Abstract
Vestibular Schwannoma (VS) is a rare tumor with varied incidence rates, predominantly affecting the 60–69 age group. In the era of artificial intelligence (AI), deep learning (DL) algorithms show promise in automating diagnosis. However, a knowledge gap exists in the automated segmentation of VS using DL. To address this gap, this meta-analysis aims to provide insights into the current state of DL algorithms applied to MR images of VS.
Methodology
Following 2020 PRISMA guidelines, a search across four databases was conducted. Inclusion criteria focused on articles using DL for VS MR image segmentation. The primary metric was the Dice score, supplemented by relative volume error (RVE) and average symmetric surface distance (ASSD).
Results
The search process identified 752 articles, leading to 11 studies for meta-analysis. A QUADAS- 2 analysis revealed varying biases. The overall Dice score for 56 models was 0.89 (CI: 0.88–0.90), with high heterogeneity (I2 = 95.9%). Subgroup analyses based on DL architecture, MRI inputs, and testing set sizes revealed performance variations. 2.5D DL networks demonstrated comparable efficacy to 3D networks. Imaging input analyses highlighted the superiority of contrast-enhanced T1-weighted imaging and mixed MRI inputs.
Discussion
This study fills a gap in systematic review in the automated segmentation of VS using DL techniques. Despite promising results, limitations include publication bias and high heterogeneity. Future research should focus on standardized designs, larger testing sets, and addressing biases for more reliable results. DL have promising efficacy in VS diagnosis, however further validation and standardization is needed.
Conclusion
In conclusion, this meta-analysis provides comprehensive review into the current landscape of automated VS segmentation using DL. The high Dice score indicates promising agreement in segmentation, yet challenges like bias and heterogeneity must be addressed in the future research.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Vestibular Schwannoma (VS), or Acoustic Schwannoma, is a rare tumor with an incidence rate ranging from 0.3 to 6.1 cases per 100,000 individuals, varying between age groups and ethnicities [1,2,3,4,5]. Mainly affecting individuals in the 60–69 age group, VS causes symptoms such as gradual hearing loss, dizziness, and tinnitus. Symptoms escalate as the tumor increases in size, leading to compression of the cranial nerves, the brainstem, and the cerebellum. This compression results in symptoms such as vomiting, headaches, nausea, or mental confusion [6,7,8].
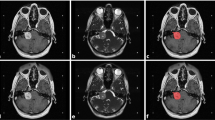
Diagnosis of VS is crucial for determining optimal interventions, which include surgery, radiotherapy, or gene therapy. Currently MR scans are considered the gold-standard, utilizing T1 weighted (T1W), T2W weighted (T2W), and contrast-enhanced T1 weighted (ceT1W) image sequences for localization and tumor segmentation assessment [9, 10].
However, as the VS follow-ups span years, manual delineation and segmentation of VS becomes an ineffective, labor-intensive process. Automated segmentation could potentially save a lot of time for physicians, improving the effectiveness of the workplace. Additionally, manual segmentation of the neoplasm is relatively subjective process, which can lead to variations in the results. Currently, challenges associated with limitations of manual segmentation include:
-
Clinical decision-making: Accurate volumetric measurements are crucial for monitoring the growth of VS to plan next treatment steps. The time-consuming manual option limits the potential of rapid decisions.
-
Superiority of volumetric analysis: 3D volume analysis allows for more comprehensive and accurate analysis than regular linear measurements of the VS. However, due to the time required for analysis, this option remains less attractive.
-
Safety of protocols: While contrast scans are still commonly used, implementation of high-resolution scans without contrast for automated segmentation could benefit patients by minimizing the risk of gadolinium accumulation in brain tissues. Additionally, cystic parts are hyperintense on these scans, allowing for better decision making.
-
Costs: Automated tools could reduce potential expenses by decreasing the total scanning length and the number of required sequences.
In the era of artificial intelligence (AI), convolutional neural networks (CNNs), a subset of deep learning (DL), demonstrated potential for the automated diagnosis of pathologies. CNNs, which are able to mimic human neurons, have demonstrated promising results in medical applications, including radiology, where DL networks have been applied for detection, tumor volume measurements, and neoplasm segmentation [11,12,13,14,15].
Despite these advancements, currently no work synthesizes information regarding the automated segmentation of VS using DL techniques. Hence, in this study we aim to conduct a systematic review and meta-analysis, describing the current state of algorithms applied to MR image segmentation of this rare neoplasm. By performing such review, we not only aim to fulfill existing gap in synthesis, but also provide insights, limitations, and guidance that may influence the future works related to the topic of automated segmentation of VS.
Methodology
This study adhered to the 2020 PRISMA guidelines for conducting a systematic review and meta-analysis [16]. The identification of eligible articles involved a comprehensive search across four databases: PubMed, Web of Science, Scopus, and Embase. Our search strategy incorporated MeSH terms and encompassed a range of keywords, including Artificial Intelligence, Deep Learning, Machine Learning, Automation, Convolutional Neural Networks, and specific terms related to vestibular schwannoma, acoustic schwannoma, and perineural fibroblastoma. The complete search strategy is provided in Appendix 1. In addition, a manual review of references was conducted to identify any studies not captured in the initial search. Two authors independently screened articles for inclusion and exclusion based on the outlined criteria, with conflicts resolved through mutual discussion with other authors. The screening process was facilitated by ZOTERO software, employed by both authors.
Inclusion and exclusion criteria
We established inclusion criteria for our study, including: (I) no restriction on publication date, encompassing articles from all years; (II) inclusion of only original articles in English; (III) exclusion of reviews, abstract-only works, preprints, and other non-original articles; (IV) focus on deep learning MR image segmentation; (V) exclusion of other artificial intelligence techniques such as conventional machine learning (ML); (VI) inclusion of studies with specified validation methods for the meta-analysis, aiming to mitigate overestimation; (VII) inclusion of studies with extractable data for meta-analysis (standard deviation or error), with the Dice score as the primary performance metric, supplemented by data on relative volume error (RVE) and average symmetric surface distance (ASSD).
Dice score, RVE, ASSD, and other extracted data
The Sørensen-Dice score, or Dice score, quantitatively measures the similarity between two sets, serving as an evaluation metric for segmentation algorithm performance. RVE provides a percentage measure of the deviation between segmented and true volumes, while ASSD gauges the alignment of segmented surfaces with ground truth surfaces (in our study in mm). Additional extracted data from each study included information on country of origin, software and hardware specifications, DL algorithm characteristics, validation types, testing set sizes, MRI inputs, patient demographics, tumor volume, MR imaging specifications, labelling, cystic analysis, training parameters, and dataset source. Independent extraction of data was followed by conflict resolution through discussion with other authors.
Statistical analysis
Each study underwent individual assessment using the Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2) tool to evaluate the potential risk of bias [17]. Robvis software facilitated the visualization of outcomes [18]. Stata v18 software was employed for statistical analysis, utilizing a random-effects REML model for forest plot construction. Distinct techniques, testing sets, DL algorithms, and MRI inputs were treated as independent results. Subgroup analyses were conducted based on 3-dimensional and 2.5-dimensional architecture algorithms, ceT1W, T2W, T1W and mixed MRI inputs, as well as different testing set sizes (< 50, 50–100, and > 100 patients). Heterogeneity was quantified using the percentage I2 index. Significance threshold was set at p < 0.05. The analysis employed 95% confidence intervals (CI), and bias was assessed through funnel plot analysis.
Results
The detailed process of our literature search is outlined in Fig. 1 [16]. We identified a total of 752 articles across four databases: 241 from PubMed, 147 from Web of Science, 201 from Embase, and 163 from Scopus. Using ZOTERO, we initially identified 371 articles to remove duplicates. During screening, two authors independently reviewed 381 records, resolving disagreements through collaboration.
Following screening, we excluded 347 works unrelated to the meta-analysis topic. In the subsequent full-text screening, 34 works were included. Unfortunately, we were unable to retrieve 2 records. For the meta-analysis, we incorporated 11 out of 34 potentially eligible works. Eleven lacked extractable data for statistical analysis, one was a preprint, one had duplicate results, another was a review, and seven were abstract-only articles. Attempts to contact authors via mail correspondence yielded no replies.
PRISMA flow diagram. Made with PRISMA template [16]
Publication bias
We performed a QUADAS-2 analysis using the robvis tool, individually assessing each of eleven studies [19,20,21,22,23,24,25,26,27,28,29]. For each study, we evaluated the patient selection process, index test, reference standard, and flow & timing of the study, with a detailed description available in the QUADAS-2 statement [17]. One article exhibited low risk of bias, seven had some concerns, while three had a high risk of bias. Major concerns were identified in the patient selection process and some in the index test, with no significant concerns in the reference standard and flow and timing section. We have identified lack (or no reports) of clear inclusion or exclusion criteria, unclear process of randomization of the datasets (patients), lack of information of blinding the index test, and two studies had concerns in the labelling process. The QUADAS-2 charts with bias assessment results are presented in Fig. 2.
QUADAS-2 chart with assessment of bias of included studies. Made with robvis [18]
Included studies
From the 11 studies included in this meta-analysis, 4 were from the UK, 3 from the USA, 1 from Taiwan, and 1 each from the Netherlands and China, with a collaborative effort between China and the UK. The publication dates ranged from 2019 to 2023. Pytorch and Tensorflow were the most commonly used software, along with NVIDIA GPU cards and Intel CPUs. However, not all studies reported details such as software versions or GPU types. All works were based on CNN platforms, with U-Net being the most popular option, featuring various modifications of modules and dimension processing. The majority of works (6) employed split validation for performance assessment and further tuning. Studies varied in MR imaging techniques, with contrast-enhanced T1-weighted imaging being the most popular, appearing in 9 studies. Detailed information about each study is available in Table 1. We have additionally extracted supplementary data from each study, related to demographics, dose radiation, tumor volume, and MRI specification which is available in the Table 2. Table 3 provides additional information about labelling, training process, and datasets [30, 31].
Overall performance analysis
Regarding the overall performance of segmentation, fifty-six models from eleven studies reported a mean Dice score for VS MR image segmentation. The pooled mean Dice score was 0.89 (CI: 0.88–0.90, p < 0.001), with a heterogeneity of 95.9% (I2). Forest plot with overall Dice results is provided in Fig. 3. The lowest reported value was 0.80, while the highest was 0.94. A funnel plot revealed asymmetry (Fig. 4).
The pooled ASSD was 0.46 (CI: 0.42–0.51, p < 0.001), with a heterogeneity (I2) of 96.1%, from 43 models. The mean relative volume error was 11.7% (CI: 9.7–13.7, p < 0.001) from 24 models, with a heterogeneity (I2) of 93.8%.
Architecture subgroup analysis
In the architecture subgroup analysis, we compared different DL architectures, mainly 2.5D networks (40 models), 3D networks (13 models), and other models that did not fit into these categories (3). Among the 2.5D DL networks, the mean Dice score was 0.89 (CI: 0.88–0.90), with a heterogeneity of 94.4% (I2). Thirty-two models reported ASSD, and the pooled result was 0.45 (CI: 0.40–0.50, I2 = 94.9%). The reported RVE was 10.3% (CI: 8.7–11.9, I2 = 86.6%) from 15 models.
For 3D networks, the pooled Dice score was 0.90 (CI: 0.88–0.91), with a heterogeneity of 94.5%. From 8 models, the average ASSD was 0.45 (CI: 0.33–0.56, I2 = 95.7%), and the pooled RVE was 13.4% (CI: 8.7–18.0, I2 = 95.8%).
For other networks, the mean Dice score was 0.82 (CI: 0.81–0.84), with I2 = 26.2%, and ASSD = 0.72 (CI: 0.55–0.88, I2 = 76.2%). Only one work pooled RVE, 18.0% (CI: 13.1–22.9).
MRI input subgroup analysis
In the MRI input subgroup analysis, models were divided into groups based on their imaging input. The mixed group was for DL networks using two or more different MRI inputs. The mean Dice score for 13 models using mixed MR imaging inputs was 0.90 (CI: 0.89–0.92, I2 = 94.9%), and the mean ASSD from 8 models was 0.37 (CI: 0.28–0.45, I2 = 94.7%). Four models reported RVE, pooling 7.8% (CI: 6.9–8.6, I2 = 0.0%).
For 22 T2W models, the mean dice score was 0.86 (CI: 0.85–0.87, I2 = 86.3%), 19 models reported ASSD 0.59 (CI: 0.54–0.64, I2 = 70.4%). Pooled RVE from 10 models was 14.0% (CI: 12.3–15.6, I2 = 57.6%).
Twenty models used ceT1W input – pooled Dice score was 0.91 (CI: 0.90–0.92, I2 = 93.5%). From 16 models reporting ASSD, mean results pooled was 0.36 (CI: 0.31–0.41, I2 = 94.5%). Ten models reported RVE, yielding 10.9% (CI: 7.1–14.6, I2 = 96.7%).
Only 1 model used T1W input and only reported a Dice score of 0.82 (CI: 0.79–0.85).
Testing set size subgroup analysis
In the final subgroup analysis, we compared models’ performance, grouping them based on their testing set size. DL algorithms were divided into < 50, 50–100, and > 100 patient set sizes. There were 41 models tested on sets smaller than 50. The pooled Dice score was 0.88 (CI: 0.87–0.90, I2 = 95.0%). Thirty-eight models reported ASSD, and the pooled result was 0.48 (CI: 0.43–0.53, I2 = 95.4%), while the mean RVE from 19 models was 11.0% (CI: 9.4–12.6, I2 = 87.1%).
Eleven models were tested on sets of size 50–100 patients. The pooled Dice score was 0.88 (CI: 0.86–0.91, I2 = 97.6%). One work reported ASSD and RVE.
Four models were tested on sets with more than 100 patients. The pooled Dice score was 0.90 (CI: 0.89–0.92, I2 = 88.2%). Pooled ASSD was 0.37 (CI: 0.32–0.41, I2 = 45.1%), while RVE was 14.2% (CI: 5.3–23.1, I2 = 98.3%). A Dice score subgroups analysis summary is available as Fig. 5, for ASSD subgroups as Fig. 6, while for RVE as Fig. 7.
Discussion
In the rapidly evolving landscape of medical imaging, our systematic review and meta-analysis aims to comprehensively summarize the current state of automated segmentation of Vestibular Schwannoma using Deep Learning techniques. As artificial intelligence, particularly Convolutional Neural Networks, continues to redefine diagnostic paradigms, understanding the application and reliability of these technologies is imperative.
The rarity of VS and its potential severity underscore the importance of accurate and timely diagnosis. Our meta-analysis emphasizes the significance of advanced imaging techniques, primarily MR scans, in evaluating tumor localization and size. Diagnosis of rare tumors might be especially challenging for inexperienced residents of radiology. The implementation of DL not only assists younger specialists but also reduces the time required for the manual segmentation process. Traditional gold standard methods, while effective, are being complemented and, in some instances, challenged by the advent of AI. Reduced workload allows for routine application of volumetric analysis [32]. Manual contouring suffers from inter-operator bias [24]. Automated segmentation offers the potential for standardized and reproducible volume measurements. Traditional linear measurements lack sensitivity, while volumetric measurements are more accurate but much more time-consuming [33, 34]. DL automation allows for efficient and accurate 3D volume calculation, facilitating better clinical decision-making. It is estimated, that automation tools could reduce even 30% time for segmentation [35]. Finally, the ability to perform segmentation using solely T2-weighted MRI addresses concerns about gadolinium contrast agents and potentially reduces costs, to an even 10-fold saving per scan [36, 37].
The integration of CNNs, a subset of DL, holds promise in automating the diagnostic process for VS. The substantial number of MRI scans associated with VS diagnosis and follow-up contributes significantly to healthcare costs. AI-based segmentation offers a novel option to address these conditions by optimizing the workflow of radiologists. Our meta-analysis attests to the increasing interest in leveraging DL for condition detection, tumor volume measurements, and neoplasm segmentation.
Despite strides in AI applications in medical imaging, our study reveals a critical knowledge gap concerning automated segmentation of VS using DL techniques. To the best of our knowledge, this is the very first systematic review and meta-analysis on this topic.
Adherence to the 2020 PRISMA guidelines and the use of the QUADAS-2 tool underscore the methodological rigor employed in our study. Comprehensive searches across major databases, incorporation of specific keywords, and manual reviews of references aimed to encompass relevant literature. Our inclusion criteria, while necessary, led to the exclusion of studies lacking extractable data, highlighting the importance of standardized reporting in future research.
The pooled mean Dice score of 0.89 indicates a commendable level of agreement between segmented and true volumes. However, the observed heterogeneity of 95.9% raises questions about methodological consistency across studies. High heterogeneity could be influenced by the different number of patients used in testing, training and validating models, population (patient) differences, different functions applied in the DL architecture, as well as technical differences in the MR images.
While the Dice score provides a comprehensive measure of segmentation accuracy, the inclusion of Relative Volume Error and Average Symmetric Surface Distance adds granularity to our assessment. The pooled RVE of 11.7% indicates the average deviation between segmented and true volumes. The ASSD of 0.46 mm highlights the alignment of segmented surfaces with ground truth surfaces, showcasing the robustness of DL algorithms in preserving anatomical details.
The subgroup analyses further illuminate the nuanced performance variations. 2.5D DL networks, with a mean Dice score of 0.89, demonstrate comparable efficacy to 3D networks (mean Dice score of 0.90) and outperform other models (mean Dice score of 0.82). Imaging input analyses reveal the superiority of contrast-enhanced T1-weighted imaging and mixed MRI inputs, emphasizing the influence of input modality on segmentation outcomes.
Studies varied in the MRI imaging protocols. Scanner field strength ranged from 1.5T to 3T. For the ceT1 scans, MPRAGE sequence was the most common option. T2W weighted scans utilized 3D CISS, TSE, and 2D Spin Echo sequences. However, notable variations in slice thickness and in-plane resolution (Table 2) could affect the effectiveness of the algorithms in other scenarios.
Despite the promising findings, limitations exist. We were unable to retrieve two records, what arises publication bias. Through QUADAS-2 tool, we have identified bias in different aspects of included studies – these risks in included works should be acknowledged, as well as high heterogeneity of the results. Additionally, articles only in the English language were included. Readers should be aware of these limitations, and future systematic reviews should address these gaps.
Regarding external validation, most of the studies did not perform external analysis of the AI, which raises the risk of overfitting to certain MRI setups. Additionally, external datasets mostly relied on a single database provided by Shapey et al. [30]. This raises question how will DL models perform in the randomized trials or prospective studies with different environments. Studies also lacked software (type and version used) and/or hardware information (GPU type), which could guide future researchers. These aspects are useful for software designers, and should be reported in the future studies.
Another aspect is the analysis of the cystic and solid parts of the VS. As the imaging properties differ, not all studies have deeply analysed them, and often only mentioned difficulties with cystic parts analysis [38]. Finally, other parameters such as size (beside volume), and proportion of the cystic cases should be provided. These aspects are crucial to be resolved in the future trials to provide more comprehensive analysis of VS for better clinical decision-making.
While the high Dice score shows promising results, which could be theoretically implemented in clinical practice, currently due to the limitations, more testing is required for full-scale clinical application. Future research should report fully validated models with more standardized designs, in order to increase homogeneity across the studies. Additionally, we believe that using bigger testing sets, would decrease overestimation of the results, as majority of DL models in this study were assessed on sets with 100 or less patients.
Conclusions
To summarize, this meta-analysis explores current knowledge of automated segmentation of VS using DL techniques. While Dice, RVE and ASSD scores show promising results for this task, the study is limited by high heterogeneity, caused by variations in MRI techniques, population differences and DL model differences, as well as bias among studies. Future research in this field should address these limitations for more standardized results.
Data Availability
Authors declare, that all relevant data was presented within the manuscript or figures. On special request, authors are able to send files with data used in this meta-analysis.
References
Rocío F-M, Wan YL, Axon P, Joannides A (2023) Incidence and presentation of vestibular schwannoma: a 3-year cohort registry study. Acta Neurochir 165:2903–2911
Park J Vestibular schwannoma (acoustic neuroma). www.uptodate.com at <
Reznitsky M, Petersen MMBS, West N, Stangerup S-E (2019) Cayé-Thomasen, P. Epidemiology of vestibular schwannomas – prospective 40-Year Data from an unselected National Cohort. Clin Epidemiol Volume 11:981–986
Cioffi G, Yeboa DN, Kelly M, Patil N, Manzoor N, Greppin K, Takaoka K, Waite K, Kruchko C, Barnholtz-Sloan (2020) J. S. Epidemiology of vestibular schwannoma in the United States, 2004–2016. Neuro-Oncology Adv 2
Marinelli JP, Beeler C, Carlson ML, Cayé-Thomasen P, Spear SA, Erbele ID (2021) Global incidence of sporadic vestibular Schwannoma: a systematic review. Otolaryngology-Head Neck Surg 167:209–214
Vestibular Schwannoma (Acoustic Neuroma) and, Neurofibromatosis NIDCD (2015) at
Ius T, Sabatino G, Panciani PP, Fontanella M, Rudà R, Castellano A, Barbagallo G, Belotti F, Boccaletti R, Catapano G, Costantino G, Alessandro Della Puppa, Francesco Di Meco, Gagliardi F, Garbossa D, Antonino Germanò M, Iacoangeli P, Mortini, Olivi A, Pessina F (2023) Surgical management of Glioma Grade 4: technical update from the neuro-oncology section of the Italian Society of Neurosurgery (SINch®): a systematic review. Journal of Neuro-oncology 162, 267–293
Stangerup S-E, Caye-Thomasen P, Tos M, Thomsen (2006) J Nat History Vestib Schwannoma Otology Neurotology 27:547–552
Lin EP, Crane BT (2017) The management and imaging of vestibular Schwannomas. Am J Neuroradiol 38:2034–2043
Benson JC, Carlson ML, Lane J (2020) I. MRI of the Internal Auditory Canal, Labyrinth, and middle ear: how we do it. Radiology 297:252–265
MathWorks. What is a Convolutional Neural Network? www.mathworks.com (2023) at https://www.mathworks.com/discovery/convolutional-neural-network-matlab.html
Hallinan JTPD, Zhu L, Yang K, Makmur A, Algazwi DAR, Thian YL, Lau S, Choo YS, Eide SE, Yap QV, Chan YH, Tan JH, Kumar N, Ooi BC, Yoshioka H, Quek ST (2021) Deep learning model for automated detection and classification of Central Canal, lateral recess, and neural Foraminal stenosis at lumbar spine MRI. Radiology 300:130–138
Lee DK, Sung DJ, Kim C-S, Heo Y, Lee JY, Park BJ, Kim MJ (2020) Three-dimensional convolutional neural network for prostate MRI segmentation and comparison of prostate volume measurements by Use of Artificial neural network and Ellipsoid Formula. Am J Roentgenol 214:1229–1238
Schouten JPE, Noteboom S, Martens RM, Mes SW, C. René Leemans, Pim de Graaf, Steenwijk M (2022) D. Automatic segmentation of head and neck primary tumors on MRI using a multi-view CNN. Cancer Imaging 22, https://doi.org/10.1186/s40644-022-00445-7
Tian J, Dong D, Liu Z, Wei J (2021) Radiomics and its clinical application. Elsevier eBooks 1–18. https://doi.org/10.1016/b978-0-12-818101-0.00004-5
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372. https://doi.org/10.1136/bmj.n71
Whiting PF (2011) QUADAS-2: a revised Tool for the Quality Assessment of Diagnostic Accuracy studies. Ann Intern Med 155:529
McGuinness LA, Higgins JPT (2020) Risk-of‐bias VISualization (robvis): an R package and Shiny web app for visualizing risk‐of‐bias assessments. Res Synthesis Methods 12. https://doi.org/10.1002/jrsm.1411
Shapey J, Kujawa A, Dorent R, Wang G, Dimitriadis A, Grishchuk D, Paddick I, Kitchen N, Bradford R, Saeed SR, Bisdas S, Ourselin S, Vercauteren T (2021) Segmentation of vestibular schwannoma from MRI, an open annotated dataset and baseline algorithm. Sci Data 8
Shapey J, Wang G, Dorent R, Dimitriadis A, Li W, Paddick I, Kitchen N, Bisdas S, Saeed SR, Ourselin S, Bradford R, Vercauteren T (2019) An artificial intelligence framework for automatic segmentation and volumetry of vestibular schwannomas from contrast-enhanced T1-weighted and high-resolution T2-weighted MRI. J Neurosurg 134:171–179
Wang G, Shapey J, Li W, Dorent R, Dimitriadis A, Bisdas S, Paddick I, Bradford R, Zhang S, Sébastien, Ourselin, Vercauteren T (2019) Automatic segmentation of vestibular Schwannoma from T2-Weighted MRI by deep spatial attention with hardness-weighted loss. 264–272. Lecture Notes in Computer Sciencehttps://doi.org/10.1007/978-3-030-32245-8_30
Wang H, Qu T, Bernstein K, Barbee D, Kondziolka D (2023) Automatic segmentation of vestibular schwannomas from T1-weighted MRI with a deep neural network. Radiat Oncol 18
Neves CA, Liu GS, Chemaly E, Bernstein T, Fu IA, F., Blevins NH (2023) Automated radiomic analysis of vestibular Schwannomas and Inner ears using contrast-enhanced T1-Weighted and T2-Weighted magnetic resonance imaging sequences and Artificial Intelligence. Otology Neurotology 44:e602
Lee W-K, Wu CC, Lee C-C, Feng Lu C, Yang H-C, Huang T-H, Lin C-Y, Chung W-Y, Wang PW, Hsiu Mei Wu, Guo, W.-Y., He Z (2020) Combining analysis of multi-parametric MR images into a convolutional neural network: Precise target delineation for vestibular schwannoma treatment planning. Artificial Intelligence in Medicine 107, 101911 https://doi.org/10.1016/j.artmed.2020.101911
Neve OM, Chen Y, Tao Q, Romeijn SR, Nick W, Grootjans, Kruit MC, Boudewijn PF, Lelieveldt, Jansen JC, Hensen EF, Verbist BM, Staring M (2022) Fully automated 3D vestibular Schwannoma segmentation with and without Gadolinium-based contrast material: a Multicenter, Multivendor Study. Radiology: Artif Intell 4. https://doi.org/10.1148/ryai.210300
Cass ND, Lindquist NR, Zhu Q, Li H, Oguz I, Tawfik K (2022) O. Machine Learning for Automated calculation of vestibular Schwannoma volumes. Otology Neurotology 43:1252–1256
Wu J, Guo D, Wang L, Yang S, Zheng Y, Shapey J, Vercauteren T, Bisdas S, Bradford R, Saeed S, Kitchen N, Ourselin S, Zhang S, Wang G (2023) TISS-Net: Brain Tumor Image Synthesis and Segmentation using Cascaded Dual-Task Networks and Error-Prediction consistency. Neurocomputing 126295. https://doi.org/10.1016/j.neucom.2023.126295
Kujawa A, Dorent R, Connor S, Oviedova A, Okasha M, Grishchuk D, Sébastien Ourselin, Paddick I, Kitchen N, Vercauteren T, Shapey J (2022) Automated Koos classification of Vestibular Schwannoma. Front Radiol 2. https://doi.org/10.3389/fradi.2022.837191
Zhai S, Wang G, Luo X, Yue Q, Li K, Zhang S (2023) PA-Seg: learning from point annotations for 3D medical image Segmentation using Contextual regularization and Cross Knowledge Distillation. IEEE Trans Med Imaging 42:2235–2246
Shapey J, Kujawa A, Dorent R Segmentation of Vestibular Schwannoma from Magnetic Resonance Imaging: An Open Annotated Dataset and Baseline Algorithm (version 2) [Data set]. The Cancer Imaging Archive doi:https://doi.org/10.7937/TCIA.9YTJ-5Q73
Dorent R crossMoDA. crossMoDA athttps://crossmoda-challenge.ml/
Varughese JK, Wentzel-Larsen T, Vassbotn FS, Moen G (2010) & Morten Lund-Johansen. Analysis of vestibular schwannoma size in multiple dimensions: a comparative cohort study of different measurement techniques. Clinical Otolaryngology 35, 97–103 https://doi.org/10.1111/j.1749-4486.2010.02099.x
Rick, Jan B, Nelemans PJ, Baumert BG, Stokroos RJ (2009) Follow-up assessment of vestibular schwannomas: volume quantification versus two-dimensional measurements. Neuroradiology 51:517–524
MacKeith S, Das T, Graves M, Patterson A, Donnelly N, Mannion R, Axon P, Tysome J (2018) A comparison of semi-automated volumetric vs linear measurement of small vestibular schwannomas. Eur Arch Otorhinolaryngol 275:867–874
Lu S-L, Xiao F-R, Cheng JC-H, Yang W-C, Cheng Y-H, Chang Y-C, Lin J-Y, Liang C-H, Lu J-T, Chen Y-F, Hsu F-M (2021) Randomized multi-reader evaluation of automated detection and segmentation of brain tumors in stereotactic radiosurgery with deep neural networks. Neurooncology 23:1560–1568
Tolisano AM, Wick CC, Hunter JB (2019) Comparing Linear and volumetric vestibular schwannoma measurements between T1 and T2 magnetic resonance imaging sequences. Otology Neurotology 40:S67–S71
Coelho DH, Tang Y, Suddarth B, Mamdani M (2017) MRI surveillance of vestibular schwannomas without contrast enhancement: clinical and economic evaluation. Laryngoscope 128:202–209
Franco Benech, Perez R, Fontanella MM, Morra B, Albera R (2005) Alessandro Ducati. Cystic versus solid vestibular schwannomas: a series of 80 grade III–IV patients. Neurosurg Rev 28:209–213
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Conflict of interest
Authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix 1
Appendix 1
Search strategy:
-
1.
PUBMED/Scopus/Web of science(AI OR ( artificial AND intelligence ) OR automation OR ( convolutional AND neural AND network ) OR cnn OR ( deep AND learning ) OR ( machine AND learning ) OR ( decision AND tree) OR ( neural AND network ) OR (support AND vector AND machine) OR lasso OR (k-means) OR knn) AND (Vestibular Schwannoma OR acoustic neuroma OR (acoustic AND neuroma) OR (Vestibular AND Schwannoma) OR acoustic neurilemmoma OR neurinoma of the acoustic nerve OR schwannoma of the acoustic nerve OR perineural fibroblastoma).
-
2.
Embase
(ai OR (artificial AND (‘intelligence’/exp OR intelligence)) OR ‘automation’/exp OR automation OR (convolutional AND neural AND (‘network’/exp OR network)) OR cnn OR (deep AND (‘learning’/exp OR learning)) OR ((‘machine’/exp OR machine) AND (‘learning’/exp OR learning)) OR ((‘decision’/exp OR decision) AND (‘tree’/exp OR tree)) OR (neural AND (‘network’/exp OR network)) OR ((‘support’/exp OR support) AND (‘vector’/exp OR vector) AND (‘machine’/exp OR machine)) OR ‘lasso’/exp OR lasso OR ‘k means’/exp OR ‘k means’ OR knn) AND (‘vestibular schwannoma’/exp OR ‘vestibular schwannoma’ OR ‘acoustic neuroma’/exp OR ‘acoustic neuroma’ OR (acoustic AND (‘neuroma’/exp OR neuroma)) OR (vestibular AND (‘schwannoma’/exp OR schwannoma)) OR ‘acoustic neurilemmoma’ OR (acoustic AND (‘neurilemmoma’/exp OR neurilemmoma)) OR 'neurinoma of the acoustic nerve’ OR ((‘neurinoma’/exp OR neurinoma) AND of AND the AND acoustic AND (‘nerve’/exp OR nerve)) OR 'schwannoma of the acoustic nerve’ OR ((‘schwannoma’/exp OR schwannoma) AND of AND the AND acoustic AND (‘nerve’/exp OR nerve)) OR ‘perineural fibroblastoma’ OR (perineural AND (‘fibroblastoma’/exp OR fibroblastoma)))
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Łajczak, P., Matyja, J., Jóźwik, K. et al. Accuracy of vestibular schwannoma segmentation using deep learning models - a systematic review & meta-analysis. Neuroradiology (2024). https://doi.org/10.1007/s00234-024-03449-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00234-024-03449-1