Abstract
Both the structure and function of host-associated microbial communities are potentially impacted by environmental conditions, just as the outcomes of many free-living species interactions are context-dependent. Many amphibian populations have declined around the globe due to the fungal skin pathogen, Batrachochytrium dendrobatidis (Bd), but enivronmental conditions may influence disease dynamics. For instance, in Panamá, the most severe Bd outbreaks have occurred at high elevation sites. Some amphibian species harbor bacterial skin communities that can inhibit the growth of Bd, and therefore, there is interest in understanding whether environmental context could also alter these host-associated microbial communities in a way that might ultimately impact Bd dynamics. In a field survey in Panamá, we assessed skin bacterial communities (16S rRNA amplicon sequencing) and metabolite profiles (HPLC-UV/Vis) of Silverstoneia flotator from three high- and three low-elevation populations representing a range of environmental conditions. Across elevations, frogs had similar skin bacterial communities, although one lowland site appeared to differ. Interestingly, we found that bacterial richness decreased from west to east, coinciding with the direction of Bd spread through Panamá. Moreover, metabolite profiles suggested potential functional variation among frog populations and between elevations. While the frogs have similar bacterial community structure, the local environment might shape the metabolite profiles. Ultimately, host-associated community structure and function could be dependent on environmental conditions, which could ultimately influence host disease susceptibility across sites.





Similar content being viewed by others


References
McFall-Ngai M, Hadfield MG, Bosch TCG, et al. (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl. Acad. Sci. U. S. A. 110:3229–3236. doi:10.2307/42583580?ref=search-gateway:7b87d5838b0343fae5e2fa5e5b3a74e2
Kaltenpoth M, Strupat K, Svatos A (2015) Linking metabolite production to taxonomic identity in environmental samples by (MA)LDI-FISH. ISME J 10:527–531. doi:10.1038/ismej.2015.122
Rebollar EA, Antwis RE, Becker MH, et al. (2016) Using “omics” and integrated multi-omics approaches to guide probiotic selection to mitigate chytridiomycosis and other emerging infectious diseases. Front. Microbiol. 7:733–719. doi:10.3389/fmicb.2016.00068
Wellington EM, Berry A, Krsek M (2003) Resolving functional diversity in relation to microbial community structure in soil: exploiting genomics and stable isotope probing. Curr. Opin. Microbiol. 6:295–301. doi:10.1016/S1369-5274(03)00066-3
Muegge BD, Kuczynski J, Knights D, et al. (2011) Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332:970–974. doi:10.1126/science.1198719
Harris RN, Lauer A, Simon MA, et al. (2009) Addition of antifungal skin bacteria to salamanders ameliorates the effects of chytridiomycosis. Dis. Aquat. Org. 83:11–16. doi:10.3354/dao02004
Hoyt JR, Cheng TL, Langwig KE, et al. (2015) Bacteria isolated from bats inhibit the growth of Pseudogymnoascus destructans, the causative agent of white-nose syndrome. PLoS One 10:e0121329–e0121312. doi:10.1371/journal.pone.0121329
Fraune S, Anton-Erxleben F, Augustin RE, et al. (2014) Bacteria-bacteria interactions within the microbiota of the ancestral metazoan Hydra contribute to fungal resistance. ISME J 9:1543–1556. doi:10.1038/ismej.2014.239
Rosenberg E, Koren O, Reshef L, et al. (2007) The role of microorganisms in coral health, disease and evolution. Nat Rev Micro 5:355–362. doi:10.1038/nrmicro1635
Sarmiento-Ramírez JM, van der Voort M, Raaijmakers JM, Diéguez-Uribeondo J (2014) Unravelling the microbiome of eggs of the endangered sea turtle Eretmochelys imbricata identifies bacteria with activity against the emerging pathogen fusarium falciforme. PLoS One 9:e95206–e95208. doi:10.1371/journal.pone.0095206
Flórez LV, Biedermann PHW, Engl T, Kaltenpoth M (2015) Defensive symbioses of animals with prokaryotic and eukaryotic microorganisms. Nat. Prod. Rep. 32:904–936. doi:10.1039/C5NP00010F
Walke JB, Becker MH, Loftus SC, et al. (2014) Amphibian skin may select for rare environmental microbes. ISME J 8:2207–2217. doi:10.1038/ismej.2014.77
Kueneman JG, Parfrey LW, Woodhams DC, et al. (2013) The amphibian skin-associated microbiome across species, space and life history stages. Mol. Ecol. 23:1238–1250. doi:10.1111/mec.12510
Belden LK, Hughey MC, Rebollar EA, et al. (2015) Panamanian frog species host unique skin bacterial communities. Front. Microbiol. 6:32–21. doi:10.3389/fmicb.2015.01171
Becker MH, Walke JB, Murrill L, et al. (2015) Phylogenetic distribution of symbiotic bacteria from Panamanian amphibians that inhibit growth of the lethal fungal pathogen Batrachochytrium dendrobatidis. Mol. Ecol. 24:1628–1641. doi:10.1111/mec.13135
Harris RN, James TY, Lauer A, et al. (2006) Amphibian pathogen Batrachochytrium dendrobatidis is inhibited by the cutaneous bacteria of amphibian species. EcoHealth 3:53–56. doi:10.1007/s10393-005-0009-1
Harris RN, Brucker RM, Walke JB, et al. (2009) Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME J 3:818–824. doi:10.1038/ismej.2009.27
Becker MH, Brucker RM, Schwantes CR, et al. (2009) The bacterially produced metabolite Violacein is associated with survival of amphibians infected with a lethal fungus. Appl. Environ. Microbiol. 75:6635–6638. doi:10.1128/AEM.01294-09
Flechas SV, Sarmiento C, Cárdenas ME, et al. (2012) Surviving chytridiomycosis: differential anti-Batrachochytrium dendrobatidis activity in bacterial isolates from three lowland species of Atelopus. PLoS One 7:e44832–e44837. doi:10.1371/journal.pone.0044832
Brucker RM, Harris RN, Schwantes CR, et al. (2008) Amphibian chemical defense: antifungal metabolites of the microsymbiont Janthinobacterium lividum on the salamander Plethodon cinereus. J. Chem. Ecol. 34:1422–1429. doi:10.1007/s10886-008-9555-7
Brucker RM, Baylor CM, Walters RL, et al. (2007) The identification of 2,4-diacetylphloroglucinol as an antifungal metabolite produced by cutaneous bacteria of the salamander Plethodon cinereus. J. Chem. Ecol. 34:39–43. doi:10.1007/s10886-007-9352-8
Daszak P, Berger L, Cunningham AA, et al. (1999) Emerging infectious diseases and amphibian population declines. Emerging Infect Dis 5:735–748. doi:10.3201/eid0506.990601
Olson DH, Aanensen DM, Ronnenberg KL, et al. (2013) Mapping the global emergence of Batrachochytrium dendrobatidis, the amphibian chytrid fungus. PLoS One 8:e56802–e56813. doi:10.1371/journal.pone.0056802
Bronstein JL, Wilson WG, Morris WF (2003) Ecological dynamics of mutualist/antagonist communities. Am. Nat. 162:S24–S39. doi:10.1086/378645
Agrawal AA, Ackerly DD, Adler F, et al. (2007) Filling key gaps in population and community ecology. Front. Ecol. Environ. 5:145–152. doi:10.1890/1540-9295(2007)5[145:FKGIPA]2.0.CO;2
Hopkins SR, Wojdak JM, Belden LK (2016) Defensive symbionts mediate host–parasite interactions at multiple scales. Trends Parasitol.:1–12. doi:10.1016/j.pt.2016.10.003
Hofstetter R, Dempsey T, Klepzig K, Ayres M (2007) Temperature-dependent effects on mutualistic, antagonistic, and commensalistic interactions among insects, fungi and mites. Community Ecology 8:47–56. doi:10.1556/ComEc.8.2007.1.7
Shnit-Orland M, Kushmaro A (2009) Coral mucus-associated bacteria: a possible first line of defense. FEMS Microbiol. Ecol. 67:371–380. doi:10.1111/j.1574-6941.2008.00644.x
Kushmaro A, Loya Y, Fine M, Rosenberg E (1996) Bacterial infection and coral bleaching. doi: 10.1038/380396a0
Boyles JG, Willis CK (2010) Could localized warm areas inside cold caves reduce mortality of hibernating bats affected by white-nose syndrome? Front. Ecol. Environ. 8:92–98. doi:10.1890/080187
James TY, Toledo LF, Rödder D, et al. (2015) Disentangling host, pathogen, and environmental determinants of a recently emerged wildlife disease: lessons from the first 15 years of amphibian chytridiomycosis research. Ecol Evol 5:4079–4097. doi:10.1002/ece3.1672
Daskin JH, Alford RA (2012) Context-dependent symbioses and their potential roles in wildlife diseases. Proceedings of the Royal Society B: Biological Sciences 279:1457–1465. doi: 10.1098/rspb.2011.2276
Crawford AJ, Lips KR, Bermingham E, Wake DB (2010) Epidemic disease decimates amphibian abundance, species diversity, and evolutionary history in the highlands of central Panama. Proc. Natl. Acad. Sci. U. S. A. 107:13777–13782. doi:10.2307/25708801?ref=search-gateway:c5e8aac1656df05dfd381d5c42b5c4df
Lips KR, Brem F, Brenes R, et al. (2006) Emerging infectious disease and the loss of biodiversity in a Neotropical amphibian community. Proceedings of the National Academy of Sciences 103:3165–3170. doi: 10.1073/pnas.0506889103
Ryan MJ, Lips KR, Eichholz MW (2008) Decline and extirpation of an endangered Panamanian stream frog population (Craugastor punctariolus) due to an outbreak of chytridiomycosis. Biol. Conserv. 141:1636–1647. doi:10.1016/j.biocon.2008.04.014
Woodhams DC, Kilburn VL, Reinert LK, et al. (2008) Chytridiomycosis and amphibian population declines continue to spread eastward in Panama. EcoHealth 5:268–274. doi:10.1007/s10393-008-0190-0
Rebollar EA, Hughey MC, Harris RN, et al. (2014) The lethal fungus Batrachochytrium dendrobatidis is present in lowland tropical forests of far eastern Panamá. PLoS One 9:e95484–e95488. doi:10.1371/journal.pone.0095484
Sapsford SJ, Alford RA, Schwarzkopf L (2013) Elevation, temperature, and aquatic connectivity all influence the infection dynamics of the amphibian chytrid fungus in adult frogs. PLoS One. doi:10.1371/journal.pone.0082425
Catenazzi A, Lehr E, Vredenburg VT (2013) Thermal physiology, disease, and amphibian declines on the eastern slopes of the Andes. Conserv. Biol. 28:509–517. doi:10.1111/cobi.12194
Gründler MC, Toledo LF, Parra-Olea G, et al. (2012) Interaction between breeding habitat and elevation affects prevalence but not infection intensity of Batrachochytrium dendrobatidis in Brazilian anuran assemblages. Dis. Aquat. Org. 97:173–184. doi:10.3354/dao02413
Kriger KM, Hero J-M (2008) Altitudinal distribution of chytrid (Batrachochytrium dendrobatidis) infection in subtropical Australian frogs. Austral Ecology 33:1022–1032. doi:10.1111/j.1442-9993.2008.01872.x
Brem F, Lips KR (2008) Batrachochytrium dendrobatidis infection patterns among Panamanian amphibian species, habitats and elevations during epizootic and enzootic stages. Dis. Aquat. Org. 81:189–202. doi:10.3354/dao01960
Jaramillo AC, Wilson LD, Ibáñez R, Jaramillo F (2010) The herpetofauna of Panama: distribution and conservation status. Conservation of Mesoamerican amphibians and reptiles. Eagle Mountain Press, Eagle Mountain, pp. 604–671
Lauer A, Simon MA, Banning JL, et al. (2007) Common cutaneous bacteria from the eastern red-backed salamander can inhibit pathogenic fungi. Copeia 630–640
Umile TP, McLaughlin PJ, Johnson KR, et al. (2014) Nonlethal amphibian skin swabbing of cutaneous natural products for HPLC fingerprinting. Anal. Methods 6:3277–3284. doi:10.1039/C4AY00566J
Lauber CL, Zhou N, Gordon JI, et al. (2010) Effect of storage conditions on the assessment of bacterial community structure in soil and human-associated samples. FEMS Microbiol. Lett. 307:80–86. doi:10.1111/j.1574-6968.2010.01965.x
Whitfield SM, Kerby J, Gentry LR, Donnelly MA (2012) Temporal variation in infection prevalence by the amphibian chytrid fungus in three species of frogs at La Selva, Costa Rica. Biotropica 44:779–784. doi:10.1111/j.1744-7429.2012.00872.x
Caporaso JG, Lauber CL, Walters WA, et al. (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. doi:10.1038/ismej.2012.8
Aronesty E (2011) ea-utils: Command-line tools for processing biological sequencing data. Expression Analysis 1–1
Caporaso JG, Kuczynski J, Stombaugh J, et al. (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Meth 7:335–336. doi:10.1038/nmeth.f.303
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. doi:10.1093/bioinformatics/btq461
DeSantis TZ, Hugenholtz P, Larsen N, et al. (2006) Greengenes, a chimera-checked 16S rRNA Gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72:5069–5072. doi:10.1128/AEM.03006-05
Caporaso JG, Bittinger K, Bushman FD, et al. (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73:5261–5267. doi:10.1128/AEM.00062-07
Bokulich NA, Subramanian S, Faith JJ, et al. (2012) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Meth 10:57–59. doi:10.1038/nmeth.2276
Boyle DG, Boyle DB, Olsen V, et al. (2004) Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real-time Taqman PCR assay. Dis. Aquat. Org. 60:141–148. doi:10.3354/dao060141
Becker MH, Walke JB, Cikanek S, et al. (2015) Composition of symbiotic bacteria predicts survival in Panamanian golden frogs infected with a lethal fungus. Proceedings of the Royal Society B: Biological Sciences 282:20142881–20142881. doi: 10.1098/rspb.2014.2881
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Bates D, Maechler M, Bolker BM, Walker SC (2014) Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67:1–48
Venables WN, Ripley BD (2002) Modern applied statistics with S. Fourth Edition. 1–1
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom. J. 50:346–363. doi:10.1002/bimj.200810425
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecology 26:32–46. doi:10.1111/j.1442-9993.2001.01070.pp.x
Okansen J, Blanchet FG, Kindt R, et al. (2016) vegan: community ecology package. R package version 2.3–5 https://CRAN.R-project.org/package=vegan. 1–1
Shade A, Handelsman J (2011) Beyond the Venn diagram: the hunt for a core microbiome. Environ. Microbiol. 14:4–12. doi:10.1111/j.1462-2920.2011.02585.x
Warnes GR, Bolker B, Bonebakker L, et al. (2016) gplots: various R programming tools for plotting data. R package version 3.0.1 https://CRAN.R-project.org/package=gplots. 1–1
Bryant JA, Lamanna C, Morlon H, et al. (2008) Colloquium paper: microbes on mountainsides: contrasting elevational patterns of bacterial and plant diversity. Proc. Natl. Acad. Sci. U. S. A. 105(Suppl 1):11505–11511. doi:10.1073/pnas.0801920105
Singh D, Takahashi K, Kim M, et al. (2012) A hump-backed trend in bacterial diversity with elevation on Mount Fuji, Japan. Microb. Ecol. 63:429–437. doi:10.1007/s00248-011-9900-1
Fierer N, McCain CM, Meir P, et al. (2011) Microbes do not follow the elevational diversity patterns of plants and animals. Ecology 92:797–804
Yang Y, Gao Y, Wang S, et al. (2014) The microbial gene diversity along an elevation gradient of the Tibetan grassland. ISME J 8:430–440. doi:10.1038/ismej.2013.146
Daskin JH, Bell SC, Schwarzkopf L, Alford RA (2014) Cool temperatures reduce antifungal activity of symbiotic bacteria of threatened amphibians—implications for disease management and patterns of decline. PLoS One 9:e100378–e100379. doi:10.1371/journal.pone.0100378
Woodhams DC, Brandt H, Baumgartner S, et al. (2014) Interacting symbionts and immunity in the amphibian skin mucosome predict disease risk and probiotic effectiveness. PLoS One 9:e96375–e96313. doi:10.1371/journal.pone.0096375
Gacheva G, Gigova L (2014) Biological activity of microalgae can be enhanced by manipulating the cultivation temperature and irradiance. Open Life Sciences 9:1–14. doi:10.2478/s11535-014-0350-x
Humair B, González N, Mossialos D, et al. (2009) Temperature-responsive sensing regulates biocontrol factor expression in Pseudomonas fluorescens CHA0. ISME J 3:955–965. doi:10.1038/ismej.2009.42
Raaijmakers JM, Vlami M, de Souza JT (2002) Antibiotic production by bacterial biocontrol agents. Antonie Van Leeuwenhoek 81:537–547
Cundliffe E (2006) Antibiotic production by actinomycetes: the Janus faces of regulation. J. Ind. Microbiol. Biotechnol. 33:500–506. doi:10.1007/s10295-006-0083-6
Krause S, Le Roux X, Niklaus PA (2014) Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Salamandra 5:1–10. doi:10.3389/fmicb.2014.00251/abstract
Webster G, Embley TM, Freitag TE (2005) Links between ammonia oxidizer species composition, functional diversity and nitrification kinetics in grassland soils. Salamandra 7:676–684. doi:10.1111/j.1462-2920.2004.00740.x
Attard E, Poly F, Commeaux C, et al. (2010) Shifts between Nitrospira- and Nitrobacter-like nitrite oxidizers underlie the response of soil potential nitrite oxidation to changes in tillage practices. Environ. Microbiol. 12:315–326. doi:10.1111/j.1462-2920.2009.02070.x
Walke JB, Becker MH, Loftus SC, et al. (2015) Community structure and function of amphibian skin microbes: an experiment with bullfrogs exposed to a chytrid fungus. PLoS One 10:e0139848. doi:10.1371/journal.pone.0139848
Jani AJ, Briggs CJ (2014) The pathogen Batrachochytrium dendrobatidis disturbs the frog skin microbiome during a natural epidemic and experimental infection. Proceedings of the National Academy of Sciences 111:E5049–E5058. doi: 10.1073/pnas.1412752111
Bresciano JC, Salvador CA, Paz-y-Miño C, et al. (2015) Variation in the presence of anti-Batrachochytrium dendrobatidis bacteria of amphibians across life stages and elevations in Ecuador. EcoHealth 12:310–319. doi:10.1007/s10393-015-1010-y
Lips KR, Diffendorfer J, Mendelson JR, Sears MW (2008) Riding the wave: reconciling the roles of disease and climate change in amphibian declines. PLoS Biol. 6:e72–e14. doi:10.1371/journal.pbio.0060072
Rodríguez-Brenes S, Rodríguez D, Ibáñez R, Ryan MJ (2016) Spread of amphibian chytrid fungus across lowland populations of Túngara frogs in Panamá. PLoS One 11:e0155745–e0155748. doi:10.1371/journal.pone.0155745
Rebollar EA, Hughey MC, Medina D, et al. (2016) Skin bacterial diversity of Panamanian frogs is associated with host susceptibility and presence of Batrachochytrium dendrobatidis. ISME J 10:1682–1695. doi:10.1038/ismej.2015.234
Hacioglu N, Tosunoglu M (2013) Determination of antimicrobial and heavy metal resistance profiles of some bacteria isolated from aquatic amphibian and reptile species. Environ. Monit. Assess. 186:407–413. doi:10.1007/s10661-013-3385-y
Acknowledgements
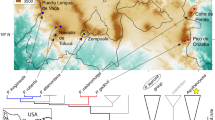
Thanks to Molly Bletz for assistance in the field, Skylar Hopkins and James Skelton for statistical advice, Eria Rebollar for help with the export and import of samples, and Jennifer Smith for producing Fig. 1. We also thank the Smithsonian Tropical Research Institute for providing logistical support and laboratory space and the Interfaces of Global Change graduate program at Virginia Tech for fellowship support to DM. Funds were provided by the National Science Foundation (NSF: DEB-1136640 to LKB; DEB-1136662 to KPCM). Samples were collected under the scientific permit SE/A-35-13 granted by MIAmbiente.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Medina, D., Hughey, M.C., Becker, M.H. et al. Variation in Metabolite Profiles of Amphibian Skin Bacterial Communities Across Elevations in the Neotropics. Microb Ecol 74, 227–238 (2017). https://doi.org/10.1007/s00248-017-0933-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-0933-y