016-1454-x
Zhang TH, Su YZ, Cui JY, Zhang ZH, Chang XX (2006) A leguminous shrub (Caragana microphylla) in semiarid sandy soils of north China. Pedosphere 16(3):319–325. https://doi.org/10.1016/s1002-0160(06)60058-1
Zhang W, Ren C, Deng J, Zhao F, Yang G, Tong X, Feng Y, Han X (2018) Plant functional composition and species diversity affect soil C, N, and P during secondary succession of abandoned farmland on the Loess Plateau. Ecol Eng 122:91–99. https://doi.org/10.1016/j.ecoleng.2018.07.031
Zhang X, Li W, Zhong Z, Zhang Q, Wang X, Han X, Ren C, Yang G (2020) Response of soil microbial community to C:N: P stoichiometry along a Caragana korshinskii restoration gradient on the Loess Plateau. China Forests 11(8):823. https://doi.org/10.3390/f11080823
Zhang X, Zhao W, Wang L, Liu Y, Liu Y, Feng Q (2019) Relationship between soil water content and soil particle size on typical slopes of the Loess Plateau during a drought year. Sci Total Environ 648:943–954. https://doi.org/10.1016/j.scitotenv.2018.08.211
Zhang Y, Cao C, Cui Z, Qian W, Liang C, Wang C (2019) Soil bacterial community restoration along a chronosequence of sand-fixing plantations on moving sand dunes in the Horqin sandy land in northeast China. J Arid Environ 165:81–87. https://doi.org/10.1016/j.jaridenv.2019.04.003
Zhang ZH, Huisingh D (2018) Combating desertification in China: monitoring, control, management and revegetation. J Clean Prod 182:765–775. https://doi.org/10.1016/j.jclepro.2018.01.233
Zhao Y, Wang L, Knighton J, Evaristo J, Wassen M (2021) Contrasting adaptive strategies by Caragana korshinskii and Salix psammophila in a semiarid revegetated ecosystem. Agric For Meteorol 300:108323. https://doi.org/10.1016/j.agrformet.2021.108323
Zheng Q, Hu Y, Zhang S, Noll L, Bockle T, Dietrich M, Herbold CW, Eichorst SA, Woebken D, Richter A, Wanek W (2019) Soil multifunctionality is affected by the soil environment and by microbial community composition and diversity. Soil Biol Biochem 136:107521. https://doi.org/10.1016/j.soilbio.2019.107521
Zhou XJ, An XL, De Philippis R, Ye CR, Ke T, Zhang YR, Chen LZ (2019) The facilitative effects of shrub on induced biological soil crust development and soil properties. Appl Soil Ecol 137:129–138. https://doi.org/10.1016/j.apsoil.2019.02.010
Zhou XJ, Ke T, Li SX, Deng SQ, An XL, Ma X, De Philippis R, Chen LZ (2020) Induced biological soil crusts and soil properties varied between slope aspect, slope gradient and plant canopy in the Hobq desert of China. CATENA 190:104559. https://doi.org/10.1016/j.catena.2020.104559
Zhou Z, Yu M, Ding G, Gao G, He Y, Wang G (2020) Effects of Hedysarum leguminous plants on soil bacterial communities in the Mu Us Desert, northwest China. Ecol Evol 10(20):11423–11439. https://doi.org/10.1002/ece3.6779
Funding
Our study was funded by the Project for High-Level Overseas Talents Returning to China ([2019] 160), Ningxia Key Research and Development Project (2019BEB04011), National Natural Science Foundation of China [42001095], and Ningxia Natural Science Foundation (2020AAC03273, 2022AAC03641).
Author information
Authors and Affiliations
Contributions
Qiu KY and Xie YZ designed the study and conceived the idea; Liu WS, Huang YY, Wang RX, Li HC, Meng WF, He Y, Li YY, Li HQ, Zhao PB, and Yang Y conducted the experiments and collected the data; and Liu WS analyzed the data. All authors contributed to the draft and facilitated the publication of the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, W., Qiu, K., Xie, Y. et al. High-Throughput Absolute Quantification Sequencing Reveals that a Combination of Leguminous Shrubs Is Effective in Driving Soil Bacterial Diversity During the Process of Desertification Reversal. Microb Ecol 86, 1145–1163 (2023). https://doi.org/10.1007/s00248-022-02151-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-022-02151-0