Abstract
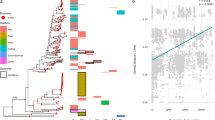
How ecological diversity is maintained and distributed within populations is a longstanding question in microbial ecology. In the thermophilic cyanobacterium Synechococcus B′, high observed levels of recombination are predicted to maintain ecological variation despite the simultaneous action of diverse selective pressures on different regions of the genome. To investigate ecological diversity in these bacteria, we directly isolated laboratory strains of Synechococcus B′ from samples collected along the thermal gradients of two geothermal environments in Yellowstone National Park. Extensive recombination was evident for a multi-locus sequence data set, and, consequently, our sample did not exhibit the sequence clustering expected for distinct ecotypes evolving by periodic clonal selection. Evidence for local selective sweeps at specific loci suggests that sweeps may be common but that recombination is effective for maintaining diversity of unlinked genomic regions. Thermal performance for strain growth was positively associated with the temperature of the environment, indicating that Synechococcus B′ populations consist of locally adapted ecological specialists that occupy specific thermal niches. Because this ecological differentiation is observed despite the absence of dispersal barriers among sites, we conclude that these bacteria may freely exchange much of the genome but that barriers to gene flow exist for loci under direct temperature selection.





Similar content being viewed by others
References
Cadillo-Quiroz H, Didelot X, Held NL, Herrera A, Darling A, Reno ML, Krause DJ, Whitaker RJ (2012) Patterns of gene flow define species of thermophilic Archaea. PLoS Biol. 10:e1001265
Shapiro BJ, Friedman J, Cordero OX, Preheim SP, Timberlake SC, Szabó G, Polz MF, Alm EJ (2012) Population genomics of early events in the ecological differentiation of bacteria. Science 336:48–51
Kashtan N, Roggensack SE, Rodrigue S, Thompson JW, Biller SJ, Coe A, Ding H, Marttinen P, Malmstrom RR, Stocker R, Follows MJ, Stepanauskas R, Chisholm SW (2014) Single-cell genomics reveals hundreds of coexisting subpopulations in wild Prochlorococcus. Science 344:416–420
Kraemer SA, Boynton PJ (2017) Evidence for microbial local adaptation in nature. Mol. Ecol. 26:1860–1876
Fraser C, Hanage WP, Spratt BG (2007) Recombination and the nature of bacterial speciation. Science 315:476–480
Cohan FM, Perry EB (2007) A systematics for discovering the fundamental units of bacterial diversity. Curr Biol 17:R373–R386
Wall CA, Koniges GJ, Miller SR (2014) Divergence with gene flow in a population of thermophilic bacteria: a potential role for spatially varying selection. Mol Ecol 23:3371–3383
Retchless AC, Lawrence JG (2007) Temporal fragmentation of speciation in bacteria. Science 317:1093–1096
Rosen MJ, Davison M, Bhaya D, Fisher DS (2015) Fine-scale diversity and extensive recombination in a quasisexual bacterial population occupying a broad niche. Science 348:1019–1023
Miller SR, Strong AL, Jones KL, Ungerer MC (2009) Bar-coded pyrosequencing reveals shared bacterial community properties along the temperature gradients of two alkaline hot springs in Yellowstone National Park. Appl Environ Microbiol 75:4565–4572
Haldane JBS (1948) The theory of a cline. J Genet 48:277–284
Endler J (1973) Differentiation of populations. Science 165:1228–1232
Wu C-I (2001) The genic view of the process of speciation. J Evol Biol 14:851–865
Wright S (1943) Isolation by distance. Genetics 28:114–138
Castenholz RW (1988) Culturing methods for cyanobacteria. Methods Enzymol 167:68–93
Miller SR, Castenholz RW (2000) Evolution of thermotolerance in hot spring cyanobacteria of the genus Synechococcus. Appl Environ Microbiol 66:4222–4229
Bhaya D, Grossman AR, Steunou A-S, Khuri N, Cohan FM, Hamamura N, Melendrez MC, Bateson MM, Ward DM, Heidelberg JF (2007) Population level functional diversity in a microbial community revealed by comparative genomic and metagenomic analyses. ISME J 1:703–713
Inskeep WP, Jay ZJ, Tringe SG, Herrgård MJ, Rusch DB, YNP Metagenome Project Steering Committee and Working Group Members (2013) The YNP metagenome project: environmental parameters responsible for microbial distribution in the Yellowstone geothermal ecosystem. Front Microbiol 4:67
Klatt CG, Inskeep WP, Herrgard MJ, Jay ZJ, Rusch DB, Tringe SG, Parenteau MN, Ward DM, Boomer SM, Bryant DA, Miller SR (2013) Community structure and function of high-temperature chlorophototrophic microbial mats inhabiting diverse geothermal environments. Front Microbiol 4:106
Huson DH (1998) SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14(1):68–73
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Bohonak AJ (2002) IBD (isolation by distance): a program for analyses of isolation by distance. J Hered 93:153–154
Huson DH, Bryant D (2005) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Allewalt JP, Bateson MM, Revsbech NP, Slack K, Ward DM (2006) Effect of temperature and light on growth of and photosynthesis by Synechococcus isolates typical of those predominating in the octopus spring microbial mat community of Yellowstone National Park. Appl Environ Microbiol 72:544–550
Ward DM, Castenholz RW, Miller SR (2012) Cyanobacteria in geothermal habitats. In: Whitton BE (ed) Ecology of cyanobacteria II. Springer, Dordrecht, pp 39–63
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003) The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4:981–994
Humbert J-F, Barbe V, Latifi A, Gugger M, Calteau A, Coursin T, Lajus A, Castelli V, Oztas S, Samson G, Longin C, Medigue C, de Marsac NT (2013) A tribute to disorder in the genome of the bloom-forming freshwater cyanobacterium Microcystis aeruginosa. PLoS One 8:e70747
Meyer KA, Davis TW, Watson SB, Denef VJ, Berry MA, Dick GJ (2017) Genome sequences of lower Great Lakes Microcystis sp. reveal strain-specific genes that are present and expressed in western Lake Erie blooms. PLoS One 12:e0183859
Acknowledgments
Field work was conducted under National Park Service research permit YELL-5482. We thank two anonymous reviewers for comments on an earlier version of the manuscript.
Funding
This work was supported by National Science Foundation award EF-0801999 to SRM.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Supplementary Fig. 1
(PDF 251 kb)
Supplementary Table 1
(DOCX 72 kb)
Rights and permissions
About this article
Cite this article
Miller, S.R., Carvey, D. Ecological Divergence with Gene Flow in a Thermophilic Cyanobacterium. Microb Ecol 78, 33–41 (2019). https://doi.org/10.1007/s00248-018-1267-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1267-0