Abstract
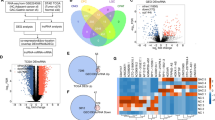
Non-small cell lung cancer (NSCLC) is one of the most lethal cancer types in the world. Currently, the molecular mechanisms and pathways underlying NSCLC oncogenesis are poorly understood. Using multiple Omics data, we systematically explored the differentially expressed circular RNAs (circRNAs) in NSCLC. We also investigated potential microRNA sponges (that absorb circRNAs) in NSCLC and downstream target genes with experimental verifications. hsa_circ_0003497 was down-regulated in NSCLC and played an inhibitory role in tumorigenesis. In contrast, miR-197-3p was up-regulated in NSCLC. hsa_circ_0003497 directly interacts with miR-197-3p and releases a target gene of miR-197-3p termed CTNND1 (a known tumor suppressor gene). Evolutionary analysis reveals fast evolution of this hsa_circ_0003497-miR-197-3p-CTNND1-NSCLC axis in mammals. This work clarified the biological functions and molecular mechanisms of how hsa_circ_0003497 suppresses NSCLC through miR-197-3p and CTNND1. We discovered molecular markers for the prognosis of NSCLC and provided potential intervention targets for its treatment.









Similar content being viewed by others
Data Availability
This study did not produce raw sequence or structure data. The microarray circRNA data of three pairs of NSCLC and normal tissues were downloaded from GEO database (http://www.ncbi.nlm.nih.gov/geo) with accession number GSE112214.
Abbreviations
- circRNA:
-
Circular RNA
- ncRNA:
-
Non-coding RNA
- miRNA:
-
MicroRNA
- NSCLC:
-
Non-small cell lung cancer
- LCLC:
-
Large cell lung cancer
- LUSC:
-
Lung Squamous Carcinoma
- LUAD:
-
Lung Adenocarcinoma
- WT:
-
Wild type
- NC:
-
Negative control
- PCR:
-
Polymerase chain reaction
References
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Chen LL (2020) The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol 21:475–490
Chu D, Wei L (2019) Nonsynonymous, synonymous and nonsense mutations in human cancer-related genes undergo stronger purifying selections than expectation. BMC Cancer 19:359
Consortium GT (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45:580–585
Di Giorgio S, Martignano F, Torcia MG, Mattiuz G, Conticello SG (2020) Evidence for host-dependent RNA editing in the transcriptome of SARS-CoV-2. Sci Adv 6:5813
Dobzhansky T (1973) Nothing in biology makes sense except in light of evolution. Am Biol Teach 35:125–129
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB (2016) Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res 44:2846–2858
Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M (2016) CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 13:34–42
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Faivre C, El Cheikh R, Barbolosi D, Barlesi F (2017) Mathematical optimisation of the cisplatin plus etoposide combination for managing extensive-stage small-cell lung cancer patients. Br J Cancer 116:344–348
Guarnerio J, Zhang Y, Cheloni, G, Panella R, Mae Katon J, Simpson M, Matsumoto A, Papa A, Loretelli C, Petri A et al (2019) Intragenic antagonistic roles of protein and circRNA in tumorigenesis. Cell Res 29:628–640
Hansen TB, Kjems J, Damgaard CK (2013) Circular RNA and miR-7 in cancer. Cancer Res 73:5609–5612
Hernandez-Martinez R, Ramkumar N, Anderson KV (2019) p120-catenin regulates WNT signaling and EMT in the mouse embryo. Proc Natl Acad Sci USA 116:16872–16881
Huang W, Yang Y, Wu J, Niu Y, Yao Y, Zhang J, Huang X, Liang S, Chen R, Chen S et al (2020) Circular RNA cESRP1 sensitises small cell lung cancer cells to chemotherapy by sponging miR-93-5p to inhibit TGF-beta signalling. Cell Death Differ 27:1709–1727
Li J, Yu CP, Li Q, Chang S, Xie LL, Wang S (2022) Large-scale omics data reveal the cooperation of mutation-circRNA-miRNA-target gene network in liver cancer oncogenesis. Future Oncol 18:163–178
Li Y, Yang X, Wang N, Wang H, Yin B, Yang X, Jiang W (2020a) GC usage of SARS-CoV-2 genes might adapt to the environment of human lung expressed genes. Mol Genet Genomics 295:1537–1546
Li Y, Yang XN, Wang N, Wang HY, Yin B, Yang XP, Jiang WQ (2020b) The divergence between SARS-CoV-2 and RaTG13 might be overestimated due to the extensive RNA modification. Future Virol 15:341–347
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L et al (2015) Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22:256–264
Liu L, Liu FB, Huang M, Xie K, Xie QS, Liu CH, Shen MJ, Huang Q (2019a) Circular RNA ciRS-7 promotes the proliferation and metastasis of pancreatic cancer by regulating miR-7-mediated EGFR/STAT3 signaling pathway. Hepatobiliary Pancreat Dis Int 18:580–586
Liu M, Wang Q, Shen J, Yang BB, Ding X (2019b) Circbank: a comprehensive database for circRNA with standard nomenclature. RNA Biol 16:899–905
Liu X, Liu X, Zhou J, Dong Y, Jiang W, Jiang W (2022) Rampant C-to-U deamination accounts for the intrinsically high mutation rate in SARS-CoV-2 spike gene. RNA 28:917–926
Liu Y, Wang S, Dong QZ, Liu N, Han Y, Zhang XP, Fan CF, Wang EH (2018) The expression pattern of p120-catenin is associated with acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors in non-small cell lung cancer. Appl Immunohistochem Mol Morphol 26:64–70
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Martignano F, Di Giorgio S, Mattiuz G, Conticello SG (2022) Commentary on “Poor evidence for host-dependent regular RNA editing in the transcriptome of SARS-CoV-2.” J Appl Genet 63:423–428
Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA (2008) Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc 83:584–594
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Panda AC, Dudekula DB, Abdelmohsen K, Gorospe M (2018) Analysis of circular RNAs using the web tool circinteractome. Methods Mol Biol 1724:43–56
Qiu M, Xia W, Chen R, Wang S, Xu Y, Ma Z, Xu W, Zhang E, Wang J, Fang T et al (2018) The circular RNA circPRKCI promotes tumor growth in lung adenocarcinoma. Cancer Res 78:2839–2851
Siegel RL, Miller KD, Jemal A (2020) Cancer statistics, 2020. CA Cancer J Clin 70:7–30
Tan S, Gou Q, Pu W, Guo C, Yang Y, Wu K, Liu Y, Liu L, Wei YQ, Peng Y (2018) Circular RNA F-circEA produced from EML4-ALK fusion gene as a novel liquid biopsy biomarker for non-small cell lung cancer. Cell Res 28:693–695
Wang H, Sun G, Jiang Y (2022) Cost-efficiency optimization serves as a conserved mechanism that promotes osteosarcoma in mammals. J Mol Evol 90:139–148
Wang Y, Gai Y, Li Y, Li C, Li Z, Wang X (2021) SARS-CoV-2 has the advantage of competing the iMet-tRNAs with human hosts to allow efficient translation. Mol Genet Genomics 296:113–118
Wei L (2022) Reconciling the debate on deamination on viral RNA. J Appl Genet. https://doi.org/10.1007/s13353-022-00698-9
Xie B, Zhao Z, Liu Q, Wang X, Ma Z, Li H (2019) CircRNA has_circ_0078710 acts as the sponge of microRNA-31 involved in hepatocellular carcinoma progression. Gene 683:253–261
Yang R, Hong YC, Ren ZZ, Tang K, Zhang H, Zhu JK, Zhao CZ (2019) A role for PICKLE in the regulation of cold and salt stress tolerance in Arabidopsis. Front Plant Sci 10:1
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, Huang N, Yang X, Zhao K, Zhou H et al (2018) Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J Natl Cancer Inst 110:1
Yao R, Yao Y, Li C, Li X, Ni W, Quan R, Liu L, Li H, Xu Y, Zhang M et al (2020) Circ-HIPK3 plays an active role in regulating myoblast differentiation. Int J Biol Macromol 155:1432–1439
Ye F, Gao G, Zou Y, Zheng S, Zhang L, Ou X, Xie X, Tang H (2019) circFBXW7 inhibits malignant progression by sponging miR-197-3p and encoding a 185-aa protein in triple-negative breast cancer. Mol Ther Nucleic Acids 18:88–98
Yu YY, Li Y, Dong Y, Wang XK, Li CX, Jiang WQ (2021) Natural selection on synonymous mutations in SARS-CoV-2 and the impact on estimating divergence time. Future Virol 16:447–450
Zhang N, Nan A, Chen L, Li X, Jia Y, Qiu M, Dai X, Zhou H, Zhu J, Zhang H et al (2020) Circular RNA circSATB2 promotes progression of non-small cell lung cancer cells. Mol Cancer 19:101
Zhang Y, Jin X, Wang H, Miao Y, Yang X, Jiang W, Yin B (2021a) Compelling evidence suggesting the codon usage of SARS-CoV-2 adapts to human after the split from RaTG13. Evol Bioinform Online 17:11769343211052012
Zhang Y, Jin X, Wang H, Miao Y, Yang X, Jiang W, Yin B (2022) SARS-CoV-2 competes with host mRNAs for efficient translation by maintaining the mutations favorable for translation initiation. J Appl Genet 63:159–167
Zhang YP, Jiang W, Li Y, Jin XJ, Yang XP, Zhang PR, Jiang WQ, Yin B (2021b) Fast evolution of SARS-CoV-2 driven by deamination systems in hosts. Future Virol 16:587–590
Zhao MM, Li CX, Dong Y, Wang XK, Jiang WQ, Chen YG (2022) Nothing in SARS-CoV-2 makes sense except in the light of RNA modification? Future Virol. https://doi.org/10.2217/fvl-2022-0043
Zhao S, Song S, Qi Q, Lei W (2021) Cost-efficiency tradeoff is optimized in various cancer types revealed by genome-wide analysis. Mol Genet Genomics 296:369–378
Zhu L, Liu X, Zhang W, Hu H, Wang Q, Xu K (2022a) MTHFD2 is a potential oncogene for its strong association with poor prognosis and high level of immune infiltrates in urothelial carcinomas of bladder. BMC Cancer 22:556
Zhu L, Wang Q, Zhang W, Hu H, Xu K (2022b) Evidence for selection on SARS-CoV-2 RNA translation revealed by the evolutionary dynamics of mutations in UTRs and CDSs. RNA Biol 19:866–876
Zong J, Zhang Y, Guo F, Wang C, Li H, Lin G, Jiang W, Song X, Zhang X, Huang F et al (2022) Poor evidence for host-dependent regular RNA editing in the transcriptome of SARS-CoV-2. J Appl Genet 63:413–421
Acknowledgements
We thank our colleagues and any people in our institution who have given precious suggestions to this work.
Funding
The authors have not declared a specific grant for this research from any funding agency in the public, commercial or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
LW, MC, XP, BL: methodology, writing—original draft. JW: conceptualization, supervision, writing—original draft and writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Ethical Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Additional information
Handling editor: Kerry Geiler-Samerotte.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, L., Cao, M., Pu, X. et al. The Sponge Interaction Between Circular RNA and microRNA Serves as a Fast-Evolving Mechanism That Suppresses Non-small Cell Lung Cancer (NSCLC) in Humans. J Mol Evol 90, 362–374 (2022). https://doi.org/10.1007/s00239-022-10067-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-022-10067-z