Abstract
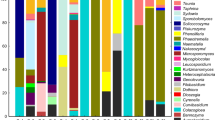
Lichens are structured associations of a fungus with a cyanobacteria and/or green algae in a symbiotic relationship, which provide specific habitats for diverse bacterial communities, including actinomycetes. However, few studies have been performed on the phylogenetic relationships and biosynthetic potential of actinomycetes across lichen species. In the present study, a total of 213 actinomycetes strains were isolated from 35 lichen samples (22 lichen genera) collected in Yunnan Province, China. 16S rRNA gene sequence analysis revealed an unexpected level of diversity among these isolates, which were distributed into 38 genera, 19 families, and 9 orders within the Actinobacteria phylum. The detailed taxa of isolates had no clear relationship to the taxonomic affiliations of the associated lichens. To the best of our knowledge, this is the first report to describe the isolation of Actinophytocola, Angustibacter, Herbiconiux, Kibdelosporangium, Kineosporia, Kitasatospora, Nakamurella, Nonomuraea, Labedella, Lechevalieria, Lentzea, Schumannella, and Umezawaea species from lichens. At least 40 isolates (18.78%) are likely to represent novel actinomycetes taxa within 15 genera. In addition, all 213 isolates were tested for antimicrobial activity and screened for genes associated with secondary metabolite production to evaluate their biosynthetic potential. These results demonstrate that the lichens of Yunnan Province represent an extremely rich reservoir for the isolation of a significant diversity of actinomycetes, including novel species, which are potential source for discovering biologically active compounds.

Similar content being viewed by others
References
Berdy J (2012) Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot 65:385–395
Bull AT, Stach JEM (2007) Marine actinobacteria: new opportunities for natural produce search and discovery. Trends Microbiol 15:491–499
Wagner M, Abdel-Mageed WM, Ebel R, Bull AT, Goodfellow M, Fiedler HP, Jaspars M (2014) Dermacozines H-J isolated from a deep-sea strain of Dermacoccus abyssi from Mariana Trench sediments. J Nat Prod 77:416–420
Li J, Jia H, Cai X, Zhong H, Feng Q, Sunagawa S, Arumugam M, Kultima JR, Prifti E, Nielsen T, Juncker AS, Manichanh C, Chen B, Zhang W, Levenez F, Wang J, Xu X, Xiao L, Liang S, Zhang D (2014) An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32:834–841
Kim KH, Ramadhar TR, Beemelmanns C, Cao S, Poulsen M, Currieb CR, Clardy J (2014) Natalamycin A, an ansamycin from a termite-associated Streptomyces sp. Chem Sci 5:4333–4338
Janso JE, Carter GT (2010) Biosynthetic potential of phylogenetically unique endophytic actinomycetes from tropical plants. Appl Environ Microbiol 76:4377–4386
Qin S, Li J, Chen HH, Zhao GZ, Zhu WY, Jiang CL, Xu LH, Li WJ (2009) Isolation, diversity, and antimicrobial activity of rare actinobacteria from medicinal plants of tropical rain forests in Xishuangbanna, China. Appl Environ Microbiol 75:6176–6186
Ezra D, Castillo UF, Strobel GA, Hess WM, Porter H, Jensen JB, Condron MAM, Teplow DB, Sears J, Maranta M, Hunter M, Weber B, Yaver D (2004) Coronamycins, peptide antibiotics produced by a verticillated Streptomyces sp. (MSU-2110) endophytic on Monstera sp. Microbiology 150:785–793
Castillo U, Harper JK, Strobel GA, Sears J, Alesi K, Ford E, Lin J, Hunter M, Maranta M, Ge H, Yaver D, Jensen JB, Porter H, Robison R, Millar D, Hess WM, Condron M, Teplow D (2003) Kakadumycins, novel antibiotics from Streptomyces sp. NRRL 30566, an endophyte of Grevillea pteridifolia. FEMS Letters 224:183–190
Castillo UF, Browne L, Strobel G, Hess WM, Ezra S, Pacheco G, Ezra D (2007) Biologically active endophytic streptomycetes from Nothofagus spp. and other plants in Patagonia. Microb Ecol 53:12–19
Okoro CK, Brown R, Jones AL, Andrews BA, Asenjo JA, Goodfellow M, Bull AT (2009) Diversity of culturable actinomycetes in hyper-arid soils of the Atacama Desert, Chile. Antonie Van Leeuwenhoek 95:121–133
Hong K, Gao AH, Xie QY, Gao H, Zhuang L, Lin HP, Yu HP, Yao XS, Goodfellow M, Ruan JS (2009) Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar Drugs 7:24–44
Gonzάlz I, Ayuso-Sacido A, Anderson A, Genilloud O (2005) Actinomycetes isolated from lichens: evaluation of their diversity and detection of biosynthetic gene sequences. FEMS Microbiol Ecol 54:401–415
Parrot D, Antony-Babu S, Intertaglia L, Grube M, Tomasi S, Suzuki M (2015) Littoral lichens as a novel source of potentially bioactive actinobacteria. Sci Rep 5:15839. doi:10.1038/srep15839
Davies J, Wang H, Taylor T, Warabi K, Huang XH, Andersen RJ (2005) Uncialamycin, a new enediyne antibiotic. Org Lett 7:5233–5236
Williams DE, Davies J, Patrick BO, Bottriell H, Tarling T, Roberge M, Andersen RJ (2008) Cladoniamides A-G, tryptophan-derived alkaloids produced in culture by Streptomyces uncialis. Org Lett 10:3501–3504
Motohashi K, Takagi M, Yamamura H, Hayakawa M, Shin-ya K (2010) A new angucycline and a new butenolide isolated from lichen-derived Streptomyces spp. J Antibiot 63:545–548
Tehler A, Wedin M (2008) Systematics of lichenized fungi. In: Nash TH (ed) Progress in botany, vol 46. Cambridge University Press, Cambridge, pp. 336–352
Cardinale M, de Castro JV, Müller H, Berg G, Grube M (2008) In situ analysis of the bacterial community associated with the reindeer lichen Cladonia arbuscula reveals predominance of Alphaproteobacteria. FEMS Microbiol Ecol 66:63–71
Cardinale M, Puglia AM, Grube M (2006) Molecular analysis of lichen-associated bacterial communities. FEMS Microbiol Ecol 57:484–495
Grube M, Cardinale M, de Castro JV, Müller H, Berg G (2009) Species-specific structural and functional diversity of bacterial communities in lichen symbioses. ISME J 3:1105–1115
Hodkinson B, Lutzoni F (2009) A microbiotic survey of lichen-associated bacteria reveals a new lineage from the Rhizobiales. Symbiosis 49:163–180
Bates ST, Cropsey GWG, Caporaso JG, Knight R, Fierer N (2011) Bacterial communities associated with the lichen symbiosis. Appl Environ Microbiol 77:1309–1314
Phongsopitanun W, Matsumoto A, Inahashi Y, Kudo T, Mori M, Shiomi K, Takahashi Y, Tanasupawat S (2016) Actinoplanes lichenis sp. nov., isolated from lichen. Int J Syst Evol Microbiol 66:468–473
Cardinale M, Grube M, Berg G (2011) Frondihabitans cladoniiphilus sp. nov., an actinobacterium of the family Microbacteriaceae isolated from lichen, and emended description of the genus Frondihabitans. Int J Syst Evol Microbiol 61:3033–3038
Yamamura H, Ashizawa H, Nakagawa Y, Hamada M, Ishida Y, Otoguro M, Tamura T, Hayakawa M (2011) Actinomycetospora rishiriensis sp. nov., isolated from a lichen. Int J Syst Evol Microbiol 61:2621–2625
Yamamura H, Ashizawa H, Nakagawa Y, Hamada M, Ishida Y, Otoguro M, Tamura T, Hayakawa M (2011) Actinomycetospora iriomotensis sp. nov., a novel actinomycete isolated from a lichen sample. J Antibiot 64:289–292
Hirsch P, Mevs U, Kroppenstedt RM, Schumann P, Stackebrandt E (2004) Crytpoendolithic actinomycetes from antarctic sandstone: Micromonospora endolithica sp. nov. and two isolates related to Micromonospora coerulea Jensen 1932. Syst Appl Microbiol 27:166–174
An SY, Xiao T, Yokota A (2009) Leifsonia lichenia sp. nov., isolated from lichen in Japan. J Gen Appl Microbiol 55:339–343
Li B, Xie CH, Yokota A (2007) Nocardioides exalbidus sp. nov., a novel actinomycete isolated from lichen in Izu-Oshima Island, Japan. Actinomycetologica 21:22–26
An S, Xiao T, Yokota A (2008) Schumannella luteola gen. nov., sp. nov., a novel genus of the family Microbacteriaceae. J Gen Appl Microbiol 54:253–258
Shen B (2003) Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms. Curr Opin Chem Biol 7:285–295
Christiansen G, Dittmann E, Ordorika LV, Rippka R, Herdman M, Börner T (2001) Nonribosomal peptide synthase genes occur in most cyanobacterial genera as evidenced by their distribution in axenic strains of the PCC. Arch Microbiol 176:452–458
Ayuso-Sacido A, Genilloud O (2005) New PCR primers for the screening of NRPS and PKS-I systems in actinomycetes: detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb Ecol 49:10–24
Ketela MM, Virpi S, Halo L, Hautala A, Hakala J, Mantsala P, Ylihonko K (1999) An efficient approach for screening minimal PKS genes from Streptomyces. FEMS Microbiol Lett 180:1–6
Demain AL, Sanchez S (2009) Microbial drug discovery: 80 years of progress. J Antibiot 62:5–16
Anderson AS, Clark DJ, Gibbons PH, Sigmund JM (2002) The detection of diverse aminoglycoside phosphotransferases within natural populations of actinomycetes. J Ind Microbiol Biotechnol 29:60–69
Sigmund JM, Clark DC, Rainey FA, Anderson AS (2003) Detection of eubacterial 3-hydroxy-3-methylglutaryl coenzyme a reductases from natural populations of actinomycetes. Microb Ecol 46:106–112
Hwang YB, Lee MY, Park HJ, Han K, Kim ES (2007) Isolation of putative polyene-producing actinomycetes strains via PCR-based genome screening for polyene-specific hydroxylase genes. Process Biochem 42:102–107
Biodiversity Committee of Chinese Academy of Sciences (2016) Catalogue of life china 2016 annual checklist edition. Science press, Beijing, China
Jiang Y, Wang XY, Li GD, Li QY, Chen X, Wang LS, Li Y, Jiang CL (2015) Diversity and anti-microbial activities of actinomycetes associated with three species of lichens. Am J BioSci 3:171–177
Hayakawa M, Nonomura H (1987) Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J Ferment Technol 65:501–509
Chun J, Goodfellow M (1995) A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int J Syst Bacteriol 45:240–245
Cui XL, Mao PH, Zeng M, Li WJ, Zhang LP, Xu LH, Jiang CL (2001) Streptomonospora salina gen. nov., sp. nov., a new member of the family Nocardiopsaceae. Int J Syst Evol Microbiol 51:357–363
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Bergquist PR, Bedford JJ (1978) The incidence of antibacterial activity in marine demospongiae; systematic and geographic considerations. Mar Biol 46:215–221
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence-analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Meier-Kolthoff JP, Göker M, Spröer C, Klenk HP (2013) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol 195:413–418
Stach JE, Maldonado LA, Masson DG, Ward AC, Goodfellow M, Bull AT (2003) Statistical approaches for estimating actinobacterial diversity in marine sediments. Appl Environ Microbiol 69:6189–6200
Robert DS, George WC, John JD, Walter H, John REH, Joseph RV, Lee W, Kenneth MS (1985) Aridicins, novel glycopeptide antibiotics II. Isolation and charactrization. J Antibiot 38:555–560
Shearer MC, Giovenella AJ, Grappel SF, Hedde RD, Mehta RJ, Oh YK, Pan CH, Pitkin DH, Nisbet LJ (1986) Kibdelins, novel glycopeptide antibiotics I. Discovery, production, and biological evaluation. J Antibiot 39:1386–1394
Choi CW, Choi JS, Ko YK, Kim CJ, Kim YH, Oh JS, Ryu SY, Yon GH (2014) A new tricyclic undecose nucleoside from Streptomyces scopuliridis RB72. Bull Kor Chem Soc 45:1215–1217
Farris MH, Duffy C, Findlay RH, Olson JB (2011) Streptomyces scopuliridis sp. nov., a bacteriocin-producing soil streptomycete. Int J Syst Evol Microbiol 61:2112–2116
Song Y, Li Q, Liu X, Chen Y, Zhang Y, Sun A, Zhang W, Zhang J, Ju J (2014) Cyclic hexapeptides from the deep south china sea-derived Streptomyces scopuliridis SCSIO zj46 active against pathogenic Gram-positive bacteria. J Nat Prod 77:1937–1941
David M, Pascal B (2015) Polypeptides isolated from Brevibacterium aurantiacum and their use for the treatment of cancer. USA Patent: US9,051,562 B2
Zhou X, Huang H, Li J, Song Y, Jiang R, Liu J, Zhang S, Hua Y, Ju J (2014) New anti-infective cycloheptadepsipeptide congeners and absolute stereochemistry from the deep sea-derived Streptomyces drozdowiczii SCSIO 10141. Tetrahedron 70:7795–7801
Aoyagi T, Suda H, Uotani K, Kojima F, Aoyama T, Horiguchi K, Hamada M, Takeuchi T (2005) Nagstatin, a new inhibitor of N-acetyl-β-d-glucosaminidase, produced by Streptomyces amakusaensis MG846-fF3, taxonomy, production, isolation, physico-chemical properties and biological activities. J Antibiot 15:374–388
Haneda K, Shinose M, Seino A, Tabata N, Tomoda H, Iwai Y, Ōmura S (1994) Cytosaminomycins, new anticoccidial agents produced by Streptomyces sp. KO-8119 I. Taxonomy, production, isolation and physico-chemical and biological properties. J Antibiot 47:756–761
Deushi T, Iwasaki A, Kamiya K, Mizoguchi T, Nakayama M, Itoh H, Mori T (1979) New aminoglycoside antibiotics, sannamycin. J Antibiot 32:1061–1065
Yang SX, Gao JM, Zhang AL, Laatsch H (2011) Sannastatin, a novel toxic macrolactam polyketide glycoside produced by actinomycete Streptomyces sannanensis. Bioorg Med Chem Lett 21:3905–3908
Li Y, Zheng D, Li J, Han L, Cui X, Lang L, Li M, Wang Z, Zhao J, Huang X (2009) Sannanine, a new cytotoxic alkaloid from Streptomyces sannanensis. J Antibiot 62:647–648
Aroonsri A, Kitani S, Ikeda H, Nihira T (2012) Kitasetaline, a novel β-carboline alkaloid from Kitasatospora setae NBRC 14216T. J Biosci Bioeng 114:56–58
Bowman EJ, Siebers A, Altendorf K (1988) Bafilomycins: a class of inhibitors of membrane ATPases from microorganisms, animal cells, and plant cells. Proc Natl Acad Sci USA 85:7972–7976
Ōmura S, Otoguro K, Nishikiori T, Oiwa R, Iwai Y (1981) Setamycin, a new antibiotic. J Antibiot 34:1253–1256
Terekhova LP, Sadikova OA, Preobrazhenskaia TP (1977) New species of Actinoplanes cyanea sp. nov. and its antagonistic properties. Antibiotiki 22:1059–1063
An DS, Im WT, Yoon MH (2008) Microlunatus panaciterrae sp. nov., a β-glucosidase-producing bacterium isolated from soil in a ginseng field. Int J Syst Evol Microbiol 58:2734–2738
Singh SB, Garrity GM, Genillourd O, Lingham RB, Martin I, NallinOmstead M, Silverman KC, Zink DL (1997) Inhibitor compounds of farnesyl-protein transferase and chemotherapeutic compositions containing the same, produced by strain ATCC 55532. USA Patent 5:627,057
Cheenpracha S, Vidor NB, Yoshida WY, Davies J, Chang LC (2010) Coumabiocins A-F, aminocoumarins from an organic extract of Streptomyces sp. L-4-4. J Nat Prod 73:880–884
Acknowledgments
This work was funded by the National Natural Science Foundation of China (Grant nos. 31460005, 31270001, 81573327, 31170023, 31400022, 31370069). We thank graduate Xin Ye (Yunnan University of Traditional Chinese Medicine, China) for his help in lichen collection.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
.
ESM 1
(DOCX 108 kb)
Rights and permissions
About this article
Cite this article
Liu, C., Jiang, Y., Wang, X. et al. Diversity, Antimicrobial Activity, and Biosynthetic Potential of Cultivable Actinomycetes Associated with Lichen Symbiosis. Microb Ecol 74, 570–584 (2017). https://doi.org/10.1007/s00248-017-0972-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-0972-4