Abstract
Purpose
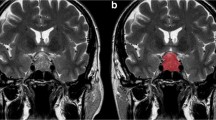
To evaluate the potential value of machine learning (ML)-based histogram analysis (or first-order texture analysis) on T2-weighted magnetic resonance imaging (MRI) for predicting consistency of pituitary macroadenomas (PMA) and to compare it with that of signal intensity ratio (SIR) evaluation.
Methods
Fifty-five patients with 13 hard and 42 soft PMAs were included in this retrospective study. Histogram features were extracted from coronal T2-weighted original, filtered and transformed MRI images by manual segmentation. To achieve balanced classes (38 hard vs 42 soft), multiple samples were obtained from different slices of the PMAs with hard consistency. Dimension reduction was done with reproducibility analysis, collinearity analysis and feature selection. ML classifier was artificial neural network (ANN). Reference standard for the classifications was based on surgical and histopathological findings. Predictive performance of histogram analysis was compared with that of SIR evaluation. The main metric for comparisons was the area under the receiver operating characteristic curve (AUC).
Results
Only 137 of 162 features had excellent reproducibility. Collinearity analysis yielded 20 features. Feature selection algorithm provided six texture features. For histogram analysis, the ANN correctly classified 72.5% of the PMAs regarding consistency with an AUC value of 0.710. For SIR evaluation, accuracy and AUC values were 74.5% and 0.551, respectively. Considering AUC values, ML-based histogram analysis performed better than SIR evaluation (z = 2.312, p = 0.021).
Conclusion
ML-based T2-weighted MRI histogram analysis might be a useful technique in predicting the consistency of PMAs, with a better predictive performance than that of SIR evaluation.





Similar content being viewed by others
References
Alimohamadi M, Sanjari R, Mortazavi A, Shirani M, Moradi Tabriz H, Hadizadeh Kharazi H, Amirjamshidi A (2014) Predictive value of diffusion-weighted MRI for tumor consistency and resection rate of nonfunctional pituitary macroadenomas. Acta Neurochir 156:2245–2252. https://doi.org/10.1007/s00701-014-2259-6
Pierallini A, Caramia F, Falcone C, Tinelli E, Paonessa A, Ciddio AB, Fiorelli M, Bianco F, Natalizi S, Ferrante L, Bozzao L (2006) Pituitary macroadenomas: preoperative evaluation of consistency with diffusion-weighted MR imaging—initial experience. Radiology 239:223–231. https://doi.org/10.1148/radiol.2383042204
Romano A, Coppola V, Lombardi M, Lavorato L, di Stefano D, Caroli E, Rossi Espagnet MC, Tavanti F, Minniti G, Trillò G, Bozzao A (2017) Predictive role of dynamic contrast enhanced T1-weighted MR sequences in pre-surgical evaluation of macroadenomas consistency. Pituitary 20:201–209. https://doi.org/10.1007/s11102-016-0760-z
Snow RB, Johnson CE, Morgello S, Lavyne MH, Patterson RH Jr (1990) Is magnetic resonance imaging useful in guiding the operative approach to large pituitary tumors? Neurosurgery 26:801–803. https://doi.org/10.1227/00006123-199005000-00011
Iuchi T, Saeki N, Tanaka M, Sunami K, Yamaura A (1998) MRI prediction of fibrous pituitary adenomas. Acta Neurochir 140:779–786
Naganuma H, Satoh E, Nukui H (2002) Technical considerations of transsphenoidal removal of fibrous pituitary adenomas and evaluation of collagen content and subtype in the adenomas. Neurol Med Chir (Tokyo) 42:202–212 discussion 213
Yang C, Zhang H, Wang H j et al (2002) Controlled pathological study and preoperative MRI evaluation of the consistency of pituitary adenomas. Ai Zheng 21:675–677
Smith KA, Leever JD, Chamoun RB (2015) Prediction of consistency of pituitary adenomas by magnetic resonance imaging. J Neurol Surg B Skull Base 76:340–343. https://doi.org/10.1055/s-0035-1549005
Suzuki C, Maeda M, Hori K, Kozuka Y, Sakuma H, Taki W, Takeda K (2007) Apparent diffusion coefficient of pituitary macroadenoma evaluated with line-scan diffusion-weighted imaging. J Neuroradiol 34:228–235. https://doi.org/10.1016/j.neurad.2007.06.007
Chakrabortty S, Oi S, Yamaguchi M et al (1993) Growth hormone-producing pituitary adenomas: MR characteristics and pre- and postoperative evaluation. Neurol Med Chir 33:81–85
Bahuleyan B, Raghuram L, Rajshekhar V, Chacko AG (2006) To assess the ability of MRI to predict consistency of pituitary macroadenomas. Br J Neurosurg 20:324–326. https://doi.org/10.1080/02688690601000717
Thotakura AK, Patibandla MR, Panigrahi MK, Mahadevan A (2017) Is it really possible to predict the consistency of a pituitary adenoma preoperatively? Neurochirurgie 63:453–457. https://doi.org/10.1016/j.neuchi.2017.06.003
Ganeshan B, Miles KA (2013) Quantifying tumour heterogeneity with CT. Cancer Imaging 13:140–149. https://doi.org/10.1102/1470-7330.2013.0015
Kocak B, Durmaz ES, Kadioglu P, Polat Korkmaz O, Comunoglu N, Tanriover N, Kocer N, Islak C, Kizilkilic O (2018) Predicting response to somatostatin analogues in acromegaly: machine learning-based high-dimensional quantitative texture analysis on T2-weighted MRI. Eur Radiol. https://doi.org/10.1007/s00330-018-5876-2
Lubner MG, Smith AD, Sandrasegaran K, Sahani DV, Pickhardt PJ (2017) CT texture analysis: definitions, applications, biologic correlates, and challenges. RadioGraphics 37:1483–1503. https://doi.org/10.1148/rg.2017170056
Tustison NJ, Avants BB, Cook PA, Yuanjie Zheng, Egan A, Yushkevich PA, Gee JC (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29:1310–1320. https://doi.org/10.1109/TMI.2010.2046908
Collewet G, Strzelecki M, Mariette F (2004) Influence of MRI acquisition protocols and image intensity normalization methods on texture classification. Magn Reson Imaging 22:81–91. https://doi.org/10.1016/j.mri.2003.09.001
Shafiq-ul-Hassan M, Zhang GG, Latifi K, Ullah G, Hunt DC, Balagurunathan Y, Abdalah MA, Schabath MB, Goldgof DG, Mackin D, Court LE, Gillies RJ, Moros EG (2017) Intrinsic dependencies of CT radiomic features on voxel size and number of gray levels. Med Phys 44:1050–1062. https://doi.org/10.1002/mp.12123
Van Griethuysen JJM, Fedorov A, Parmar C et al (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res 77:e104–e107. https://doi.org/10.1158/0008-5472.CAN-17-0339
Koo TK, Li MY (2016) A guideline of selecting and reporting intraclass correlation coefficients for reliability research. J Chiropr Med 15:155–163. https://doi.org/10.1016/j.jcm.2016.02.012
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G, Marquéz JRG, Gruber B, Lafourcade B, Leitão PJ, Münkemüller T, McClean C, Osborne PE, Reineking B, Schröder B, Skidmore AK, Zurell D, Lautenbach S (2013) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography (Cop) 36:27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x
Kohavi R, John GH (1997) Wrappers for feature subset selection. Artif Intell 97:273–324. https://doi.org/10.1016/S0004-3702(97)00043-X
Bermejo P, Gamez JA, Puerta JM (2011) Improving incremental wrapper-based subset selection via replacement and early stopping. Int J Pattern Recognit Artif Intell 25:605–625. https://doi.org/10.1142/S0218001411008804
Mwangi B, Tian TS, Soares JC (2014) A review of feature reduction techniques in neuroimaging. Neuroinformatics 12:229–244. https://doi.org/10.1007/s12021-013-9204-3
Cawley GC, Talbot NLC (2010) On over-fitting in model selection and subsequent selection bias in performance evaluation. J Mach Learn Res 11:2079–2107
Varma S, Simon R (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics 7:91. https://doi.org/10.1186/1471-2105-7-91
Ahmad A, Dey L (2005) A feature selection technique for classificatory analysis. Pattern Recogn Lett 26:43–56. https://doi.org/10.1016/j.patrec.2004.08.015
Demšar J (2006) Statistical comparisons of classifiers over multiple data sets. J Mach Learn Res 7:1–30
Rui W, Wu Y, Ma Z, Wang Y, Wang Y, Xu X, Zhang J, Yao Z (2019) MR textural analysis on contrast enhanced 3D-SPACE images in assessment of consistency of pituitary macroadenoma. Eur J Radiol 110:219–224. https://doi.org/10.1016/j.ejrad.2018.12.002
Hagiwara A, Inoue Y, Wakasa K, Haba T, Tashiro T, Miyamoto T (2003) Comparison of growth hormone–producing and non-growth hormone-producing pituitary adenomas: imaging characteristics and pathologic correlation. Radiology 228:533–538. https://doi.org/10.1148/radiol.2282020695
Abe N, Kinoshita Y, Mahmoud OM et al (2010) Role of PROPELLER diffusion-weighted imaging and apparent diffusion coefficient in the evaluation of pituitary adenomas. Eur J Radiol 80:412–417. https://doi.org/10.1016/j.ejrad.2010.05.023
Boxerman JL, Rogg JM, Donahue JE, Machan JT, Goldman MA, Doberstein CE (2010) Preoperative MRI evaluation of pituitary macroadenoma: imaging features predictive of successful transsphenoidal surgery. AJR Am J Roentgenol 195:720–728. https://doi.org/10.2214/AJR.09.4128
Yamamoto J, Korogi Y, Takahashi M et al (2013) Tumor consistency of pituitary macroadenomas: predictive analysis on the basis of imaging features with contrast-enhanced 3D FIESTA at 3T. Am J Neuroradiol 35:297–303. https://doi.org/10.3174/ajnr.a3667
Funding
No funding was received for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
We considered all ethical standards in this study.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in the studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Requirement for informed consent was waived by IRB of Istanbul University-Cerrahpasa, Cerrahpasa Medical Faculty.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeynalova, A., Kocak, B., Durmaz, E.S. et al. Preoperative evaluation of tumour consistency in pituitary macroadenomas: a machine learning-based histogram analysis on conventional T2-weighted MRI. Neuroradiology 61, 767–774 (2019). https://doi.org/10.1007/s00234-019-02211-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-019-02211-2