Abstract
Membrane fusion plays a lead role in the transport of vesicles, neurotransmission, mitochondrial dynamics, and viral infection. There are fusion proteins that catalyze and regulate the fusion. Interestingly, various types of fusion proteins are present in nature and they possess diverse mechanisms of action. We have highlighted the importance of the functional domains of intracellular heterotypic fusion, homotypic endoplasmic reticulum (ER), homotypic mitochondrial, and type-I viral fusion. During intracellular heterotypic fusion, the SNAREs and four-helix bundle formation are prevalent. Type-I viral fusion is controlled by the membrane destabilizing properties of fusion peptide and six-helix bundle formation. The ER/mitochondrial homotypic fusion is controlled by GTPase activity and the membrane destabilization properties of the amphipathic helix(s). Although the mechanism of action of these fusion proteins is diverse, they have some similarities. In all cases, the lipid composition of the membrane greatly affects membrane fusion. Next, examples of lipidation of the fusion proteins were discussed. We suggest that the fatty acyl hydrophobic tail not only acts as an anchor but may also modulate the energetics of membrane fusion intermediates. Lipidation is also important to design more effective peptide-based fusion inhibitors. Together, we have shown that membrane lipid composition and lipidation are important to modulate membrane fusion.
Graphical Abstract

Similar content being viewed by others
Introduction
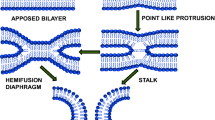
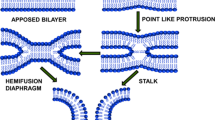
Lipids are one of the essential molecules in biology. For a long period, lipids were not considered as fashionable as proteins (catalyze reactions), and nucleic acids (store genetic information). However, the traditional view on lipids has changed substantially during the last few decades. It is now well-recognized that protein-lipid interaction plays an important role in the survival of cells. Lipids form supramolecular assemblies; like micelle, bilayer, and hexagonal phase in water. The bilayer assemblies (membrane) were chosen by nature during evolution to form compartments (Izgu et al. 2016; Sarkar et al. 2020). These compartments were important to the proto-cell development, and evolution of life. The ‘Lipid World’ model of evolution also suggests that prebiotic amphiphile and lipid bilayer membrane catalyzed biological reactions during early evolution (Segré et al. 2001; Lancet et al. 2019). Amino acids and lipidated amino acids further stabilize the membrane compartments to support the evolution (Cornell et al. 2019; Joshi et al. 2021). It was proposed that communication between the compartments was essential for establishing networks. We propose that the prebiotic communication between the membrane compartments may be recognized as the primitive version of membrane fusion. It occurs between two separate lipid bilayers that merge into a continuous bilayer and the internal contents of two separated membranes mingle (Fig. 1). Membrane fusion is a lipid-centric event and immensely important in living systems, from viruses to eukaryotic cells. It contributes to virus infection, neurotransmitter release, cell fertilization, and intracellular transport (Jahn et al. 2003; Han et al. 2017). Fusion starts with the close apposition of two bilayers. However, due to the hydration repulsion between the proximal leaflets of the membranes, fusion is not a spontaneous process. In today’s world, the fusion proteins catalyze and regulate the fusion. There are various proposals on how membrane fusion proteins catalyze the fusion in living systems. Among them, the widely accepted lipid-centric simple model suggests that the conformational energy of the protein (pre-fusion to a post-fusion structure) is harnessed to generate the hemifusion intermediate (through arrays of non-lamellar lipid intermediate structures), and then the fusion pore (Fig. 1) (Collins et al. 2012). More precisely, the fusion starts with pre-fusion contact/docking. Then a hemifusion intermediate(s) were formed, which lead to pore formation (Fig. 1). Every step has an activation barrier due to the membrane curvature deformations and unfavorable lipid-water arrangements (Chakraborty et al. 2012). Studies have shown that lipid composition can influence every step: from docking to hemifusion to fusion pore (Chernomordik and Kozlov 2003; Tamm et al. 2003; Meher and Chakraborty 2019). Lipid composition can also modulate the structure, organization, and dynamics of fusion proteins. In addition, the lipid-binding domain of several fusion proteins binds to regulatory lipids and thus controls membrane fusion. Since the membrane fusion proceeds via lipidic intermediates; the lipid composition of the membrane and the lipidation of the fusion proteins may perturb fusion. Lipidation is a covalent attachment of lipid molecules to the protein/peptide (Jiang et al. 2018). In the present review, we focus on how the lipid composition of the membrane and lipidation of fusion proteins modulate several types of membrane fusion.
Membrane Fusion Proteins
There are two types of fusion in biology: homotypic and heterotypic. When two same-type membranes merge together, it is termed homotypic fusion. This type of fusion occurs in vacuoles, mitochondria, and the endoplasmic reticulum. In heterotypic fusion, two different types of membranes fuse with each other. Examples include virus and cell/endosomal membrane fusion, synaptic vesicles-plasma membrane fusion, late endosome-lysosome fusion etc. (Fig. 2) (Martens and McMahon 2008).
During synaptic vesicle fusion (neuronal transmission) and intracellular heterotypic fusion, soluble N-ethylmalemide sensitive factor attachment protein receptors (SNAREs) play the most important role in fusion (Jahn and Scheller 2006; Rizo and Xu 2015). The SNARE proteins that are responsible for the synaptic vesicle fusion comprise R-SNARE (attached to synaptic vesicles) and Q-SNARE (attached to the target membrane). Under the appropriate condition, the R-SNARE and Q-SNAREs interact with each other, bring two membranes to close proximity, and facilitate fusion (Fig. 3A). The post-fusion structure of the SNARE complex suggests that they form a four-helix bundle (coiled-coil motif) structure that usually contains one R-SNARE with two or three Q-SNAREs. It is well understood that the energy released by the four-helix bundle initiates and drives the fusion (Jahn et al. 2003). SNAREs also control the yeast homotypic vacuole fusion via a similar coiled-coil four-helix bundle structure (Wickner 2010).
A SNARE-mediated fusion via four-helix bundle formation. B Type-I viral fusion protein-mediated fusion via six-helix bundle (6-HB) formation, FP is the fusion peptide, RBD is the Receptor Binding Domain. C Mitofusin induced fusion of mitochondrial outer membrane occur via GTPase activity and helix-bundles
The fusion between the viral membrane and the host membrane is triggered by viral fusion proteins. In the case of the type-I fusion protein, a six-helix bundle (coiled-coil motif) formation between the heptad repeat 1 and 2 (HR1 and HR2) formation is crucial. The fusion protein also contains a 15–30 amino acid segment, known as the fusion peptide. During fusion, the fusion peptide binds to the target membrane, and then the six-helix bundle formation by HR1 and HR2 domains supplies the energy required for fusion (Fig. 3B). Therefore both the fusion peptide and the six-helix bundle have an important role to catalyze the type-I viral fusion (Table 1) (Meher and Chakraborty 2019). Studies have shown that the fusion peptide may influence the membrane interface and induce non-bilayer structures to catalyze the fusion.
The homotypic fusion of the endoplasmic reticulum membrane is triggered by dynamin-like fusogenic GTPases, atlastins, and the homotypic fusion of mitochondrial outer membranes is triggered by GTPase mitofusins (MFNs), optic atrophy-1 (OPA1) (Moon and Jun 2020). MFNs consist of an N-terminal GTPase domain followed by HR domains, transmembrane domains, and a C-terminal HR domain (Fig. 3C). In most cases, MFNs have two transmembrane domains and thus both HR domains reside at the cytoplasmic site (Fig. 3C) (Li et al. 2019). The exact mechanism by which the HR domains of MFNs facilitate fusion remains unclear (Cohen and Tareste 2018; Moon and Jun 2020). However, the present model suggests that the MFNs of two fusing membrane forms a homodimer via the GTPase domain and the interaction of the HR2 domain. Upon GTP hydrolysis, a drastic conformational change results in the merger of the membranes (Cao et al. 2017). A recent report suggests that the HR1 domain of MFN also possesses membrane destabilization properties (Daste et al. 2018). The ER fusion protein family, atlastin contains an N-terminal GTPase domain followed by a helical domain, transmembrane domains, and a C-terminal domain (Bian et al. 2011; Yan et al. 2015). The GTPase dimerization, GTP hydrolysis, and HR2-membrane interaction control the atlastin-mediated fusion. It is noteworthy to mention that the fusion catalyzing proteins have different mechanisms of action in heterotypic intracellular fusion or viral fusion or ER/mitochondrial homotypic fusion. In SNAREs, the formation of a highly stable four-helix bundle is essential. The viral fusion is controlled by a six-helix bundle and fusion peptide-membrane interaction. The ER fusion/mitochondrial fusion is controlled by GTP hydrolysis, HR bundle, and HR-membrane interaction (Table 1).
Membrane Lipid Composition and Fusion
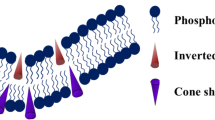
Membrane lipid may modulate fusion in two different pathways. We term the first pathway as intrinsic, where the lipid head group or tail structure affects the energetics of lipidic hemifusion intermediates. The second pathway is extrinsic, where lipid—fusion protein interaction modulates the energetics of fusion. In the intrinsic pathway, pioneering works were reported on the modulation of membrane curvature, thickness, hydration, and dipole moment. The biological membrane is not homogeneous and the lipid composition of different membranes is tuned to match their specific biological role. Phospholipids, sphingolipids, cholesterol (yeast: ergosterol), and diacylglycerol are major components of the biological membrane. Phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylserine (PS), phosphatidic acid (PA), phosphatidylglycerol (PG), phosphatidylinositol (PI), phosphatidylinositol phosphate (PIP) are the major phospholipids present in biological membranes (Lauwers et al. 2016). Among them, PC is cylindrical in shape and thus assembles into a planar-like bilayer (Fig. 4). PE has an inverted cone-shaped molecular structure, which promotes negative curvature to the membrane and facilitates the hemifusion structure to catalyze fusion (Fig. 4) (Fuller and Rand 2001; Meher and Chakraborty 2019). Diacylglycerol, cholesterol, and oleic acid also induce negative curvature and promote hemifusion. It is well-documented that the transition from bilayer to hemifusion state is associated with lipid-water protrusion and hexagonal phase formation (Chakraborty et al. 2012). Lysolipids like lysophosphatidylcholine and lysophosphoglycan have a large headgroup and the conical shape induces positive curvature to the membrane (Fig. 4). The positive curvature in the outer leaflet of the bilayer generally increases the activation barrier for the hemifusion intermediate formation and thus inhibits fusion. PS, PA, PG, PI, PIP are negatively charged and they bind to the polybasic patches of the protein (van den Bogaart et al. 2011; Tarafdar et al. 2015). However, negatively charged lipids like PS, PG inhibit model membrane fusion because of charge-charge repulsion (Tarafdar et al. 2012). The fusion may be promoted in presence of Ca2+, which brings PS-doped membranes close to each other (Tarafdar et al. 2012). Apart from phospholipids, cholesterol is one of the major lipid components of the cellular and sub-cellular membrane (Ingólfsson et al. 2014). Cholesterol induces negative curvature in lipid monolayers (Wang et al. 2007), and alters the membrane thickness, and fluidity to modulate the fusion. It generally promotes fusion by increasing membrane fluidity and negative curvature. Sphingomyelin (SM), a major sphingolipid mostly stabilizes the membrane and reduces membrane fusion (Haque et al. 2001). The unsaturated fatty acyl groups in phospholipids may change the overall geometry of the lipid, and increase the fluidity of the lipid membrane, to control membrane fusion (Pinot et al. 2014). Membrane fusion may also be affected by the presence of the liquid-ordered (Lo) phase. Cholesterol, with its unique structure, reduces the transgauche isomerizations of the neighboring fatty acyl chains and therefore facilitates the formation of Lo phase. Growing evidence suggests that HIV fusion peptides preferentially target the liquid-ordered/ liquid-disordered (Ld) boundary regions and promote fusion at the interfaces between Lo and Ld domains (Yang et al. 2015).
In the extrinsic pathway, lipids interact with the fusion machinery components to control the fusion. For example, PS and PIP recruit the synaptic fusion machinery components and facilitate the reorganization of fusion proteins and their domains (van den Bogaart et al. 2011; Pérez-Lara et al. 2016). During the pre-fusion state (synaptic fusion), the negatively charged PS and phosphatidylinositol biphosphate (PIP2) cause electrostatic repulsion between the membranes and prevent spontaneous fusion (van den Bogaart et al. 2011; Tarafdar et al. 2012). However, the PIP2 (highly negatively charged) interacts with the polybasic membrane juxtaposed region of syntaxin-1 to form clusters and prepare the effective docking sites (Fig. 5A) (van den Bogaart et al. 2011; Milovanovic et al. 2016). PIP2 and PS are also known to promote the interaction of synaptotagmin (a cofactor of synaptic fusion) to Ca2+ (Bai et al. 2004). In addition, the conserved polybasic patch located on the C2B domain of synaptotagmin binds to negatively charged multivalent lipids like PIP2 (Chapman 2008; Südhof 2013), and promotes the interactions either between SNAREpin and synaptotagmin or syntaxin and synaptotagmin (Pérez-Lara et al. 2016). These interactions are necessary for fast Ca2+-dependent exocytosis.
Membrane cholesterol also influences SNARE-mediated membrane fusion as the depletion of cholesterol from the membrane leads to a decrease in exocytosis (Yang et al. 2016; Han et al. 2017). Cholesterol alters the physical properties of the membrane and their interaction with fusion proteins (SNAREs) modulates the formation of cholesterol-dependent clusters (raft-like domain) that control docking at the fusion sites (Murray and Tamm 2009, 2011). Apart from SNARE-mediated exocytosis, membrane cholesterol also modulates viral infections by affecting the fusion between viral and host cell membranes. Cholesterol changes the fluidity, thickness, compressibility, water penetration, and intrinsic curvature of lipid bilayers to catalyze viral glycoprotein-mediated membrane fusion (Yang et al. 2016). In the case of influenza, increased cholesterol content decreases the energy to form hemifusion stalk (Chlanda et al. 2016) by altering the host membrane curvature (Liu and Boxer 2020). For HIV, cholesterol helps fusion between cholesterol-poor disordered and cholesterol-rich domains of target membranes (Yang et al. 2015). Cholesterol-transmembrane domain interactions of the Ebola virus glycoprotein (GP) affect structural features of GP to facilitate fusion (Lee et al. 2021). Increasing the cholesterol levels in the membrane improves the chances of full fusion events, whereas cholesterol-depleted virus-like particles show reduced cell entry (Lee et al. 2021). Cholesterol may also alter the oligomerization status of the N-terminal fusion peptide of SARS-CoV to modulate the membrane organization and dynamics (Meher et al. 2019). In the case of the SARS-CoV-2 virus, it was suggested that the depletion of cholesterol from the viral membrane affects membrane fusion and infection (Sanders et al. 2021). Similar studies with SARS-CoV-2 internal fusion peptide also showed that it induces more hemifusion in cholesterol-rich membranes (Pattnaik et al. 2021).
In yeast vacuole fusion, various lipids such as phosphatidic acid (PA), phosphatidylinositol phosphates (PIP), sphingolipids, diacylglycerol (DAG), and ergosterol (yeast cholesterol) regulate the homotypic membrane fusion (Wickner 2010; Wickner and Rizo 2017; Hurst and Fratti 2020). During atlastin-mediated ER homotypic fusion the specific lipid composition of ER membranes was essential to maintain the homeostasis. Sey1p (yeast ortholog of atlastin)-mediated in vitro liposome fusion studies were highly susceptible to the omission of ergosterol or PE (Sugiura and Mima 2016; Lee et al. 2019). Sey1p-dependent fusion was also reduced by omitting three acidic lipids (PS, PA, PI) from the liposome bilayer (Sugiura and Mima 2016). The C-terminal tail of atlastin was reported to be amphipathic and it is likely that the polybasic patches interact with the acidic membrane lipids, thereby affecting the local curvature and the stability of the membrane (Drin and Antonny 2010; Liu et al. 2012).
The role of lipids in MFN-mediated mitochondrial outer membrane fusion is widely documented (Moon and Jun 2020). The GTPase domain, HR1, HR2, and TMDs are essential for fusion. The HR1 domains of MFN bind to the lipid bilayer via amphipathic alpha-helix (Daste et al. 2018). Upon GTP hydrolysis, the dimeric HR1 domain interacts with the respective lipid membrane (Figs. 3C, 5B), which brings outer mitochondrial membranes in closer proximity. The binding of the amphipathic helix with the membrane perturbs the membrane structure and promotes fusion (Daste et al. 2018). Apart from the binding, the individual lipid component (PA and PE content) may introduce local negative curvature into the mitochondrial membrane and thus facilitate the MFN-mediated fusion (Frohman 2015; Cohen and Tareste 2018). Therefore, the lipid composition of the mitochondrial membrane guides the membrane fusion. It may be noted that the mitochondrial membrane has ~ 20% PE, and 5% PA. Since the overexpression of phospholipase A1, which degrades PA to lysophosphatidic acid triggers mitochondrial fragmentation (Baba et al. 2014), it has been suggested that mitochondrial fusion may depend on the PA content of the membrane. This led to the speculation that PA and HR1 (of MFN) interaction might facilitate the mitochondrial outer-membrane fusion (Fig. 5B) (Cohen and Tareste 2018).
The mitochondrial inner-membrane fusion is regulated by OPA1. Cardiolipin (CL) is an important mitochondrial inner-membrane lipid that comprises ~ 25% of the total lipid content (Horvath and Daum 2013; Moon and Jun 2020). CL contains two phosphate head groups and four acyl chains. The inactivation of CL synthase causes morphological defects in mitochondria (Matsumura et al. 2018). Therefore, the CL content of the membrane is crucial in mitochondrial inner-membrane fusion. CL may also enhance the GTPase activity of the yeast ortholog of OPA1 (Mgm1) and regulates the mitochondrial morphology (Frohman 2015; Ban et al. 2017; Ge et al. 2020).
Lipidation of Protein and Membrane Fusion
Membrane fusion is an event related to lipid bilayer, membrane attachment of the fusion proteins is essential. Viral membrane fusion proteins attach to the target membrane have transmembrane domain and fusion peptide(s) to anchor the lipid bilayer. The intracellular fusion proteins and the machinery components anchor membrane via transmembrane domains, Ca2+-binding domain, PS/PIP binding domains, or via acylation. Various types of acylation are reported in the literature including S-palmitoylation, S-prenylation, N-myristoylation, lysine-acylation etc. (Jiang et al. 2018; Suazo et al. 2021).
During neurotransmission, the SNARE complex in the presynaptic neuron was composed of syntaxin-1, vesicle-associated membrane protein-2 (VAMP-2), and SNAP-25 (Fig. 3A). VAMP-2 is located on the synaptic vesicle and syntaxin-1 and SNAP-25 are located on the presynaptic plasma membrane. VAMP-2 and syntaxin-1 contain a transmembrane domain (TMD), which anchors them into the lipid membrane. The SNAP-25 is connected to the membranes via palmitoylation, where covalent bonds between multiple cysteine side chains of the protein and palmitoyl side chains of the membrane and multiple cysteine side chains of the protein (Fig. 6A) (Gonzalo et al. 1999; Prescott et al. 2009). Protein palmitoylation is a post-translational modification that modulates the proper function of a variety of eukaryotic fusion proteins. The palmitoylation of the target protein is catalyzed by a family of palmitoyl acyltransferases (PAT), which are localized in the endoplasmic reticulum, Golgi, and plasma membrane. It is widely believed that the S-palmitoylation of SNAP-25 plays an important role in membrane anchoring and targeting. The thioester bonds which formed after palmitoylation are labile and thus resemble other reversible regulatory post-translational protein modifications (Sobocinska et al. 2018). It was proposed that the palmitoyl chains of SNAP-25 are required for membrane targeting (Gonzalo et al. 1999). However, recent results suggest that the lipid linker of SNAP-25 plays a more active role in calcium-triggered exocytosis. The lipid linker might be responsible for the fast exocytosis (rate enhancement) during neurotransmitter release (Nagy et al. 2008). Recently, Shaaban et al. showed that the palmitoylated linker segments interact with neighboring lipids to facilitate the triggering of fusion (Shaaban et al. 2019). We suggest that role of lipid linkers is not restricted only to membrane targeting, rather they take part actively in fusion. Apart from SNAP-25, other intracellular fusion proteins e.g. SNAP-23(Pallavi and Nagaraj 2003), syntaxin-11 (Kinoshita et al. 2019), Ykt6 were reported to be palmitoylated for the membrane association and localization. The exact role of how the palmitoyl chain of these proteins affects fusion was not explored in detail (Hong and Lev 2014).
In the case of a viral fusion protein, such S-palmitoylation was not abundant. As mentioned earlier, the viral fusion protein has TMDs and it attaches to the target membrane via a hydrophobic aminoacid-rich domain, the fusion peptide. Therefore acylation is not directly necessary to bind the target membrane. However, many recent reports suggest a more potential role of S-acylation (with cysteine) of the viral fusion protein in virus entry and infection. In case of the influenza virus, the cytoplasmic tail of hemagglutinin (HA) fusion protein is highly conserved with three cysteine residues. The thiols are post-translationally modified by covalently bound fatty acids. Recently Chlanda et al. mutated three cysteine residues to understand the role of S-palmitoylation (Chlanda et al. 2017). Deacylated HA of different influenza virus strains imparted the virus assembly and membrane fusion. The palmitoylation contributes to the local curvature of the membrane to control fusion and pore expansion (Chlanda et al. 2017). S-palmitoylation of mouse hepatitis virus (MHV) fusion protein (spike) was proposed to mediate its association with lipid microdomains, promote syncytia formation, and increase infectivity (Thorp et al. 2006). In the case of SARS-CoV-2, viral fusion is initiated by a similar glycoprotein (spike) that exists as a trimer (Fig. 6B). The receptor-binding domain of the spike initiates the binding to the angiotensin-converting enzyme 2 (ACE2) receptor, and subsequently, after the protease action, the fusion peptide of spike protein gets exposed and interacts with the target membrane to promote viral fusion. S-acylation of the spike protein by ZDHHC20 is essential for SARS-CoV-2 infectivity, stabilizing the spike protein trimer and facilitating its interaction with lipid bilayers (Wu et al. 2021). It was also found that the S-acylation of spike protein triggers cholesterol/sphingolipid-rich raft-like domains to form, enabling membrane fusion (Vilmen et al. 2021; Mesquita et al. 2021). The palmitoylation and its association to a raft-like domain protect it from degradation (Vilmen et al. 2021). The palmitoylation of SARS‐CoV‐2 spike protein is critical for S‐mediated syncytia formation and SARS‐CoV‐2 pseudo-virus particle entry (Li et al. 2022). Apart from SARS-CoV-2, the spike protein of SARS-CoV has also been shown to promote spike-mediated fusion (Petit et al. 2007; McBride and Machamer 2010). Since palmitoylation is essential for fusion and infectivity of coronavirus, it has been suggested that the fatty acid synthase (FAS) may be a useful target to control fusion and viral infection. The inhibitors of FAS and PAT may reduce viral replication and entry to the host cells (Gadalla and Veit 2020; Chu et al. 2021).
Lipid Anchor and Virus Entry Inhibitor
Conventional antiviral drugs have been developed to target viral proteins and host cell proteins that are involved in the infection process. The type-I viral fusion protein follows a common mechanism and forms a six-helix bundle (6HB) to drive the membrane fusion (Fig. 3B). Many peptides inhibitors were developed that interfere with the formation of the 6HB. T20, a 36-amino acid peptide-based fusion inhibitor developed to inhibit HIV entry. T1249, a 39 amino acid long sequence of gp41 also effectively blocks HIV entry. The covalent binding of a cholesterol moiety to a classical HIV-1 fusion inhibitor peptide, C34 increases its antiviral activities. Conjugating cholesterol to the inhibitory peptide enhances the membrane partitioning of the peptide to the model membrane as well as in human blood cells. Therefore lipidation (cholesterol-conjugated peptide) may exhibit a better inhibitory effect than the parent peptide (Park and Gallagher 2017). GPI-anchored 2P23 (GPI-2P23) derived from the C-terminal HR domain of gp41 completely protected cells (TZM-bl) from infections of divergent HIV-1, HIV-2, and SIV isolates as well as a panel of T20-resistant mutants (Tang et al. 2019). Peptides derived from influenza hemagglutinin (HA) are generally inefficient to inhibit viral entry. However, the C-terminal cholesterol conjugation of these peptides acts as potent inhibitors for influenza entry. Cholesterol-conjugated peptides such as S-KKWK and its derivatives exhibited potent and broad anti-influenza A virus activity in vitro and in vivo. S-KKWK binds to a conserved hydrophobic pocket on the HA2 subunit, thereby preventing the six-helix bundle formation (Wu et al. 2015; Lin et al. 2017). In the case of SARS-CoV, it was shown that the N-terminal HR-derived peptide does not show any inhibitory effect. A C-terminal HR-derived peptide EK1 targets the N-terminal HR region of human coronavirus (HCoV) spike protein. Thus EK1 inhibits infection of different HCoVs, including SARS-CoV-2, MERS-CoV, and SARS-CoVs (Outlaw et al. 2020). Yuxian He and co-workers have synthesized a lipo-peptide EK1V1 by modifying EK1 with cholesterol, which exhibited significantly improved antiviral activity. EK1V1 displayed potent cross inhibitory activities against divergent HIV-1, HIV-2, and simian immunodeficiency virus (SIV) (Yu et al. 2021). Another EK1 derivative, the lipo-peptide EK1C4 increases the inhibitory activity against the SARS-CoV-2 and HIV-1 infection (Xia et al. 2020). The cholesterol-conjugated peptide fusion inhibitor, IPB02 has a high ability to inhibit SARS-CoV-2 and SARS-CoV infections (Zhu et al. 2020). IPB02 was modified further and a new lipo-peptide IPB02V1 was created by introducing a flexible PEG8 linker into IPB02 between the peptide sequence and cholesterol molecule. It was shown that the cholesterol conjugated IPB02V1 exhibited high potency to inhibit all the tested pseudoviruses (Zhu et al. 2021). Recently, Matteo Porotto and co-workers developed dimeric cholesterol-conjugated peptide ([SARSHRC-PEG4]2-chol) fusion inhibitor (de Vries et al. 2021). The peptide robustly inhibited S-mediated fusion of several emerging SARS-CoV-2 variants (de Vries et al. 2021).
Membrane fusion inhibitors that target the lipid bilayer’s physical properties may have a broad spectrum of antiviral efficacy, and they can be useful for drug-resistant mutation. Surfactins are cyclic lipo-peptides naturally produced by various strains of Bacillus subtilis, and they have a broad spectrum of bioactivities. Surfactin showed antiviral activity against several enveloped viruses such as herpes-simplex-virus (HSV-1, HSV-2), Vesicular stomatitis virus (VSV), Simian immunodeficiency virus (SIV), Newcastle disease virus (NDV), and porcine epidemic diarrhea virus (PEDV). The antiviral activity of surfactin is due to the inhibition of membrane fusion events between the virus and host cell (Yuan et al. 2019). Surfactin can be inserted into the cell membrane and stabilize the positive curvature of the outer leaflet (Yuan et al. 2018). Apart from surfactin, the pulmonary surfactants, and the surfactant production stimulants appear to be effective in the treatment of COVID-19 (Mandato and Vajro 2021). It was hypothesized that these surfactants probably interfere with ACE2 receptor binding, however, the positive curvature induction by these molecules cannot be ruled out. The rigid amphipathic fusion inhibitors (RAFIs) are synthetic compounds of inverted cone molecular geometry. The inverted cone molecular geometry of the RAFIs increases the energy barrier for the hemifusion stalk, and the perylene group exposed to visible light may induce viral lipid peroxidation, inhibiting fusion (st. Vincent et al. 2010; Vigant et al. 2014). Recently, our group explored the mechanism of mycobacterium evasion of phagosome-lysosome fusion and tested the antiviral potential of designed lipo-peptides as an effective fusion inhibitor. We have shown that the lipo-dipeptide (myr-WD) acts as a potent fusion inhibitor, whereas the WD peptide (no lipidation) has a marginal effect to inhibit fusion (Sardar et al. 2021). The myr-WD peptide can tackle H1N1 and murine coronavirus infections (Sardar et al. 2021). The lipidation facilitates the peptide binding to the membrane to exert the fusion inhibitory effect (Sardar et al. 2021). The lipidated peptide (myr-WD) increases the membrane order at the interface and also decreases the water penetration. The membrane physical property tuning ability of the lipo-peptide may be due to the specific peptide sequence (WD) coupled with the lipid linker. The fatty acyl part of the lipid linker may directly modulate the energetics of fusion and further studies are necessary in this regard. However, we suggest that lipidation is an important strategy to develop useful antiviral drugs.
Conclusion
Membrane fusion is an important biological process in the life of eukaryotic cells. Fusion is a step-wise process that involves the formation of non-bilayer hemifusion intermediates. The shapes of lipids are important and a non-bilayer forming lipid (that induces negative curvature) generally promotes fusion. In addition, the charge-charge repulsion of negatively charged lipids also contributes to the fusion. The fusion proteins, which catalyze the fusion interact with the lipids, and the specific lipid composition of the membrane may be desirable. The mechanism of action of the fusion proteins is really diverse. During intracellular heterotypic fusion, the SNAREs and the four-helix bundle play the predominant role. During ER/mitochondrial fusion, the GTP hydrolysis (by GTPase domain) and amphipathic helix-membrane interaction catalyze fusion. In the case of viral fusion, the fusion peptide bind to the membrane, and its membrane destabilizing property is highly important apart from the dominant role of six-helix bundle formation. However, we found that the lipid composition of the membrane controls the fusion of all types of fusion proteins. Therefore, the lipid composition has really great role in biological fusion. Some of the fusion proteins get palmitoylated (with the help of PAT, lipidation). The lipidation helps them to anchor to the membrane, improves localization, provides stability, and promotes fusion by stabilizing the hemifusion intermediates. The membrane anchoring ability of lipidation was also helpful to design peptide-based fusion inhibitors that block the six-helix bundle formation in type-I viral protein-mediated fusion. Together, the ‘Lipid World’, be it the lipid composition or the lipidation of the fusion protein has an immense role to control biological fusion.
Data Availability
All data generated or analyzed during this study are included in this article.
References
Baba T, Kashiwagi Y, Arimitsu N et al (2014) Phosphatidic acid (PA)-preferring phospholipase A1 regulates mitochondrial dynamics. J Biol Chem 289:11497–11511. https://doi.org/10.1074/JBC.M113.531921
Bai J, Wang CT, Richards DA et al (2004) Fusion pore dynamics are regulated by synaptotagmin•t-SNARE interactions. Neuron 41:929–942. https://doi.org/10.1016/S0896-6273
Ban T, Ishihara T, Kohno H et al (2017) Molecular basis of selective mitochondrial fusion by heterotypic action between OPA1 and cardiolipin. Nat Cell Biol 19:856–863. https://doi.org/10.1038/ncb3560
Bian X, Klemm RW, Liu TY et al (2011) Structures of the atlastin GTPase provide insight into homotypic fusion of endoplasmic reticulum membranes. Proc Natl Acad Sci USA 108:3976–3981
Cao YL, Meng S, Chen Y et al (2017) MFN1 structures reveal nucleotide-triggered dimerization critical for mitochondrial fusion. Nature 542:372–376. https://doi.org/10.1038/nature21077
Chakraborty H, Tarafdar PK, Bruno MJ et al (2012) Activation thermodynamics of poly(ethylene glycol)-mediated model membrane fusion support mechanistic models of stalk and pore formation. Biophys J 102:2751–2760. https://doi.org/10.1016/J.BPJ.2012.04.053
Chapman ER (2008) How does synaptotagmin trigger neurotransmitter release? Annu Rev Biochem 77:615–641. https://doi.org/10.1146/ANNUREV.BIOCHEM.77.062005.101135
Chernomordik LV, Kozlov MM (2003) Protein-lipid interplay in fusion and fission of biological membranes. Annu Rev Biochem 72:175–207. https://doi.org/10.1146/ANNUREV.BIOCHEM.72.121801.161504
Chlanda P, Mekhedov E, Waters H et al (2016) The hemifusion structure induced by influenza virus haemagglutinin is determined by physical properties of the target membranes. Nat Microbiol 1:1–8. https://doi.org/10.1038/nmicrobiol.2016.50
Chlanda P, Mekhedov E, Waters H et al (2017) Palmitoylation contributes to membrane curvature in influenza A virus assembly and hemagglutinin-mediated membrane fusion. J Virol 91:e00947-e1017. https://doi.org/10.1128/JVI.00947-17
Chu J, Xing C, Du Y et al (2021) Pharmacological inhibition of fatty acid synthesis blocks SARS-CoV-2 replication. Nat Metab 3:1466–1475. https://doi.org/10.1038/s42255-021-00479-4
Cohen MM, Tareste D (2018) Recent insights into the structure and function of mitofusins in mitochondrial fusion. F1000Research 7:1–13. https://doi.org/10.12688/F1000RESEARCH.16629.1
Collins RN, Holz RW, Zimmerberg J (2012) 5.14 The biophysics of membrane fusion. Compr Biophys 5:273. https://doi.org/10.1016/B978-0-12-374920-8.00523-3
Cornell CE, Black RA, Xue M et al (2019) Prebiotic amino acids bind to and stabilize prebiotic fatty acid membranes. Proc Natl Acad Sci USA 116:17239–17244. https://doi.org/10.1073/PNAS.1900275116/-/DCSUPPLEMENTAL
Daste F, Sauvanet C, Bavdek A et al (2018) The heptad repeat domain 1 of Mitofusin has membrane destabilization function in mitochondrial fusion. EMBO Rep 19:e43637. https://doi.org/10.15252/EMBR.201643637
de Vries RD, Schmitz KS, Bovier FT et al (2021) Intranasal fusion inhibitory lipopeptide prevents direct-contact SARS-CoV-2 transmission in ferrets. Science 371:1379–1382. https://doi.org/10.1126/SCIENCE.ABF4896/SUPPL_FILE/ABF4896_S1.MP4
Drin G, Antonny B (2010) Amphipathic helices and membrane curvature. FEBS Lett 584:1840–1847. https://doi.org/10.1016/J.FEBSLET.2009.10.022
Frohman MA (2015) Role of mitochondrial lipids in guiding fission and fusion. J Mol Med 93:263–269. https://doi.org/10.1007/S00109-014-1237-Z
Fuller N, Rand RP (2001) The influence of lysolipids on the spontaneous curvature and bending elasticity of phospholipid membranes. Biophys J 81:243–254. https://doi.org/10.1016/S0006-3495(01)75695-0
Gadalla MR, Veit M (2020) Toward the identification of ZDHHC enzymes required for palmitoylation of viral protein as potential drug targets. Expert Opin Drug Discov 15:159–177. https://doi.org/10.1080/17460441.2020.1696306
Ge Y, Shi X, Boopathy S et al (2020) Two forms of Opa1 cooperate to complete fusion of the mitochondrial inner-membrane. Elife 9:e50973. https://doi.org/10.7554/ELIFE.50973
Gonzalo S, Greentree WK, Linder ME (1999) SNAP-25 is targeted to the plasma membrane through a novel membrane-binding domain. J Biol Chem 274:21313–21318. https://doi.org/10.1074/JBC.274.30.21313
Han J, Pluhackova K, Böckmann RA (2017) The multifaceted role of SNARE proteins in membrane fusion. Front Physiol 8:5. https://doi.org/10.3389/FPHYS.2017.00005/BIBTEX
Haque E, McIntosh TJ, Lentz BR (2001) Influence of lipid composition on physical properties and peg-mediated fusion of curved and uncurved model membrane vesicles: “nature’s own” fusogenic lipid bilayer. Biochemistry 40:4340–4348. https://doi.org/10.1021/BI002030K
Hong WJ, Lev S (2014) Tethering the assembly of SNARE complexes. Trends Cell Biol 24:35–43. https://doi.org/10.1016/J.TCB.2013.09.006
Horvath SE, Daum G (2013) Lipids of mitochondria. Prog Lipid Res 52:590–614. https://doi.org/10.1016/J.PLIPRES.2013.07.002
Hurst LR, Fratti RA (2020) Lipid rafts, sphingolipids, and ergosterol in yeast vacuole fusion and maturation. Front Cell Dev Biol 8:539. https://doi.org/10.3389/FCELL.2020.00539/TEXT
Ingólfsson HI, Melo MN, van Eerden FJ et al (2014) Lipid organization of the plasma membrane. J Am Chem Soc 136:14554–14559. https://doi.org/10.1021/JA507832E/SUPPL_FILE/JA507832E_SI_003.XLSX
Izgu EC, Björkbom A, Kamat NP et al (2016) N-Carboxyanhydride-mediated fatty acylation of amino acids and peptides for functionalization of protocell membranes. J Am Chem Soc 138:16669–16676. https://doi.org/10.1021/JACS.6B08801/ASSET/IMAGES/LARGE/JA-2016-08801F_0005.JPEG
Jahn R, Scheller RH (2006) SNAREs—engines for membrane fusion. Nat Rev Mol Cell Biol 7:631–643. https://doi.org/10.1038/nrm2002
Jahn R, Lang T, Südhof TC (2003) Membrane fusion. Cell 112:519–533. https://doi.org/10.1016/S0092-8674(03)00112-0
Jiang H, Zhang X, Chen X et al (2018) Protein lipidation: occurrence, mechanisms, biological functions, and enabling technologies. Chem Rev 118:919–988. https://doi.org/10.1021/ACS.CHEMREV
Joshi MP, Sawant AA, Rajamani S (2021) Spontaneous emergence of membrane-forming protoamphiphiles from a lipid–amino acid mixture under wet–dry cycles. Chem Sci 12:2970–2978. https://doi.org/10.1039/D0SC05650B
Kinoshita D, Sakurai C, Morita M et al (2019) Syntaxin 11 regulates the stimulus-dependent transport of Toll-like receptor 4 to the plasma membrane by cooperating with SNAP-23 in macrophages. Mol Biol Cell 30:1085–1097. https://doi.org/10.1091/MBC.E18-10-0653/ASSET/IMAGES/LARGE/MBC-30-1085-G008.JPEG
Lancet D, Segrè D, Kahana A (2019) Twenty years of “lipid world”: a fertile partnership with david deamer. Life 9:77. https://doi.org/10.3390/LIFE9040077
Lauwers E, Goodchild R, Verstreken P (2016) Membrane lipids in presynaptic function and disease. Neuron 90:11–25. https://doi.org/10.1016/J.NEURON.2016.02.033
Lee M, Moon Y, Lee S et al (2019) Ergosterol interacts with Sey1p to promote atlastin-mediated endoplasmic reticulum membrane fusion in Saccharomyces cerevisiae. FASEB J 33:3590–3600. https://doi.org/10.1096/FJ.201800779RR
Lee J, Kreutzberger AJB, Odongo L et al (2021) Ebola virus glycoprotein interacts with cholesterol to enhance membrane fusion and cell entry. Nat Struct Mol Biol 28:181–189. https://doi.org/10.1038/s41594-020-00548-4
Li YJ, Cao YL, Feng JX et al (2019) Structural insights of human mitofusin-2 into mitochondrial fusion and CMT2A onset. Nat Commun 10:1–14. https://doi.org/10.1038/s41467-019-12912-0
Li D, Liu Y, Lu Y et al (2022) Palmitoylation of SARS-CoV-2 S protein is critical for S-mediated syncytia formation and virus entry. J Med Virol 94:342–348. https://doi.org/10.1002/JMV.27339
Lin D, Luo Y, Yang G et al (2017) Potent influenza A virus entry inhibitors targeting a conserved region of hemagglutinin. Biochem Pharmacol 144:35–51. https://doi.org/10.1016/J.BCP.2017.07.023
Liu KN, Boxer SG (2020) Target membrane cholesterol modulates single influenza virus membrane fusion efficiency but not rate. Biophys J 118:2426–2433. https://doi.org/10.1016/J.BPJ.2020.03.021
Liu TY, Bian X, Sun S et al (2012) Lipid interaction of the C terminus and association of the transmembrane segments facilitate atlastin-mediated homotypic endoplasmic reticulum fusion. Proc Natl Acad Sci USA 109:E2146–E2154. https://doi.org/10.1073/PNAS.1208385109/SUPPL_FILE/PNAS.201208385SI.PDF
Mandato C, Vajro P (2021) Pulmonary lipid modulation: a possible therapeutic target for SARS-CoV-2 infection. Med Hypotheses 149:110529. https://doi.org/10.1016/J.MEHY.2021.110529
Martens S, McMahon HT (2008) Mechanisms of membrane fusion: disparate players and common principles. Nat Rev Mol Cell Biol 9:543–556. https://doi.org/10.1038/nrm2417
Matsumura A, Higuchi J, Watanabe Y et al (2018) Inactivation of cardiolipin synthase triggers changes in mitochondrial morphology. FEBS Lett 592:209–218. https://doi.org/10.1002/1873-3468.12948
McBride CE, Machamer CE (2010) Palmitoylation of SARS-CoV S protein is necessary for partitioning into detergent-resistant membranes and cell–cell fusion but not interaction with M protein. Virology 405:139–148. https://doi.org/10.1016/J.VIROL.2010.05.031
Meher G, Chakraborty H (2019) Membrane composition modulates fusion by altering membrane properties and fusion peptide structure. J Membr Biol 252:261–272. https://doi.org/10.1007/S00232-019-00064-7
Meher G, Bhattacharjya S, Chakraborty H (2019) Membrane cholesterol modulates oligomeric status and peptide-membrane interaction of severe acute respiratory syndrome coronavirus fusion peptide. J Phys Chem B 123:10654–10662. https://doi.org/10.1021/acs.jpcb.9b08455
Mesquita FS, Abrami L, Sergeeva O et al (2021) S-acylation controls SARS-CoV-2 membrane lipid organization and enhances infectivity. Dev Cell 56:2790–2807. https://doi.org/10.1016/J.DEVCEL.2021.09.016
Milovanovic D, Platen M, Junius M et al (2016) Calcium promotes the formation of syntaxin 1 mesoscale domains through phosphatidylinositol 4,5-bisphosphate. J Biol Chem 291:7868–7876. https://doi.org/10.1074/JBC.M116.716225
Moon Y, Jun Y (2020) The effects of regulatory lipids on intracellular membrane fusion mediated by dynamin-like GTPases. Front Cell Dev Biol 8:518. https://doi.org/10.3389/FCELL.2020.00518/BIBTEX
Murray DH, Tamm LK (2009) Clustering of syntaxin-1A in model membranes is modulated by phosphatidylinositol 4,5-bisphosphate and cholesterol. Biochemistry 48:4617–4625. https://doi.org/10.1021/bi9003217
Murray DH, Tamm LK (2011) Molecular mechanism of cholesterol- and polyphosphoinositide-mediated syntaxin clustering. Biochemistry 50:9014–9022. https://doi.org/10.1021/BI201307U
Nagy G, Milosevic I, Mohrmann R et al (2008) The SNAP-25 linker as an adaptation toward fast exocytosis. Mol Biol Cell 19:3769–3781. https://doi.org/10.1091/MBC.E07-12-1218
Outlaw VK, Bovier FT, Mears MC et al (2020) Inhibition of coronavirus entry in vitro and ex vivo by a lipid-conjugated peptide derived from the SARS-CoV-2 spike glycoprotein HRC domain. mBio 11:1–14. https://doi.org/10.1128/MBIO.01935-20
Pallavi B, Nagaraj R (2003) Palmitoylated peptides from the cysteine-rich domain of SNAP-23 cause membrane fusion depending on peptide length, position of cysteines, and extent of palmitoylation. J Biol Chem 278:12737–12744. https://doi.org/10.1074/JBC.M208598200
Park JE, Gallagher T (2017) Lipidation increases antiviral activities of coronavirus fusion-inhibiting peptides. Virology 511:9–18. https://doi.org/10.1016/J.VIROL.2017.07.033
Pattnaik GP, Bhattacharjya S, Chakraborty H (2021) Enhanced cholesterol-dependent hemifusion by internal fusion peptide 1 of SARS coronavirus-2 compared to its N-terminal counterpart. Biochemistry 60:559–562. https://doi.org/10.1021/ACS.BIOCHEM.1C00046/SUPPL_FILE/BI1C00046_SI_001.PDF
Pérez-Lara Á, Thapa A, Nyenhuis SB et al (2016) PtdInsP2 and PtdSer cooperate to trap synaptotagmin-1 to the plasma membrane in the presence of calcium. Elife 5:e15886. https://doi.org/10.7554/ELIFE.15886
Petit CM, Chouljenko VN, Iyer A et al (2007) Palmitoylation of the cysteine-rich endodomain of the SARS-coronavirus spike glycoprotein is important for spike-mediated cell fusion. Virology 360:264–274. https://doi.org/10.1016/J.VIROL.2006.10.034
Pinot M, Vanni S, Pagnotta S et al (2014) Lipid cell biology. Polyunsaturated phospholipids facilitate membrane deformation and fission by endocytic proteins. Science 345:693–697. https://doi.org/10.1126/SCIENCE.1255288
Prescott GR, Gorleku OA, Greaves J, Chamberlain LH (2009) Palmitoylation of the synaptic vesicle fusion machinery. J Neurochem 110:1135–1149. https://doi.org/10.1111/J.1471-4159.2009.06205.X
Rizo J, Xu J (2015) The synaptic vesicle release machinery. Annu Rev Biophys 44:339–367. https://doi.org/10.1146/ANNUREV-BIOPHYS-060414-034057
Sanders DW, Jumper CC, Ackerman PJ et al (2021) SARS-CoV-2 requires cholesterol for viral entry and pathological syncytia formation. Elife 10:e65962. https://doi.org/10.7554/ELIFE.65962
Sardar A, Lahiri A, Kamble M et al (2021) Translation of mycobacterium survival strategy to develop a lipo-peptide based fusion inhibitor. Angew Chem Int Ed Engl 60:6101–6106. https://doi.org/10.1002/ANIE.202013848
Sarkar S, Das S, Dagar S et al (2020) Prebiological membranes and their role in the emergence of early cellular life. J Membr Biol 253:589–608. https://doi.org/10.1007/S00232-020-00155-W
Segré D, Ben-Eli D, Deamer DW, Lancet D (2001) The lipid world. Orig Life Evol Biosph 31:119–145. https://doi.org/10.1023/A:1006746807104
Shaaban A, Dhara M, Frisch W et al (2019) The SNAP-25 linker supports fusion intermediates by local lipid interactions. Elife 8:e41720. https://doi.org/10.7554/ELIFE.41720
Sobocinska J, Roszczenko-Jasinska P, Ciesielska A, Kwiatkowska K (2018) Protein palmitoylation and its role in bacterial and viral infections. Front Immunol 8:1–19. https://doi.org/10.3389/FIMMU.2017.02003
St. Vincent MR, Colpitts CC, Ustinov AV et al (2010) Rigid amphipathic fusion inhibitors, small molecule antiviral compounds against enveloped viruses. Proc Natl Acad Sci USA 107:17339–17344. https://doi.org/10.1073/PNAS.1010026107
Suazo KF, Park K-Y, Distefano MD (2021) A not-so-ancient grease history: click chemistry and protein lipid modifications. Chem Rev 121:7178–7248. https://doi.org/10.1021/ACS.CHEMREV.0C01108/ASSET/IMAGES/LARGE/CR0C01108_0025.JPEG
Südhof TC (2013) Neurotransmitter release: the last millisecond in the life of a synaptic vesicle. Neuron 80:675–690. https://doi.org/10.1016/J.NEURON.2013.10.022
Sugiura S, Mima J (2016) Physiological lipid composition is vital for homotypic ER membrane fusion mediated by the dynamin-related GTPase Sey1p. Sci Rep 6:1–9. https://doi.org/10.1038/srep20407
Tamm LK, Crane J, Kiessling V (2003) Membrane fusion: a structural perspective on the interplay of lipids and proteins. Curr Opin Struct Biol 13:453–466. https://doi.org/10.1016/S0959-440X(03)00107-6
Tang X, Jin H, Chen Y et al (2019) A membrane-anchored short-peptide fusion inhibitor fully protects target cells from infections of human immunodeficiency virus type 1 (HIV-1), HIV-2, and simian immunodeficiency virus. J Virol 93:1177–1196. https://doi.org/10.1128/JVI.01177-19
Tarafdar PK, Chakraborty H, Dennison SM, Lentz BR (2012) Phosphatidylserine inhibits and calcium promotes model membrane fusion. Biophys J 103:1880–1889. https://doi.org/10.1016/J.BPJ.2012.09.030
Tarafdar PK, Chakraborty H, Bruno MJ, Lentz BR (2015) Phosphatidylserine-dependent catalysis of stalk and pore formation by synaptobrevin JMR-TMD peptide. Biophys J 109:1863–1872. https://doi.org/10.1016/J.BPJ.2015.08.051
Thorp EB, Boscarino JA, Logan HL et al (2006) Palmitoylations on murine coronavirus spike proteins are essential for virion assembly and infectivity. J Virol 80:1280–1289. https://doi.org/10.1128/JVI.80.3.1280-1289.2006
van den Bogaart G, Meyenberg K, Risselada HJ et al (2011) Membrane protein sequestering by ionic protein-lipid interactions. Nature 479:552–555. https://doi.org/10.1038/NATURE10545
Vigant F, Hollmann A, Lee J et al (2014) The rigid amphipathic fusion inhibitor dUY11 acts through photosensitization of viruses. J Virol 88:1849–1853. https://doi.org/10.1128/JVI.02907-13
Vilmen G, Banerjee A, Freed EO (2021) Rafting through the palms: S-acylation of SARS-CoV-2 spike protein induces lipid reorganization. Dev Cell 56:2787–2789. https://doi.org/10.1016/J.DEVCEL.2021.10.002
Wang W, Yang L, Huang HW (2007) Evidence of cholesterol accumulated in high curvature regions: implication to the curvature elastic energy for lipid mixtures. Biophys J 92:2819–2830. https://doi.org/10.1529/BIOPHYSJ.106.097923
Wickner W (2010) Membrane fusion: five lipids, four SNAREs, three chaperones, two nucleotides, and a rab, all dancing in a ring on yeast vacuoles. Annu Rev Cell Dev Biol 26:115–136. https://doi.org/10.1146/annurev-cellbio-100109-104131
Wickner W, Rizo J (2017) A cascade of multiple proteins and lipids catalyzes membrane fusion. Mol Biol Cell 28:707–711. https://doi.org/10.1091/MBC.E16-07-0517
Wu W, Wang J, Lin D et al (2015) Super short membrane-active lipopeptides inhibiting the entry of influenza A virus. Biochim Biophys Acta 1848:2344–2350. https://doi.org/10.1016/J.BBAMEM.2015.06.015
Wu Z, Zhang Z, Wang X et al (2021) Palmitoylation of SARS-CoV-2 S protein is essential for viral infectivity. Signal Transduct Target Ther 6:1–4. https://doi.org/10.1038/s41392-021-00651-y
Xia S, Liu M, Wang C et al (2020) Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res 30:343–355. https://doi.org/10.1038/S41422-020-0305-X
Yan L, Sun S, Wang W et al (2015) Structures of the yeast dynamin-like GTPase Sey1p provide insight into homotypic ER fusion. J Cell Biol 210:961–972. https://doi.org/10.1083/JCB.201502078
Yang ST, Kiessling V, Simmons JA et al (2015) HIV gp41-mediated membrane fusion occurs at edges of cholesterol-rich lipid domains. Nat Chem Biol 11:424–431. https://doi.org/10.1038/NCHEMBIO.1800
Yang ST, Kreutzberger AJB, Lee J et al (2016) The role of cholesterol in membrane fusion. Chem Phys Lipid 199:136–143. https://doi.org/10.1016/J.CHEMPHYSLIP.2016.05.003
Yang ST, Kiessling V, Tamm L (2017) Line tension at lipid phase boundaries as driving force for HIV fusion peptide-mediated fusion. Nat Commun 7:1–14. https://doi.org/10.1038/ncomms11401
Yu D, Zhu Y, Yan H et al (2021) Pan-coronavirus fusion inhibitors possess potent inhibitory activity against HIV-1, HIV-2, and simian immunodeficiency virus. Emerg Microbes Infect 10:810–821. https://doi.org/10.1080/22221751.2021.1917309
Yuan L, Zhang S, Wang Y et al (2018) Surfactin inhibits membrane fusion during invasion of epithelial cells by enveloped viruses. J Virol 92:e008092-e8118. https://doi.org/10.1128/JVI.00809-18
Yuan L, Zhang S, Peng J et al (2019) Synthetic surfactin analogues have improved anti-PEDV properties. PLoS ONE 14:e0215227. https://doi.org/10.1371/JOURNAL.PONE.0215227
Zhu Y, Yu D, Yan H et al (2020) Design of potent membrane fusion inhibitors against SARS-CoV-2, an emerging coronavirus with high fusogenic activity. J Virol 94:e00635-e720. https://doi.org/10.1128/JVI.00635-20
Zhu Y, Yu D, Hu Y et al (2021) SARS-CoV-2-derived fusion inhibitor lipopeptides exhibit highly potent and broad-spectrum activity against divergent human coronaviruses. Signal Transduct Target Ther 6:1–3. https://doi.org/10.1038/s41392-021-00698-x
Acknowledgements
We thank IISER Kolkata for its generous support. PKT acknowledges support from DST-Inspire and the Science and Engineering Research Board (SERB). AS thanks UGC; ND thanks CSIR; BP and DB thank DST-Inspire, for fellowship.
Funding
PKT sincerely thanks the DST-Inspire (Grant No: IFA13-CH-120, 2014) and the Science and Engineering Research Board (SERB) Early Career Award (Grant No: ECR/2016/001935) for the financial support. Funding was provided by University Grants Commission (UGC-Fellowship), CSIR-India (CSIR Fellowship), and Department of Science and Technology, Ministry of Science and Technology, India (Inspire Fellowship).
Author information
Authors and Affiliations
Contributions
AS, ND, BP, DB, and PKT wrote the manuscript. AS, ND, and PKT conceptualized the idea. AS, ND, BP, and DB made the figures.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sardar, A., Dewangan, N., Panda, B. et al. Lipid and Lipidation in Membrane Fusion. J Membrane Biol 255, 691–703 (2022). https://doi.org/10.1007/s00232-022-00267-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-022-00267-5