Abstract
Understanding how SARS-CoV-2 (Severe Acute Respiratory Syndrome Coronavirus 2) efficiently reproduces itself by taking resources from the human host could facilitate the development of drugs against the virus. SARS-CoV-2 translates its own proteins by using the host tRNAs, so that its GC or codon usage should fit that of the host cells. It is necessary to study both the virus and human genomes in the light of evolution and adaptation. The SARS-CoV-2 virus has significantly lower GC content and GC3 as compared to human. However, when we selected a set of human genes that have similar GC properties to SARS-CoV-2, we found that these genes were enriched in particular pathways. Moreover, these human genes have the codon composition perfectly correlated with the SARS-CoV-2, and were extraordinarily highly expressed in human lung tissues, demonstrating that the SARS-CoV-2 genes have similar GC usage as compared to the lung expressed human genes. RSCU (relative synonymous codon usage) and CAI (codon adaptation index) profiles further support the matching between SARS-CoV-2 and lungs. Our study indicates that SARS-CoV-2 might have adapted to the human lung environment by observing the high correlation between GC usage of SARS-CoV-2 and human lung genes, which suggests the GC content of SARS-CoV-2 is optimized to take advantage of human lung tissues.
Similar content being viewed by others
Introduction
The recent outbreak of SARS-CoV-2 (Severe Acute Respiratory Syndrome Coronavirus 2) has caused severe damage to China especially the Hubei province (Cowling and Leung 2020; Hui et al. 2020; Wang et al. 2020). It is urgent to find ways to control its transmission and cure the infected patients (Edelstein and Heymann 2015; Sheahan and Baric 2018; Wang et al. 2020). Particularly, understanding how SARS-CoV-2 achieved rapid reproduction in the human host might be helpful in developing antivirus drugs. Notably, SARS-CoV-2 translates its own proteins using the host tRNAs, which indicates that the codon composition of SARS-CoV-2 should have adapted to that of the human hosts. Therefore, it is rational to study both the virus and human genomes from the perspective of evolution and adaptation.
It is well established that the viruses themselves do not survive alone, and they could only translate and reproduce their own proteins by using the resources from host cells (Taghinezhad et al. 2017). Proteins commonly consist of 20 amino acids, which are encoded by specific codons. The host cells do not equally use each amino acid and codon (Plotkin and Kudla 2011). Instead, there are different GC biases or codon biases in different organisms (Harrison and Charlesworth 2011; Wei 2020). The GC content across the genome almost determines which set of codons (G/C ending or A/T ending) are favored by this organism. For SARS-CoV-2 or other coronaviruses, the GC content in its coding sequence is around 38% (Berkhout and van Hemert 2015). In eukaryotes, such as vertebrates, including humans, the GC content could be around 60%. It seems that SARS-CoV-2 has a very different GC preference as compared to the human genome. This raises a question that how the virus could efficiently reproduce itself in an environment with very different resources.
However, human is a multiorgan/tissue organisms. The genes express differentially in specific organs/tissues/cells. The GC preference could also vary across tissues according to the tissue-specifically expressed genes. For SARS-CoV-2, although it exhibits a different GC preference as compared to the entire human genome, it might adapt to the environment of particular tissues (e.g., lung). The aim of this study is to test whether SARS-CoV-2 has developed a particular GC usage similar to that of the human lung tissues. If this is the case, then this idea might help understand how the virus adapt to its living environment. In the multiple domains of life, using the genomic big data to deal with the relationship between organisms and environmental adaptation has already been proposed (Argueso et al. 2019), and analyzing the codon usage might be part of the key issues (Feng et al. 2013; Wang and Hickey 2007).
RSCU (relative synonymous codon usage) and CAI (codon adaptation index) are classic parameters measuring the extent of codon bias (Sharp and Li 1987). Codons with higher RSCU values are more frequently used by the genome and are usually GC enriched. Genes with higher CAI values tend to utilize more optimal codons. Since CAI of a gene is calculated as the geometric mean of RSCU, and RSCU is largely based on the set of genes of interest, this allows us to compare the codon bias of virus and human genes from different aspects.
We found the following patterns. The SARS-CoV-2 virus has significantly lower GC content and GC3 compared to human. However, when we selected a set of human genes which have similar GC properties to SARS-CoV-2, we found that these genes were enriched in particular pathways that might be related to virus invasion. Moreover, these human genes were extraordinarily highly expressed in human lung tissues, suggesting that the SARS-CoV-2 genes have similar GC usage compared to the lung expressed human genes. When we further employed the RSCU and CAI, the matching between SARS-CoV-2 and human lungs is consolidated.
Our study indicates that SARS-CoV-2 might have adapted to the human lung environment. The GC usage of SARS-CoV-2 is optimized to take advantage of human lungs. These results might help understand how the virus adapt to its living environment.
Materials and methods
Data collection
We downloaded the SARS-CoV-2 genome from the NCBI website (https://www.ncbi.nlm.nih.gov/genome/). The coding sequence of human genome was downloaded from the Ensembl website of version hg19 (ftp://ftp.ensembl.org/pub/release-75/fasta/homo_sapiens/cds/). The RNA-seq data used in this study were downloaded with accession numbers GSM1120308 (polyA RNA sequencing of STL001 Lung Cells) and GSE93482 (polyA mRNA RNA-seq from lung). These are data from normal lung tissues.
GC content calculation
GC content (GC%) used in this study is the percentage of guanosines and cytidines in the coding sequence of genomes. GC3 is the percentage of guanosines and cytidines at the third codon position. GC12 is the percentage of guanosines and cytidines at the first and second codon positions.
RNA-seq mapping
We mapped the RNA-seq reads to the human coding sequence with software bowtie2 (Langmead and Salzberg 2012). Two mismatches per read were allowed. Only the longest coding sequence of each gene was chosen to be the reference file. The uniquely mapped reads were maintained. The gene expression was measured by RPKM (reads per kilobase per million mapped reads) = reads count of a gene/gene length (Kb)/library size (million reads). In the RSCU and CAI analyses, the highly expressed genes in lungs represent the top 1000 genes with highest RPKM in the combined GSM1120308 and GSE93482 samples.
Functional annotation
Functional annotation of human genes was accomplished on the DAVID website (Jiao et al. 2012). The gene ontology (GO) terms with multiple testing corrected p value (FDR, the column “Benjamini”) < 0.05 were maintained.
RSCU (relative synonymous codon usage) and CAI (codon adaptation index)
To calculate RSCU we first need to know which set of genes should be used. We tried using different sets of genes to calculate RSCU. The different sets of genes are: (1) All human genes; (2) lung genes: top 1000 (RPKM) genes expressed in lung (combined GSM1120308 and GSE93482); (3) housekeeping genes: among the 1000 lung genes we extracted the ribosomal genes; (4) lung-non-house genes: the 1000 lung genes excluding ribosomal genes; (5) virus genes: the eleven non-redundant ORFs of SARS-CoV-2. Among each set of genes, a number of codon appearance were counted. RSCU is the codon count weighted by the mean count across synonymous codons. By this way, if an amino acid has N synonymous codons, then the sum of RSCU values of these codons should be equal to N. RSCU greater than one represents frequently used codons, and vice versa. CAI of a gene is the geometric mean of RSCU values of codons in that gene. The ribosomal genes were selected based on the gene names. In the gene annotation file (hg19 version), all ribosomal genes are named with prefix RPL (ribosome protein large subunit) or RPS (ribosome protein small subunit). We extracted these genes by matching the string “^RPL|^RPS”.
Statistical analyses
We used R language to conduct statistical analyses including correlation tests and KS tests.
Results
GC content of SARS-CoV-2 genes is lower than that of human genes
We downloaded the coding sequences of SARS-CoV-2 virus and human (“Materials and methods”). We define GC3 as the percentage of guanosines and cytidines at the third codon position, and GC12 as the percentage of guanosines and cytidines at the first and second codon positions (“Materials and methods”). We found moderately positive correlations between the GC3 and GC12 values among the 12 SARS-CoV-2 genes (Fig. 1a, Spearman’s correlation = 0.39) and the twenty thousand human genes (Fig. 1b, Spearman’s correlation = 0.62). However, we found a significantly higher GC content in human than SARS-CoV-2 (Fig. 1c).
GC content of SARS-CoV-2 and human genes. a Dot plot showing the GC3 and GC12 of the coding sequences of 12 genes of SARS-CoV-2. Spearman’s correlation = 0.39. b Dot plot showing the GC3 and GC12 of the coding sequences of all human genes. Spearman’s correlation = 0.62. c Boxplot displaying the GC3 and GC12 of SARS-CoV-2 (purple) and human genes (brown). “3” means GC3 and “1 + 2” mean GC12. “***”Indicates p value < 1e−3. The p values were calculated by Wilcoxon rank sum tests
Human genes with similar GC content/codon bias to that of SARS-CoV-2 are highly expressed in lungs
The distinctly different GC contents between the SARS-CoV-2 and human genomes raise a question that how the virus could efficiently reproduce itself in an environment with very different GC resources. We first defined the 25% and 75% quantiles of GC contents of SARS-CoV-2 genes. 25% quantile GC3 = 0.252; 75% quantile GC3 = 0.334; 25% quantile GC12 = 0.396; 75% quantile GC12 = 0.437. We selected the human genes within this range of GC content (Fig. 2a) and totally obtained 324 human genes. We performed gene ontology (GO) enrichment analysis (“Materials and methods”) on these 324 genes and found an enrichment in cilium morphogenesis and double-strand break repair (Fig. 2b). Although without direct evidence, we could surmise that these genes might be related to virus invasion since they entail the processes similar to immune or stress response. At this stage, this point remains speculative and untested. Note that the purpose of this part is to seek for potential human genes with similar GC profile with SARS-CoV-2. It does not mean the human genes adapted to SARS-CoV-2 genes. It should be the other way around: evolution rate in coronaviruses are orders of magnitude higher than that of eukaryotic cells, so it is the SARS-CoV-2 genes that have adapted to particular human genes (or the virus’ living environment).
Human genes with similar GC properties to that of SARS-CoV-2 genes. a Dot plot showing the set of human genes (colored purple, 324 genes) with similar GC properties to those of SARS-CoV-2 genes (the human genes with GC3 and GC12 values within the 25% ~ 75% quantiles of the GC3 and GC12 of SARS-CoV-2). b Gene ontology enrichment of the 324 genes which have similar GC properties to those of SARS-CoV-2 genes. “Benjamini” is the p value after multiple testing correction. c Dot plots displaying the correlation between codon frequency (proportion) in SARS-CoV-2 and human genes. The selected 324 genes are in purple. The other genes are in gray. Pearson’s correlation coefficients are shown
Moreover, when we calculated the codon frequency (proportion) of the 61 sense codons among different genes, we found that the codon proportion among the 324 human genes was significantly positively correlated with the codon proportion among the SARS-CoV-2 genes (Fig. 2c, left), while the codon proportion among the other human genes did not show such a significant correlation (Fig. 2c, right). This result is not surprising because the codon identity could be affected by the GC status.
Because these 324 human genes have the most similar GC contents to the SARS-CoV-2 and the SARS-CoV-2 invades human lungs, we intuitively wonder whether these 324 genes are highly expressed in lungs. We downloaded two sets of RNA-seq data of human lungs (“Materials and methods”). We discovered that the 324 genes indeed had significantly higher expression levels than the remaining genes (Fig. 3a). In other words, this result tells us that the SARS-CoV-2 genes have similar GC usage as compared to the human lung expressed genes.
Gene expression in human lungs and the GC content of coding sequences of different viruses. a Boxplots displaying the gene expression levels (RPKM) of the 324 genes versus other genes in two sets of lung RNA-seq data. The p values were calculated by KS tests. b The overlapping between the top 1000 genes (highest RPKM) of the two datasets
We have shown that the human genes with similar GC profile with SARS-CoV-2 genes are highly expressed in lungs. Next, we use a more direct way to test the correlation between the codon usage between SARS-CoV-2 and human genes. We will change the angle and directly use the top 1000 highly expressed genes in lung instead of the “324 candidate genes”. The overlapping between the top 1000 genes in the two datasets > 86% (Fig. 3b), so in the following analyses we combined GSM1120308 and GSE93482.
RSCU (relative synonymous codon usage) profile in virus and human genes
We want to better understand the effect of GC content on the synonymous codon usage. The RSCU provides quantitative insight into the synonymous codon profile (“Materials and methods”). We first calculated the RSCU values using three different sets of genes: (1) All human genes; (2) lung expressed genes: top 1000 (RPKM) genes expressed in lung; (3) the SARS-CoV-2 genes. We see that the RSCU calculated from all human genes and lung expressed genes are similar but they differ from RSCU calculated from SARS-CoV-2 genes (Fig. 4a, b).
The RSCU (relative synonymous codon usage) profile calculated from different sets of genes. a The RSCU values of each codon except stop codons. “virus” means the RSCU calculated from SARS-CoV-2 genes. “human” means the RSCU calculated from all human genes. “lung” means the RSCU calculated from top 1000 genes expressed in lung. b Correlation between the RSCU values shown in (a)
CAI (codon adaptation index) profile of virus and human genes
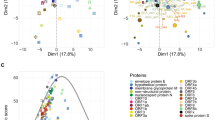
The purpose of knowing RSCU is to calculate the CAI of genes. Given a particular set of RSCU values, one can calculate the geometric mean to get a CAI value for a gene (“Materials and methods”). Since we already have three different sets of RSCU values, we would obtain three versions of CAI values for a single gene. We inspect the CAI of the following genes: the CAI of 12 SARS-CoV-2 ORFs separately, the median CAI of 12 SARS-CoV-2 ORFs, and the median CAI of all human genes (Fig. 5a).
The CAI (codon adaptation index) profile of different genes. a The CAI values of 12 SARS-CoV-2 ORFs, mean of 12 SARS-CoV-2 ORFs, and mean of all ~ 20,000 human genes. b Pairwise spearman correlation between the RSCU values calculated from each single gene. The horizontal axis is the ~ 20,000 human genes. The vertical axis is the 12 SARS-CoV-2 ORFs
We see an interesting result. For human genes, using the virus RSCU would lead to an obviously lower CAI profile. Using the all human gene RSCU or lung gene RSCU does not seem to affect the CAI of human genes (Fig. 5a). For virus genes, the choice of all human gene RSCU or virus gene RSCU does not strongly affect the CAI of virus genes, either. However, using lung gene RSCU would produce remarkably higher CAI values (Fig. 5a).
The N gene (nucleocapsid) has the largest difference between CAI values calculated from human RSCU and virus RSCU (Fig. 5a). It simply means that nucleocapsid has the codon usage most distant from the host. Another indirect illustration of this issue is the pairwise Spearman correlation between the RSCU values of 12 SARS-CoV-2 ORFs and the ~ 20,000 human genes (Fig. 5b). N gene seems to have a different correlation profile, suggesting its potentially different “interactome” with human genes. A possible explanation for the outstanding feature of gene N is that the proteins interacting with host antibodies have a tend to show codon usage very distant from the host so that to allow lower exposition to the immune system. So far, these are the observation and deduction. We currently do not have experimental data to prove this evolutionary notion.
Are SARS-CoV-2 genes more adapted to lung cells than the lung genes?
An interesting but confusing issue is whether SARS-CoV-2 adapted to lung cells or the lung genes? First, we think the definition of “lung genes” partially comes from the observed highly expressed genes in lungs. But the highly expressed genes in lungs should include several housekeeping genes which are not “lung genes”. Anyway, “highly expressed genes in lung cells” and “lung genes” are not independent. They might overlap with each other for a large proportion. Next, if we have to tell them apart, as we understand, the difference between “highly expressed genes in lung cells” and “lung genes” is that the former contains some housekeeping genes (such as ribosomal genes).
Apart from the lowly expressed genes (most genes in a sample) in lung (combined GSM1120308 and GSE93482), we divide the “highly expressed genes in lung cells” (top one thousand RPKM) into two categories, one is the ribosomal genes (termed “house”), another is the remaining genes (termed “lung-non-house”). We draw the median values of the gene expression of each set of genes (Fig. 6a). Then we calculate the RSCU values of codons based the house genes and lung-non-house genes, respectively. With these two sets of RSCU values, we next calculated the CAI of 12 SARS-CoV-2 genes. The CAI from RSCU of lung-non-house genes is always higher than the CAI from house genes (Fig. 6b). We can preliminarily conclude that SARS-CoV-2 genes more adapted to human lung genes (represented by lung-non-house genes) when compared with lung cells (represented by house genes).
The expression and CAI of housekeeping genes (ribosomal genes) and other highly expressed genes (lung-non-house) in lung. a Median RPKM value of three sets of genes. “house” means the ribosomal genes among the top 1000 genes in lung. “lung-non-house” means the top 1000 genes in lung excluding ribosomal genes. “low expression” means all genes excluding top 1000 genes in lung. b CAI of 12 SARS-CoV-2 genes calculated from two sets of RSCU values described in (a)
Discussion
The mutation (Zhao et al. 2004) and modification (Carpenter et al. 2009; Fung and Liu 2018) of virus sequences usually make it difficult to trace the origin and evolution of viruses (Li et al. 2020). However, the codon composition would be less affected by the noises. The SARS-CoV-2 needs to utilize the resources from host cells to reproduce itself. In theory, the virus should choose a host environment where the GC and codon usages are most similar to the virus genome itself. We have found that the GC/codon usage of SARS-CoV-2 is optimized to take advantage of human lungs, supporting the idea that its GC usage fits that of the host cells. Indeed, we did not perform functional verifications to prove any relevance between the GC content of virus and human. However, we hope our ideas could be a novel aspect to decipher the needs and demands relationship between virus and host cells. In the light of evolution and adaptation, the host and the parasite should co-evolve so that their genomic features like GC or codon usage should be shaped to fit each other. The actual process might be, the host genome evolves allowing to escape from the parasite invasion and the parasite genome evolves allowing to take advantage of host genomes, the results of which could be the similarity between their genomic features. Frankly speaking, the correlation of codon usage between host and parasite only serves as supporting evidence but not decisive evidence to connect the two organisms. The most solid evidence is still the other medical-related methodologies.
Indeed, the vast majority of codon usage studies would calculate the virus CAI by using the human RSCU. We added additional analyses on the CAI based on the virus RSCU. One certain thing is that the traditional calculation of virus CAI using host RSCU is the standard approach, but our comparison of CAI calculated from different sets of RSCU could reflect the difference of the codon usage profile between human and SARS-CoV-2.
All we have talked above is to emphasize the importance of adapting to the codon profile of the host. Inversely, one may propose whether it is possible that low GC content in SARS-CoV-2 is an adaptation for not competing with the cell in order to use tRNA that are not the most frequently used by the cell? This leads to a dilemma. (1) If the codon profile of the parasite is highly correlated with the codon usage in host, then theory could be like: the parasite has adapted to take advantage of the host; (2) If the codon profile of the parasite is poorly correlated with the codon usage in host, then theory could be like: the parasites avoid competing with hosts so that they choose to use the codons that are rarely used by the host. From another aspect, there is an opinion that the correlated codon usage between two species is a “necessary bit not sufficient” prerequisite for the host–parasite relationship. Without this prerequisite, the two species even have no change to become host–parasite. Therefore, our current work is to display the existence of this prerequisite, and support the possibility of SARS-CoV-2 adapting to the human lung tissues. Anyway, this question remains open and we tend to believe the former hypothesis which assumes the codon usage of parasites should be as close as possible to their host.
Hopefully, more detailed analyses in the future could reveal whether the codon usage of virus is optimized towards a set of codons which are excessively used in human lungs. These results could help understand the interaction between virus and hosts, and hint us about how the virus adapt to its living environment. Our study indicates that SARS-CoV-2 might have adapted to the human lung environment. The GC usage and codon usage of SARS-CoV-2 are optimized to take advantage of human lungs. These results might help understand how the virus adapt to its living environment.
Data availability
We downloaded the SARS-CoV-2 genome from the NCBI website (https://www.ncbi.nlm.nih.gov/genome/). The coding sequence of human genome was downloaded from the Ensembl website of version hg19 (ftp://ftp.ensembl.org/pub/release-75/fasta/homo_sapiens/cds/). The RNA-seq data used in this study were downloaded with accession numbers GSM1120308 (polyA RNA sequencing of STL001 Lung Cells) and GSE93482 (polyA mRNA RNA-seq from lung).
References
Argueso CT, Assmann SM, Birnbaum KD, Chen S, Dinneny JR, Doherty CJ, Eveland AL, Friesner J, Greenlee VR, Law JA et al (2019) Directions for research and training in plant omics: big questions and big data. Plant Direct 3:e00133
Berkhout B, van Hemert F (2015) On the biased nucleotide composition of the human coronavirus RNA genome. Virus Res 202:41–47
Carpenter JA, Keegan LP, Wilfert L, O'Connell MA, Jiggins FM (2009) Evidence for ADAR-induced hypermutation of the Drosophila sigma virus (Rhabdoviridae). BMC Genet 10:75
Cowling BJ, Leung GM (2020) Epidemiological research priorities for public health control of the ongoing global novel coronavirus (2019- nCoV) outbreak. Euro Surveill 25(6):2000110. https://doi.org/10.2807/1560-7917.ES.2020.25.6.2000110
Edelstein M, Heymann DL (2015) What needs to be done to control the spread of Middle East respiratory syndrome coronavirus? Future Virol 10:497–505
Feng C, Xu CJ, Wang Y, Liu WL, Yin XR, Li X, Chen M, Chen KS (2013) Codon usage patterns in Chinese bayberry (Myrica rubra) based on RNA-Seq data. BMC Genomics 14
Fung TS, Liu DX (2018) Post-translational modifications of coronavirus proteins: roles and function. Future Virol 13:405–430
Harrison RJ, Charlesworth B (2011) Biased gene conversion affects patterns of codon usage and amino acid usage in the Saccharomyces sensu stricto group of yeasts. Mol Biol Evol 28:117–129
Hui DS, Azhar EI, Madani TA, Ntoumi F, Kock R, Dar O, Ippolito G, McHugh TD, Memish ZA, Drosten C et al (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 91:264–266
Jiao X, Sherman BT, da Huang W, Stephens R, Baseler MW, Lane HC, Lempicki RA (2012) DAVID-WS: a stateful web service to facilitate gene/protein list analysis. Bioinformatics 28:1805–1806
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–U354
Li Y, Yang X, Wang N, Wang H, Yang X, Jiang W (2020) The divergence between SARS-CoV-2 and RaTG13 might be overestimated due to the extensive RNA modification. Future Virol. https://doi.org/10.2217/fvl-2020-0066
Plotkin JB, Kudla G (2011) Synonymous but not the same: the causes and consequences of codon bias. Nat Rev Genet 12:32–42
Sharp PM, Li WH (1987) The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15:1281–1295
Sheahan TP, Baric RS (2018) Is regulation preventing the development of therapeutics that may prevent future coronavirus pandemics? Future Virol 13:143–146
Taghinezhad S, Razavilar V, Keyvani H, Razavi MR, Nejadsattari T (2017) Codon optimization of Iranian human papillomavirus Type 16 E6 oncogene for Lactococcus lactis subsp cremoris MG1363. Future Virol 12:499–511
Wang C, Horby PW, Hayden FG, Gao GF (2020) A novel coronavirus outbreak of global health concern. Lancet 395:470–473
Wang HC, Hickey DA (2007) Rapid divergence of codon usage patterns within the rice genome. BMC Evol Biol 7:S6
Wei L (2020) Selection on synonymous mutations revealed by 1135 genomes of Arabidopsis thaliana. Evol Bioinform Online 16:1176934320916794
Zhao Z, Li H, Wu X, Zhong Y, Zhang K, Zhang YP, Boerwinkle E, Fu YX (2004) Moderate mutation rate in the SARS coronavirus genome and its implications. BMC Evol Biol 4:21
Acknowledgements
We thank the members in our group that have given suggestions to our project. At this SARS-CoV-2 time, we should especially thank all the medical workers fighting against SARS-CoV-2.
Funding
This study was funded by Qingdao Haici Medical Group (Grant number 2019-WJZD051).
Author information
Authors and Affiliations
Contributions
The corresponding author designed and supervised this research. All authors contributed to the big data analyses. All authors contributed to writing this article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Y., Yang, X., Wang, N. et al. GC usage of SARS-CoV-2 genes might adapt to the environment of human lung expressed genes. Mol Genet Genomics 295, 1537–1546 (2020). https://doi.org/10.1007/s00438-020-01719-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01719-0